Abstract
Bacterial contamination of ready-to-eat (RTE) street foods is a major concern worldwide. Dissemination of antibiotic resistant pathogens from food is an emerging public-health threat. To investigate the prevalence of antibiotic resistance genes and ceftazidime resistance-associated efflux pumps in foodborne pathogens, 270 RTE street foods samples were collected in three densely populated Taiwanese cities. Among 70 ceftazidime non-susceptible isolates, 21 Stenotrophomonas maltophilia, 12 Pseudomonas spp., 22 Acinetobacter spp., and 15 Enterobacteriaceae isolates were identified. Phylogenetic analyses revealed high levels of genetic diversity between all of the different strains. Multi-drug resistance was observed in 86.4% (19/22) of Acinetobacter spp., 100% (12/12) of Pseudomonas spp., 71.4% (15/21) of S. maltophilia, and 93.3% (14/15) of Enterobacteriaceae. Of 70 ceftazidime non-susceptible isolates, 13 contained ESBLs or plasmid-mediated ampC genes and 23 contained ceftazidime resistance-associated efflux pumps, with Acinetobacter spp. identified as predominant isolate (69.6%; 16/23). AdeIJK pump RNA expression in Acinetobacter isolates was 1.9- to 2-fold higher in active efflux strains. Nine clinically resistant genes were detected: catIII and cmlA (chloramphenicol); aacC1, aacC2, aacC3, and aacC4 (gentamicin); tet(A), tet(C), and tet(D) (tetracycline). The scope and abundance of multidrug-resistant bacteria described in this report underscores the need for ongoing and/or expanded RTE monitoring and control measures.
Similar content being viewed by others
Introduction
Taiwan is one of many countries with a vibrant street food culture, with ready-to-eat (RTE) foods and snacks easily found at almost any time of day, and with night markets and food stands being major attractions for both domestic and international tourists1,2,3,4. According to one estimate, more than 150,000 types of food and drink items are sold on the streets and in night markets across the country5. Street food culture has always been associated with food poisoning and other gastrointestinal maladies that can also be transmitted in restaurants and various food service venues. According to data gathered by the Republic of China Food and Drug Administration, there was a significant increase in reported food poisoning cases between 1991–2000 (38,938) and 2000–2010 (43,404)6. Taiwanese citizens are clearly aware of this problem, and have organized efforts to get government agencies to deal with it.
RTE meats and salads have long been described as hidden vectors of microbial foodborne pathogens such as Enterobacteriaceae and Listeria monocytogenes 7,8,9. Many researchers have reported on the role of street foods as vectors of pathogenic bacterial transmission to humans. Whereas most studies focus on microbiological and hygienic quality, fewer efforts have been made to identify antibiotic resistance mechanisms and the dissemination of antibiotic resistance genes—important information in the fight against foodborne pathogens10,11,12,13. The literature contains several studies that assessed foodborne pathogens in RTE food in other countries. For example, among 154 foodborne Staphylococcus aureus isolates analyzed in Turkey, 39 (25.3%) were found to have multi-drug resistance14. In one study of vegetables collected in Switzerland, 78.3% (47/60) of multidrug-resistant extended-spectrum β-lactamase (ESBL)-producing Enterobacteriaceae isolates were identified15. ESBLs are enzymes that hydrolyze penicillin as well as aztreonam and first-, second-, and third-generation cephalosporins16. Although, third-generation cephalosporins are broad-spectrum antimicrobial agents against bacterial infection and useful for different type of clinical situations17, the problem of ESBL-producing bacteria has been identified in medical systems and communities worldwide16,18. A primary reason for this rapid spread is that ESBL-producing bacteria can be transmitted by contaminated food or water9,19,20.
In Taiwan, at least two research teams have focused on microbiological quality in RTE foods, but no efforts have been made to determine antibiotic resistance levels and associated resistance mechanisms in these foods10,12. Bacterial isolates containing ESBLs have been detected in hospital patients and livestock, though transmission mechanisms remain unidentified21,22,23,24. To determine the potential contribution of food to the dissemination of antibiotic resistant pathogens, we collected and analyzed 270 RTE street food samples in the densely populated cities of Kaohsiung, Taichung and Taipei, including spring rolls, cold noodles and fruit platters. One of our primary motivations was to determine the prevalence and mechanisms of antibiotic resistance, specifically the contribution of efflux pumps to ceftazidime resistance.
Results
Prevalence of ceftazidime non-susceptible bacterial isolates
Of the 270 food samples (three RTE food types: spring rolls, cold noodles, and fruit platters) we tested, ceftazidime non-susceptible Enterobacteriaceae, S. maltophilia, Pseudomonas spp. and Acinetobacter spp. were selected by eosin-methylene blue (EMB) agar containing ceftazidime (8 μg/ml) and identified using 16S rRNA gene analysis (Fig. 1). Characterization of 70 bacterial isolates identified 21 instances of S. maltophilia, 12 Pseudomonas spp., 22 Acinetobacter spp. and 15 Enterobacteriaceae (Fig. 2A). The highest percentage of ceftazidime non-susceptible bacteria was isolated in cold noodle samples (43.3%; 39 of 90); the spring rolls and fruit platters we tested had similar levels of isolates (16.7% and 17.8%) (Fig. 2B). The lowest percentages were in spring rolls collected in Taipei (1 Pseudomonas aeruginosa and 1 S. maltophilia) and fruit platters collected in Kaohsiung (1 Enterobacter cloacae and 2 Acinetobacter baumannii) (Fig. 1).
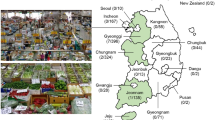
Map showing population densities of geographic locations throughout Taiwan, including the three cities where the RTE street food samples used in this study were collected: Kaohsiung, Taichung and Taipei. The map of population densities in Taiwan was modified from Kanguole (2013) via Wikimedia Commons (https://upload.wikimedia.org/wikipedia/commons/8/88/Population_density_of_Taiwan_by_district.svg). This file is licensed under the Creative Commons Attribution-Share Alike 3.0 Unported licensing agreement (https://creativecommons.org/licenses/by-sa/3.0/).
Antimicrobial susceptibility testing
As shown in Fig. 3, we observed high levels of antibiotic resistance (73.3–100%) among 15 Enterobacteriaceae isolates (Fig. 3A). The lowest level of resistance was for levofloxacin (20%) and the highest for trimethoprim-sulfamethoxazole (53.3%). We observed 27.3–90.9% antibiotic resistance in 22 Acinetobacter spp. isolates (Fig. 3B). Resistance levels for ticarcillin and ticarcillin-clavulanic acid were 68.2% and 40.9%, respectively. According to this finding, combinations of these antibiotics are capable of increased efficacy against ticarcillin-resistant Acinetobacter spp. Among the 12 Pseudomonas spp. isolates, high levels of antibiotic resistance (75–100%) were found in 9 (Fig. 3C); all 12 were susceptible to levofloxacin. Among the 21 S. maltophilia isolates, we noted high resistance levels (71.4–100%) against 3 antibiotics (Fig. 3D). The ticarcillin-clavulanic acid resistance level for S. maltophilia was 66.7%; only 1 isolate was found to be levofloxacin-resistant.
Percentages of antibiotic resistance in 70 bacterial isolates analyzed in this study. CAZ, ceftazidime; CTX, cefotaxime; CHL, chloramphenicol; CL, colistin; GM, gentamicin; LVX, levofloxacin; MEM, meropenem; TIC, ticarcillin; TIM, ticarcillin-clavulanic acid; TE, tetracycline; SXT, trimethoprim-sulfamethoxazole. Shown are bacterial isolate and resistance percentage data for (A) Enterobacteriaceae (N = 15), (B) Acinetobacter spp. (N = 22), (C) Pseudomonas spp. (N = 12) and (D) S. maltophilia (N = 21). Further details are shown in Supplementary Table S2.
For this study, multidrug resistance was defined as resistance to a minimum of 3 antibiotics. Our data indicate the presence of multidrug resistance in 86.4% (19/22) of Acinetobacter spp., 100% (12/12) of Pseudomonas spp., 71.4% (15/21) of S. maltophilia, and 93.3% (14/15) of Enterobacteriaceae isolates (Fig. 3E). These high percentages indicate that RTE street foods in Taiwan may be a significant reservoir for antibiotic resistant pathogens.
Investigation of ceftazidime non-susceptible bacterial isolates
Phylogenetic typing identified 20 pulsotypes in 22 Acinetobacter spp. isolates. Two isolates sharing the same pulsotype, TP36 and TP47, were identified in two fruit platter samples collected in Taipei (Supplementary Fig. S1). PFGE analysis of 12 Pseudomonas spp., 21 S. maltophilia, and 15 Enterobacteriaceae isolates indicate that they were non-clonal (Supplementary Figs S2–S4). The high levels of genetic diversity and multi-drug resistance identified among these bacterial isolates suggest that they developed antibiotic resistance mechanisms independently in the three cities where samples were collected.
Ceftazidime resistance mechanism
Among the 70 ceftazidime non-susceptible isolates that were the focus of this study, 13 contained ESBLs or plasmid-mediated ampC genes (Table 1). bla PER-1 was found in 5 Acinetobacter spp. isolates and bla CTX-M-9 was detected in E. coli (n = 1), S. maltophilia (n = 1), and A. baumannii (n = 1). This suggests that bla CTX-M-9 is horizontally transferred among various bacterial species.
To clarify the contribution of efflux pumps to ceftazidime resistance, we used the efflux pump inhibitor CCCP to analyze the 70 ceftazidime non-susceptible isolates. Bacterial isolates containing ceftazidime resistance-associated efflux pumps were identified as any CCCP- inoculated strain exhibiting a minimum 4-fold decrease in minimum inhibitory concentration (MIC). Our data indicate that 72.7% (16/22) of the Acinetobacter spp. and 25% (3/12) of the Pseudomonas spp. isolates contained efflux pumps conferring resistance to ceftazidime (Fig. 4). A ceftazidime-induced effect on efflux activity was observed in only 3 S. maltophilia isolates (14.3%) and 1 Enterobacteriaceae isolate (6.7%). These findings suggest a significant prevalence of ceftazidime resistance-associated efflux pumps in Acinetobacter spp.
Efflux pump contributions to ceftazidime resistance in 70 non-susceptible bacterial isolates. Ceftazidime resistance was defined as a minimum four-fold decrease in MIC level in the presence of carbonyl cyanide-m-chlorophenyl hydrazine (CCCP). Number of strains are bacterial isolates with ceftazidime resistance-associated efflux pumps.
RNA expression of two efflux pumps in Acinetobacter spp
Previous studies have determined a link between the active efflux pumps adeDE and adeIJK and resistance to third-generation cephalosporins in Acinetobacter spp.25,26 We analyzed expression of adeDE and adeIJK in four representative isolates exhibiting efflux pump activity (strains KA19, KA82, TP36, TP68) relative to a control strain that did not (strain TC24). Our data indicate that adeE expression was not significantly different from that of the control strain (Fig. 5A), while adeI expression was 1.9- to 2.2-fold (p < 0.05) higher than in the control strain (Fig. 5B). These results suggest that ceftazidime resistance in Acinetobacter spp. may be attributed to adeI expression levels.
Antimicrobial resistance determinants
Among the 70 isolates, 4 contained chloramphenicol resistance genes, 7 gentamicin resistance genes, and 3 tetracycline resistance genes (Table 2). Among the 4 with chloramphenicol resistance genes, 1 S. maltophilia and 3 Pseudomonas spp. isolates encoded catIII and cmlA genes, respectively. Among the 7 containing gentamicin resistance genes, aacC2 was identified in 3 (2 Acinetobacter spp. and 1 Pseudomonas spp.) and aacC4 in 2 (1 Acinetobacter spp. and 1 Enterobacteriaceae). Two Acinetobacter spp. isolates also carried the gentamicin resistance genes aacC1 and aacC3. Three tetracycline resistance genes (tet(A), tet(C) and tet(D)) were identified in three Enterobacteriaceae isolates.
Discussion
The literature contains studies of the microbiological quality of various RTE foods in countries around the world. Oliveira et al. tested 162 minimally processed leafy vegetable samples in Brazil and reported that 53.1% contained E. coli, 3.7% Listeria spp., and 1.2% Salmonella spp.13. In Argentina, the foodborne pathogens Bacillus cereus, Clostridium perfringens, S. aureus and Salmonella spp. were found in 101 RTE cooked food samples collected from the central kitchen facility of a school district27. The researchers focused on B. cereus (found in 63.4% of their samples) and reported bacterial counts of <4 log CFU/g. In a Portuguese study, Campos et al. described poor microbiological quality in a number of RTE food types and food products prepared by vendors working out of trailers: 100% tested positive for Enterobacteriaceae and coliforms, 20% for E. coli (4 hamburgers, 4 other trailer foods), and 20% for L. monocytogenes (2 hamburgers/2 hotdogs, 3 other trailer foods)7. In an Italian study of whole vegetables and RTE salads, Salmonella spp., L. monocytogenes, E. coli O157:H7, hepatitis A, and noroviruses were detected in 964 samples28. Of these, only 2 and 3 tested positive for Salmonella spp. and L. monocytogenes, respectively.
In one of two studies conducted in Taiwan, Fang et al. reported the following levels of contamination in RTE food products stored at 18 °C: 7.9% E. coli, 49.8% B. cereus, 17.9% S. aureus, and 42.7% Pseudomonas spp.10. Among the 4 major food types they tested (all stored at 18 °C), coliforms were found in 88% of ham samples, 80% of seafood, 72.7% of other meats, and 62.2% of vegetables. In a separate study, Wei et al. reported S. aureus contamination in 19% of commercial RTE food products purchased from traditional markets, 12.7% from supermarkets, and 9.5% from warehouse stores12. They also identified E. coli in 5–7% of all food samples obtained from supermarkets and warehouse stores. Neither one of these studies looked at antimicrobial susceptibility patterns or resistance mechanisms.
In the present study, the highest percentages of ceftazidime-resistant bacteria were isolated in RTE street food samples of cold noodles (43.3%), followed by fruit platters (17.8%) and spring rolls (16.7%) (Fig. 2B). Results from efforts to determine antimicrobial resistance patterns indicate unexpectedly high levels of multi-drug resistance (71.4–100%) (Fig. 3E). Low resistance was determined for levofloxacin (Fig. 3A–D), indicating that it should be considered for treating severe cases of food poisoning. While meropenem resistance levels were high in Enterobacteriaceae (13/15; 86.7%), Acinetobacter spp. (20/22; 90.9%), and Pseudomonas spp. (12/12; 100%) (Supplementary Table S2), carbapenemases were not detected in these isolates. Carbapenemases such as bla KPC-2 and bla IMP-8 tend to be detected in hospitalized patients, with low potential for widespread dissemination to communities or via RTE foods29,30.
Researchers studying different types of salads in Portugal and Switzerland have reported that RTE food pathogens share the same ESBL genes as clinical isolates collected from hospital patients, suggesting the potential for commensal bacteria to act as antibiotic-resistant gene reservoirs, especially in hospital settings9,20. Further, ESBL-producing Enterobacteriaceae isolates have been reported in beef burger, seafood, and raw vegetable samples15,31,32. Based on these reports, ESBL gene sources in human food chains include both RTE and non-RTE food products. In the present study we identified various ESBLs and plasmid-mediated ampC genes commonly found among bacteria isolates collected in Taiwanese hospitals21,22,23. These genes were especially abundant in ceftazidime non-susceptible non-fermenting gram-negative bacilli, mostly Acinetobacter spp. (Table 1). The data confirm the presence of antibiotic-resistant gene reservoirs in RTE street foods sold in Taiwan’s three largest cities, and that stepped-up surveillance efforts are required to monitor their movement.
Efflux pump activity has been associated with ceftazidime resistance25,26. Quantitative RT-PCR results indicate that the AdeIJK efflux pump contributed to the ceftazidime resistance that we observed in Acinetobacter spp. (Fig. 5). Specifically, among the 22 ceftazidime non-susceptible Acinetobacter spp. isolates we examined, 3 contained ESBL genes, 5 contained ESBL genes plus efflux pump activity, 11 efflux pump activity only, and 3 unknown resistance mechanisms. To our knowledge, this is the first report to demonstrate a link between the AdeIJK efflux pump and ceftazidime resistance in foodborne pathogens. One previous study reported that efflux pumps contributed to 94.4% of ceftazidime-resistant (17/18) and 72.7% of chloramphenicol-resistant (16/22) Burkholderia cepacia complex isolates21. Combined, these reports and data from the present study indicate that efflux pump activity plays an important role in antibiotic resistance mechanisms. The low ceftazidime resistance levels that we observed in S. maltophilia and Enterobacteriaceae isolates suggest that the ceftazidime resistance noted in this study is attributable to an unknown ESBL gene or different efflux pump type. We will attempt to confirm this finding and identify the gene and pump types in a future study.
Our results confirm the capacity of ready-to-eat street foods found in Taiwan, especially cold noodles, to act as antimicrobial-resistant bacteria reservoirs. Additional research efforts are required to quantify the antimicrobial-resistant bacteria that were isolated and analyzed in the present study. The potential for the horizontal transfer of resistance genes and the transmission of multidrug-resistant bacteria warrant further monitoring and control measures throughout Taiwan, but especially in densely populated urban areas. Unfortunately, one study concluded that special training in hygienic practices in Taiwan did not result in a sufficient increase in knowledge or effective practices among food vendors33. Clearly a renewed effort needs to be made to establish strict education standards for hygienic practices among street food vendors and food preparers, with local health officials monitoring such effort to ensure RTE food product safety.
Methods
Sample collection
A total of 270 food samples (90 spring rolls, 90 servings of cold noodles, and 90 fruit platters) were collected from randomly selected vendors operating in Kaohsiung, Taichung, and Taipei between June and November of 2014. All samples were transported to a single laboratory in their original packaging within 2 h (Kaohsiung and Taichung), or within 4 h at a slightly chilled temperature of 16 °C (Taipei).
Microbiological analysis
Ten grams of each sample were homogenized in a stomacher blender, enriched using brain-heart infusion broth (BD, East Rutherford, NJ), and incubated overnight at 37 °C. Individual loopfuls of bacterial suspension were added to eosin-methylene blue (EMB) agar containing ceftazidime (8 μg/ml) and incubated overnight at 37 °C. When different colony morphologies were observed in the same food sample, all colonies were subcultured on EMB agar containing 8 μg/ml ceftazidime. Bacterial identification was performed using 16S rRNA gene sequencing as described in a previous study30. ESBL production was confirmed by tests recommended by the Clinical and Laboratory Standards Institute, using cefotaxime (30 mg) and ceftazidime (30 mg) disks alone and in combination with clavulanic acid (10 mg) (Becton, Dickinson, MD)34. Positive phenotypes were defined as growth-inhibitory zones from either cefotaxime or ceftazidime disks plus clavulanic acid ≥5 mm of growth-inhibitory zones from either cefotaxime or ceftazidime disks.
Antimicrobial susceptibility
These tests were performed using agar dilution methods according to Clinical and Laboratory Standards Institute guidelines34. Minimum inhibitory concentration (MIC) was defined as the lowest concentration of an antibiotic preventing bacterial growth after 16 to 20 h of incubation at 37 °C. We tested 11 antimicrobial agents: cefotaxime, ceftazidime, chloramphenicol, colistin, gentamicin, levofloxacin, meropenem, ticarcillin, ticarcillin-clavulanic acid, tetracycline, and trimethoprim-sulphamethoxazole.
Pulsed-field gel electrophoresis (PFGE)
PFGE typing of restriction enzyme-digested DNA (New England BioLabs, Ipswich, MA) was prepared as previously described21. Enterobacteriaceae and S. maltophilia were digested with XbaI; Pseudomonas spp. and Acinetobacter spp. were digested with SpeI and ApaI, respectively. Restriction fragments ranging from 50 to 500 kb were separated using a CHEF Mapper apparatus (Bio-Rad) for 20 h at 200 V and 14 °C. Gels were stained with ethidium bromide and photographed under UV light. Dice similarity indices were used to construct pulsotype relationship dendrogrammes by an unweighted pair group method using arithmetic averages (UPGMA) with BioNumerics software (v.6.5, Applied Maths). Based on dendrogramme results, pulsotypes were assigned to the same cluster if they exhibited at least 80% similarity.
Detection of antibiotic resistance genes
Plasmid DNA was extracted using a QIAGEN Plasmid Mini Kit. ESBLs (bla SHV, bla TEM , bla CTX-M-G1, bla CTX-M-G2, bla CTX-M-G9, bla PER, bla VEB); OXA genes (bla OXA-1, bla OXA-2, bla OXA-9, and bla OXA-10 genes); plasmid-mediated AmpC genes (bla DHA and bla CMY)23,30,35,36; carbapenemases (bla KPC, bla NDM, bla VIM, bla IMP, bla NMC, bla SME, bla SPM-1, bla GIM-1, bla SIM-1, bla IMI, bla GES, and bla OXA-48)30; chloramphenicol resistance genes (catI, catII, catIII and cmlA)37; tetracycline resistance genes (tet(A), tet(B), tet(C), tet(D), tet(E) and tet(G)); and gentamicin resistance genes (aacC1, aacC2, aacC3, and aacC4)38 were detected by PCR, using primer sets listed in previous reports (Supplementary Table S1). Positive controls for these resistance genes were included in all PCR analyses.
Efflux pump activity assays
To evaluate the contributions of efflux pump activity to ceftazidime resistance among the isolates, MIC resistance patterns were determined via Mueller-Hinton agar dilution with and without 12.5 μM of the efflux pump inhibitor carbonyl cyanide-m-chlorophenyl hydrazone (CCCP; Sigma- Aldrich, St. Louis, MO)39. Bacterial isolates containing ceftazidime resistance-associated efflux pumps were identified as any CCCP-inoculated strain exhibiting a minimum 4-fold decrease in MIC.
Quantitative RNA expression for efflux pumps
RNA extraction and cDNA synthesis were performed as described in our previous study21. PCR reactions took place in buffer containing 1X FastStart Universal SYBR Green Master (Roche), 300 nM primers, and 2 μl cDNA in an ABI7000 machine following manufacturer instructions (96-well plates). AdeIJK and AdeDE efflux pumps have previously been described as contributing to ceftazidime resistance in Acinetobacter spp.26,40. Quantitative RNA expression of adeI and adeE was analyzed using the primers listed in Supplementary Table S1 40. Relative fold changes in the transcript levels of indicated genes were normalized to the 16S rDNA gene (internal control) and calculated using the 2−ΔΔCT method. Non-parametric Mann-Whitney U tests were used to examine differences in quantitative RNA expressions of adeI and adeE between test (ceftazidime resistance-associated efflux pumps) and control strains (no efflux pump activity). Statistical significance was established as p < 0.05.
References
Chang, J., Min, J. C. H., Pearl Lin, Y. H. & Chiang, C. H. Profiling Japanese tourists visiting night markets in Taiwan. J Qual Assur Hospit Tourism 8, 25–44, https://doi.org/10.1300/J162v08n02_02 (2007).
Hsieh, A. T. & Chang, J. Shopping and tourist night markets in Taiwan. Tour Manag 27, 138–145, https://doi.org/10.1016/j.tourman.2004.06.017 (2006).
Koo, L. C., Fredrick, K. C. T. & John, H. C. Y. Preferential segmentation of restaurant attributes through conjoint analysis. Int J Contemp Hosp M 11, 242–253, https://doi.org/10.1108/09596119910272784 (1999).
Tinker, I. Street foods into the 21st century. Agric Human Values 16, 327–333, https://doi.org/10.1023/a:1007569728512 (1999).
DGBAS (Directorate-General of Budget, A., and Statistics of Taiwan). 2003 annual survey and analysis report on Taiwan vendors. Taipei, Taiwan: Executive Yuan National Statistics. Retrieved November 9th, 2010, from www.dgbas.gov.tw/public/Data/943015213971.doc. (2005).
Dong, T. T. M. The knowledge, attitude, and practice of consumers towards food safety issues: A review of Taiwan. Int J Res Stud Manage 4, 13–22, https://doi.org/10.5861/ijrsm.2015.976 (2015).
Campos, J., Gil, J., Mourão, J., Peixe, L. & Antunes, P. Ready-to-eat street-vended food as a potential vehicle of bacterial pathogens and antimicrobial resistance: An exploratory study in Porto region, Portugal. Int J Food Microbiol 206, 1–6, https://doi.org/10.1016/j.ijfoodmicro.2015.04.016 (2015).
El-Shenawy, M., El-Shenawy, M., Manes, J. & Soriano, J. M. Listeria spp. in street-vended ready-to-eat foods. Interdiscip Perspect. Infect Dis 2011, 968031, https://doi.org/10.1155/2011/968031 (2011).
Campos, J. et al. Microbiological quality of ready-to-eat salads: an underestimated vehicle of bacteria and clinically relevant antibiotic resistance genes. Int J Food Microbiol 166, 464–470, https://doi.org/10.1016/j.ijfoodmicro.2013.08.005 (2013).
Fang, T. J., Wei, Q. K., Liao, C. W., Hung, M. J. & Wang, T. H. Microbiological quality of 18 degrees C ready-to-eat food products sold in Taiwan. Int J Food Microbiol 80, 241–250, https://doi.org/10.1016/s0168-1605(02)00172-1 (2003).
Gormley, F. J., Little, C. L., Grant, K. A., de Pinna, E. & McLauchlin, J. The microbiological safety of ready-to-eat specialty meats from markets and specialty food shops: a UK wide study with a focus on Salmonella and Listeria monocytogenes. Food microbiol 27, 243–249, https://doi.org/10.1016/j.fm.2009.10.009 (2010).
Wei, Q. K., Hwang, S. L. & Chen, T. R. Microbiological quality of ready-to-eat food products in southern Taiwan. J Food Drug Anal 14, 68–73 (2006).
Oliveira, M. A. d., M. de Souza, V., Morato Bergamini, A. M. & De Martinis, E. C. P. Microbiological quality of ready-to-eat minimally processed vegetables consumed in Brazil. Food Control 22, 1400–1403, https://doi.org/10.1016/j.foodcont.2011.02.020 (2011).
Aydin, A. et al. Prevalence and antibiotic resistance of foodborne Staphylococcus aureus isolates in Turkey. Foodborne Pathog Dis 8, 63–69, https://doi.org/10.1089/fpd.2010.0613 (2011).
Zurfluh, K. et al. Extended-spectrum-β-lactamase-producing Enterobacteriaceae isolated from vegetables imported from the Dominican Republic, India, Thailand, and Vietnam. Appl Environ Microbiol 81, 3115–3120, https://doi.org/10.1128/AEM.00258-15 (2015).
El Salabi, A., Walsh, T. R. & Chouchani, C. Extended spectrum β-lactamases, carbapenemases and mobile genetic elements responsible for antibiotics resistance in Gram-negative bacteria. Crit Rev Microbiol 39, 113–122, https://doi.org/10.3109/1040841X.2012.691870 (2013).
Klein, N. C. & Cunha, B. A. Third-generation cephalosporins. Med Clin North Am 79, 705–719 (1995).
Song, J. H., Thamlikitkul, V. & Hsueh, P. R. Clinical and economic burden of community-acquired pneumonia amongst adults in the Asia-Pacific region. Int J Antimicrob Agents 38, 108–117, https://doi.org/10.1016/j.ijantimicag.2011.02.017 (2011).
Mesa, R. J. et al. Extended-spectrum β-lactamase-producing Enterobacteriaceae in different environments (humans, food, animal farms and sewage). J Antimicrob Chemother 58, 211–215, https://doi.org/10.1093/jac/dkl211 (2006).
Nuesch-Inderbinen, M., Zurfluh, K., Peterhans, S., Hachler, H. & Stephan, R. Assessment of the prevalence of extended-spectrum β-lactamase-producing Enterobacteriaceae in ready-to-eat salads, fresh-cut fruit, and sprouts from the Swiss market. J Food Prot 78, 1178–1181, https://doi.org/10.4315/0362-028X.JFP-15-018 (2015).
Tseng, S. P. et al. The contribution of antibiotic resistance mechanisms in clinical Burkholderia cepacia complex isolates: an emphasis on efflux pump activity. PloS One 9, e104986, https://doi.org/10.1371/journal.pone.0104986 (2014).
Yan, J. J. et al. Complexity of Klebsiella pneumoniae isolates resistant to both cephamycins and extended-spectrum cephalosporins at a teaching hospital in Taiwan. J Clin Microbiol 42, 5337–5340, https://doi.org/10.1128/JCM.42.11.5337-5340.2004 (2004).
Yan, J. J., Tsai, S. H., Chuang, C. L. & Wu, J. J. OXA-type β-lactamases among extended-spectrum cephalosporin-resistant Pseudomonas aeruginosa isolates in a university hospital in southern Taiwan. J Microbiol Immunol Infect 39, 130–134 (2006).
Yan, J. J. et al. Dissemination of bla CMY-2 among Escherichia coli isolates from food animals, retail ground meats, and humans in southern Taiwan. Antimicrob Agents Chemother 48, 1353–1356, https://doi.org/10.1128/AAC.48.4.1353–1356.2004 (2004).
Chau, S. L., Chu, Y. W. & Houang, E. T. Novel resistance-nodulation-cell division efflux system AdeDE in Acinetobacter genomic DNA group 3. Antimicrob Agents Chemother 48, 4054–4055, https://doi.org/10.1128/AAC.48.10.4054-4055.2004 (2004).
Damier-Piolle, L., Magnet, S., Bremont, S., Lambert, T. & Courvalin, P. AdeIJK, a resistance-nodulation-cell division pump effluxing multiple antibiotics in Acinetobacter baumannii. Antimicrob Agents Chemother 52, 557–562, https://doi.org/10.1128/AAC.00732-07 (2008).
Tessi, M. A. et al. Microbiological quality and safety of ready-to-eat cooked foods from a centralized school kitchen in Argentina. J Food Prot 65, 636–642 (2002).
De Giusti, M. et al. The evaluation of the microbial safety of fresh ready-to-eat vegetables produced by different technologies in Italy. J Appl Microbiol 109, 996–1006, https://doi.org/10.1111/j.1365-2672.2010.04727.x (2010).
Chiu, S. K. et al. National surveillance study on carbapenem non-susceptible Klebsiella pneumoniae in Taiwan: The emergence and rapid dissemination of KPC-2 carbapenemase. PloS One 8, e69428, https://doi.org/10.1371/journal.pone.0069428 (2013).
Tseng, S. P. et al. First report of bla IMP-8 in Raoultella planticola. Antimicrob Agents Chemother 58, 593–595, https://doi.org/10.1128/AAC.00231-13 (2014).
Fernandes, S. A. et al. Prevalence of extended-spectrum β-lactamases CTX-M-8 and CTX-M-2-producing Salmonella serotypes from clinical and nonhuman isolates in Brazil. Microb Drug Resist 23, 580–589, https://doi.org/10.1089/mdr.2016.0085 (2017).
Nguyen, D. T. et al. Prevalence, antibiotic resistance, and extended-spectrum and AmpC β-lactamase productivity of Salmonella isolates from raw meat and seafood samples in Ho Chi Minh City, Vietnam. Int J Food Microbiol 236, 115–122, https://doi.org/10.1016/j.ijfoodmicro.2016.07.017 (2016).
Sun, Y. M., Wang, S. T. & Huang, K. W. Hygiene knowledge and practices of night market food vendors in Tainan City, Taiwan. Food Control 23, 159–164, https://doi.org/10.1016/j.foodcont.2011.07.003 (2012).
Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. Twenty-third informational supplement. Document M100-S23 CLSI, Wayne, PA. (2013).
Fallah, F. et al. Prevalence of bla NDM, bla PER, bla VEB, bla IMP, and bla VIM genes among Acinetobacter baumannii isolated from two hospitals of Tehran, Iran. Scientifica (Cairo) 2014, 245162, https://doi.org/10.1155/2014/245162 (2014).
Peymani, A. et al. Emergence of OXA-type extended-spectrum β-lactamases among Enterobacter cloacae isolates collected From hospitals of Tehran, Karaj and Qazvin, Iran. Biotech Health Sci 1, e22085, https://doi.org/10.17795/bhs-22085 (2014).
Ahmed, M. O., Clegg, P. D., Williams, N. J., Baptiste, K. E. & Bennett, M. Antimicrobial resistance in equine faecal Escherichia coli isolates from North West England. Ann Clin Microbiol Antimicrob 9, 12, https://doi.org/10.1186/1476-0711-9-12 (2010).
Saenz, Y. et al. Mechanisms of resistance in multiple-antibiotic-resistant Escherichia coli strains of human, animal, and food origins. Antimicrob Agents Chemother 48, 3996–4001, https://doi.org/10.1128/AAC.48.10.3996-4001.2004 (2004).
Pournaras, S. et al. Outbreak of multiple clones of imipenem-resistant Acinetobacter baumannii isolates expressing OXA-58 carbapenemase in an intensive care unit. J Antimicrob Chemother 57, 557–561, https://doi.org/10.1093/jac/dkl004 (2006).
Chu, Y. W., Chau, S. L. & Houang, E. T. Presence of active efflux systems AdeABC, AdeDE and AdeXYZ in different Acinetobacter genomic DNA groups. J Med Microbiol 55, 477–478, https://doi.org/10.1099/jmm.0.46433-0 (2006).
Acknowledgements
This study was supported by grants from the Ministry of Science and Technology of Taiwan (MOST 104-2511-S-214-003-MY3 and MOST 105-2320-B-037-003).
Author information
Authors and Affiliations
Contributions
Conceptualization: L.L., S.P.T. Data curation: S.F.W., T.Y.Y., M.Y.C. Formal analysis: T.Y.Y., W.C.H., M.Y.C. Funding acquisition: L.L., S.P.T. Investigation: W.C.H. Writing original draft: L.L., S.P.T. Writing review & editing: L.L., S.P.T.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, L., Wang, SF., Yang, TY. et al. Antimicrobial resistance and genetic diversity in ceftazidime non-susceptible bacterial pathogens from ready-to-eat street foods in three Taiwanese cities. Sci Rep 7, 15515 (2017). https://doi.org/10.1038/s41598-017-15627-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15627-8
This article is cited by
-
Optimizing the Antibacterial Activity of Iron Oxide Nanoparticles Using Central Composite Design
Journal of Inorganic and Organometallic Polymers and Materials (2022)
-
Microbiological safety of street-vended foods in Bangladesh
Journal of Consumer Protection and Food Safety (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.







