Abstract
To understand the dynamics behind the worldwide spread of the mcr-1 gene, we determined the population structure of Escherichia coli and of mobile genetic elements (MGEs) carrying the mcr-1 gene. After a systematic review of the literature we included 65 E. coli whole genome sequences (WGS), adding 6 recently sequenced travel related isolates, and 312 MLST profiles. We included 219 MGEs described in 7 Enterobacteriaceae species isolated from human, animal and environmental samples. Despite a high overall diversity, 2 lineages were observed in the E. coli population that may function as reservoirs of the mcr-1 gene, the largest of which was linked to ST10, a sequence type known for its ubiquity in human faecal samples and in food samples. No genotypic clustering by geographical origin or isolation source was observed. Amongst a total of 13 plasmid incompatibility types, the IncI2, IncX4 and IncHI2 plasmids accounted for more than 90% of MGEs carrying the mcr-1 gene. We observed significant geographical clustering with regional spread of IncHI2 plasmids in Europe and IncI2 in Asia. These findings point towards promiscuous spread of the mcr-1 gene by efficient horizontal gene transfer dominated by a limited number of plasmid incompatibility types.
Similar content being viewed by others
Introduction
Antimicrobial resistance (AMR) represents a growing threat to global health1. With barely any new antimicrobial drugs in development2, limiting the spread of AMR is key in order to maintain current treatment options3.
Colistin is an antibiotic of the polymyxin class, discovered in 1950 and effective against Gram-negative bacteria4. The emergence of multidrug-resistant Gram-negative bacteria, especially those producing carbapenemases, has reintroduced colistin as a last resort antibiotic for the treatment of severe infections5. In contrast to its limited use in humans, colistin is widely used in food-producing animals6. While colistin resistance was long thought to be caused by chromosomal mutations only7, the emergence of plasmid-mediated resistance, conferred by the mobilized colistin resistance (mcr-1) gene, was recently reported8. This gene encodes for a protein of the phosphoethanolamine transferase enzyme family, and its expression results in the addition of a phosphoethanolamine to lipid A, the target of colistin, decreasing the interaction between colistin and the bacterial lipopolysaccharide8. Since its discovery in 2015 in China, this gene has been described in several bacterial species that were isolated from animals, animal food products, humans and environmental samples from around the world9,10,11,12,13. Our previous study in travellers indicated acquisition of mcr-1 carrying bacteria by healthy individuals during travel to destinations around the world, potentially related to food exposure, as well as rapid clearance after return14. It has been suggested that mcr-1 has spread from food animals to humans8,15,16,17, but there is a lack of comparison of mcr-1 carrying isolates on a global level to support this hypothesis.
We studied the global population structure as well as the geographic and host distribution of mcr-1-carrying Escherichia coli, and mobile genetic elements (MGEs), to establish the population structure and to assess whether the spread of the mcr-1 gene is linked to clonal dissemination or transmission of MGEs from animal, human, or environmental sources within geographic regions.
Results
Literature search
A systematic review of the literature on mcr-1, published until 1 January 2017 resulted in the inclusion of 95 articles, representing a total of 410 entries (whole genome sequences, MLST profiles, and/or plasmid types) for analysis (See detailed methods and results in Supplementary data, Supplementary Figure 1 and Supplementary Table 1).
Population structure
Whole genome sequencing (WGS)
The genomes of 65 mcr-1-carrying E. coli were analysed, including 6 genomes from E. coli isolated from travellers that were sequenced for the purpose of the present study. Isolates originated from Asia (n = 36; 55.4%), Europe (n = 20; 30.8%), North-America (n = 4; 6.2%), South-America (n = 4; 6.2%) and Africa (n = 1; 1.5%). 45 were of animal origin (69.2%), 19 of human origin (29.2%) and one strain (1.5%) was isolated from water (Supplementary Table 1).
The average size of the genomes (all contigs in each assembly, representing chromosomes and plasmids) of these 65 isolates was 4.9 Mbp, with a median number of genes identified of 4785 (ranging from 4266 to 7083), representing a pangenome of 23248 genes and a core genome (defined by genes present in at least 99% of the isolates) of 2216 genes. An unbiased analysis of the population structure was performed using a Bayesian approach with the BAPS software18, based on the nucleotide alignment of the core genome sequences. It revealed the presence of 5 distinct phylogenetic clusters (Fig. 1; Supplementary Figure 2; Supplementary Table 1). The largest cluster (cluster 1) consisted of 26 isolates from 16 different STs (26/65; 40.0%) and the second cluster consisted of 24 isolates from 15 different STs (36.9%). No significant relationship between clustering (BAPS) and geographical origin or isolation source was observed (χ2-test) (Fig. 1) except that all 5 isolates that belong to BAPS cluster 3 are from Europe. Twenty isolates showed less than 10 SNPs/Mbp difference with at least 1 other isolate and were considered clonally related (Supplementary Tables 2 and 3).
Maximum-likelihood tree based on concatenated core genome sequences of 65 mcr-1-carrying E. coli isolates. Branch colours indicate phylogenetic clusters as determined by BAPS. Isolates from ST10, ST165 and closely related isolates are all grouped in the BAPS cluster 2 (dark blue). Leaf (isolates identifiers) colours indicate geographical region of origin. Isolation source is indicated in brackets: A = animal or meat; H = human; E = environment. The 6 travellers’ isolates that were sequenced for this study are highlighted in bold and names start with CBT. Tree scale in number of substitutions per site. *Number of isolates.
Multilocus sequence typing (MLST)
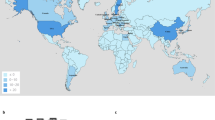
For 312 E. coli isolates originating from 69 studies, a MLST profile was published or could be deduced from the corresponding WGS. Of these, 206 were isolated from animals or animal products (66.0%), 101 were isolated from humans (32.4%), including the 6 travel acquired isolates, and 5 from the environment (1.6%). 141 Isolates from 25 studies (141/312; 45.2%) originated from Asia and 125 isolates from 25 studies (40.1%) from Europe, together accounting for 85.3% of all included isolates. The isolates represented 112 unique sequence types (STs) with ST10 being most common, comprising 40/312 (12.8%) isolates originating from Africa, Asia, Europe and South-America.
eBURST analysis19 was performed on all isolates included in the study, to identify their genetic relatedness based on their MLST profiles. Three main clusters were identified, for which the predicted founders, i.e. the ST in a cluster from which all other SLVs and DLVs in the cluster have most likely diversified19, were ST10, ST1114 and ST410. The largest cluster contained all 40 ST10 isolates and an additional 46 isolates in 21 STs that were single (SLV) or double locus variants (DLV) of ST10 (86/312; 27.6%) (Supplementary Figure 3). The predicted founder of the second largest cluster was ST1114, a SLV of ST165 and ST100, and included 19 isolates belonging to 7 different STs (5.4%), while the third cluster was centred on ST410 and included 14 isolates from 3 different STs (4.5%).
A maximum-likelihood tree based on concatenated MLST gene sequences showed a main clade of 128 isolates (represented by blue branches in Fig. 2 and Supplementary Figure 4; bootstrap value of the main branch = 0.98), including most, but not all, isolates from the eBURST clusters of ST10 and ST1114 (Supplementary Figure 4A). All isolates from these 2 eBURST clusters for which a WGS was available were grouped in BAPS cluster 2. Similarly, all the isolates from the eBURST cluster ST410 grouped into BAPS cluster 1, along with 6 isolates from ST155. Seven isolates belonged to the globally successful extra-intestinal pathogenic E. coli clone ST131 (Supplementary Figure 4A).
Phylogeny of the mcr-1-carrying E. coli isolates. Maximum-likelihood tree based on concatenated MLST gene sequences, mid-point rooted. Inner coloured circle: isolation source; outer circle: region of origin. Stars indicate the isolates from which a whole genome sequence was available. The 6 travellers’ isolates that were sequenced for this study are highlighted in green. Bootstrap values between 0.9 and 1 are indicated by red triangles (size proportional to bootstrap value). The blue branches represent the main clade of 128 isolates including most isolates from ST10. Tree scale in number of substitutions per site. See Supplementary Figure 4A for additional information on the relationship between STs, eBURST clustering and WGS BAPS clustering.
As observed in the WGS analysis, animal isolates were interspersed with isolates from humans and the environment throughout the tree, as were isolates from different continents indicating a lack of clustering by isolation source or geographical origin (Fig. 2). Similarly, no clustering by health status of the host was observed (Supplementary Figure 4B).
Mobile genetic elements
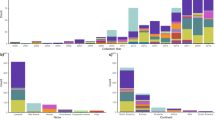
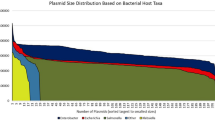
The plasmid incompatibility group of the mcr-1-carrying plasmids could be determined for 217 Enterobacteriaceae isolates from 7 different species (Escherichia sp., Salmonella sp., Klebsiella sp., Cronobacter sp., Enterobacter sp., Kluyvera sp. and Shigella sp.), representing a total of 219 plasmids since 2 isolates carried 2 different plasmids (Table 1). These plasmids were described in 71 studies (1 to 33 plasmids per study, average = 3.1). In addition, the gene was integrated in the chromosome of 6 isolates. The incompatibility group could not be determined for 27 of the 65 isolates for which WGS was available. Similarly the plasmid type was not available for 182 of the 312 isolates included in the MLST analysis. A total of 14 different plasmid incompatibility groups were identified. 198/219 (90.4%) of the identified plasmids belonged to one of 3 incompatibility groups: IncX4 (77/219 plasmids, 35.2%), IncI2 (76/219 plasmids, 34.7%) and IncHI2 (45/219 plasmids, 20.5%). 50/76 IncI2 plasmids (65.8%) originated from Asia and 33/45 IncHI2 plasmids (73.3%) from Europe. IncX4 plasmids were more evenly distributed: 44/77 (57.1%) were recovered from Europe, 29 from Asia (37.7%) and 4 from other regions (5.2%). Observed proportions were significantly different from expected for IncI2 (χ2-test, p < 0.001) and IncHI2 (p < 0.001) but not for IncX4. The distribution of these 3 plasmid types was not significantly different from expected between animal (χ2-test, p = 0.24), human (p = 0.88) and environmental sources (p = 0.38). Isolates from the BAPS groups 1 and 2 carried plasmids from the 3 major types in similar proportions (Supplementary Figure 5A; Supplementary Table 1). Isolates from the eBURST clusters of ST10 carried plasmids belonging to 7 different incompatibility groups, including IncHI2, IncI2 and IncX4. No clustering of plasmid type with MLST phylogeny was observed either (Supplementary Figure 5B; Supplementary Table 1).
Figure 3 shows the alignment of the complete sequences or contigs from IncI2 (panel A), IncX4 (B) and IncHI2 (C) incompatibility group plasmids. IncHI2 plasmids had the largest size, with sequence lengths up to 267486 bp.
Alignment of mcr-1-containing plasmids and contigs. Panel A: IncI2 plasmids (n = 29); panel B: IncX4 (n = 24); panel C: IncHI2 (n = 9). Black outer ring: plasmid used as reference for the alignment; name and size of the reference indicated in the middle of each panel. Plasmid names followed by “_mcr1_contig” refer to assembled contigs from whole genome sequences. Other names refer to plasmid sequences deposited in online databases. The mcr-1 gene and ISA-pl1 location are underlined in red. Plasmids indicated with an asterisk are from the 6 travellers’ isolates that were sequenced for this study. Panel C: Putative MDR cassette is highlighted in orange.
The ISApl1 transposon element situated upstream of the mcr-1 gene was present in 7/9 (77.8%) IncHI2 plasmids, but only in 11/29 (37.9%) of IncI2 plasmids and completely missing in all of the 24 reported IncX4 plasmids (Fig. 3). In the isolates from travellers the ISApl1 transposon was identified in 3 out of our 6 mcr-1-carrying contigs including one isolate from a traveler to Asia (ST101, IncI2 incompatibility group), one to Africa (ST744, IncHI2) and one to South-America (ST744, incompatibility group not identified).
Antimicrobial resistance genes
All sequences of the mcr-1 gene collected in the present study were 100% identical to the original sequence described by Liu et al.8.
Multiple resistance genes were detected in most of the studied isolates (Supplementary Results). The florfenicol resistance gene floR was present in 32 (49.2%) isolates; in 22 of 45 isolates from animals (48.9%) and 10 of 19 isolates from humans (52.6%). The baeR and baeS genes, encoding novobiocin resistance, were found in 64 (98.5%) and 65 (100%) isolates, respectively.
Plasmid analysis from the WGS data showed that 4 of the 29 mcr-1-carrying IncI2 plasmids (13.8%) contained an additional ESBL gene. IncHI2 plasmids (n = 9) carried between 0 and 12 additional AMR genes. In particular, 4 plasmids carried CTX-M ESBL genes and 2 carried the floR gene. In 4 out of the 9 IncHI2 plasmids analysed in this study, the mcr-1 gene was shown to be integrated alongside a large multi-drug resistance (MDR) gene cassette (Fig. 3C). None of the IncX4 plasmids carried additional AMR genes.
Discussion
Analysis of all reported WGS of mcr-1-carrying isolates shows that the population of E. coli is highly diverse, but is dominated by 2 large groups of related isolates. Most of the isolates from BAPS group 2 grouped into a MLST cluster centred on ST10. An overrepresentation of isolates related to ST10 and ST165 (a SLV of ST1114) in mcr-1-carrying E. coli isolates was previously reported at a smaller scale in isolates from European farm animals20. E. coli ST10 and closely related STs are frequently recovered from food and human intestinal samples and studies have shown a higher prevalence of plasmid-carried AMR genes in ST10, including CTX-M ESBL genes, compared to other STs21,22,23,24.
The other BAPS group of interest in our study (group 1) included isolates belonging to ST155. This ST has been described as a major vector of spread of ESBL genes from animals to humans25. It is thus possible that zoonotic transmission leads to the spread of the mcr-1 gene, as has been suggested in studies from China and Vietnam16,17, notably through the 2 main phylogenetic clusters identified in this study.
Additionally, we found clonally related isolates, including some belonging to ST744, a SLV of ST10, carrying the mcr-1 gene on different plasmid backbones and recovered from different continents (see Supplementary Results). These results point towards a worldwide dissemination of mcr-1 driven mainly by highly promiscuous plasmids rather than the worldwide spread of one or more mcr-1-carrying clones. We hypothesize that several populations of E. coli isolates, notably those related to ST10 or ST155, acquired the mcr-1 gene due to their intrinsic ability of acquiring AMR genes and their high prevalence in humans and food animals. These populations of commensal isolates then may play a crucial role as a reservoir for this gene, which can explain their over-representation in the present study.
In the timeframe of our literature search, 3 E. coli strains carrying the mcr-2 gene were isolated from animals in Belgium. These isolates belonged to ST10 (2 isolates) and ST167 which is a SLV of ST10 and carried the gene on an IncX4-type plasmid. No WGS data was available from these isolates26. More data about the mcr-2 gene is needed to assess its spread and determine if E. coli ST10 plays a similar role in its dissemination as it does for mcr-1.
More than 90% of published plasmid types carrying mcr-1 genes belonged to either IncI2, IncX4 or IncHI2. Almost 75% of the isolates carrying an IncHI2 plasmid originated from Europe: 26 from animals and 7 from humans (Table 1 and Supplementary Table 1). In a traveller’s isolate acquired in Tunisia, the mcr-1-carrying plasmid was identified as an IncHI2-type backbone of the ST4 pMLST subtype which co-carried a CTX-M-1 ESBL gene (Fig. 3C). This traveller reported consumption of beef, chicken and eggs during travel to Tunisia which can potentially be the source for the acquisition of the mcr-1 positive isolate. When investigating the presence of the mcr-1 gene in cephalosporin resistant E. coli isolates from chicken farms in Tunisia, Grami et al.27 found that all mcr-1-carrying plasmids from their study (n = 37) also belonged to the IncHI2-type, ST4 subtype and harboured CTX-M-1 genes. PFGE typing of the isolates harbouring this plasmid showed various bacterial genetic backgrounds. Interestingly, these chickens were all imported from France, either as adults or chicks. Other studies showed the presence of this IncHI2, CTX-M-1 and mcr-1 combination in Salmonella enterica Typhimurium isolates from meat samples in Portugal from 201128,29 and diarrhoeic veal calves in France30. The IncHI2 subtype ST4 was also detected in an E. coli isolate from retail chicken breast in Germany31 and the faecal sample of a veal calve from the Netherlands32, suggesting widespread dissemination of this particular plasmid in European farm animals and possible transmission to humans.
The high prevalence of novobiocin baeR and baeS and florfenicol floR resistance genes33,34 in the genomes of isolates of human and animal origin together with the fact that florfenicol and novobiocin are used almost exclusively in veterinary medicine further supports the potential role of food animals as an important reservoir of mcr-1 containing bacteria and MGEs15.
In contrast with the IncHI2 plasmids, 65.8% of all IncI2 plasmids recovered so far originated from Asia, with a much lower prevalence in mcr-1 carrying Enterobacteriaceae from other regions. Taken together, these elements point toward a more regional circulation and dissemination of the mcr-1-carrying plasmids IncI2 and IncHI2.
We found the ISApl1 transposon element associated with the mcr-1 gene, as originally described by Liu et al.8 to be present in a minority of studied plasmids and contigs. However, since some of the mcr-1-carrying contigs were obtained by assembly of Illumina short reads from WGS data, we cannot exclude that some of these gaps are explained by an incomplete assembly of (plasmid) sequences. The ISApl1 transposon element is considered to be the main driver of horizontal gene transfer of the mcr-1 gene and has been shown to be highly unstable in IncI2 plasmids35,36,37. The absence of the ISApl1 transposon element in mcr-1-carrying IncX4 plasmids as described here has recently been proposed to be essential for the maintenance of the mcr-1 gene in this particular backbone, but the exact mechanism still requires further investigation38.
WGS analysis provided in-depth information about the mcr-1-carrying E. coli isolates and their phylogenetic relationship, but the number of available genomes was limited. On the other hand, whilst MLST data have a lower resolution, the higher number of available profiles allowed analysis of the isolates’ origin (geographical, source of isolation, diseased status of the host, etc.).
A limitation of our study is the potential for bias. The overrepresentation of isolates originating from Asia and Europe could be explained by a higher prevalence of mcr-1 genes on these continents, but the effect of publication bias cannot be excluded. Isolates from North-America only represented 2.2% of the collection. Noteworthy, colistin, except for ophthalmic ointment, has never been marketed for use in animals in the United States39,40.
Sampling bias should also be considered when several isolates with an identical ST are presented from a single study, as is the case for ST100 and ST752. Additionally, in the absence of a control population of mcr-1-negative isolates obtained from similar sources as the mcr-1 positive isolates, results of analysis of population structures should be interpreted with caution. Because many studies screened existing collections of (resistant) isolates for colistin resistance or presence of mcr-1, selection bias has probably been introduced.
The findings in this study suggests that the mcr-1 gene has locally and globally disseminated through MGEs that are mainly IncHI2, IncI2 and IncX4 plasmids and provides additional support for the hypothesis of the animal reservoir, that is driven by the use of colistin in livestock, as a source of mcr-1 in humans. A global ban of colistin use in animals to preserve colistin for use in human medicine seems therefore justified.
Material and Methods
Selection of isolates for whole genome sequencing
We subjected 6 mcr-1 positive isolates that were collected as part of a prospective study (COMBAT study) aimed at studying acquisition of extended-spectrum β-lactamase (ESBL) -producing Enterobacteriaceae during travel to whole-genome sequencing14,41. Additionally, we included 22 whole genome sequences of isolates from Vietnamese chickens and humans that were still unpublished when performing our literature search17.
Literature search
Relevant papers that published on mcr-1 and mcr-2 were identified in PubMed, Web of Science, Scopus, ScienceDirect and Google Scholar using the query ‘mcr-1 OR mcr1 OR mcr-2 OR mcr2 OR (mcr AND colistin)’ (see Supplementary Material for full search strategies). To be able to study the associations between phylogeny, geographic distribution and isolation source we only included sequences from papers that provided sufficient metadata. As a consequence, plasmids and genomes sequences that were deposited in online databases without metadata were not included in the analysis.
Whole genome sequencing of mcr-1-positive E. coli isolates
Bacterial DNA was extracted from fresh pure cultures using the Qiagen DNeasy Blood and Tissue kit (Qiagen, Hilden, Germany). Library preparation was done according to manufacturer’s instruction (Illumina, San Diego, CA, USA) and sequenced using Illumina MiSeq technology with 150nt paired-end settings. Sequences have been deposited in the European Nucleotide Archive under the accession numbers SAME104030441 to SAME104030446.
Bio-informatic analysis
MLST analysis
For each mcr-1-carrying E. coli isolate for which the ST or the whole genome sequence was available, the sequences of the corresponding alleles were downloaded from the E. coli MLST genes repository of the University of Warwick (http://mlst.warwick.ac.uk/mlst/dbs/Ecoli/handlers/getFileData/home/cbailster/mlst/zope/Extensions/gadfly/Ecoli/DB/) and concatenated. When STs of isolates were not described in literature, the ST was determined from available whole genomes using the online service provided by the Center for Genomic Epidemiology (https://cge.cbs.dtu.dk/services/MLST/) according to the Achtman MLST scheme42,43. MLST clusters (STs and their single locus or double locus variants) were defined using e-burst V3 (http://eburst.mlst.net/v3/enter_data/single/)19 and goeBURST v1.2.144 using only profiles from this study.
WGS and plasmid analysis
Raw sequence reads in fastq format or pre-assembled sequences in fasta format were downloaded from online databases for all available isolates (Supplementary Table 1). Additional sequences not yet deposited in online databases were requested from their respective authors. The quality of the raw sequence reads was checked using fastqc (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/), quast45 and KmerFinder 2.0 (https://cge.cbs.dtu.dk/services/KmerFinder/) (see Supplementary Methods for more details). Reads were trimmed using Trimmomatic V0.3346. De-novo genome assembly was performed with SPAdes 3.947 for Illumina short reads and with Canu v1.3 for PacBio long reads48. Contigs of less than 500 bp long were removed from the genomes to improve the overall quality of the assembly. Size of the genomes was calculated by adding the length of all remaining contigs. Identification of open reading frames (ORFs) and gene contents in the assembled genomes (de-novo assemblies and pre-assembled sequences) was performed using Prokka v1.1149. Core genome analysis was performed with Roary v3.6.850. Clustering of isolates was performed using the hierBAPS module of the Bayesian Analysis of Population Structure (BAPS) software v6.018. The core genome alignment output provided by Roary was used as input for BAPS with 2 levels of hierarchy and a maximum number of cluster (K) of 10. The estimated number of clusters was 5 for both levels of hierarchy.
Sequences (concatenated MLST loci or concatenated core genes) were aligned using mafft v6.864b51. The resulting alignment was used as input for calculation of distances and tree building using RAxML v8.1.652. MLST and WGS trees were visualized using iTOL v3.3.1 (http://itol.embl.de/)53.
Identification of plasmid incompatibility group and typing of IncHI2 plasmids were performed on assembled sequences (de-novo or pre-assembled) via the CGE online services PlasmidFinder v1.3 (https://cge.cbs.dtu.dk/services/PlasmidFinder/) and pMLST v1.4 (https://cge.cbs.dtu.dk/services/pMLST/)54. Alignment and visualization of plasmids was performed with BRIG v0.9555. The majority of the isolates and plasmids described in this study were sequenced using a short read technology (Illumina). This technology does not allow for a high quality assembly of the plasmids due to the high number of repeat regions present in these MGEs. Therefore no phylogenetic analysis of the mcr-1-carrying plasmids was conducted in this study.
Two different databases were used for identification of other antibiotic resistance genes: ResFinder (https://cge.cbs.dtu.dk/services/ResFinder/56) was used to detect acquired resistance genes commonly located on mobile genetic elements (MGEs) and CARD Resistance Gene Identifier (https://card.mcmaster.ca/analyze/rgi57) was used to detect chromosomal genes.
Statistics
The distribution of isolates and plasmids from different geographical origins and isolation sources was determined by a χ2-test comparing the expected distribution (proportions of the total studied population) to the observed proportions using GraphPad Prism6 (La Jolla, CA, USA).
Change history
14 February 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Appelbaum, P. C. 2012 and beyond: Potential for the start of a second pre-antibiotic era? J. Antimicrob. Chemother. 67, 2062–2068 (2012).
Brown, E. D. & Wright, G. D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Laxminarayan, R. et al. Antibiotic resistance-the need for global solutions. Lancet Infect. Dis. 13, 1057–1098 (2013).
Koyama, Y., Kurosasa, A., Tsuchiya, A. & Takakuta, K. A new antibiotic ‘colistin’produced by spore-forming soil bacteria. J Antibiot 3, 457–458 (1950).
Lim, L. M. et al. Resurgence of colistin: a review of resistance, toxicity, pharmacodynamics, and dosing. Pharmacotherapy 30, 1279–91 (2010).
Catry, B. et al. Use of colistin-containing products within the European Union and European Economic Area (EU/EEA): development of resistance in animals and possible impact on human and animal health. Int. J. Antimicrob. Agents 46, 297–306 (2015).
Olaitan, A. O., Morand, S. & Rolain, J. Mechanisms of polymyxin resistance: acquired and intrinsic resistance in bacteria. Front. Microbiol. 5, 1–18 (2014).
Liu, Y.-Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 3099, (2015).
Skov, R. L. & Monnet, D. L. Plasmid-mediated colistin resistance (mcr-1gene): three months later, the story unfolds. Eurosurveillance 21, (2016).
Schwarz, S. & Johnson, A. P. Transferable resistance to colistin: a new but old threat. J. Antimicrob. Chemother. 71, 2066–2070 (2016).
Leangapichart, T. et al. Acquisition of mcr-1 Plasmid-Mediated Colistin Resistance in Escherichia coli and Klebsiella pneumoniae during Hajj 2013 and 2014. Antimicrob. Agents Chemother. 60, 6998–6999 (2016).
Vading, M. et al. Frequent acquisition of low-virulence strains of ESBL-producing Escherichia coli in travellers. J. Antimicrob. Chemother. 71, 3548–3555 (2016).
Bernasconi, O. J. et al. Travelers Can Import Colistin-Resistant Enterobacteriaceae, Including Those Possessing the Plasmid-Mediated mcr-1 Gene. Antimicrob. Agents Chemother. 60, 5080–5084 (2016).
Arcilla, M. S. et al. Dissemination of the mcr-1 colistin resistance gene. Lancet Infect. Dis. 16, 147–149 (2016).
Poirel, L. & Nordmann, P. Emerging plasmid-encoded colistin resistance: the animal world as the culprit? J. Antimicrob. Chemother. 71, 2326–2327 (2016).
Wang, Y. et al. Comprehensive resistome analysis reveals the prevalence of NDM and MCR-1 in Chinese poultry production. Nat. Microbiol. 2, 16260 (2017).
Trung, N. V. et al. Zoonotic Transmission of mcr-1 Colistin Resistance Gene from Small-Scale Poultry Farms, Vietnam. Emerg. Infect. Dis. J. 23, 529 (2017).
Cheng, L., Connor, T. R., Sirén, J., Aanensen, D. M. & Corander, J. Hierarchical and spatially explicit clustering of DNA sequences with BAPS software. Mol. Biol. Evol. 30, 1224–1228 (2013).
Feil Edward, J., Li, B. C., Aanensen, D. M., Hanage, W. P. & Spratt, B. G. eBURST: inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J. Bacteriol. 186, 1518–1530 (2004).
El Garch, F. et al. mcr-1 is borne by highly diverse Escherichia coli isolates since 2004 in food-producing animals in Europe. Clin. Microbiol. Infect. 23, 51e1–51e4 (2017).
Manges, A. R. & Johnson, J. R. Food-Borne Origins of Escherichia coli Causing Extraintestinal Infections. Clin. Infect. Dis. 55, 712–719 (2012).
Oteo, J. et al. Extended-spectrum β-lactamase-producing Escherichia coli in Spain belong to a large variety of multilocus sequence typing types, including ST10 complex/A, ST23 complex/A and ST131/B2. Int. J. Antimicrob. Agents 34, 173–176 (2009).
Fam, N. et al. CTX-M-15-producing Escherichia coli clinical isolates in Cairo (Egypt), including isolates of clonal complex ST10 and clones ST131, ST73, and ST405 in both community and hospital settings. Microb. Drug Resist. 17, 67–73 (2011).
Manges, A. R. et al. Clonal distribution and associated characteristics of Escherichia coli clinical and surveillance isolates from a military medical center. Diagn. Microbiol. Infect. Dis. 87, 382–385 (2017).
Skurnik, D. et al. Emergence of antimicrobial-resistant Escherichia coli of animal origin spreading in humans. Mol. Biol. Evol. 33, 898–914 (2016).
Xavier, B. B. et al. Identification of a novel plasmid-mediated colistinresistance gene, mcr-2, in Escherichia coli, Belgium, june 2016. Eurosurveillance 21, (2016).
Grami, R. et al. Impact of food animal trade on the spread of mcr-1-mediated colistin resistance, tunisia, july 2015. Eurosurveillance 21, 1–5 (2016).
Tse, H. & Yuen, K.-Y. Dissemination of the mcr-1 colistin resistance gene. Lancet Infect. Dis. 3099, 3099 (2015).
Figueiredo, R., Henriques, A., Sereno, R., Mendonça, N. & da Silva, G. J. Antimicrobial Resistance and Extended-Spectrum β-Lactamases of Salmonella enterica Serotypes Isolated from Livestock and Processed Food in Portugal: An Update. Foodborne Pathog. Dis. 12, 110–117 (2014).
Haenni, M. et al. Co-occurrence of extended spectrum β lactamase and MCR-1 encoding genes on plasmids. Lancet Infect. Dis. 3099, 3099 (2016).
Falgenhauer, L. et al. Chromosomal Locations of mcr-1 and bla(CTX-M-15) in Fluoroquinolone-Resistant Escherichia coli ST410. Emerg. Infect. Dis. 22, 1689–1691 (2016).
Veldman, K. et al. Location of colistin resistance gene mcr-1 in Enterobacteriaceae from livestock and meat. J. Antimicrob. Chemother. 71, 2340–2342 (2016).
Lilic, M., Jovanovic, M., Jovanovic, G. & Savic, D. J. Identification of the CysB-regulated gene, hslJ, related to the Escherichia coli novobiocin resistance phenotype. FEMS Microbiol. Lett. 224, 239–246 (2003).
Baranova, N. & Nikaido, H. The BaeSR Two-Component Regulatory System Activates Transcription of the yegMNOB (mdtABCD) Transporter Gene Cluster in Escherichia coli and Increases Its Resistance to Novobiocin and Deoxycholate. J. Bacteriol. 184, 4168–4176 (2002).
Stoesser, N., Mathers, A. J., Moore, C. E., Day, N. P. & Crook, D. W. Colistin resistance gene mcr-1 and pHNSHP45 plasmid in human isolates of Escherichia coli and Klebsiella pneumoniae. Lancet Infect. Dis. 3099, 3099 (2016).
Falgenhauer, L. et al. Colistin resistance gene mcr-1 in extended-spectrum β-lactamase-producing and carbapenemase-producing Gram-negative bacteria in Germany. Lancet Infect. Dis. 3099, 3099 (2016).
Snesrud, E. et al. Analysis of Serial Isolates of mcr-1- positive Escherichia coli Reveals a Highly Active ISApl1 Transposon. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.00056-17 (2017).
Sun, J. et al. Genetic Analysis of the IncX4 Plasmids: Implications for a Unique Pattern in the mcr-1 Acquisition. Sci. Rep. 7, 424 (2017).
FDA. 2015 Summary Report On Antimicrobials Sold or Distributed for Use in Food-Producing Animals. at https://www.fda.gov/downloads/ForIndustry/UserFees/AnimalDrugUserFeeActADUFA/UCM534243.pdf (2016).
European Medicines Agency. Updated advice on the use of colistin products in animals within the European Union: development of resistance and possible impact on human and animal health. 44 (2016).
Arcilla, M. S. et al. Import and spread of extended-spectrum beta-lactamase-producing Enterobacteriaceae by international travellers (COMBAT study): a prospective, multicentre cohort study. Lancet Infect. Dis. 17, 78–85 (2017).
Larsen, M. V. et al. Multilocus sequence typing of total-genome-sequenced bacteria. J. Clin. Microbiol. 50, 1355–1361 (2012).
Wirth, T. et al. Sex and virulence in Escherichia coli: An evolutionary perspective. Mol. Microbiol. 60, 1136–1151 (2006).
Francisco, A. P., Bugalho, M., Ramirez, M. & Carriço, Ja Global optimal eBURST analysis of multilocus typing data using a graphic matroid approach. BMC Bioinformatics 10, 152 (2009).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bankevich, A. et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 19, 455–477 (2012).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. bioRxiv at http://biorxiv.org/content/early/2017/01/04/071282.abstract (2017).
Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Page, A. J. et al. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Katoh, K. & Toh, H. Recent developments in the MAFFT multiple sequence alignment program. Brief. Bioinform. 9, 286–298 (2008).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Letunic, I. & Bork, P. Interactive tree of life (iTOL)v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Carattoli, A. et al. In Silico Detection and Typing of Plasmids using PlasmidFinder and Plasmid Multilocus Sequence Typing. Antimicrob. Agents Chemother. 58, 3895–3903 (2014).
Alikhan, N.-F., Petty, N. K., Ben Zakour, N. L. & Beatson, S. A. BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genomics 12, 402 (2011).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644 (2012).
McArthur, A. G. & Wright, G. D. Bioinformatics of antimicrobial resistance in the age of molecular epidemiology. Curr. Opin. Microbiol. 27, 45–50 (2015).
Acknowledgements
The authors would like to thank the team of the Department of Genome analysis of the AMC (in particular Dr. F. Baas and L. Koster) for their support in the sequencing bacterial genomes. Financial support for this study was provided by the European COMPARE project (http://www.compare-europe.eu/) under the European Union’s Horizon 2020 research and innovation programme, grant agreement No. 643476. The VIBRE and COMBAT studies were supported by The Netherlands Organisation for Health Research and Development/The Netherlands Organisation for Scientific Research (ZonMw) under grants number 205100012 and 50–51700–98–120 respectively.
Author information
Authors and Affiliations
Contributions
S.M. and J.v.H. contributed equally to this work. S.M and J.v.H. performed the systematic review, performed the experiments, interpreted the results and wrote the manuscript. N.W. contributed to bio-informatics analyses. M.A., D.M., J.P., M.d.J., the COMBAT consortium members (M.C.J.B., P.J.v.G., A.G., M.P.G., N.M., A.M.L.O.L., E.E.S. and H.A.V.), N.V., N.T.H. and C.S. contributed isolates and genome sequence data and associated meta data. M.d.J. and C.S. designed the study. All authors contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Matamoros, S., van Hattem, J.M., Arcilla, M.S. et al. Global phylogenetic analysis of Escherichia coli and plasmids carrying the mcr-1 gene indicates bacterial diversity but plasmid restriction. Sci Rep 7, 15364 (2017). https://doi.org/10.1038/s41598-017-15539-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15539-7
This article is cited by
-
Low prevalence of mcr-1 in Escherichia coli from food-producing animals and food products in China
BMC Veterinary Research (2024)
-
Convergence of resistance and evolutionary responses in Escherichia coli and Salmonella enterica co-inhabiting chicken farms in China
Nature Communications (2024)
-
Nanopore R10.4 metagenomic detection of blaCTX-M/blaDHA antimicrobial resistance genes and their genetic environments in stool
Nature Communications (2024)
-
Quinolone-resistant Escherichia coli at the interface between humans, poultry and their shared environment- a potential public health risk
One Health Outlook (2023)
-
Regulatory fine-tuning of mcr-1 increases bacterial fitness and stabilises antibiotic resistance in agricultural settings
The ISME Journal (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.