Abstract
Efflux pumps of the ATP-binding cassette transporters superfamily (ABC transporters) are frequently involved in the multidrug-resistance (MDR) phenomenon in cancer cells. Herein, we describe a new atomistic model for the MDR-related ABCG2 efflux pump, also named breast cancer resistance protein (BCRP), based on the recently published crystallographic structure of the ABCG5/G8 heterodimer sterol transporter, a member of the ABCG family involved in cholesterol homeostasis. By means of molecular dynamics simulations and molecular docking, a far-reaching characterization of the ABCG2 homodimer was obtained. The role of important residues and motifs in the structural stability of the transporter was comprehensively studied and was found to be in good agreement with the available experimental data published in literature. Moreover, structural motifs potentially involved in signal transmission were identified, along with two symmetrical drug-binding sites that are herein described for the first time, in a rational attempt to better understand how drug binding and recognition occurs in ABCG2 homodimeric transporters.
Similar content being viewed by others
Introduction
Multidrug resistance (MDR) to anticancer agents is a major health concern worldwide due to the long-lasting physical and psychological outcomes1. Herein, MDR is a multi-factorial phenomenon often connected to over-expression of ATP Binding Cassette (ABC) transporters at the cell surface, acting by decreasing the intracellular accumulation of cytotoxic drugs and impairing the success of chemotherapeutic regimens2. Of the 48 members of the human ABC family, P-glycoprotein (P-gp, ABCB1)3, Multidrug-Resistance Protein 1 (MRP1, ABCC1)4 and Breast Cancer Resistance Protein (BCRP, ABCG2)5,6,7 are the most important transporters in cancer MDR, although a new member of the ABCB family (ABCB5)8,9 is also gaining relevance in melanoma resistance to anticancer agents.
ABCG2 in MDR was identified almost simultaneously by several groups in MCF-7/AdrVp cell lines (BCRP or MXR)5,7 and in placenta (ABCP)6. In both MCF-7/AdrVp3000 (over-expressing ABCG2) and full length ABCG2 cDNA-transfected MCF-7 breast cancer cells, the over-expression of this transporter conferred resistance to several xenobiotics including mitoxantrone, doxorubicin and daunorubicin, also reducing the intracellular accumulation of Rhodamine-123 (R123) by an ATP-dependent mechanism5,7. Although studied due to its important role in MDR, ABCG2 also participates in normal detoxification mechanisms that can be found in previoulsy published literature10,11,12.
ABCG2 is characterized as a “half-transporter”, hypothetically comprising a transmembrane domain (TMD) with 6 α-helices that spawns the membrane bilayer and a nucleotide-binding domain (NBD) where ATP binds and hydrolyzes. However, unlike P-gp that shows a TMD-NBD arrangement in both halves, ABCG2 shows a distinct domain organization where the NBD precedes the TM domain (in a NBD-TMD arrangement, Fig. 1)13. Moreover, ABCG2 includes a second canonical ABC signature (positions 352–356, LSGGE) also coupled with ATP binding and/or hydrolysis but not related with substrate specificity14. Although higher orders of oligomerization were also described, namely tetrameric and dodecameric forms, to become a fully functional transporter ABCG2 forms a homodimer15,16. Furthermore, with only a minority dimer population being registered, it was recently determined that tetrameric complexes are the most predominant ones in plasma membranes17. The possibility that such higher oligomeric forms may serve as a regulator for functional dimeric ABCG2 transporters13 has become increasingly accepted.
Membrane topology model for ABCG2. Cysteine residues involved in disulfide bridges are depicted as circles and N-glycosilated residue as a triangle. The de novo intracellular C2 helices are depicted in green. TMD, transmembrane domain; NBD, nucleotide-binding domain (adapted from Taylor et al.)18.
The role of specific cysteines in ABCG2 oligomerization has also been thoroughly studied since the discovery of ABCG2 homodimers. While an early study by Kage et al. claimed that ABCG2 homodimerization was due to the formation of an intermolecular disulfide bond at position 60315, it was later concluded that such disulfide bond is not essential for protein expression and function13,16,19,20,21. Although present in the absence of reducing agents15, and as the dimeric ABCG2 was found only in isolated membranes, the presence of a -S-S- bridge can also derive from oxidation processes during cell lysis and membrane preparation13. Furthermore, it was also shown that only 35% of ABCG2 was cross-linked as dimers13 and that non-covalent protein-protein interactions between transmembrane helices 5 and 6 (TM5-loop-TM6) could also be involved in ABCG2 oligomerization22.
Similarly, the role of an intramolecular disulfide bond between residues 592 and 608 was initially also thought to be crucial for the activity of the transporter20. However, while in one study ABCG2 variants without cysteines in the extracellular loop (C592, C603 and C608) showed similar cellular location and drug-stimulated activity comparable to the wild-type (WT) protein23, other two studies concluded that i) protein trafficking to the plasma membrane may be impaired in the presence of C592A/C603A or C603A/C608A mutations but not C592A/C608A20 and (ii) the intramolecular disulfide bond is an important “checkpoint” that determines the fate of de novo synthesized ABCG2 proteins24. Therefore, it seems that further studies are needed to fully understand the role of inter- and intramolecular disulfide bonds in ABCG2 expression and function.
Regarding ABCG2 function, it was shown that mutation of an arginine by a threonine or glycine residue at position 482 increased efflux function towards R123 and anthracyclines as daunorubicin16, not interfering directly with substrate binding but affecting other important factors as ATP-related energy coupling, which is intimately linked to the conformational changes, and/or signal transduction mechanisms25,26. As for P-gp, the mutation of the aspartate residue in Walker B motif (Mg2+-chelating motif, D210N) led to a non-functional protein that was still able to be expressed at the cell surface. Moreover, as it was also found that ABCG2 prefers detergent-resistant cell membranes characterized by high cholesterol content, cholesterol depletion induces a marked decrease on ABCG2 transport activity without affecting the localization of the transporter within the cell membrane27. Interestingly, 30 molar-% cholesterol content was found to provide the best conditions for optimal ABCG2 ATPase activity. As such, cholesterol is now considered an essential activator of ABCG2 function, probably acting as a slowly transported substrate, an allosteric co-activator or a co-transporter28. Indeed, at least five cholesterol recognition/interaction amino acid consensus (CRAC)29 were identified in the ABCG2 transporter, but while mutation in Y413 and Y570 induced higher levels of ATPase activity that were not dependent of cholesterol presence, only one (Y413) was able to modulate ATPase activity in the presence of cholesterol30.
To date, P-gp is the most studied ABC transporter involved in MDR. Since its publication in 200931, numerous structure-based studies allowed a deeper understanding of the molecular mechanisms involved not only in drug binding and recognition32,33,34,35 but also about conformational changes intimately related with the efflux mechanism itself36,37,38,39. Quite recently, the publication of the first structural models of the bovine Multidrug Resistance Protein 1 (MRP1, ABCC1; PDB ID: 5UJA)40 and human Breast Cancer Resistance Protein (BCRP, ABCG2; PDB ID: 5NJ3)18 provided new templates for novel structure-based studies, aiming for the development of novel MDR inhibitors but also for studies concerning a greater comprehension about the efflux mechanism in these ABC transporters.
Preceding the publication of the human ABCG2 structure, the structure of a heterodimeric sterol transporter ABCG5/G8 41 revealed a new transmembrane organization that was thought to be characteristic of the ABCG family. Indeed, while sharing low amino acid identity (28%), ABCG5 and ABCG8 revealed a high degree of structural conservation with a relatively low root mean square deviation (RMSD = 2.0 Å) between both halves41. Likewise, ABCG5 and ABCG8 share similar identities (27% and 26% respectively) and similarities (48% and 44% respectively) with the newly published ABCG2 structure, thus being a suitable template for studying not only the dynamics of ABCG5/G8 heterodimer but also, by homology modeling, the ABCG2 homodimer. This hypothesis is supported by the fact that i) despite their low identities and similarities, the structural conservation between ABCG5 and ABCG2 monomers remains high (RMSD = 2.4 Å) while between ABCG8 and ABCG2 is lower (RMSD = 3.8 Å), ii) in the original ABCG5/G8 publication a homology model of the Drosophila white/brown heterodimer was obtained following a CLANS network analysis showing that the TMDs of both transporters shared a high pairwise similarity relationship based on their FASTA sequences and iii) the same pairwise relationship can be observed between ABCG5/G8 and ABCG2 41. To that matter, computational approaches as molecular dynamics (MD) and docking are valuable tools that can be used, in a similar approach as for P-gp36, to further refine a homology model of the ABCG2 homodimer and to evaluate the structural dynamics of the ABCG2 transporter, also unveiling new information on hypothetical drug-binding locations that can be used to better understand the ABCG2 role on MDR.
Results
ABCG2 model validation
The publication of the ABCG5/G8 crystallographic structure showed that despite the low sequence identity between ABCG5 and ABCG8 (28% amino acid identity), a high degree of structural conservation could still be found41. Thus, as ABCG5 and ABCG2 share 27% amino acid identity, a high pairwise similarity between both FASTA sequences and a high degree of structural conservation (as seen by their RMSD), the homodimeric ABCG5 transporter is a more suitable template for ABCG2. After obtaining the full-length ABCG2 homology model, structure validation servers were used to assess the quality of the model (Table 1). Furthermore, an evaluation of the recently published human ABCG2 cryo-EM structure18 was also performed to compare and validate the herein developed ABCG2 model.
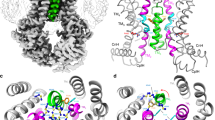
By the superposition of the structural data of the human ABCG2 transporter with our homology model, obtained by duplication of the ABCG5 half transporter (Fig. 2A), all major structural features are found to overlap (monomer RMSD, 2.81 Å; homodimer RMSD, 4.30 Å), with the larger deviations being found at the extracellular coils, the de novo modeled intracellular C2 helices and at the top of the TM5a, before the 5b/5c helical bundle found to be characteristic of the ABCG family. Moreover, and while a slight backbone shift of TM1b and TM3 were observed, an inward-facing cavity was found to be present in our homology model, similar to the one identified in the ABCG2 crystal structure due to the backbone shift of TM helices 2 and 5a, but absent in a previously published homology model42 obtained from the whole ABCG5/G8 heterodimeric transporter41.
In opposition to a previous model published recently42, a significant improvement was observed when compared with the ABCG5/G8 crystallographic structure. When compared with the human ABCG2 crystal structure, our refined model still behaves quite well and, despite the lower scores for the G2MODEL and G2FULL in the SwissModel structure assessment tools, the values are comparable with the ones obtained for the crystallographic structure and are in agreement with the stereochemical quality expected for membrane proteins because QMEAN Z-Score uses solvent accessibility as one of its scoring components, which confers poorer scores for membrane proteins and predicts these as lower quality models43,44. Furthermore, for membrane proteins Z-scores above −5 are indicative of a good quality model.
When considering the ABCG2 structures obtained after the 200 ns MD runs, all three models (models 1, 2 and 3) showed better scores when compared with the previously published one. Hereafter, as both models 2 and 3 were evaluated as the best ranked models, the discussion will be focused on the results obtained for model 3 unless stated otherwise. Herein, and when compared with the hABCG2 cryo-EM data, the equilibration of the structure induced a distinct asymmetry between both halves, mainly due to the backbone shift of TM4 and TM5a in one half towards TM1b’ and TM2’ of the opposite monomer. However, it is also evident that the overall architecture for the ABCG2 transporter is maintained (RMSD of 3.18 and 3.48 Å between each of the monomers with the monomeric hABCG2) and that the modeling of the missing A-loop (Y44-L64), the connecting helices C2a-C2b (CnH, G311-S354) and the highly charged linker between the connecting helices CnH and TM1a (G355-Y369) did not have any impact on the structural stability of either the nucleotide-binding domains or the first transmembrane helix (TM1a and TM1b).
By comparing the Ramachandran plots between the hABCG2 crystal structure, the initial homology model and after the 200 ns MD simulation, and despite the overlapping plot for the initial homodimers, the overall structure of the ABCG2 is maintained (Fig. 3) with 87.1% of all residues in the most favorable regions, 10.7% on allowed regions and only 2.2% in disfavored regions, in accordance with previous results obtained for the MD simulation of another ABC transporter (P-glycoprotein)36. Still, from the 27 outliers, 12 residues can easily shift into the allowed region of the plot (they overlap the contour line that defines the boundaries for such regions) and the remaining 15 are mainly located in coils, except for Phe402 and Ser413 that are located at the second transmembrane helix in chain B.
The root-mean-square deviation (RMSD) is another important parameter for assessing structural deviations from a given reference structure, in this case the initial homology model (Fig. 4). The RMSD value for the whole transporter increased in the first 50 ns, reaching a plateau at ~4 Å that was maintained until the end of the simulation time. When this parameter is plotted for each ABCG2 domain separately (Fig. 4B), it is clear that in both chains the de novo modeled C2a/C2b/Linker domains are the major contributors for the total RMSD. Moreover, in the recently published G5G8 structure, the ABCG8 subunit have a similar structural organization to ABCG2 (two α-helices followed by a long linker)41. However, even in the most recent ABCG2 structure such domains were unable to be fully determined by crystallography, which is in agreement with the increased flexibility of these particular regions found in the MD simulations. Figure 4B also clearly shows that the other structural domains (TMD and NBD) remains stable in both subunits, within a maximum RMSD of about ~3 Å.
(A) Final structure with liquorice representation of the cholesterol residues within the lipid membrane and lipid heads in CPK (red, phosphate; blue, nitrogen); (B) RMSD of residue α-carbons grouped by domains in both chains A and B (NBD, nucleotide-binding domain; TMD, transmembrane domain; C2a/C2b/Linker, de novo modeled segments.
To further evaluate the overall fold quality of the homology model, we also fitted our 200 ns ABCG2 structure to i) a previously published low-resolution 3D structure, obtained by electron cryo-microscopy from two-dimensional crystals and in the absence of nucleotides and substrates49 and ii) to the recently published three-dimensional cryo-EM density map from which hABCG2 was obtained18 (data not shown). In the first case, by overlapping our model into the cryo-EM electron density map while changing the contour level to a value closer to the mean density (σ = 0) we were able to obtain a good fitting with the electron density, in better agreement than the previous homology models built using P-gp as a template50,51. Moreover, and by using the most recent cryo-EM electron densities from hABCG2, while the transmembrane domains were found to fit within the electron densities corresponding to the membrane-embedded alpha-helical domains deeply buried into the lipidic nanodiscs, the de novo modeled segments were found to correspond to the electron density next to both nucleotide-binding domains attributed to helical segments C2a and C2b. Interestingly, and unlike C2a/C2b/Linker motifs, it was observed that the position of the helix 1a remained stable at the membrane interface throughout the whole MD simulation while simultaneously showing the ability to “slide” through the interface, accompanying the positional shifts of helix 1b, which may mean that this particular segment may have an important function in the efflux conformational dynamics of ABCG2.
Disulfide bridges, cholesterol binding and ATP hydrolysis
In this homology model, no disulfide bonds were modeled because i) intermolecular disulfide bond between cysteine residues in position 603 are not required for its transport function19,21, ii) intramolecular disulfide bonds may exist but, while not essential for ABCG2 localization and transport activity20,23, are a key factor in determining the fate of de novo synthesized proteins24 and iii) cysteine-less transporters can still be expressed and targeted to membranes, with internal cysteines being more important than the ones located at each ECL52.
We measured the Cα-Cα distance between cysteines 592 and 608 (intramolecular) or between cysteines 603 in both chains (intermolecular) to assess if the distances are compatible with the existence of disulfide bonds. In addition, due to the absence of physical bonds between both ABCG2 monomers, we also assessed their relative free energy of binding between subunits with g_mmpbsa added with the implicit membrane correction. While considering intramolecular bridges (C592-C608), Cα-Cα distances ranged from 0.60 ± 0.07 nm (model 1) up to 1.26 ± 0.07 nm (model 3), and are therefore compatible with the physiological existence of disulfide bonds between both residues. For the intermolecular bridge, Cα-Cα distances were found to be 0.95 ± 0.10 nm (model 1), 1.36 ± 0.15 nm (model 2) and 1.05 ± 0.23 nm (model 3), again compatible with intermolecular disulfide bond formation when in an oxidative environment and also in agreement with previous cited experimental data.
Similarly, for cysteine residues at position 284, also thought to be involved in intermolecular bridges responsible for a dimeric protein with slow mobility52, mean Cα-Cα distances ranged from 1.53 ± 0.17 nm (model 3) up to 1.76 ± 0.07 and 1.83 ± 0.08 nm in models 1 and 2 respectively, in good agreement with the above experimental results. Moreover, the location of the remaining cysteines do not allow the formation of either intra or intermolecular disulfide bonds, but nonetheless any mutations in these residues are expected to affect NBD:TMD communication (C438, located close to the coupling helices in TM helix 2) or signal propagation (C374, at the end of the ‘linker’) without compromising substrate specificity52. Finally, free energies of binding between both ABCG2 monomers for the last 50 ns of the final MD simulation were calculated to be −1124 ± 63 kcal.mol−1, lower than the value calculated for the G5G8 heterodimer (−902 ± 59 kcal.mol−1, unpublished data).
Several motifs in ABCG2 were also described to be important for its cholesterol sensitivity30, ATP binding and hydrolysis14,25,26 or substrate binding53. Regarding cholesterol, Gál et al.30 showed that while mutations in the LxxL motif (aa 555–558) resulted in an apparent cholesterol insensitivity, only one of the five identified CRAC domains (Y413) was sensitive to the presence of cholesterol. Interestingly, in ABCG2 both domains (LVNL and VIGAIYFGLK) are located close to each other (Fig. 5A) and next to an additional cholesterol insensitive CRAC domain (Y570, LSWLQYFSIPR)30. More relevant, a cholesterol molecule was found to be located in this particular region, close to all three motifs and making a hydrogen bond with Q569. Herein, the relative free energy of binding for cholesterol was calculated to be −57 ± 5 kcal.mol−1, against only −40 ± 6 kcal.mol−1 when located closer to the dimer interface (Fig. 5B). In addition, another cholesterol molecule placed in the symmetrical location (near the Y570 CRAC motif in the other ABCG2 monomer, Fig. 5C) also showed a similar relative free energy of binding of −56 ± 6 kcal.mol−1. Therefore, our results clearly show that such regions are involved in cholesterol binding and are also in agreement with the electron density data published by Lee et al.41 for the G5G8 transporter in which some features in the electron density map suggest that cholesterol is bound at this location.
When considering ATP binding and hydrolysis, mutations at position R48225,26 or in a second signature motif (C2-sequence, LSGGE)14 were also proved to disturb ATP hydrolysis. However, while R482 is located in transmembrane helix 3, and thus connected to Walker A through the NBD coupling loop (TMH2-loop-TMH3, Fig. 6A), the C2-sequence is located in the beginning of the linker, anchoring the preceding helix to the NBD and also in contact with the internal helical domain containing the Walker A motif (Fig. 6B). Thus, structural modifications in these two regions will have a direct impact on the signal propagation between nucleotide-binding and transmembrane domains, with little impact on drug binding.
Finally, considering drug binding and transport, mutations in proline and methionine residues at positions 485 (P485A) and 549 (M549A) respectively were found to be related with differences in the ability of ABCG2 to transport several substrates53. While P485 is located in the same helix as R482, M549 is located in the vicinity of the LxxL and CRAC motifs, described above as part of a binding site for cholesterol. Therefore, such location can also be a possible drug-binding site for ABCG2 substrates (more details, in the Docking section below). Moreover, the A540F mutation in the ABCG5, which is closely related with the M549A in the ABCG2 structure, also impaired cholesterol transport without affecting G5G8 heterodimer expression41.
Protein global motions
Being ABCG2 a homodimeric transporter, we were particularly interested on how conformational changes could propagate through the structure. In a previous study of another ABC transporter (P-gp), it was found that the ‘linker’ domains acts as a ‘damper’ in order to reduce NBD fluctuations and that, in the presence of substrates, a clear change to efflux-like motions could be observed36. In ABCG2, however, and due to the absence of a linker connecting both halves, signal transmission between monomers occurs through the external helical domains (residues 305–351), acting as a ‘spring’ in transmitting the motions from one NBD to the other (Fig. 7).
In the most predominant mode 1 (Fig. 7A) it is possible to verify that motions in one NBD are propagated to the opposite NBD by a direct interaction with its external helices (although with a minimal impact on its core), while mode 2 shows that motions originated at the NBD:NBD interface are also propagated through these helical segments, reaching the membrane-anchored portion of the linker (Fig. 7B).
Thus, if specifically involved in signal propagation, it is expected that these domains (hereby named signal-propagating domains, or SP domains) should be extremely mobile, quickly shifting between conformations in order to better propagate the signal between monomers. Interestingly, similar domains are also incomplete in the crystallographic ABCG8 structure, which corroborates these assumptions. It was previously proposed that the A-loop, also missing in both ABCG5 and ABCG8 structures, could act as a ‘filter’ at the entry of the substrate cavity or by contributing as the first step of allosteric communication between drug binding and ATP binding42. However our results show that, in both monomers, the A-loop is anchored at the membrane interface through residues 48–55 (LKSGFLPC) and, therefore, they seem unable to participate in allosteric interactions with substrates.
However, in the presence of ATP (Fig. 8), mode 1 of the principal component analysis distinctly shows an upward motion of one NBD in which the A-loop moves closer to the opposite NBD, i.e. closing the access to the ATP-binding site and promoting interactions between the Walker A and Signature motifs, together with a coordinate movement of the SP domains that propagate the signal forward to the transmembrane domains (Fig. 8A). Moreover, active modes 2 and 3 additionally shows distinct efflux-like motions, characterized by NBDA:NBDB distance fluctuations (active mode 2) and rotation of the TM helical bundle (active mode 3, Fig. 8B), similar to that observed for P-gp36 or G5G8 heterodimer (unpublished data). Therefore, A-loop and SP domains seem to be crucial in the ATP-dependent efflux cycle: while the A-loop act as a “cover” for the ATP-binding site, favoring ATP binding and enhancing the contact between Walker A and Signature motifs upon NBD dimerization, SP domains are involved in signal transmission, either between the nucleotide-binding domains (in its apo form) or after ATP binding to the NBD (holo form).
Molecular docking
Kinetic and equilibrium data by Clark et al.54, suggest the presence of at least two symmetric drug-binding sites on ABCG2, one in each monomer, although displaying allosteric communication between them. Due to the large substrate overlapping with P-gp, it is also expected that ABCG2 extrudes its substrates from the lipid bilayer after membrane partitioning from the cytoplasm55. Therefore, in our study the whole TM domains (embedded within the membrane) were used to define the docking box and a known substrate, mitoxantrone (MX), was used to sample possible drug-binding sites within the ABCG2 transporter (Fig. 9). Three drug-binding sites could be identified from the best docking poses for mitoxantrone: although the last one (Fig. 9B, black arrow, −5.7 kcal.mol−1) is in agreement with the electron density reported for cholesterol in the recently published human ABCG2 cryo-EM structure18, the other two sites are herein described for the first time (Fig. 8A,B, purple arrows, −6.3 and −5.8 kcal.mol−1 respectively). The latter two, located in each monomer, comprise a surface ‘groove’ immediately below a short helical segment bearing a cholesterol-insensitive CRAC domain (residues 565–575) that is part of the long extracellular loop connecting helices 5 and 6 (named “plug” in the hABCG2 structure).
Quite interestingly, these particular sites are also deeply buried within the membrane55, show a distinct symmetry between monomers54 and share common residues with the above described cholesterol-binding site56, suggesting that the drug-binding sites in ABCG2 may encompass the large majority of the membrane-facing surface of the transmembrane helical segments to allow binding of larger molecules than cholesterol. Indeed, a close inspection of the top-ranked binding poses for other ABCG2 substrates as pheophorbide A (−7.7 kcal.mol−1), flavopiridol (−7.7 kcal.mol−1) or 9-aminocamptothecin (−7.8 kcal.mol−1) show that these sites are located in a larger ‘surface cleft’ with approximately 1900 A3 (as estimated by EPOSBP) spawning from the CRAC-insensitive helical segment above described to the center of the dimer interface (reported for cholesterol by Taylor et al.18) and also in close agreement with the sterol-binding sites proposed for ABCG5/G8 41. Each drug-binding site is flanked by TM helices 1-2 of the opposite monomer, by TM helices 4–6 and, on top, by a small helical domain that is part of the large extracellular loop between TM helices 5 and 6 (Fig. 9C). Interestingly, it was also verified that, in our model, the second transmembrane helix of one of the monomers is involved in both drug-binding sites, suggesting some degree of allosteric communication between both sites as suggested by Clark et al.54 The identification of these large ‘surface clefts’ within the ABCG2 structure as the hypothetical drug-binding sites led us to thoroughly characterize its lining residues, mean relative polarities and volumes through EPOSBP software. Unlike P-gp, where M-site and R/H sites are characterized by a greater number of aromatic and polar residues respectively32, ABCG2 binding clefts have instead a larger number of hydrophobic residues as valine, alanine, isoleucine and leucine (Fig. 10). In each monomer, hydrophobic residues account for 60 ± 3% of all residues against 29 ± 2% of polar and only 11 ± 1% of aromatic residues, but while hydrophobic residues are scattered through the whole cleft, polar residues are more abundant in helices close to the dimer interface (most particularly in TM helices 1 and 2) whereas aromatic residues are found at the top of the pocket, in the short α-helix that precedes the ECL between TM helices 5 and 6.
Moreover, both binding clefts have similar mean polarities ( +0.32 and +0.29) when compared with those found for P-gp ( +0.32 and +0.33 for R and H sites respectively). Yet, some differences could still be found within the drug-binding sites due to the asymmetrical residue distribution (Fig. 11). First, while the center of the cleft is more polar ( +0.35 or +0.32 in each monomer), the uppermost region of the site is more lipophilic (ranging from +0.25 to +0.28). Second, the spatial position of the first transmembrane helix affects the polarity of the drug-binding site near the dimer interface. As in monomer A TM helix 1 of the opposite monomer is closer to TM helix 5, the dimer interface is less hydrophilic, with a mean polarity value similar to the one observed at the top of the drug-binding site ( +0.26, Fig. 11A).
However, a different conformation in monomer B induced by the lateral shift of TM helix 1b exposes the more hydrophilic core of TM helix 5 and creates an additional site with a mean polarity similar to the DBS center ( +0.32) and providing an additional binding site where cholesterol could be found18. Thus, as in P-gp32, these distinct polarities within the binding cleft may be one possible explanation for the results obtained by Clark et al.54 in which, although using the R482G variant, several substrates appear to bind to distinct locations (Fig. 12A). By combining this information with the results for cholesterol (obtained from both the MD simulations and molecular docking) and with Rhodamine-123 (R123) docking poses in monomer B (at Site 2, −6.7 kcal.mol−1 against −7.4 kcal.mol−1 for the top-ranked binding pose), a new schematic representation of the ABCG2 drug-binding sites, based on the one previously proposed by Clark et al., could be generated (Fig. 12B). Hence, while mitoxantrone (MX, −6.3 kcal.mol−1) and Hoechst33342 (H33342, −8.9 kcal.mol−1) bind near the top of the drug-binding site, daunorubicin (DAU, −7.9 kcal.mol−1), doxorubicin (DOX, −7.8 kcal.mol−1) and prazosin (PRAZ, −7.4 kcal.mol−1) are found to bind closer to the center of the ‘groove’.
(A) Top-ranked docking poses for substrates: mitoxantrone (MX, pink), Hoechst 33342 (H3342, green), daunorubicin (DAU, cyan), doxorubicin (DOX, orange) and prazosin (PRAZ, yellow) at the drug-binding site in monomer A, (B) Proposed schematic representation of ABCG2 drug-binding sites, based on a previous model by Clark et al. and (C) Top-ranked docking poses for modulators: cyclosporine (CYC, brown), fumitremorgin C (FTC, purple) and KO143 (dark green). The mesh represents the drug-binding pocket surface and is colored by the electrostatic potential (blue, electron donor; gray, neutral and red, electron acceptor).
Concerning R123, although a specific site is proposed to exist close to the dimer interface (as for mitoxantrone) in the wild-type protein, our data together with the recently reported cholesterol localization in the human ABCG2 crystal structure suggests that this location would alternatively be part of the translocation pathway (Fig. 12B, red dashed line). Therefore, one possible explanation for the increased R123 efflux described for the R482G variant16,42 may be that the Arg → Gly mutation at position 482 induces a permanent shift on the position of TMH 3 towards TMH 4 while moving away from TMH 1, which would allow the formation of the above identified cavity between TMH1 and TMH5 (as seen in one of the monomers, Fig. 11B) where R123 was found in our docking procedure. Finally, a distinct site for cholesterol could also be identified in the vicinity of the active Y413 CRAC motif (Figs. 5A-B, docking, −7.9 kcal.mol−1), in agreement with experimental studies in which cholesterol influence the binding process of ABCG2 substrates27 as an allosteric co-activator or through co-transport with substrates28 and in agreement with the results found for the heterodimeric sterol transporter ABCG5/G8 41.
Membrane data
As cholesterol is a crucial component of lipid bilayers and is important for the activity of the ABCG2 transporter, we measured important parameters as the protein’s angle of insertion (tilt), area per lipid (A L), thickness (D HH) and cholesterol distribution around the ABCG2 transporter to assess the influence of the transporter on the surrounding lipid environment. Regarding the membrane insertion angle (tilt), in our systems ABCG2 is found to have a 3 ± 1° tilt, which is in agreement with the low tilt angle predicted by OPM database (6°). As P-gp, another member of the ABC transporter family that is known to reshape the surrounding lipid environment57,58, ABCG2 was found to have a strong influence on the membrane. When comparing the area per lipid (A L), and taking into account the values for pure DMPC (0.602 nm2) and with 20 molar-% cholesterol (0.531 nm2; partial areas of 0.563 and 0.396 nm2 for DMPC and cholesterol respectively), it was found that the presence of ABCG2 increases the A L up to values similar to pure DMPC membranes (0.591 nm2), corresponding to a 10% increase on the membrane’s mean areas per lipid (partial areas of 0.661 and 0.316 nm2 for DMPC and cholesterol, respectively). Similarly, although membrane thickness also decreases (3.62 nm) when compared with 20 molar-% CHOL:DMPC membranes (3.98 nm), it does not reach the values for pure DMPC (3.27 nm). We revisited our previous studies on P-glycoprotein-membrane systems36 and observed that while A L values shifts from 0.636 nm2 (pure POPC) to 0.593 nm2 (20% CHOL:POPC, −7%), in P-gp/bilayer systems comprising cholesterol (system built from the refined model but not included in ref15) the A L is 0.612 nm2 (3% increase). Regarding thickness, in the presence of P-gp the calculated thickness of a 20% CHOL:POPC membrane was 4.01 nm, against 3.79 nm for 20% CHOL:POPC membrane ( +5%) and 3.72 ( +7%) for pure POPC. Therefore, these data show a stronger effect by ABCG2 in the surrounding lipid environment when compared to P-gp.
As it is known that cholesterol reduces membrane fluidity by increasing the orientational order of the hydrophobic chains, reducing its area per lipid and by increasing its thickness59,60, this led us to hypothesize that the modulation of cholesterol content within the membrane27,61 affects the ABCG2 dimer cohesion due to a decrease on the membrane’s lateral tension. This hypothesis also provides a suitable explanation for the irreversible dissociation of ABCG2 dimers by detergents, as previously shown by Telbisz et al.28. This also suggests that ABCG2 has a higher dependency on the biophysical properties of the membrane.
In addition to the above results, we also generated volumetric maps interpolating the z coordinates of selected atoms (phosphate atoms) into a regular orthogonal grid in the xy plane and superimposing them with the ABCG2 transporter62 (Fig. 13). Quite interestingly, from the average thickness map (calculated from the whole membrane; Fig. 13A), it is possible to identify regions of thicker membrane patches next to the cholesterol recognition/interaction amino acid consensus, corresponding to an increase in the number of cholesterol molecules and closer to the previously identified drug-binding sites, along with thinner patches on the vicinity of TM helix 6. However, by observing the deformation maps for the periplasmic (Fig. 13B) and cytoplasmic (Fig. 13C) leaflets it could be identified two symmetrical membrane ‘patches’ with increased thickness, both having a funnel-like shape, from which molecules can access, for each monomer, to the surface clefts where the drug-binding sites are located.
Volumetric thickness maps for the ABCG2/membrane systems. (A) average thickness map (B) deformation map for the upper leaflet and (C) deformation map for the lower leaflet. ABCG2 is depicted in cartoon representation63. Negative and positive deformations are depicted in a blue-red scale for membrane contraction and expansion, respectively.
Conclusion
Since the identification of the ABC transporters role in multidrug resistance (MDR) that efflux pumps as P-glycoprotein, MRP1, BCRP (also known as ABCG2) and, more recently ABCB5, are being thoroughly studied in order to better understand their mechanism of action, aiming the development of potent and selective MDR modulators to avoid chemotherapy failures.
Unlike P-glycoprotein64, the lack of suitable crystallographic structures for ABCC1 and ABCG2 has severely compromised the knowledge on these two types of ABC transporters. The recent publication of a crystallographic structure of the ABCG family, the ABCG5/G8 heterodimeric transporter, revealed for the first time a new transmembrane arrangement that is characteristic of the ABCG transporters. Although incomplete and despite the low structural identity with ABCG2, it could still be a proper template for modeling ABCG2, thus allowing new insights on the structural dynamics for this class of transporters. Herein, we report on a new ABCG2 homology model from a ABCG5 homodimer (obtained by duplication of the C chain of the G5G8 crystallographic structure)41. This was performed to avoid possible asymmetries due to the utilization of a heterodimeric transporter as a homology template. Indeed, we also provided evidences that the herein newly obtained model is comparable to the recently published human ABCG2 cryo-EM structure and that it performed better than a previously published one42, when evaluated using available structure assessment and validation computational tools. It is also important to refer that i) our model and all the above results were obtained prior to the release of the cryo-EM hABCG2 structure and ii) due to the release of the cryo-EM structure, we felt the need to rewrite some parts of this paper in order to validate our in silico approach for the development of a ABCG2 homology model. Both our approach and our model are now supported through a thorough comparison with the structural information that could be retrieved from the novel cryo-EM structure and, more important, our paper provides new and valuable information that can be further used to better understand i) the structural dynamics of the ABCG transporters family, ii) possible drug-binding sites within the ABCG2 structure and iii) the intimate relationships between the surrounding lipid bilayer and the transporter.
In our homology model no disulfide bonds were modeled, because neither intra- or intermolecular disulfide bonds are required for the ABCG2 transport activity19,20,21,22,23,24. Even so, Cα-Cα distances between cysteine residues involved in intra- (592 and 608) or intermolecular (603) bridges were found to be stable, and at distances compatible with the physiological formation of disulfide bonds in ABCG2 homodimeric complexes. Thus, our results combined with previous published experimental data suggests that inter- and intramolecular disulfide bridges involving C592, C603 and C608 are more important for specific mechanisms as membrane targeting20 or monoclonal antibody binding65, rather than directly affecting the expression, function or activity of the transporter.
Signal propagation through the homodimeric structure was also assessed through a principal component analysis, in order to better understand how conformational changes can lead to substrate efflux. Our simulations demonstrated for the first time that signal transmission occurs through the herein de novo modeled external helices (absent in the crystallographic structure) that connect the nucleotide-binding domain with the first transmembrane helix. Accordingly, we suggest that drug binding and/or ATP binding is able to induce spring-like movements that propagate conformational changes through these specific domains i) into the opposite NBD or ii) into the TM domain of the same monomer. Thus, the development of small molecules targeting these specific motifs may become a promising alternative to develop novel and specific ABCG2 efflux modulators able to tackle MDR in cancer by impairing drug-induced signal transmission, similar to what was recently suggested for P-gp66.
Molecular docking revealed two symmetric drug-binding clefts, one in each monomer, in agreement with a previous study by Clark et al.54. These two membrane-exposed clefts are flanked by TMH1-2 of the opposite monomer and by TM helices 4–6 and occupy almost all the buried TM surface, with a mean volume of ~1900Å3 and regions of distinct polarities, which may explain why substrates seem to bind to distinct locations within this region. Thus, a new schematic representation of the ABCG2 drug-binding sites is herein proposed, based on a previous scheme by Clark and co-workers54, where a distinct region for cholesterol binding (based on MD and docking results) and a translocation pathway can be proposed. Interestingly, as our results suggest the existence of distinct drug-binding sites for substrates, modulators and cholesterol, it theoretically would be possible to develop specific ABCG2 efflux modulators that i) could specifically compete with cholesterol (thus reducing the activity of the pump), ii) binds in a different location with high affinity while simultaneously being able to block the substrate-binding site and iii) could specifically bind to the whole “surface cleft” in order to impair conformational changes by increasing the structural cohesion of the transmembrane helical domains (as observed for tariquidar in P-gp)32,67,68.
Finally, like other ABC transporters that also reshape the surrounding lipid environment2, the biophysical properties of the membrane patch surrounding ABCG2 were also assessed. Important changes in area per lipid, thickness and membrane deformation profiles were observed that may explain the strong dependency of ABCG2 for cholesterol-enriched membranes. Interestingly, a thicker “funnel-like” shape region within the membrane was identified from which molecules can access, for each monomer, to the surface clefts where the drug-binding sites are located. As the presence of detergents are often associated to function impairment in ABC transporters28,69,70, our results also corroborate that reducing its cholesterol content and increasing membrane fluidity may have a deleterious effect not only in protein function but also regarding drug access to ABCG2 from the lipid bilayer.
Material and Methods
Initial Structures and Software
The ABCG5/G8 heterodimeric crystal structure (PDB ID: 5DO7)41 was obtained from the Protein Data Bank (www.rcsb.org)71 and parameterized according to the GROMOS9672,73 force field with the 54A774,75 parameter set. A lipid membrane comprising dimiristoylphosphatidylcholine (DMPC) with 20% cholesterol (CHOL) was obtained from the Computational Molecular Biophysics Group at the Georg-August-Universität Göttingen (http://cmb.bio.uni-goettingen.de/cholmembranes.html)76,77. For DMPC, the parameterization developed by Poger et al.78,79 was used due to its ability to accurately reproduce properties of the lipid bilayers as area and volume per lipid ratios78,80,81, fluid-phase78,79,82,83 and solvation78,79,83 properties. Cholesterol and ATP parameterization was prepared online through the Automated Topology Builder (ATB) and Repository84,85 or PRODRG86 servers, manually curated and added with Merz-Kollman partial charges, assigned through ab initio calculations at the Hartree-Fock level of theory using the 6–31 G(d) basis set in Gaussian0387 program. Protein manipulation, protonation and homology modeling was performed in MOE 2015.100188. The GROMACS simulation package 5.0.789,90,91,92 was used for the MD simulations and protein insertion into the lipid membranes through the g_membed 93 module. Areas per lipid (A L) and thickness (D HH) of all systems were calculated with the MembPlugin62 extension in VMD94. Principal Component analysis (PCA) for the evaluation of the protein’s motion patterns was performed using the ProDy95,96 software through the NMWiz plugin in VMD94. Free energies of binding were calculated using g_mmpbsa 97, with polar solvation energies corrected by generating ion-accessibility and dielectric maps incorporating the membrane environment (dielectric slab constant is set to 2.0 using the draw_membrane2 program) through in-house python scripts98. Both VMD and MOE were used for molecular inspection and visualization.
Systems Construction
Due to a higher identity and similarity between ABCG5 and ABCG2 (27% and 48% respectively and better than ABCG8), and to avoid homology errors that may arise from crystallographic asymmetries in the ABCG5/G8 heterodimer, one of the initial steps was the assembly of an ABCG5 homodimer by superimposing a copy of the ABCG5 (chain C) structure on the ABCG8 transporter (chain D), being chain C chosen over chain A due to its higher quality (as described in the Full wwPDB X-ray Structure Validation Report99, available at http://ftp.wwpdb.org/pub/pdb/validation_reports/do/5do7/ 5do7_full_validation.pdf).
Missing sequences
In order to obtain a full-length ABCG5, three sequences that were missing in the initial crystallographic structure were built: the A-loop (residues 47–66), the linker (residues 349–354) and the long extracellular loop (ECL, residues 590–598), based on the Chou-Fassman Secondary Structure Prediction server100. While the linker and ECL sequences were assessed as disordered, the A-loop was predicted to be a beta hairpin101, similar to that found in P-gp. Therefore, this segment was constructed using the Protein Builder module in MOE, followed by protonation and energy minimization, keeping the distance between the amine and carboxyl groups to the experimental one found in the crystallographic structure (4.5 Å). The resulting structure was then solvated, neutralized by adding an adequate number of counter ions and further studied in GROMACS through a series of eight MD simulations, each one with 50 ns duration and initial random velocities assigned from the temperature-related Maxwell-Boltzmann distribution. All obtained structures were evaluated regarding the final partial energies of the system, Ramachandran102 plots, and visual inspection. The most suitable conformation was added to the ABCG5 crystallographic structure to be used as a template for the ABCG2 homology model generation. The remaining sequences (linker and ECL) were modeled as coils.
Homology modeling
To minimize possible clashes between atoms of the crystallographic structure and the added missing peptide sequences, prior to homology modeling a short energy minimization step took place while keeping the crystallographic structures’ heavy atoms spatially restrained. After loading the ABCG2 fasta sequence (retrieved from UniProt, accession number Q9UNQ0), a homology model was built in MOE (Homology Model module), creating 100 mainchain models with 10 sidechain samplings at 300 K, in a total of 1000 models (Amber:EHT force-field). All other options were set as default. However, visual evaluation of the final model detected several problems, mainly related to mismodeled helical domains. Therefore, the secondary structure of these domains was predicted in CFSSP and built de novo in the homology model: while TM helices 4 and 5 were partially rebuilt as fully helical only from the distorted portions up to the nearest loop, the connecting helical domains following the NPXDF motif (two α-helices that precede the coil linking the NDB to the TMD, named CnH in ABCG5/G8 and C2a/C2b in hABCG2) were completely rebuilt as fully helical, keeping its relative location similar to the original spatial coordinates in the ABCG5 experimental structure.
Linker equilibration
Since the linker connecting the NBD-TMD (residues 354–375) is longer than in ABCG5, we performed an additional computational study of this structure in GROMACS in the presence of a DMPC membrane. To that matter, after building the coil in MOE, a system comprising residues 332–389 (helix-linker-helix) was ported into GROMACS, inserted in an adequate membrane patch, solvated and neutralized with an adequate number of counter ions. Then, the system was energy minimized and equilibrated through a NVT run (10 ps, 303 K), followed by a short 4 ns NpT run (for membrane equilibration, 1 bar) and a 50 ns NpT run while keeping the flanking helices 332–353 (cytosolic) and 374–389 (anchored at the membrane interface) spatially restrained to allow the equilibration of the connecting coil. At the end, the system was evaluated and the most favorable conformation was inserted into the ABCG2 homology model. The quality of the model was assessed through online validation servers as ERRAT103, MolProbity104,105, PROCHECK106,107 and SwissModel Structure assessment108,109,110 tools.
Construction of membrane systems and refinement protocol
The relative position of the lipid bilayer was taken from the OPM111 database. Accordingly, a DMPC membrane with 20 molar-% cholesterol was chosen because i) cholesterol is required for optimal ABCG2 activity27 probably acting as an allosteric regulator112, ii) in the original publication the protein was also reconstituted in DMPC bicelles in the presence of cholesterol prior to the crystallization step and ii) it provides an optimal hydrophobic environment to accommodate the hydrophobic thickness for the ABCG2 transmembrane domains (PPM server prediction:113 28.6 ± 0.9 Å). After membrane insertion, the system was water soaked (59.982 molecules) and neutralized with 22 chlorine ions, originating a system with 221.728 atoms. The system was then energy minimized and a 10 ps NVT run at 303 K followed, above the DMPC gel-fluid phase transition114,115. Then, the DMPC:CHOL membrane was allowed to correctly adjust to the protein structure through a 20 ns NpT run where all the protein’s heavy atoms were kept restrained. Finally, in order to better equilibrate all the de novo modeled domains, a refinement protocol was applied by progressively removing the heavy atoms’ spatial restrictions through a series of three sequential 10 ns NpT runs for the modeled helices (residues 310–354), linker (residues 355–375) and both helix-linker (residues 310–375) respectively. The resulting system was the starting point for a 200 ns unrestrained NpT run. Two additional systems were obtained by removing one DMPC molecule, found to be located between both ABCG2 monomers, at the beginning (200 ns MD run) or after 100 ns of the 200 ns NpT run (for another 100 ns), for a total of 500 ns simulation time. At the end, the quality of all three models was again assessed by the previously used validation servers.
ATP systems
Using the final conformation of the 200 ns NpT run and to assess the effect of ATP binding on the ABCG2 homodimer, two ATP-Mg2+ complexes were placed in the nucleotide-binding domains (one for each monomer), with the magnesium ion in the vicinity of D210 (Mg2+-chelating residue), the ribose moiety close to the Walker A motif and the adenine next to the A-loop, followed by a short energy minimization step in MOE to allow the residues sidechains to adjust to the presence of ATP. Then, in GROMACS, the protein was further allowed to equilibrate to the presence of the nucleotide through a 10 ns NpT run where ATP-Mg2+ coordinates were kept spatially restrained. Afterwards, a 100 ns unrestrained NpT run was performed.
Simulation parameters
For all simulations, periodic boundary conditions (PBC) were applied. Simple energy minimizations were performed using the steepest descent method. All NVT runs were performed at 303 K using a Velocity-rescale (V-rescale)116 thermostat. NpT runs used the Nosé-Hoover117,118 thermostat and the Parrinello-Rahman117,119 barostat for temperature (303 K) and pressure (1 bar) coupling, respectively. Due to the presence of membranes, pressure equilibration was achieved through a semi-isotropic pressure coupling, with the systems’ compressibility defined as 4.5 × 10−5 bar−1 and the initial box was defined with dimensions xyz of 17.37 × 11.58 × 13.50 nm3. The Particle Mesh Ewald (PME) with cubic interpolation120,121 was employed, with cut-off radius of 12 Å for both electrostatic and van der Waals interactions and an FFT grid spacing of 0.16 nm for long-range electrostatics. Group-based or Verlet122 cut-off schemes were applied for the calculation of non-bonded interactions done on CPU or GPU respectively. The SETTLE123 (for water molecules) or LINCS124,125 algorithms were used to constrain all bond lengths.
Molecular Docking
Substrates (N = 69) and modulators (N = 19) were selected according to the publication by Mo et al.10 Molecular docking was made with all three ABCG2 homology models obtained at the end of the MD simulations. MarvinSketch v17.2.20126 was used for drawing and minimizing structures. PDBQT files were generated with AutoDockTools127 for further utilization in AutoDock VINA v1.1.2128,129 docking software. The binding location was defined by a docking box including the whole transmembrane domains, with dimensions xyz of 16.87 × 14.06 × 16.87 Å3 (xy corresponds to the membrane plane). Due to the large search space volume (over 27.000 Å3), ‘exhaustiveness’ parameter was manually set to 50. Visual inspection of the docking poses was made in MOE to allow the identification of individual docking zones. For each site, the docked molecules were overlapped with the cavity search results by EPOSBP 130,131, thus allowing the identification of lining atoms (within a distance of 5 Å from the pocket probes), mean pocket volumes and polarities (ratio of the sum of N, O, and S atoms to the sum of N, O, S, and C atoms). Graphical images of the docking poses and molecular maps of the pocket surfaces were generated in MOE.
Data availability
The final configuration of the MD refined ABCG2 homology structure and the ATP systems are available for download at our website (http://chemistrybits.com/).
Change history
17 April 2018
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Ferlay, J. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer 136, E359–386 (2015).
Ferreira, R. J., dos Santos, D. J. V. A. & Ferreira, M.-J. U. P-glycoprotein and membrane roles in multidrug resistance. Future Med Chem 7, 929–946 (2015).
Juliano, R. L. & Ling, V. A surface glycoprotein modulating drug permeability in Chinese hamster ovary cell mutants. Biochim Biophys Acta 455, 152–162 (1976).
Cole, S. P. et al. Overexpression of a transporter gene in a multidrug-resistant human lung cancer cell line. Science 258, 1650–1654 (1992).
Doyle, L. A. et al. A multidrug resistance transporter from human MCF-7 breast cancer cells. Proc Natl Acad Sci USA 95, 15665–15670 (1998).
Allikmets, R., Schriml, L. M., Hutchinson, A., Romano-Spica, V. & Dean, M. A human placenta-specific ATP-binding cassette gene (ABCP) on chromosome 4q22 that is involved in multidrug resistance. Cancer Res 58, 5337–9 (1998).
Miyake, K. et al. Molecular cloning of cDNAs which are highly overexpressed in mitoxantrone-resistant cells: demonstration of homology to ABC transport genes. Cancer Res 59, 8–13 (1999).
Frank, N. Y. et al. Regulation of Progenitor Cell Fusion by ABCB5 P-glycoprotein, a Novel Human ATP-binding Cassette Transporter. J Biol Chem 278, 47156–47165 (2003).
Frank, N. Y. et al. ABCB5-mediated doxorubicin transport and chemoresistance in human malignant melanoma. Cancer Res 65, 4320–4333 (2005).
Mo, W. & Zhang, J.-T. Human ABCG2: structure, function, and its role in multidrug resistance. Int J Biochem Mol Biol 3, 1–27 (2012).
Stacy, A. E., Jansson, P. J. & Richardson, D. R. Molecular Pharmacology of ABCG2 and Its Role in Chemoresistance. Mol Pharmacol 84, 655–669 (2013).
Jani, M. et al. Structure and function of BCRP, a broad specificity transporter of xenobiotics and endobiotics. Arch Toxicol 88, 1205–1248 (2014).
Xu, J., Liu, Y., Yang, Y., Bates, S. E. & Zhang, J.-T. Characterization of oligomeric human half-ABC transporter ATP-binding cassette G2. J Biol Chem 279, 19781–9 (2004).
Macalou, S. et al. The linker region of breast cancer resistance protein ABCG2 is critical for coupling of ATP-dependent drug transport. Cell Mol Life Sci 73, 1927–1937 (2016).
Kage, K. et al. Dominant-negative inhibition of breast cancer resistance protein as drug efflux pump through the inhibition of S-S dependent homodimerization. Int J Cancer 97, 626–630 (2002).
Litman, T. et al. Use of peptide antibodies to probe for the mitoxantrone resistance-associated protein MXR/BCRP/ABCP/ABCG2. BBA-Biomembranes 1565, 6–16 (2002).
Wong, K., Briddon, S. J., Holliday, N. D. & Kerr, I. D. Plasma membrane dynamics and tetrameric organisation of ABCG2 transporters in mammalian cells revealed by single particle imaging techniques. Biochim Biophys Acta 1863, 19–29 (2016).
Taylor, N. M. I. et al. Structure of the human multidrug transporter ABCG2. Nature 546, 504–509 (2017).
Kage, K., Fujita, T. & Sugimoto, Y. Role of Cys-603 in dimer/oligomer formation of the breast cancer resistance protein BCRP/ABCG2. Cancer Sci 96, 866–872 (2005).
Henriksen, U., Fog, J. U., Litman, T. & Gether, U. Identification of intra- and intermolecular disulfide bridges in the multidrug resistance transporter ABCG2. J Biol Chem 280, 36926–36934 (2005).
Mitomo, H. et al. A functional study on polymorphism of the ATP-binding cassette transporter ABCG2: critical role of arginine-482 in methotrexate transport. Biochem J 373, 767–74 (2003).
Xu, J. et al. Oligomerization Domain of the Multidrug Resistance-Associated Transporter ABCG2 and Its Dominant Inhibitory Activity. Cancer Res 67, 4373–4381 (2007).
Bhatia, A., Schäfer, H.-J. & Hrycyna, C. A. Oligomerization of the human ABC transporter ABCG2: evaluation of the native protein and chimeric dimers. Biochemistry 44, 10893–904 (2005).
Wakabayashi, K. et al. Intramolecular Disulfide Bond Is a Critical Check Point Determining Degradative Fates of ATP-binding Cassette (ABC) Transporter ABCG2 Protein. J Biol Chem 282, 27841–27846 (2007).
Ejendal, K. F. K., Diop, N. K., Schweiger, L. C. & Hrycyna, C. A. The nature of amino acid 482 of human ABCG2 affects substrate transport and ATP hydrolysis but not substrate binding. Protein Sci 15, 1597–1607 (2006).
Pozza, A., Perez-Victoria, J. M., Sardo, A., Ahmed-Belkacem, A. & Di Pietro, A. Purification of breast cancer resistance protein ABCG2 and role of arginine-482. Cell Mol Life Sci 63, 1912–1922 (2006).
Storch, C. H., Ehehalt, R., Haefeli, W. E. & Weiss, J. Localization of the Human Breast Cancer Resistance Protein (BCRP/ABCG2) in Lipid Rafts/Caveolae and Modulation of Its Activity by Cholesterol in Vitro. J Pharmacol Exp Ther 323, 257–264 (2007).
Telbisz, Á., Özvegy-Laczka, C., Hegedűs, T., Váradi, A. & Sarkadi, B. Effects of the lipid environment, cholesterol and bile acids on the function of the purified and reconstituted human ABCG2 protein. Biochem J 450, 387–395 (2013).
Li, H. & Papadopoulos, V. Peripheral-type benzodiazepine receptor function in cholesterol transport. Identification of a putative cholesterol recognition/interaction amino acid sequence and consensus pattern. Endocrinology 139, 4991–4997 (1998).
Gál, Z. et al. Mutations of the central tyrosines of putative cholesterol recognition amino acid consensus (CRAC) sequences modify folding, activity, and sterol-sensing of the human ABCG2 multidrug transporter. Biochim Biophys Acta 1848, 477–487 (2015).
Aller, S. G. et al. Structure of P-glycoprotein reveals a molecular basis for poly-specific drug binding. Science 323, 1718–1722 (2009).
Ferreira, R. J., Ferreira, M.-J. U. & dos Santos, D. J. V. A. Molecular docking characterizes substrate-binding sites and efflux modulation mechanisms within P-glycoprotein. J Chem Inf Model 53, 1747–1760 (2013).
Jara, G. E., Vera, D. M. A. & Pierini, A. B. Binding of Modulators to Mouse and Human Multidrug Resistance P-glycoprotein. A Computational Study. J Mol Graph Model 46, 10–21 (2013).
Dolghih, E., Bryant, C., Renslo, A. R. & Jacobson, M. P. Predicting binding to p-glycoprotein by flexible receptor docking. PLoS Comput Biol 7, e1002083 (2011).
McCormick, J. W., Vogel, P. D. & Wise, J. G. Multiple Drug Transport Pathways through Human P-Glycoprotein. Biochemistry 54, 4374–4390 (2015).
Ferreira, R. J., Ferreira, M.-J. U. & dos Santos, D. J. V. A. Insights on P-Glycoprotein’s Efflux Mechanism Obtained by Molecular Dynamics Simulations. J Chem Theory Comput 8, 1853–1864 (2012).
O’Mara, M. L. & Mark, A. E. The Effect of Environment on the Structure of a Membrane Protein: P-Glycoprotein under Physiological Conditions. J Chem Theory Comput 8, 3964–3976 (2012).
Ma, J. & Biggin, P. C. Substrate versus inhibitor dynamics of P-glycoprotein. Proteins 81, 1653–1668 (2013).
Prajapati, R. & Sangamwar, A. T. Translocation mechanism of P-glycoprotein and conformational changes occurring at drug-binding site: Insights from multi-targeted molecular dynamics. BBA-Biomembranes 1838, 2882–2898 (2014).
Johnson, Z. L. & Chen, J. Structural Basis of Substrate Recognition by the Multidrug Resistance Protein MRP1. Cell 168, 1075–1085 (2017).
Lee, J.-Y. et al. Crystal structure of the human sterol transporter ABCG5/ABCG8. Nature 533, 561–564 (2016).
László, L., Sarkadi, B. & Hegedűs, T. Jump into a New Fold—A Homology Based Model for the ABCG2/BCRP Multidrug Transporter. PLoS One 11, e0164426 (2016).
Benkert, P., Biasini, M. & Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 27, 343–350 (2011).
Domicevica, L. & Biggin, P. C. Homology modelling of human P-glycoprotein. Biochem Soc Trans 43, 952–958 (2015).
Bordoli, L. et al. Protein structure homology modeling using SWISS-MODEL workspace. Nat. Protoc. 4, 1–13 (2008).
Zhou, H. & Zhou, Y. Distance-scaled, finite ideal-gas reference state improves structure-derived potentials of mean force for structure selection and stability prediction. Protein Sci 11, 2714–2726 (2009).
Morris, A. L., MacArthur, M. W., Hutchinson, E. G. & Thornton, J. M. Stereochemical quality of protein structure coordinates. Proteins 12, 345–364 (1992).
Engh, R. A. & Huber, R. Accurate bond and angle parameters for X-ray protein structure refinement. Acta Cryst 47, 392–400 (1991).
Rosenberg, M. F. et al. Three-dimensional structure of the human breast cancer resistance protein (BCRP/ABCG2) in an inward-facing conformation. Acta Crystallogr D Biol Crystallogr 71, 1725–1735 (2015).
Hazai, E. & Bika, Z. Homology modeling of breast cancer resistance protein (ABCG2). J Struct Biol 162, 63–74 (2008).
Ni, Z., Bikadi, Z., F. Rosenberg, M. & Mao, Q. Structure and Function of the Human Breast Cancer Resistance Protein (BCRP/ABCG2). Curr Drug Metab 11, 603–617 (2010).
Liu, Y., Yang, Y., Qi, J., Peng, H. & Zhang, J.-T. Effect of Cysteine Mutagenesis on the Function and Disulfide Bond Formation of Human ABCG2. J Pharmacol Exp Ther 326, 33–40 (2008).
Haider, A. J. et al. Identification of residues in ABCG2 affecting protein trafficking and drug transport, using co-evolutionary analysis of ABCG sequences. Biosci Rep 35, e00241 (2015).
Clark, R., Kerr, I. D. & Callaghan, R. Multiple drug binding sites on the R482G isoform of the ABCG2 transporter. Br J Pharmacol 149, 506–515 (2006).
Xu, Y. et al. Allocrite Sensing and Binding by the Breast Cancer Resistance Protein (ABCG2) and P-Glycoprotein (ABCB1). Biochemistry 54, 6195–6206 (2015).
Telbisz, A., Hegedus, C., Varadi, A., Sarkadi, B. & Ozvegy-Laczka, C. Regulation of the Function of the Human ABCG2 Multidrug Transporter by Cholesterol and Bile Acids: Effects of Mutations in Potential Substrate and Steroid Binding Sites. Drug Metab Dispos 42, 575–585 (2014).
Romsicki, Y. & Sharom, F. J. The membrane lipid environment modulates drug interactions with the P-glycoprotein multidrug transporter. Biochemistry 38, 6887–6896 (1999).
Oleinikov, V. A., Fleury, F., Ianoul, A., Zaitsev, S. & Nabiev, I. P-glycoprotein effect on the properties of its natural lipid environment probed by Raman spectroscopy and Langmuir-Blodgett technique. FEBS Lett 580, 4953–4958 (2006).
Yeagle, P. L. Modulation of membrane function by cholesterol. Biochimie 73, 1303–1310 (1991).
Yeagle, P. L. Cholesterol and the cell membrane. BBA-Rev Biomembr 822, 267–287 (1985).
Telbisz, Á. et al. Membrane cholesterol selectively modulates the activity of the human ABCG2 multidrug transporter. BBA-Biomembranes 1768, 2698–2713 (2007).
Guixa-Gonzalez, R. et al. MEMBPLUGIN: studying membrane complexity in VMD. Bioinformatics 30, 1478–1480 (2014).
Frishman, D. & Argos, P. Knowledge-based protein secondary structure assignment. Proteins Struct Funct Genet 23, 566–579 (1995).
Ferreira, R. J., Ferreira, M.-J. U. & dos Santos, D. J. V. A. Reversing cancer multidrug resistance: insights into the efflux by ABC transports from in silico studies. WIREs Comput Mol Sci 5, 27–55 (2015).
Hegedu, C., Litman, T., Goda, K., Dokholyan, N. V. & Sorrentino, B. P. Interaction with the 5D3 Monoclonal Antibody Is Regulated by Intramolecular Rearrangements but Not by Covalent Dimer Formation of the Human ABCG2 Multidrug Transporter. J Biol Chem 283, 26059–26070 (2008).
Ferreira, R. J., Bonito, C. A., Ferreira, M. J. U. & dos Santos, D. J. V. A. About P-glycoprotein: a new drugable domain is emerging from structural data. WIREs Comput Mol Sci 7, e1316 (2017).
Loo, T. W. & Clarke, D. M. Mapping the Binding Site of the Inhibitor Tariquidar That Stabilizes the First Transmembrane Domain of P-glycoprotein. J Biol Chem 290, 29389–29401 (2015).
Li, X.-Q. et al. Reversal of P-gp and BCRP-mediated MDR by Tariquidar derivatives. Eur J Med Chem 101, 560–572 (2015).
Hegedüs, C., Telbisz, Á., Hegedüs, T., Sarkadi, B. & Özvegy-Laczka, C. Lipid Regulation of the ABCB1 and ABCG2 Multidrug Transporters. in Advances in Cancer Research (eds. Schuetz, J. D. and Ishikawa, T.) 97–137 (Academic Press, 2015).
Sharom, F. J. Complex Interplay between the P-Glycoprotein Multidrug Efflux Pump and the Membrane: Its Role in Modulating Protein Function. Front Oncol 4, 41 (2014).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res 28, 235–242 (2000).
Bonvin, A. M., Mark, A. E. & van Gunsteren, W. F. The GROMOS96 benchmarks for molecular simulation. Comput Phys Commun 128, 550–557 (2000).
Scott, W. R. et al. The GROMOS Biomolecular Simulation Program Package. J Phys Chem A 103, 3596–3607 (1999).
Schmid, N. et al. Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur Biophys J 40, 843–56 (2011).
Reif, M. M., Hünenberger, P. H. & Oostenbrink, C. New Interaction Parameters for Charged Amino Acid Side Chains in the GROMOS Force Field. J Chem Theory Comput 8, 3705–3723 (2012).
Hub, J. S., Winkler, F. K., Merrick, M. & de Groot, B. L. Potentials of Mean Force and Permeabilities for Carbon Dioxide, Ammonia, and Water Flux across a Rhesus Protein Channel and Lipid Membranes. J Am Chem Soc 132, 13251–13263 (2010).
Wennberg, C. L., van der Spoel, D. & Hub, J. S. Large Influence of Cholesterol on Solute Partitioning into Lipid Membranes. J Am Chem Soc 134, 5351–5361 (2012).
Poger, D., Van Gunsteren, W. F. & Mark, A. E. A new force field for simulating phosphatidylcholine bilayers. J Comput Chem 31, 1117–1125 (2010).
Poger, D. & Mark, A. E. On the Validation of Molecular Dynamics Simulations of Saturated and cis -Monounsaturated Phosphatidylcholine Lipid Bilayers: A Comparison with Experiment. J Chem Theory Comput 6, 325–336 (2010).
Chiu, S.-W. et al. Incorporation of Surface Tension into Molecular Dynamics Simulation of Interface: A Fluid Phase Lipid Bilayer Membrane. Biophys J 69, 1230–1245 (1995).
Kukol, A. Lipid Models for United-Atom Molecular Dynamics Simulations of Proteins. J Chem Theory Comput 5, 615–626 (2009).
Chandrasekhar, I. et al. A consistent potential energy parameter set for lipids: dipalmitoylphosphatidylcholine as a benchmark of the GROMOS96 45A3 force field. Eur Biophys J 32, 67–77 (2003).
Chandrasekhar, I., Oostenbrink, C. & van Gunsteren, W. F. Simulating the Physiological Phase of Hydrated DPPC Bilayers: The Ester Moiety. Soft Matter 2, 27–45 (2004).
Malde, A. K. et al. An Automated Force Field Topology Builder (ATB) and Repository: Version 1.0. J Chem Theory Comput 7, 4026–4037 (2011).
Koziara, K. B., Stroet, M., Malde, A. K. & Mark, A. E. Testing and validation of the Automated Topology Builder (ATB) version 2.0: prediction of hydration free enthalpies. J Comput Aided Mol Des 28, 221–33 (2014).
Schüttelkopf, A. W. & van Aalten, D. M. F. PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr D Biol Crystallogr 60, 1355–63 (2004).
Frisch, M. J. et al. Gaussian 03, Revision D.01 (2004).
Molecular Operating Environment (MOE) v2015.1001 (2015).
Berendsen, H. J., van der Spoel, D. & van Drunen, R. GROMACS: A message-passing parallel molecular dynamics implementation. Comput Phys Commun 91, 43–56 (1995).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: Algorithms for Highly Efficient, Load-Balanced, and Scalable Molecular Simulation. J Chem Theory Comput 4, 435–447 (2008).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J Comput Chem 26, 1701–18 (2005).
Wolf, M. G., Hoefling, M., Aponte-Santamaría, C., Grubmüller, H. & Groenhof, G. g_membed: Efficient insertion of a membrane protein into an equilibrated lipid bilayer with minimal perturbation. J Comput Chem 31, 2169–74 (2010).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J Mol Graph 14, 33–38 (1996).
Bakan, A., Meireles, L. M. & Bahar, I. ProDy: protein dynamics inferred from theory and experiments. Bioinformatics 27, 1575–1577 (2011).
Bahar, I., Lezon, T. R., Bakan, A. & Shrivastava, I. H. Normal mode analysis of biomolecular structures: functional mechanisms of membrane proteins. Chem Rev 110, 1463–1497 (2010).
Kumari, R., Kumar, R. & Lynn, A. G-mmpbsa -A GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54, 1951–1962 (2014).
Ferreira, R. J., Ferreira, M.-J. U. & dos Santos, D. J. V. A. Do adsorbed drugs onto P-glycoprotein influence its efflux capability? Phys Chem Chem Phys 17, 22023–22034 (2015).
Montelione, G. T. et al. Recommendations of the wwPDB NMR Validation Task Force. Structure 21, 1563–1570 (2013).
Chou, P. Y. & Fasman, G. D. Prediction of protein conformation. Biochemistry 13, 222–45 (1974).
Milner-White, E. J. & Poet, R. Four classes of beta-hairpins in proteins. Biochem J 240, 289–292 (1986).
Lovell, S. C. et al. Structure validation by Calpha geometry: phi,psi and Cbeta deviation. Proteins 50, 437–50 (2003).
Colovos, C. & Yeates, T. O. Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Sci 2, 1511–1519 (1993).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res 35, W375–W383 (2007).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr 66, 12–21 (2010).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Cryst 26, 283–291 (1993).
Laskowski, R. A., Rullmannn, J. A., MacArthur, M. W., Kaptein, R. & Thornton, J. M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8, 477–486 (1996).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42, W252–W258 (2014).
Benkert, P., Künzli, M. & Schwede, T. QMEAN server for protein model quality estimation. Nucleic Acids Res 37, W510–W514 (2009).
Lomize, M. A., Lomize, A. L., Pogozheva, I. D. & Mosberg, H. I. OPM: orientations of proteins in membranes database. Bioinformatics 22, 623–5 (2006).
Pal, A. et al. Cholesterol Potentiates ABCG2 Activity in a Heterologous Expression System: Improved in Vitro Model to Study Function of Human ABCG2. J Pharmacol Exp Ther 321, 1085–1094 (2007).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res 40, D370–D376 (2012).
Needham, D. & Evans, E. Structure and Mechanical Properties of Giant Lipid (DMPC) Vesicle Bilayers from 20 °C below to 10 °C above the Liquid Crystal-Crystalline Phase Transition at 24 °C. Biochemistry 27, 8261–8269 (1988).
Petrache, H. I., Dodd, S. W. & Brown, M. F. Area per Lipid and Acyl Length Distributions. Biophys J 79, 3172–3192 (2000).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J Chem Phys 126, 14101–14107 (2007).
Nosé, S. & Klein, M. L. Constant pressure molecular dynamics for molecular systems. Mol Phys 50, 1055–1076 (1983).
Hoover, W. Canonical dynamics: Equilibrium phase-space distributions. Phys Rev A 31, 1695–1697 (1985).
Parrinello, M. Polymorphic transitions in single crystals: A new molecular dynamics method. J Appl Phys 52, 7182–90 (1981).
Essmann, U. et al. A smooth particle mesh Ewald method. J Chem Phys 103, 8577–8593 (1995).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. J Chem Phys 98, 10089–92 (1993).
Páll, S. & Hess, B. A flexible algorithm for calculating pair interactions on SIMD architectures. Comput Phys Commun 184, 2641–2650 (2013).
Miyamoto, S. & Kollman, P. A. Settle: An analytical version of the SHAKE and RATTLE algorithm for rigid water models. J Comput Chem 13, 952–962 (1992).
Hess, B., Bekker, H., Berendsen, H. J. & Fraaije, J. G. E. M. LINCS: A linear constraint solver for molecular simulations. J Comput Chem 18, 1463–1472 (1997).
Hess, B. P-LINCS: A Parallel Linear Constraint Solver for Molecular Simulation. J Chem Theory Comput 4, 116–122 (2008).
ChemAxon. MarvinSketch v.17.2.20. (2016).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem 30, 2785–91 (2009).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31, 455–61 (2010).
Sanner, M. F. Python: a programming language for software integration and development. J Mol Graph Model 17, 57–61 (1999).
Brady, G. P. & Stouten, P. F. Fast prediction and visualization of protein binding pockets with PASS. J Comput Aided Mol Des 14, 383–401 (2000).
Kohlbacher, O. & Lenhof, H. P. BALL–rapid software prototyping in computational molecular biology. Biochemicals Algorithms Library. Bioinformatics 16, 815–24 (2000).
Acknowledgements
This project received funding from European Structural & Investment Funds through the COMPETE Programme and from National Funds through FCT – Fundação para a Ciência e a Tecnologia under the Programme grants PTDC/QEQMED/0905/2012, UID/DTP/04138/2013 and SAICTPAC/0019/2015 (FFUL). Ricardo J. Ferreira acknowledges the Ph.D grant from FCT, SFRH/BD/84285/2012. This work additionally received financial support from FCT through national funds, and co-financed by the European Union (FEDER funds) under the Partnership Agreement PT2020, through projects UID/QUI/50006/2013 and POCI/01/0145/FEDER/007265 (LAQV@REQUIMTE).
Author information
Authors and Affiliations
Contributions
R.J.F. and D.J.V.A.S. conceived the experiment(s), R.J.F. conducted the experiment(s), R.J.F., C.A.B. and D.J.V.A.S. analyzed the results. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ferreira, R.J., Bonito, C.A., Cordeiro, M.N.D.S. et al. Structure-function relationships in ABCG2: insights from molecular dynamics simulations and molecular docking studies. Sci Rep 7, 15534 (2017). https://doi.org/10.1038/s41598-017-15452-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15452-z
This article is cited by
-
New uracil analog as inhibitor/modulator of ABC transporters or/and NF-κB in taxol-resistant MCF-7/Tx cell line
Journal of Cancer Research and Clinical Oncology (2024)
-
Restriction of access to the central cavity is a major contributor to substrate selectivity in plant ABCG transporters
Cellular and Molecular Life Sciences (2023)
-
PBK/TOPK inhibitor OTS964 resistance is mediated by ABCB1-dependent transport function in cancer: in vitro and in vivo study
Molecular Cancer (2022)
-
Ins and outs of AlphaFold2 transmembrane protein structure predictions
Cellular and Molecular Life Sciences (2022)
-
The transport pathway in the ABCG2 protein and its regulation revealed by molecular dynamics simulations
Cellular and Molecular Life Sciences (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.