Abstract
The PhoPQ two-component regulatory system coordinates the response of Salmonella enterica serovar Typhimurium to diverse environmental challenges encountered during infection of hosts, including changes in Mg2+ concentrations, pH, and antimicrobial peptides. Moreover, PhoPQ-dependent regulation of gene expression promotes intracellular survival of Salmonella in macrophages, and contributes to the resistance of this pathogen to reactive nitrogen species (RNS) generated from the nitric oxide produced by the inducible nitric oxide (NO) synthase of macrophages. We report here that Salmonella strains with mutations of phoPQ are hypersensitive to killing by RNS generated in vitro. The increased susceptibility of ∆phoQ Salmonella to RNS requires molecular O2 and coincides with the nitrotyrosine formation, the oxidation of [4Fe-4S] clusters of dehydratases, and DNA damage. Mutations of respiratory NADH dehydrogenases prevent nitrotyrosine formation and abrogate the cytotoxicity of RNS against ∆phoQ Salmonella, presumably by limiting the formation of peroxynitrite (ONOO−) arising from the diffusion-limited reaction of exogenous NO and endogenous superoxide (O2 •−) produced in the electron transport chain. The mechanism underlying PhoPQ-mediated resistance to RNS is linked to the coordination of Mg2+ homeostasis through the PhoPQ-regulated MgtA transporter. Collectively, our investigations are consistent with a model in which PhoPQ-dependent Mg2+ homeostasis protects Salmonella against nitrooxidative stress.
Similar content being viewed by others
Introduction
Mutations in phoPQ attenuate Salmonella virulence by at least 10,000-fold1,2,3. The attenuated phenotype of phoPQ mutants has been associated with poor intracellular survival in macrophages, defective activation of Salmonella pathogenicity island 2 (SPI2) transcription, and hypersensitivity to defensins, antimicrobial peptides, divalent cations, iron, acid and bile salts1,4,5,6,7,8,9,10. PhoPQ signaling also boosts antioxidant defenses through the positive regulation of the sodCI-encoded superoxide dismutase, the posttranslational stabilization of the alternative σS factor, and the limitation in the availability of free iron7,11,12. In addition, PhoPQ lessens the cytotoxicity of reactive nitrogen species (RNS) generated by inducible nitric oxide synthase (iNOS) in the innate response of mononuclear phagocytic cells13.
The antimicrobial activity of NO is best demonstrated in IFNγ-activated phagocytes; however, very little anti-Salmonella activity is derived from iNOS expressed through the innate recognition of Salmonella lipopolysaccharide by host-cell Toll-like receptor 414,15,16,17,18,19. There are several possible explanations underlying the marked resistance of Salmonella to the nitrosative species synthesized by iNOS during the innate response of professional phagocytes. The low NO fluxes generated in the innate response dramatically limit the synthesis of autooxidative products such as dinitrogen trioxide (N2O3), which has been associated with sustained anti-Salmonella activity of IFNγ-primed macrophages17. On the other hand, the SPI2 type III secretion system, the Hmp flavohemoprotein, and low-molecular weight thiols protect Salmonella against moderate NO rates generated in the innate immune response20,21,22. As just mentioned, we have recently shown that PhoPQ signaling enhances the intracellular fitness of Salmonella by antagonizing the innate host response associated with NO13. The mechanism by which the PhoPQ two-component regulatory system defends Salmonella against the antimicrobial actions of NO congeners remains unknown. The investigations presented herein have revealed that the PhoPQ two-component regulatory system enhances the resistance of Salmonella against the nitrooxidative stress generated in the interaction of exogenous NO with endogenously produced O2 •− through its regulation of intracellular Mg2+ concentrations.
Results
PhoPQ-deficient Salmonella are hypersusceptible to NO
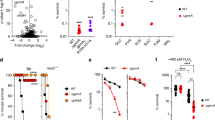
The PhoPQ signaling cascade coordinates important aspects of the antioxidant and antinitrosative defenses of Salmonella 12,13. The PhoPQ two component regulatory system is involved in Salmonella defense against Fenton-mediated oxidative stress7, however, it is unclear how PhoPQ signaling promotes resistance to RNS. To learn more about the role of PhoPQ in resistance of Salmonella to RNS, we investigated the survival of a ∆phoQ mutant exposed to the NO generator spermine NONOate (sperNO). Most wild-type Salmonella survived 6 h after challenge with 250 µM sperNO, while ∼99% of ∆phoQ Salmonella were killed upon sperNO treatment (Fig. 1A). The NO-mediated killing of ∆phoQ Salmonella was already noted after 4 h of challenge. The susceptibility of ∆phoQ Salmonella to sperNO appears to rely on the generation of NO, because the polyamine spermine control lacked antimicrobial activity (Fig. 1B). A dose-dependent inhibition of growth by sperNO was observed for both ∆phoP or ∆phoQ strains when inoculated in LB broth or minimal E salts medium supplemented with malic acid (Fig. 1C and Fig. S1), which is consistent with the notion that the PhoP response regulator boosts the antinitrosative potential of Salmonella in conjunction with its cognate PhoQ sensor kinase. The growth of ∆phoQ Salmonella in E salts was completely inhibited by 250 µM sperNO, which corresponded to the dose of sperNO that resulted in significant lethality when cells were challenged in PBS (Fig. 1 and Fig. S1). The hypersensitivity of ∆phoQ Salmonella to sperNO does not appear to be due defects in viability, as wild-type and ∆phoQ Salmonella strains grew with similar kinetics in LB and various minimal E salts media in the absence of sperNO (Fig. S2). Complementation of ∆phoQ Salmonella with a plasmid encoding a wild-type allele of phoQ (pPhoQ) restored wild-type levels of growth following sperNO treatment (Fig. 1C). Collectively, these data indicate that the PhoPQ two-component regulatory system contributes to the protection of Salmonella against the cytotoxic activity associated with RNS.
The PhoPQ two-component regulatory system protects Salmonella against RNS cytotoxicity. The susceptibility of wild-type (WT) and ∆phoQ Salmonella to killing by 250 µM spermine NONOate (sperNO) in PBS at 37 °C was compared after 0, 2, 4 and 6 h (A). The bactericidal capacity of 250 µM sperNO was compared to the polyamine base spermine following 6 h of incubation at 37 °C (B). Results represent the mean % survival ± SD of 4 independent observations collected from two separate experiments. *P < 0.01 compared to untreated controls. To determine effects of RNS on the growth of Salmonella, strains were cultured in LB broth in the presence or absence of 2.5 mM sperNO at 37 °C. Growth was determined by measuring the optical density (OD600nm) over time in 96-well microtiter plates using a Biotek Cytation 5 multi-mode plate reader (C). Data represent the mean optical density of 3 biological replicates.
Oxygen is required for the NO-dependent killing of phoQ-deficient Salmonella
RNS including nitrogen dioxide (NO2 •), N2O3 and ONOO− produced in the reaction of NO with O2 and O2 •− indirectly mediate NO cytotoxicity23. To determine whether killing of ∆phoQ Salmonella by sperNO is mediated by NO itself or by a variety of RNS, Salmonella were exposed to 250 µM sperNO in the presence or absence of O2. To generate a hypoxic environment, PBS was flushed with N2 for 10 min and the experiments were carried out in sealed tubes. The viability of wild-type Salmonella was not (P > 0.05) affected by sperNO in either normoxic or hypoxic conditions (Fig. 2A). In contrast, the NO-dependent killing of ∆phoQ Salmonella was completely abrogated in hypoxic cultures (Fig. 2B). These findings suggest that the PhoPQ two-component regulatory system protects Salmonella against nitrooxidative products formed in the reaction of NO and O2 metabolites.
Oxygen is required for the NO-dependent killing of phoQ-deficient Salmonella. The survival of wild-type (WT) and ∆phoQ Salmonella strains after incubation at 37 °C for 6 h in the presence or absence of 250 µM sperNO was compared under normoxic (+O2) and low oxygen (−O2) conditions (A,B). The contribution of the PmrA response regulator, the CorA metal transporter, the copper-zinc superoxide dismutase (SodCI) and the alternative sigma factor RpoS to the resistance of Salmonella to 250 µM sperNO is shown in (C,D and E), respectively. *P < 0.05 compared to WT. **P < 0.05 compared to the ∆phoQ strain.
The antioxidant defenses associated with PhoPQ signaling rely on the expression of a functional PmrAB two-component regulatory system and the CorA metal transporter7,24. The susceptibility of ∆phoQ Salmonella to NO appears, however, to be independent of pmrA (Fig. 2C), which is consistent with previous investigations that reported that a pmrA mutant is as resistant to sperNO as wild-type controls24. The hypersusceptibility of phoP mutants to Fe2+-mediated oxidative stress can be prevented by a mutation in the corA metal transporter7. However, a mutation in corA did not prevent the killing of ∆phoQ Salmonella by sperNO (Fig. 2C). In addition to contributing to iron homeostasis, PhoPQ can activate Salmonella’s antioxidant defenses through the positive regulation of SodCI expression and the stabilization of RpoS11,12. However, neither SodCI or RpoS appear to contribute to the increased susceptibility of ∆phoQ Salmonella under the experimental conditions tested here (Fig. 2D and E). Interestingly, a strain of Salmonella lacking both phoQ and rpoS was even more susceptible to the RNS-dependent cytotoxicity than the phoQ mutant, suggesting that in the absence of PhoPQ the alternative sigma factor RpoS assumes a critical role in the regulation of the antinitrosative defenses of Salmonella.
Salmonella exposed to NO undergoes nitrooxidative stress
To determine whether wild-type and ∆phoQ Salmonella experience different degrees of nitrooxidative stress upon exposure to sperNO, we monitored the formation of N2O3 (a reactive species generated upon autooxidation of NO in the presence of O2) and nitrotyrosine (an oxidative signature of the reaction of ONOO− or other RNS with tyrosyl residues). Similar concentrations of N2O3 were generated after treatment of wild-type or ∆phoQ Salmonella with 250 µM sperNO (Fig. 3A). Substantial nitrotyrosine formation was also detected within 30 min after Salmonella were challenged with sperNO (Fig. 3B). Moreover, the profiles and kinetics of nitrotyrosine formation were similar in both wild-type and ∆phoQ Salmonella strains (Fig. 3B). Nitrotyrosine formation was not observed in low O2 cultures (Fig. 3C), suggesting that ONOO− arising from the diffusion-limited reaction of exogenous NO with endogenous O2 •− is a likely candidate for the covalent oxidation of tyrosine residues in our experiments. In addition to tyrosine residues, [4Fe-4S] clusters of dehydratases are among the most avid targets (k = 1.4 × 105 M−1 s−1) of ONOO− 25. We therefore monitored the enzymatic activity of the [4Fe-4S] cluster-containing aconitase as a surrogate marker of ONOO− mediated oxidative stress. Wild-type and ∆phoQ Salmonella harbored comparable basal levels of aconitase activity (Fig. 3D). Moreover, the aconitase activity of both wild-type and ∆phoQ Salmonella was similarly inhibited 6 h after exposure to sperNO (Fig. 3D). Together, these findings indicate that 250 µM sperNO exert substantial nitrooxidative stress on Salmonella. However, wild-type and ∆phoQ Salmonella appear to be exposed to similar levels of nitrooxidative species under the experimental conditions tested here. This idea is further substantiated by the degree of NO-dependent genotoxicity seen in these two strains of Salmonella. DNA damage was indirectly measured by following the expression of a transcriptional lacZY fusion to the SOS response recA gene. The recA::lacZY transcriptional fusion was similarly induced in both wild-type and ∆phoQ Salmonella after exposure to 12 J/m2 UV light or treatment with 2.5 mM sperNO (Fig. 3E).
Generation of RNS in NO-treated Salmonella. N2O3 was quantified by following the generation of naphthyltriazol (NAT) in 5 × 105 CFU ml−1 of wild-type (WT) and ∆phoQ Salmonella strains cultured in PBS at 37 °C in the presence or absence of 250 µM sperNO (A). The results are expressed as the mean arbitrary units (A.U.) ± S.D. representing 3 biological replicates from 2 independent experiments. Nitrotyrosine formation in whole-cell lysates isolated from ~ 2 × 108 CFU ml−1 of WT and ∆phoQ Salmonella strains exposed to 250 µM sperNO in PBS at 37 °C was measured by Western blotting (B). A cropped image of a western blot showing the effect of O2 on the generation of nitrotyrosine in WT Salmonella exposed to 250 µM sperNO for 6 h is shown in (C). The NO-dependent damage of [Fe-S] cluster-containing dehydratases was monitored by following aconitase activity from cell lysates harvested from WT and phoQ-deficient Salmonella strains after exposure to 250 µM sperNO for 6 h at 37 °C in PBS (D). The expression of a recA::lacZY transcriptional fusion was quantified using β-galactosidase activity assays in wild-type (WT) and ∆phoQ Salmonella strains following exposure to 12 J/m2 ultraviolet light (UV) for 30 s or to 2.5 mM sperNO (E). Data in D & E represent the mean ± S.D. of 3 biological replicates. Statistical analysis was performed using a two-way ANOVA for data. *P < 0.001 compared to untreated controls; ns, no statistically significant difference compared to untreated controls.
Mutations in NADH dehydrogenases NDH-I and NDH-II protect ∆phoQ Salmonella from NO-dependent cytotoxicity and protein nitration
We examined in more detail the mechanism underlying the cytotoxicity of RNS against ∆phoQ Salmonella. NADH dehydrogenases of the electron transport chain can be a sizable source of oxidative stress in the cell26. We tested whether the sperNO-mediated, O2-dependent killing of ∆phoQ Salmonella was the result of the synergism between exogenous NO and O2 •− arising from the adventitious reduction of O2 by NADH dehydrogenases of the electron transport chain. To test this hypothesis, the ∆phoQ::km mutant allele was introduced into the ∆nuo ∆ndh mutant strain AV0438 lacking both NDH-I and NDH-II NADH dehydrogenases. As noted for H2O2 and ONOO− 27, the complex I-deficient ∆nuo ∆ndh strain AV0438 was resistant to 250 µM sperNO (Fig. 4A). Strikingly, strain AV0810 harboring mutations in phoQ, nuo and ndh was also resistant to NO (Fig. 4A). Similar to the complex I-deficient isogenic strain AV0438, strain AV0810 lacking phoQ, nuo and ndh appear to be protected from ONOO− as indicated by a lack of nitrotyrosine formation 6 h after exposure to 250 µM sperNO (Fig. 4B). Collectively, these data indicate that ONOO− dependent nitrooxidative stress engendered upon reaction of exogenous NO with O2 •− produced by NADH dehydrogenases of the electron transport chain contributes to the NO-mediated killing of ∆phoQ Salmonella.
Defects in NADH dehydrogenases protect phoQ-deficient Salmonella from NO-dependent cytotoxicity. The survival of WT, ΔphoQ, Δndh Δnuo, and Δndh Δnuo ΔphoQ::km strains after incubation with 250 µM sperNO in PBS at 37 °C for 6 h (A). The data in A are represented by the mean % survival ± S.D. of 6 independent observations from 2 separate experiments. Statistical analysis was performed using a two-way ANOVA for data. *P < 0.001 compared to untreated controls. The presence of nitrotyrosine in whole-cell lysates of the indicated Salmonella strains incubated in the presence or absence of 250 µM sperNO in PBS at 37 °C for 6 h was determined by Western blot (B). The data presented in B are presented as cropped panels for each individual strain tested, and are representative of two independent experiments.
Exogenous Mg2+ rescues ∆phoQ Salmonella from RNS-dependent killing
Previously, Salmonella was shown to exhibit increased susceptibility to oxidative stress following disruptions in Mg2+ uptake through mutations of phoP or the PhoPQ-regulated Mg2+ transporters mgtA and mgtB 7. Therefore, we investigated whether the increased susceptibility of ∆phoQ Salmonella to RNS was due to disruptions in Mg2+ homeostasis. The increased susceptibility of the ∆phoQ Salmonella strain to killing by 250 µM sperNO in PBS was prevented by the addition of 10 mM MgSO4, but had no effect on the survival of the wild-type strain or the ∆phoQ strain complemented with a pPhoQ plasmid (Fig. 5A). Moreover, the sperNO-dependent inhibition of growth of ∆phoQ Salmonella was alleviated by the addition of 10 mM MgSO4 when cultured in LB with 2.5 mM sperNO (Fig. 5B). This protective effect of exogenous MgSO4 was also observed in ∆phoQ Salmonella challenged with 250 µM sperNO in minimal E salts media supplemented with glucose, malic acid, or fumarate (Fig. S2). Moreover, the addition of MgCl2 also restored the growth of ∆phoQ Salmonella challenged with sperNO, while the addition of CaCl2 had no effect (Fig. S3). Collectively, these data suggest that the hypersensitivity of ∆phoQ Salmonella to RNS is due to disruptions in Mg2+ homeostasis.
PhoPQ-dependent regulation of Mg2+ homeostasis protects Salmonella from RNS cytotoxicity. The survival of WT, ΔphoQ, and ΔphoQ pWSK29::phoQ (pPhoQ) Salmonella strains was determined after incubation of strains with 250 µM sperNO in the presence or absence of 10 mM MgSO4 in PBS at 37 °C for 6 h (A). The data, which are represented as mean ± S.D., are from 8 replicates collected from 2 separate experiments. *P < 0.01 compared to sperNO-treated WT controls. The OD600nm was measured over time as described in Fig. 1 to determine the growth of Salmonella strains cultured in LB + 2.5 mM sperNO, or LB + 2.5 mM sperNO +10 mM MgSO4 at 37 °C. The survival of WT, ΔmgtA::km, ΔmgtBC, and ΔmgtBC ΔmgtA::km Salmonella strains was determined following incubation with 250 µM sperNO in the presence or absence of 10 mM MgSO4 in PBS at 37 °C for 6 h (C). The data are represented as mean % survival ± SD from 8 replicates collected from 2 separate experiments. *P < 0.01 compared to sperNO-treated WT controls. The OD600nm was measured over time as described in Fig. 1 to determine the growth of Salmonella strains cultured in LB + 2.5 mM sperNO or LB + 2.5 mM sperNO + 10 mM MgSO4 at 37 °C (D). Data represent the mean OD600nm of 3 biological replicates.
We hypothesized that the increased susceptibility of the ∆phoQ Salmonella strain to RNS was linked to its inability to upregulate the expression of Mg2+ transporters encoded by mgtA and mgtB. Therefore, we compared the susceptibility of Salmonella strains lacking mgtA and/or mgtBC to killing by 250 µM sperNO. The ∆mgtA-deficient Salmonella strain showed a significantly increased susceptibility to killing by sperNO compared to both wild-type and ∆mgtBC strains (Fig. 5C). A strain harboring mutations in both mgtA and mgtBC was no more susceptible to killing by RNS as the ∆mgtA mutant strain suggesting that the PhoPQ-dependent regulation of mgtA promotes resistance to RNS. As was the case for the ∆phoQ Salmonella strain (Fig. 5A), the addition of 10 mM MgSO4 prevented sperNO-dependent killing of the ∆mgtA Salmonella (Fig. 5D). These data suggested that the increased susceptibility of ∆phoQ Salmonella to RNS was due to its inability to upregulate the expression of mgtA, which is required for proper Mg2+ homeostasis. To test this hypothesis, we introduced a pBAD/HisA plasmid encoding mgtA (pBmgtA) under an arabinose-inducible promoter into the ∆phoQ Salmonella strain, and compared its ability to grow in LB broth in the presence or absence of sperNO. The wild-type, ∆phoQ, and ∆phoQ pMgtA strains showed similar growth kinetics in LB broth (Fig. 5D). As described earlier, ΔphoQ Salmonella were unable to grow in LB broth in the presence of 2.5 mM sperNO (Figs 1C and 5D). In contrast, the introduction of a pMgtA plasmid to the ∆phoQ strain restored growth in LB broth in the presence of 2.5 mM sperNO, albeit with an increased lag compared to the wild-type control (Fig. 5D). This lag was eliminated by the addition of 10 mM MgSO4, suggesting that the expression of mgtA from the pMgtA plasmid could only partially restore Mg2+ homeostasis in the ∆phoQ strain. Collectively, these data support the hypothesis that PhoPQ promotes resistance to nitrooxidative stress in Salmonella through the regulation of Mg2+ homeostasis.
Discussion
The PhoP regulon controls the antioxidant defenses of Salmonella, Yersinia pestis and Enterococcus faecalis 12,28,29, and work from our laboratory indicates that this two-component regulatory system also is contributes to the antinitrosative defenses of Salmonella 13. Elegant investigations by Dr. Groisman’s group have elucidated that the PhoP regulon defends Salmonella against oxidative stress engendered in the reduction of H2O2 by the Fenton catalyst Fe2+7. Little is known, however, about the nitrosative chemistry antagonized by this two-component regulatory system. We therefore deemed it important to investigate the newly described function of PhoPQ in the antinitrosative defenses of Salmonella. The investigations presented here are consistent with a model in which the PhoPQ two-component regulatory system antagonizes the antimicrobial activity of ONOO−. In support of this model, the sperNO-mediated killing of ∆phoQ Salmonella is restricted to aerobic cultures, and coincides with the formation of nitrotyrosine, and the inactivation of the TCA cycle enzyme aconitase. These observations can be explained if one takes into account that the generation of ONOO− requires the reaction of O2 •− and NO. O2 •− is formed adventitiously at the flavin or quinone-binding sites of NADH dehydrogenases of the electron transport chain and its production requires O2. The ONOO− produced in the reaction of endogenous O2 •− and exogenous NO is a powerful nitrating and oxidizing agent that could explain the formation of nitrotyrosine residues in cytoplasmic proteins and the oxidation of the [4Fe-4S] clusters of dehydratases. The protection afforded by mutations in NADH dehydrogenases against ONOO– dependent cytotoxicity could be explained by three independent and complementary mechanisms. First, the accumulation of NADH in ∆ndh ∆nuo Salmonella effectively scavenges NO2 • and OH• radicals caged in peroxynitrous acid (ONO-OH), which is the dominant ONOO− congener at the neutral pH of the bacterial cytoplasm27. Second, NADH fuels the enzymatic detoxification of ONOO− by the AhpCF alkylhydroperoxidase27,30. And third, a lack of NADH dehydrogenases diminishes ONOO− synthesis by limiting the flow of electrons through the respiratory chain that is required for the generation of O2 •−. Our investigations suggest that ONOO− is necessary but not sufficient for the NO-mediated antimicrobial activity, because wild-type Salmonella and the phoQ mutant bacteria suffer a similar degree of nitrotyrosine formation and inactivation of aconitase upon exposure to sperNO, but are differently killed by the oxidative congeners of this diatomic radical.
[4Fe-4S] clusters of dehydratases can be directly nitrosylated by NO at a rate constant of 106 M−2 sec−1 31. Because the nitrosylation of [4Fe-4S] clusters is second order for NO, this chemistry is less likely to occur at the low NO fluxes sustained in the course of our investigations. The inactivation of [4Fe-4S] clusters by ONOO−, which occurs at the fast rate of 1.4 × 105 M−1 sec−1 25, is first order for ONOO−. The speed of the reaction indicates that the oxidation of [4Fe-4S] clusters by ONOO− is limited by the production of this RNS. NO and O2 •− react with a second order rate constant of 109 M−1 sec−1 to form ONOO− 32. However, high concentrations of NO readily consume ONOO−. Therefore, generation of ONOO− in the Salmonella cytoplasm is most likely to be maximal at the low rates of NO synthesis supported during the innate immune response. In the presence of a functional PhoPQ two-component regulatory system the ONOO− produced endogenously appears to be tolerated by Salmonella.
The Hmp-mediated and cytochrome bd-mediated detoxification of NO, the stringent response, low-molecular weight thiols, along with DNA repair systems minimize the cytotoxicity of NO produced in the innate response to Salmonella 23,33,34,35. Our investigations identify PhoPQ-dependent regulation of Mg2+ homeostasis as an additional antinitrosative defense that shields Salmonella from the cytotoxicity of low NO fluxes. While the hypersensitivity of ∆phoQ Salmonella to RNS is tied to disrupted Mg2+ homeostasis, this phenotype appears to be independent of both PmrAB-dependent and CorA-dependent resistance Fe2+ toxicity. This conclusion is supported by the fact that 1) pmrA or corA single mutant strains do not exhibit increased sensitivity to RNS compared to wild-type strains, and 2) phoQ pmrA or phoQ corA double mutant Salmonella strains were as sensitive to killing by sperNO as ∆phoQ Salmonella. Cellular concentrations of Mg2+ in Salmonella are capable of reaching 100 mM36, with the majority of Mg2+ bound to ribosomes and nucleotide triphosphates37. Therefore, the reduced cytoplasmic Mg2+ concentrations in ∆phoQ Salmonella may increase the susceptibility to killing by RNS due to several factors including 1) reduced protein synthesis resulting from the dissociation of Mg2+ from ribosomes, and 2) leaching of Mg2+ from nucleotide triphosphates preventing their use as substrates for enzymatic reactions necessary to repair RNS-induced cellular damage. This is supported by the fact that the addition of 10 mM MgSO4 rescued both ∆phoQ and ∆mgtA Salmonella strains from RNS-dependent killing.
Independent of the regulation of antioxidant defenses or targets of nitrooxidative stress, a functional PhoPQ two-component regulatory system is likely to promote antinitrosative defenses through the activation of SPI2 transcription38,39,40, because the SPI2 type III secretion system has been shown to minimize fusion of Salmonella-containing vacuoles with vesicles harboring iNOS41. In contrast to the concerted and rich repertoire of antinitrosative defenses that protect Salmonella against the low NO fluxes produced in the innate response, no antinitrosative defenses are known to protect this facultative intracellular pathogen against the massive nitrosative stress unleashed in IFNγ-activated macrophages. Paradoxically, N2O3 and other high oxidation NO congeners generated by IFNγ-primed macrophages exert profound anti-Salmonella activity by repressing SPI2 transcription and PhoPQ signaling13,16,17. In turn, RNS-dependent repression of PhoPQ signaling and SPI2 transcription promotes the maturation of the Salmonella phagosome along the degradative pathway for fusion with lysosomes.
In summary, this study has revealed that in addition to its known roles in protecting Salmonella from acid pH, bile salts, antimicrobial peptides, and oxidative stress1,4,5,6,7,8,9,10, the PhoPQ two-component regulatory system contributes to the resistance of Salmonella against the nitrooxidative stress generated in the reaction of exogenous NO and endogenously produced O2 •− by maintaining Mg2+ homeostasis (Fig. 6).
Model for PhoPQ-mediated resistance to nitrooxidative stress. Our investigations herein and elsewhere indicate that the PhoPQ signaling cascade protects Salmonella against the antimicrobial activity of nitric oxide (NO) produced by the inducible NO synthase (iNOS). NADH dehydrogenases (NDH) at the inner membrane (IM) transfer electrons (e−) from NADH to the quinone pool in the electron transport chain. Although this process is remarkably efficient, some electrons adventitiously reduce molecular oxygen (O2) to generate the superoxide radical (O2 •−). The diffusion limited reaction of O2 •− and exogenous NO. gives rise to peroxynitrite (ONOO−) generating nitrooxidative stress. The PhoPQ two-component regulatory system protects Salmonella against nitrooxidative stress by maintaining optimal cytoplasmic concentrations of Mg2+ through the regulation of the Mg2+ transporter MgtA.
Methods
Bacterial Strains
Salmonella enterica serovar Typhimurium strain ATCC 14028 s was used throughout this study as wild-type, and as a background for the construction of mutations and a recA::lacZY transcriptional fusion (Table 1). The mutations were generated following the one-step, λ-Red-mediated gene replacement method of Datsenko and Wanner42. Briefly, primers encoding 40–42 nucleotides homologous to the target gene followed by 20 nucleotides homologous to the pKD13 template plasmid were used for the PCR amplification of the Flp recombinant target (FRT)-flanked kanamycin resistance cassette. The resulting PCR products were DpnI digested and electroporated into S. Typhimurium strain TT22236 carrying the pTP2223 plasmid that expresses the λ-Red recombinase under Ptac control. Mutations were moved between strains by P22-mediated transduction and pseudolysogens eliminated by streaking on Evans blue uranine agar plates. Nonpolar deletions were generated by recombining the two FRT sites flanking the kanamycin resistance cassette with the Flp recombinase encoded by the pCP20 plasmid43. The mutations were confirmed by PCR analysis. A recA::lacZY transcriptional fusion was constructed by the pCP20-mediated integration of pCE36 encoding a promoterless lacZY operon into the unique FRT scar engineered immediately downstream of the recA stop codon. The pMgtA plasmid was generated by cloning a wild-type copy of the magnesium transporter mgtA into the pBAD/HisA vector using the primers listed in Table 2. The pMgtA plasmid was electroporated into Salmonella strain AV0475 carrying a ∆phoQ::FRT allele to produce the TJB1301 strain.
Susceptibility of Salmonella to reactive nitrogen species
Salmonella strains were inoculated from frozen stocks and grown in Luria-Bertani (LB) broth with shaking at 325 r.p.m. at 37 °C for 20 h. Strains were then diluted in PBS to a concentration of ~5 × 105 cells ml−1. The bacteria were challenged at 37 °C with 250 μM of the NO donor spermine NONOate (Cayman Chemical, Ann Arbor, MI). Selected groups of bacteria were challenged at 37 °C with 250 μM spermine NONOate in PBS that had been depleted of O2 after 10 min of flushing with N2. Percent survival was calculated by recording the number of bacteria capable of forming a CFU on LB agar plates. Alternatively, stationary phase cultures of the Salmonella strains were subcultured 1:200 in fresh LB broth in the presence or absence of 2.5 mM spermine NONOate. Bacterial suspensions were seeded in 96-well plates, and were grown at 37 °C with shaking at 282 r.p.m. with the optical density measured at 600 nm (OD600nm) using a Cytation 5 multi-mode plate reader (BioTek, Winooski, VT).
Aconitase enzymatic assay
Salmonella strains grown in Luria-Bertani (LB) broth with shaking at 325 r.p.m. at 37 °C for 20 h were pelleted by centrifugation and resuspended in PBS to an OD600nm of 0.5. Soluble cytoplasmic proteins were isolated from bacteria incubated at 37 °C for 6 h in PBS in the presence or absence of 250 μM spermine NONOate. Briefly, the bacteria were washed in 20 mM Tris-citrate buffer pH 8.0, and the cytoplasmic proteins extracted by sonication. Bacterial debris was removed by centrifugation at 14,000 RPM in a microcentrifuge for 30 s. Aconitase activity contained in the cytoplasmic extracts was estimated spectrophotometrically at 240 nm by following the formation of cis-aconitate in Tris-citrate buffer containing 20 mM isocitrate44. The protein concentration in the cytoplasmic extracts was measured by the BCA protein assay (Pierce, Rockford, IL). Aconitase activity is expressed as the mean OD240nm/min/mg protein ± SD of 2 independent experiments.
Estimation of DNA damage
The accumulation of single strand and double strand DNA damage was indirectly estimated by measuring the expression of a lacZY transcriptional fusion of the SOS response recA gene. Selected groups of bacteria were irradiated with 12 J/m2 UV using a TL-2000 Ultraviolet Translinker (Ultraviolet Products, Upland, CA), or treated with 2.5 mM spermine NONOate at 37 °C for 6 h. Expression of the recA::lacZY transcriptional fusion was quantified spectrophotometrically as β-galactosidase enzymatic activity using the substrate o-nitrophenyl-β-D-galactopyranoside. β-galactosidase activity is expressed as Miller units using the equation 1,000 × [(OD420nm − 1.75 × OD550nm)/(T(min) × V(ml) × OD600nm)].
N2O3 quantification
The generation of N2O3 in Salmonella strains exposed to 250 μM spermine NONOate was determined indirectly by following the formation of the N-nitrosonapthalen derivative of 2,3-diaminonaphthalen (Sigma-Aldrich) as described45. A 100 mM stock of 2,3-diaminonaphthalen prepared in dimethylformamide was used at a final concentration of 200 μM in PBS. Accumulation of N-nitrosonapthalen was recorded for 30 min following treatment of Salmonella strains with spermine NONOate. Fluorescence was measured on a Synergy HT fluorometer (BioTek) set at λex = 375 nm and λem = 460 nm.
Detection of nitrotyrosine formation by Western blot analysis
Salmonella strains grown in Luria-Bertani (LB) broth with shaking at 325 r.p.m. at 37 °C for 20 h were pelleted by centrifugation, and resuspended in PBS to an OD600 of 0.5. Bacteria were incubated at 37 °C in the presence or absence of 250 μM spermine NONOate. At the specified timepoints, the bacteria were pelleted by centrifugation and resuspended in 200 μL of alkaline lysis buffer (25 mM Tris, 100 mM SDS, and 128 mM NaOH). The specimens were separated in 10% SDS-PAGE gels, transferred to nitrocellulose membranes, and probed with an anti-nitrotyrosine polyclonal antibody (Upstate, Lake Placid, NY) followed by a horseradish peroxidase-conjugated, anti-rabbit IgG secondary antibody. Detection was carried out using the Enhanced Chemiluminescence Kit (GE Healthcare, Piscataway, NJ) on a Molecular Imager Fx (BioRad, Hercules, CA).
Statistical analysis
Data are presented as mean ± standard deviation (SD). To determine statistical significance between multiple comparisons, two-way analysis of variance (ANOVA) were performed, followed by a Bonferroni posttest. Data were considered statistically significant when P was < 0.05.
References
Fields, P. I., Swanson, R. V., Haidaris, C. G. & Heffron, F. Mutants of Salmonella-Typhimurium That Cannot Survive within the Macrophage Are Avirulent. Proceedings of the National Academy of Sciences of the United States of America 83, 5189–5193, https://doi.org/10.1073/pnas.83.14.5189 (1986).
Galan, J. E. & Curtiss, R. 3rd Virulence and vaccine potential of phoP mutants of Salmonella typhimurium. Microb Pathog 6, 433–443 (1989).
Miller, S. I., Kukral, A. M. & Mekalanos, J. J. A two-component regulatory system (phoP phoQ) controls Salmonella typhimurium virulence. Proc Natl Acad Sci USA 86, 5054–5058 (1989).
Bader, M. W. et al. Recognition of antimicrobial peptides by a bacterial sensor kinase. Cell 122, 461–472, https://doi.org/10.1016/j.cell.2005.05.030 (2005).
Bearson, B. L., Wilson, L. & Foster, J. W. A low pH-inducible, PhoPQ-dependent acid tolerance response protects Salmonella typhimurium against inorganic acid stress. J Bacteriol 180, 2409–2417 (1998).
Bijlsma, J. J. & Groisman, E. A. The PhoP/PhoQ system controls the intramacrophage type three secretion system of Salmonella enterica. Mol Microbiol 57, 85–96, https://doi.org/10.1111/j.1365-2958.2005.04668.x (2005).
Chamnongpol, S. & Groisman, E. A. Mg2+ homeostasis and avoidance of metal toxicity. Mol Microbiol 44, 561–571, doi:2917 (2002).
Garcia Vescovi, E., Soncini, F. C. & Groisman, E. A. Mg2+ as an extracellular signal: environmental regulation of Salmonella virulence. Cell 84, 165–174, doi:S0092-8674(00)81003-X [pii] (1996).
Groisman, E. A., Parra-Lopez, C., Salcedo, M., Lipps, C. J. & Heffron, F. Resistance to host antimicrobial peptides is necessary for Salmonella virulence. Proc Natl Acad Sci USA 89, 11939–11943 (1992).
van Velkinburgh, J. C. & Gunn, J. S. PhoP-PhoQ-regulated loci are required for enhanced bile resistance in Salmonella spp. Infect Immun 67, 1614–1622 (1999).
Golubeva, Y. A. & Slauch, J. M. Salmonella enterica serovar Typhimurium periplasmic superoxide dismutase SodCI is a member of the PhoPQ regulon and is induced in macrophages. J Bacteriol 188, 7853–7861, https://doi.org/10.1128/JB.00706-06 (2006).
Tu, X., Latifi, T., Bougdour, A., Gottesman, S. & Groisman, E. A. The PhoP/PhoQ two-component system stabilizes the alternative sigma factor RpoS in Salmonella enterica. Proc Natl Acad Sci USA 103, 13503–13508, https://doi.org/10.1073/pnas.0606026103 (2006).
Bourret, T. J., Song, M. & Vazquez-Torres, A. Codependent and independent effects of nitric oxide-mediated suppression of PhoPQ and Salmonella pathogenicity island 2 on intracellular Salmonella enterica serovar typhimurium survival. Infect Immun 77, 5107–5115, https://doi.org/10.1128/IAI.00759-09 (2009).
Vazquez-Torres, A., Jones-Carson, J., Mastroeni, P., Ischiropoulos, H. & Fang, F. C. Antimicrobial actions of the NADPH phagocyte oxidase and inducible nitric oxide synthase in experimental salmonellosis. I. Effects on microbial killing by activated peritoneal macrophages in vitro. J Exp Med 192, 227–236 (2000).
Vazquez-Torres, A. et al. Toll-like receptor 4 dependence of innate and adaptive immunity to Salmonella: importance of the Kupffer cell network. J Immunol 172, 6202–6208 (2004).
McCollister, B. D., Bourret, T. J., Gill, R., Jones-Carson, J. & Vazquez-Torres, A. Repression of SPI2 transcription by nitric oxide-producing, IFNgamma-activated macrophages promotes maturation of Salmonella phagosomes. J Exp Med 202, 625–635, https://doi.org/10.1084/jem.20050246 (2005).
McCollister, B. D. et al. N(2)O(3) enhances the nitrosative potential of IFNgamma-primed macrophages in response to Salmonella. Immunobiology 212, 759–769, https://doi.org/10.1016/j.imbio.2007.09.019 (2007).
Webb, J. L., Harvey, M. W., Holden, D. W. & Evans, T. J. Macrophage nitric oxide synthase associates with cortical actin but is not recruited to phagosomes. Infect Immun 69, 6391–6400, https://doi.org/10.1128/IAI.69.10.6391-6400.2001 (2001).
Shiloh, M. U. et al. Phenotype of mice and macrophages deficient in both phagocyte oxidase and inducible nitric oxide synthase. Immunity 10, 29–38 (1999).
Chakravortty, D., Hansen-Wester, I. & Hensel, M. Salmonella pathogenicity island 2 mediates protection of intracellular Salmonella from reactive nitrogen intermediates. J Exp Med 195, 1155–1166 (2002).
Bang, I. S. et al. Maintenance of nitric oxide and redox homeostasis by the salmonella flavohemoglobin hmp. J Biol Chem 281, 28039–28047, https://doi.org/10.1074/jbc.M605174200 (2006).
De Groote, M. A., Testerman, T., Xu, Y., Stauffer, G. & Fang, F. C. Homocysteine antagonism of nitric oxide-related cytostasis in Salmonella typhimurium. Science 272, 414–417 (1996).
Henard, C. A. & Vazquez-Torres, A. Nitric oxide and salmonella pathogenesis. Front Microbiol 2, 84, https://doi.org/10.3389/fmicb.2011.00084 (2011).
Wosten, M. M., Kox, L. F., Chamnongpol, S., Soncini, F. C. & Groisman, E. A. A signal transduction system that responds to extracellular iron. Cell 103, 113–125, doi:S0092-8674(00)00092-1 (2000).
Castro, L., Rodriguez, M. & Radi, R. Aconitase is readily inactivated by peroxynitrite, but not by its precursor, nitric oxide. J Biol Chem 269, 29409–29415 (1994).
Treberg, J. R., Quinlan, C. L. & Brand, M. D. Evidence for two sites of superoxide production by mitochondrial NADH-ubiquinone oxidoreductase (complex I). J Biol Chem 286, 27103–27110, https://doi.org/10.1074/jbc.M111.252502 (2011).
Husain, M. et al. Nitric oxide evokes an adaptive response to oxidative stress by arresting respiration. J Biol Chem 283, 7682–7689, https://doi.org/10.1074/jbc.M708845200 (2008).
Muller, C. et al. Characterization of two signal transduction systems involved in intracellular macrophage survival and environmental stress response in Enterococcus faecalis. J Mol Microbiol Biotechnol 14, 59–66, https://doi.org/10.1159/000106083 (2008).
Oyston, P. C. et al. The response regulator PhoP is important for survival under conditions of macrophage-induced stress and virulence in Yersinia pestis. Infect Immun 68, 3419–3425 (2000).
Bryk, R., Griffin, P. & Nathan, C. Peroxynitrite reductase activity of bacterial peroxiredoxins. Nature 407, 211–215, https://doi.org/10.1038/35025109 (2000).
Duan, X., Yang, J., Ren, B., Tan, G. & Ding, H. Reactivity of nitric oxide with the [4Fe-4S] cluster of dihydroxyacid dehydratase from Escherichia coli. Biochem J 417, 783–789, https://doi.org/10.1042/BJ20081423 (2009).
Huie, R. E. & Padmaja, S. The reaction of no with superoxide. Free Radic Res Commun 18, 195–199 (1993).
Henard, C. A. & Vazquez-Torres, A. DksA-dependent resistance of Salmonella enterica serovar Typhimurium against the antimicrobial activity of inducible nitric oxide synthase. Infect Immun 80, 1373–1380, https://doi.org/10.1128/IAI.06316-11 (2012).
Jones-Carson, J., Husain, M., Liu, L., Orlicky, D. J. & Vazquez-Torres, A. Cytochrome bd-Dependent Bioenergetics and Antinitrosative Defenses in Salmonella Pathogenesis. MBio 7, https://doi.org/10.1128/mBio.02052-16 (2016).
Song, M. et al. Low-molecular-weight thiol-dependent antioxidant and antinitrosative defences in Salmonella pathogenesis. Mol Microbiol 87, 609–622, https://doi.org/10.1111/mmi.12119 (2013).
Alatossava, T., Jutte, H., Kuhn, A. & Kellenberger, E. Manipulation of intracellular magnesium content in polymyxin B nonapeptide-sensitized Escherichia coli by ionophore A23187. J Bacteriol 162, 413–419 (1985).
Papp-Wallace, K. M. & Maguire, M. E. Magnesium Transport andMagnesium Homeostasis. EcoSal Plus 3, https://doi.org/10.1128/ecosalplus.5.4.4.2 (2008).
Worley, M. J., Ching, K. H. & Heffron, F. Salmonella SsrB activates a global regulon of horizontally acquired genes. Mol Microbiol 36, 749–761 (2000).
Deiwick, J., Nikolaus, T., Erdogan, S. & Hensel, M. Environmental regulation of Salmonella pathogenicity island 2 gene expression. Mol Microbiol 31, 1759–1773 (1999).
Kim, C. C. & Falkow, S. Delineation of upstream signaling events in the salmonella pathogenicity island 2 transcriptional activation pathway. J Bacteriol 186, 4694–4704, https://doi.org/10.1128/JB.186.14.4694-4704.2004 (2004).
Garvis, S. G., Beuzon, C. R. & Holden, D. W. A role for the PhoP/Q regulon in inhibition of fusion between lysosomes and Salmonella-containing vacuoles in macrophages. Cell Microbiol 3, 731–744 (2001).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97, 6640–6645, https://doi.org/10.1073/pnas.120163297 (2000).
Cherepanov, P. P. & Wackernagel, W. Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene 158, 9–14, doi:037811199500193A (1995).
Bernofsky, C. & Swan, M. An improved cycling assay for nicotinamide adenine dinucleotide. Anal Biochem 53, 452–458 (1973).
Espey, M. G., Miranda, K. M., Pluta, R. M. & Wink, D. A. Nitrosative capacity of macrophages is dependent on nitric-oxide synthase induction signals. J Biol Chem 275, 11341–11347 (2000).
Acknowledgements
This work was made possible by grants from the National Institute for General Medical Science (NIGMS; 5P20GM103427), a component of the National Institutes of Health (NIH) and by the National Institutes of Health (AI54959 and T32 AI52066), the Veterans Administration (1I01BX0020073), and the Burroughs Wellcome Fund.
Author information
Authors and Affiliations
Contributions
T.B. and A.V. wrote the main manuscript text. T.B., J.S., M.H. and L.L. performed the experiments. T.B. prepared the figures and tables. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bourret, T.J., Liu, L., Shaw, J.A. et al. Magnesium homeostasis protects Salmonella against nitrooxidative stress. Sci Rep 7, 15083 (2017). https://doi.org/10.1038/s41598-017-15445-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15445-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.