Abstract
Recent genome-wide association studies (GWAS) have identified a strong association signal of microRNA137 host gene (MIR137) with schizophrenia. MIR137 dysfunction results in downregulation of presynaptic target gene complexin 1 (CPLX1) and impairs synaptic plasticity in the hippocampus. In this study, we aimed to investigate whether the variants of MIR137 and CPLX1 confer susceptibility to schizophrenia in Han Chinese. This study employed 736 patients with schizophrenia patients and 751 well-matched healthy subjects for genetic analysis, and genotyped 12 SNPs within MIR137 and CPLX1. SZDB database was used to performed brain eQTL analysis. There were no significant differences of CPLX1 expression in hippocampus, prefrontal cortex or stratum between the schizophrenia patients and control subjects. No significant differences were observed in allele and genotype frequencies in studied SNPs between the case and control groups. Gene interaction analysis showed that MIR137 SNP rs1625579 did not affect schizophrenia susceptibility in interaction with the CPLX1 polymorphic variants. Our findings do not support MIR137 and CPLX1 conferring susceptibility to schizophrenia in Han Chinese.
Similar content being viewed by others
Introduction
Schizophrenia is a severe and disabling mental illness with clinical symptoms typically manifesting in a late adolescence or early adulthood onset. Although its etiology and pathophysiology remain unknown, the underlying cause of schizophrenia is suspected to a disruption of early brain development resulted from genetic predisposition and prenatal/perinatal environment factors1. A variety of genetic risks identified in schizophrenia are genes expressing proteins involved in the regulation of synaptic plasticity2.
Recent genome-wide association studies (GWAS) have identified a strong association signal of microRNA137 host gene (MIR137) with schizophrenia3,4,5. MicroRNAs (miRNAs) are small noncoding single-stranded RNAs that function as post-transcriptional regulators of gene expression6. In the central nervous system, miRNAs may play an important role in neurodevelopment and maturation including synaptic development, dendritic protein synthesis and neural plasticity7. MicroRNA137 is a brain-enriched miRNA in human with high expression in cortical brain regions and hippocampus, and has a critical regulatory role in brain function8,9,10. At the molecular level, a single nucleotide polymorphism (SNP) rs1625579 in MIR137 has been reported to confer susceptibility to schizophrenia in populations of European ancestry. However, the association of rs1625579 with schizophrenia is inconsistent in Asian populations11,12,13,14,15,16,17. Pu and Xiao18 thereby performed a meta-analysis and provided unsupportive evidence for the association of rs1625579 with schizophrenia in Asians.
A recent study has pointed out that MIR137 dysfunction results in downregulation of presynaptic target gene complexin-1 (CPLX1) and impairs synaptic plasticity in the hippocampus in vitro and in vivo 19. Complexin has a regulatory role in synaptic vesicle exocytosis20 and complexin 1 modulates vesicle release21. A postmortem study reported that patients with schizophrenia have a significant decrease of CPLX1 protein in prefrontal cortex, when compared with healthy subjects22. However, an early genetic study scanned the haplotype-tagging23 SNPs in CPLX1 in a small sample of Japanese patients with schizophrenia, whereas no significant association of CPLX1 with schizophrenia was observed24. As abovementioned, CPLX1 is downregulated by miRNA137 gain of function, causing impairment in synaptic vesicle trafficking and alterations in synaptic plasticity19. Therefore, we hypothesized that the potential interaction effect of MIR137 and CPLX1 may influence the genetic risk for schizophrenia.
In this study, we aimed to investigate whether the variants of MIR137 and CPLX1 confer susceptibility to schizophrenia in Han Chinese. Here, we first used a public database to detect whether CPLX1 differentially expressed in brain between patients with schizophrenia and healthy controls. Second, we totally genotyped twelve SNPs of MIR137 and CPLX1 in our samples. Meanwhile, we also detected the effect of the two genes interaction in the susceptibility of schizophrenia, because a specific individual genetic variant has a minor marginal effect in such a complex psychiatric disease and gene-gene interaction has importance to describe such effect25.
Results
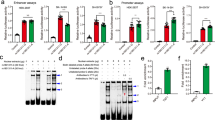
We extracted brain CPLX1 expression data between schizophrenia patients and healthy controls from SZDB database26. Figure 1 showed that there were no significant differences of CPLX1 expression in hippocampus, prefrontal cortex or stratum between the schizophrenia patients and control subjects (corrected P = 0.26, 0.64, 0.84, respectively).
Differential expression of CPLX1 in brain between patients with schizophrenia and healthy controls. Each bar represents the average level of CPLX1 expression. Error bars represent the standard deviation of the mean value. Data was extracted from the SZDB database (http://www.szdb.org/).
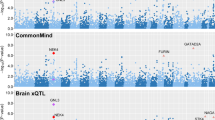
None of the genotypic distributions showed deviation from the Hardy-Weinberg equilibrium. There were no significant differences in allele and genotype frequencies in any SNP of either MIR137 or CPLX1 between the case and control groups (Table 1). After calculating LD for all pairs of SNP markers in CPLX1, we found two strong LDs between rs11248043 and rs7376690, as well as rs6832751 and rs10155482 (Supplementary Figure S1). Supplementary Table S3 listed all P values corresponding to haplotypes, with rare haplotypes (<3%) being dropped. The haplotypes showed no significant association with schizophrenia. Next, we extracted the schizophrenia genetic association data from the PGC database and observed no significant association of CPLX1 with schizophrenia either (Supplementary Figure S2). As shown in Table 2, we did not find the MIR137 SNP rs1625579 affecting schizophrenia susceptibility in interaction with the CPLX1 polymorphic variants (Ps > 0.05).
On the basis of the genotype data, the statistical power of all SNPs was more than 80% (α = 0.05) for our samples under the assumption of a modest effect size (OR = 1.5) and a log additive model and the disease prevalence of 1%.
Discussion
In this study, our results did not support the involvement of MIR137 and CPLX1 in the pathophysiology of schizophrenia, at least in Han Chinese population. Although a recent meta-analysis showed that MIR137 SNP rs1625579 significantly increases the risk of schizophrenia27, another meta-analysis indicated that the association of rs1625579 with schizophrenia did not exhibit in Asian ancestry resulted from potential genetic heterogeneity between European and Asian populations18. Our results provided further evidence to support this conclusion. On the other hand, we failed to find any positive association signals between CPLX1 and schizophrenia in Han Chinese. This is in line with early literature that Kishi et al.24 scanned CPLX1 in Japanese population and observed negative association between CPLX1 and schizophrenia. Given the modulatory effect of microRNA137 on complexin 119, we hypothesized a gene interaction between the MIR137 and CPLX1 may confer susceptibility to schizophrenia. However, our explorative analysis did not support this hypothesis.
A recent postmortem study found that transcript level for CPLX1 is significantly decreased in the anterior cingulate cortex (ACC) of schizophrenia patients28, whereas our results showed CPLX1 expression does not alter in hippocampus, prefrontal cortex or stratum of schizophrenia patients. It is generally accepted that ACC contributes to cognitive control, decision-making, empathy and emotion29,30. Animal experiment showed that CPLX1 knockout mice have pronounced deficits in social behaviors31. It is known that schizophrenia is characterized by persistent cognitive deficits, positive and negative symptoms and its etiological heterogeneity is manifested32,33,34,35. Therefore, although no association of CPLX1 with schizophrenia susceptibility was observed in our samples, we could not fully exclude the possible involvement of CPLX1 in the development of cognitive dysfunction in schizophrenia.
MicroRNA137 is enriched in hippocampal and cortical neurons that play important roles in neuronal maturation and dendritic spine morphogenesis36. It is known or predicted to regulate hundreds of genes, whose targets include many schizophrenia susceptibility genes, such as BDNF, ZNF804A, TCF4 and CACNA1C 37,38. Therefore, MIR137 associated risk for schizophrenia may be implicated with its downstream genetic effects38. Although we did not find the evidence for the involvement of MIR137 and CPLX1 in schizophrenia, further investigations are required to detect the interplay of MIR137 with its target genes in the susceptibility to schizophrenia.
This study has several limitations that should be taken into account. First, this is an exploratory study performed on a subset of the general Chinese Han population. The sample size is modest and precludes us from making any definitive statements on the associations of MIR137 and CPLX1 with schizophrenia in Han Chinese. Second, cross-sectional association studies always have the potential for population stratification. In this study, our samples were collected from Eastern China and may not be representative of the Han Chinese population in general, nor other closely related populations in the area. Third, this study was designed based on the “Common Disease-Common Variant” hypothesis, and we did not sequence the genes to assess the influence of more rare variant(s) on schizophrenia. Future targeted deep sequencing may help to undercover fundamental characteristics of pathogenic MIR137 and CPLX1 mutations and any potential association with schizophrenia.
In conclusion, our findings do not support MIR137 and CPLX1 conferring susceptibility to schizophrenia in Han Chinese. Further investigations are warranted to validate our results and identifying the polygenic effects of MIR137 with its downstream target genes in the pathophysiology of schizophrenia.
Methods
Subjects
All procedures were reviewed and approved by Institutional Review Boards of Shanghai Mental Health Center and other participating institutions. This study was performed in accordance with the guidelines laid out in the Declaration of Helsinki as revised in 1989. All subjects provided written informed consent before any study-related procedures were performed.
A total of 736 schizophrenia patients were recruited three mental hospitals in Eastern China, including Shanghai Mental Health Center, Shanghai Jiao Tong University School of Medicine, Jinhua Second Hospital and Wenzhou Kangning Hospital. The inclusion criteria for this study were according to our previous publications16,39,40. All patients (1) met the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV) criteria for schizophrenia; (2) were not first-episode; (3) had no chronic physical disease or other psychiatric disorder aside from schizophrenia. Prior to analysis, all diagnosis and review of psychiatric case records were independently checked and verified by two senior psychiatrists. The schizophrenia patients were matched with 751 control subjects enrolled from the hospital staff and students of the School of Medicine in Shanghai, all of which were interviewed by a specialized psychiatrist using the Structured Clinical Interview for DSM-IV-TR Axis I Disorders-Patient Edition (SCID-P) to determine that they had no psychiatric disorders34,41. Any healthy controls found to have any psychiatric disorder or chronic physical disease were excluded from this analysis. The patient and control groups were matched demographically, except education. Detailed participant information was summarized in Supplementary Table S1. All subjects in both the patient and control group were of Han Chinese origin.
SNP selection
We retrieved CHB data from the HapMap database (http://www.hapmap.org) and defined linkage disequilibrium (LD) blocks using Haploview 4.2 (Broad Institute, Cambridge, MA, USA) to set inclusion criteria for tagging SNPs. Haplotype-tagging single nucleotide polymorphisms (htSNPs) with r 2 cutoff >0.8 and minor allele frequency (MAF) >0.1 were selected. In total, eleven tag SNPs of CPLX1 were captured for genotyping, including rs2242237, rs2306251, rs11722977, rs7677766, rs17165034, rs9328758, rs11248042, rs11248043, rs7376690, rs6832751 and rs10155482 (Supplemental Table S2).
Genotyping
Genomic DNA of all participants was extracted from peripheral blood using a Tiangen DNA Isolation Kit (Tiangen Biotech, Beijing, China). SNP rs1625579 in MIR137 and 11 htSNPs in CPLX1 were genotyped with the Improved Multiplex Ligase Detection Reaction (iMLDR) method described in our previous study42, with technical support from the Center for Human Genetics Research, Shanghai Genesky Biotech Co., Ltd. The technicians performing genotyping were blind to the study participants. Ten percent of the samples were later randomly selected for duplicate genotyping, which produced 100% concordance.
Brain eQTL analysis for CPLX1 expression
It is known that schizophrenia originates from brain structural and functional abnormalities43,44, and dysregulation of gene expression has a key role in the pathogenesis of this disease. In this study, we performed an eQTL analysis to detect whether CPLX1 is differentially expressed in brain between patients with schizophrenia and healthy controls, using SZDB database (http://www.szdb.org/), a newly developed comprehensive resource for schizophrenia research26.
PGC data analysis
To further validate the association between the CPLX1 and schizophrenia, we extracted the schizophrenia genetic association data from the Psychiatric Genomics Consortium (PGC, http://www.broadinstitute.org/mpg/ricopili/) database4 and reanalyzed the data set as an independent sample.
Statistical analysis
Demographic data were analyzed using chi-squared or t-test as appropriate. Single marker and gene interaction analyses were conducted using SHEsisPlus (http://shesisplus.bio-x.cn/)45,46. The level of significance was corrected for multiple testing. Pair-wise linkage disequilibrium (LD) and haplotype analyses were calculated using Haploview 4.2, and the extent of LD was measured by the standardized D’. The haplotypes with a frequency under 3% were ignored. Power analysis was performed using Quanto 1.2.3 (http://hydra.usc.edu/GxE). All statistical analyses were carried out by using the SPSS 17.0 (SPSS, Inc., Chicago, IL, USA). Criterion for statistical significance was set at α = 0.05 and results were two-tailed.
References
Iannitelli, A., Quartini, A., Tirassa, P. & Bersani, G. Schizophrenia and neurogenesis: A stem cell approach. Neurosci Biobehav Rev 80, 414–442 (2017).
Kirov, G. et al. De novo CNV analysis implicates specific abnormalities of postsynaptic signalling complexes in the pathogenesis of schizophrenia. Mol Psychiatry 17, 142–153 (2012).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Schizophrenia Psychiatric Genome-Wide Association Study (GWAS) Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat Genet 43, 969–976 (2011).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet 45, 1150–1159 (2013).
Guarnieri, D. J. & DiLeone, R. J. MicroRNAs: a new class of gene regulators. Ann Med 40, 197–208 (2008).
Stark, K. L. et al. Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat Genet 40, 751–760 (2008).
Willemsen, M. H. et al. Chromosome 1p21.3 microdeletions comprising DPYD and MIR137 are associated with intellectual disability. J Med Genet 48, 810–818 (2011).
Lagos-Quintana, M., Rauhut, R., Lendeckel, W. & Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001).
Mahmoudi, E. & Cairns, M. J. MiR-137: an important player in neural development and neoplastic transformation. Mol Psychiatry 22, 44–55 (2017).
Guan, F. et al. MIR137 gene and target gene CACNA1C of miR-137 contribute to schizophrenia susceptibility in Han Chinese. Schizophr Res 152, 97–104 (2014).
Wang, S. et al. Association of microRNA137 gene polymorphisms with age at onset and positive symptoms of schizophrenia in a Han Chinese population. Int J Psychiatry Med 47, 153–168 (2014).
Ma, G. et al. Association of a miRNA-137 polymorphism with schizophrenia in a Southern Chinese Han population. Biomed Res Int 2014, 751267 (2014).
Yuan, J. et al. Lack of association between microRNA-137 SNP rs1625579 and schizophrenia in a replication study of Han Chinese. Mol Genet Genomics 290, 297–301 (2015).
Sun, Y. J. et al. Association between single nucleotide polymorphisms in MiR219-1 and MiR137 and susceptibility to schizophrenia in a Chinese population. FEBS Open Bio 5, 774–778 (2015).
Li, Z. et al. Loci with genome-wide associations with schizophrenia in the Han Chinese population. Br J Psychiatry 207, 490–494 (2015).
Saito, T. et al. Replication and Cross-Phenotype Study Based Upon Schizophrenia GWASs Data in the japanese Population: Support for Association of MHC Region with Psychosis. Am J Med Genet B 165, 421–427 (2014).
Pu, X. F. & Xiao, X. No evidence of an association between MIR137 rs1625579 and schizophrenia in Asians: a meta-analysis in 30843 individuals. Psychiat Genet 26, 203–210 (2016).
Siegert, S. et al. The schizophrenia risk gene product miR-137 alters presynaptic plasticity (vol 18, pg 1008, 2015). Nat Neurosci 19, 1115–1115 (2016).
Egbujo, C. N., Sinclair, D. & Hahn, C. G. Dysregulations of Synaptic Vesicle Trafficking in Schizophrenia. Curr Psychiatry Rep 18, 77 (2016).
Gispert, S. et al. Complexin-1 and Foxp1 Expression Changes Are Novel Brain Effects of Alpha-Synuclein Pathology. Mol Neurobiol 52, 57–63 (2015).
Sawada, K. et al. Altered immunoreactivity of complexin protein in prefrontal cortex in severe mental illness. Mol Psychiatry 7, 484–492 (2002).
Bekinschtein, P. et al. BDNF is essential to promote persistence of long-term memory storage. P Natl Acad Sci USA 105, 2711–2716 (2008).
Kishi, T. et al. No association of complexin1 and complexin2 genes with schizophrenia in a Japanese population. Schizophrenia Research 82, 185–189 (2006).
Burmeister, M., McInnis, M. G. & Zollner, S. Psychiatric genetics: progress amid controversy. Nat Rev Genet 9, 527–540 (2008).
Wu, Y., Yao, Y. G. & Luo, X. J. SZDB: A Database for Schizophrenia Genetic Research. Schizophr Bull 43, 459–471 (2017).
Ou, M. L. et al. Association between miR-137 polymorphism and risk of schizophrenia: a meta-analysis. Genet Mol Res 15 (2016).
McMeekin, L. J. et al. Cortical PGC-1alpha-Dependent Transcripts Are Reduced in Postmortem Tissue From Patients With Schizophrenia. Schizophr Bull 42, 1009–1017 (2016).
Holroyd, C. B. & Umemoto, A. The research domain criteria framework: The case for anterior cingulate cortex. Neurosci Biobehav Rev 71, 418–443 (2016).
Decety, J. & Jackson, P. L. The functional architecture of human empathy. Behav Cogn Neurosci Rev 3, 71–100 (2004).
Drew, C. J. G., Kyd, R. J. & Morton, A. J. Complexin 1 knockout mice exhibit marked deficits in social behaviours but appear to be cognitively normal. Hum Mol Genet 16, 2288–2305 (2007).
Wang, P. et al. The NCAN gene: schizophrenia susceptibility and cognitive dysfunction. Neuropsychiatr Dis Treat 12, 2875–2883 (2016).
Zhang, C. et al. A comprehensive analysis of NDST3 for schizophrenia and bipolar disorder in Han Chinese. Transl Psychiatry 6, e701 (2016).
Zhang, C. et al. Genetic modulation of working memory deficits by ankyrin 3 gene in schizophrenia. Prog Neuropsychopharmacol Biol Psychiatry 50, 110–115 (2014).
Zhang, C. et al. Effectiveness of Antipsychotic Drugs for 24-Month Maintenance Treatment in First-Episode Schizophrenia: Evidence From a Community-Based “Real-World” Study. J Clin Psychiatry 77, e1460–e1466 (2016).
Smrt, R. D. et al. MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase mind bomb-1. Stem Cells 28, 1060–1070 (2010).
Hill, M. J. et al. Transcriptional consequences of schizophrenia candidate miR-137 manipulation in human neural progenitor cells. Schizophr Res 153, 225–230 (2014).
Cosgrove, D. et al. MiR-137-derived polygenic risk: effects on cognitive performance in patients with schizophrenia and controls. Transl Psychiatry 7, e1012 (2017).
Zhu, Y. L. et al. Genetic variant in NDUFS1 gene is associated with schizophrenia and negative symptoms in Han Chinese. J Hum Genet 60, 11–16 (2015).
Tang, W. X. et al. Association study of common variants within the G protein-coupled receptor kinase 6 gene and schizophrenia susceptibility in Han Chinese. Hum Psychopharm Clin 29, 100–103 (2014).
Zhang, C. et al. Identification of ANKK1 rs1800497 Variant in Schizophrenia: New Data and Meta-Analysis. Am J Med Genet B 165, 564–571 (2014).
Zhang, C., Wu, Z. G., Hong, W., Peng, D. H. & Fang, Y. R. Evaluating the association between the SHANK3 gene and bipolar disorder. Psychiat Res 244, 284–288 (2016).
Fornito, A. & Bullmore, E. T. Reconciling abnormalities of brain network structure and function in schizophrenia. Curr Opin Neurobiol 30, 44–50 (2015).
Zhang, C., Lv, Q., Fan, W., Tang, W. & Yi, Z. Influence of CFH gene on symptom severity of schizophrenia. Neuropsychiatr Dis Treat 13, 697–706 (2017).
Shi, Y. Y. & He, L. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res 15, 97–98 (2005).
Li, Z. et al. A partition-ligation-combination-subdivision EM algorithm for haplotype inference with multiallelic markers: update of the SHEsis (http://analysis.bio-x.cn). Cell Res 19, 519–523 (2009).
Acknowledgements
We are deeply grateful to all participants. This work was supported by the National Natural Science Foundation of China (81471358), the Shanghai Science and Technology Commission Foundation (14411969000), the Shanghai Municipal Education Commission—Gaofeng Clinical Medicine Grant Support (20152530), the Shanghai Municipal Commission of Health and Family Planning Foundation (201540029) and the Shanghai Mental Health Center Foundation (2014-FX-03).
Author information
Authors and Affiliations
Contributions
W.L. and C.Z. contributed to the overall design of the study. Y.Z. and X.F. selected the SNPs and wrote the protocol for the genotyping. W.L., Y.Z., W.F., W.T., J.C. and L.S. got involved sample collection. W.L. undertook the statistical analysis and interpretation of data. W.L. carried out the majority of the study under the supervision of C.Z. W.L. and C.Z. wrote the manuscript. All authors contributed to have approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, W., Zhang, Y., Fang, X. et al. Genetic association analysis of microRNA137 and its target complex 1 with schizophrenia in Han Chinese. Sci Rep 7, 15084 (2017). https://doi.org/10.1038/s41598-017-15315-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15315-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.