Abstract
Some lactobacilli have protective effects against some heavy metals in mammals, but the underlying mechanism is not fully understood. To evaluate the remediation potency and the mechanism of Lactobacillus against chromium (Cr) in mice, Lactobacillus plantarum TW1-1 was orally administrated to Kunming mice for 7 weeks during exposure to 1 mM K2Cr2O7 in drinking water. Results showed that TW1-1 helped to decrease Cr accumulation in tissues and increase Cr excretion in feces, and may also attenuate alterations in oxidative stress and histopathological changes caused by Cr exposure. Moreover, the chromate reduction ability of fecal bacteria doubled after administration of TW1-1 upon Cr induction. MiSeq sequencing of fecal bacterial 16S rRNA genes revealed that the overall structures of gut microbiota was shifted by Cr exposure and partially restored by TW1-1. The abundances of 49 of the 79 operational taxonomic units altered by Cr were reversed by TW1-1. Based on these, we proposed a working model of TW1-1 against Cr: TW1-1 helps to remove Cr from the host and meanwhile acts as a regulator of gut microbiota, which aids in chromate reduction and provide protection against Cr. We call this process of remediation of heavy metal in the gut “gut remediation”.
Similar content being viewed by others
Introduction
Contamination of agricultural products by heavy metals is a serious problem globally. Most heavy metals in the greater environment derive from mining activities; irrigation; solid-waste disposal; pesticides and fertilizers; and atmospheric deposition1. Heavy metals in the soil are difficult to remove with current remediation methods. Crops growing in soils contaminated with metals will subsequently be contaminated as well, and their consumption by humans may result in a suite of health problems that range in severity2. It has been reported that there are up to 2.5 million potentially contaminated sites in Europe alone3; for example, 700 km2 of the Campine region of Belgium and the Netherlands have been diffusely contaminated by atmospheric deposition of cadmium (Cd), zinc, and lead (Pb)4. In China, a total of 2.88 × 106 ha of land has been contaminated with heavy metals as a result of mining, with an additional mean area of 46,700 ha polluted annually5. A classic example of the health effects of consumption of crops grown in areas polluted with heavy metals is the ‘itai-itai’ disease in Japan, which was traced to the consumption of rice and soybean grown in soil heavily polluted with Cd6. Such examples highlight the extent of the challenge and the importance of mitigating heavy metal pollution of crops.
A number of countermeasures for remediation of heavy metal contaminated soils have been introduced over the past several decades, including physical, chemical, and microbial techniques, and phytoremediation. Removal of heavy metals using living organisms is a particularly useful approach7. Microbial remediation has certain advantages, such as greater public acceptance, lower costs, and minimal site disruption8,9. Phytoremediation is another effective technique10, as it eliminates the need for soil excavation and transport11. However, the total area remediated by phytoremediation and microbial remediation is far smaller than the total area of contamination, and thus alternative approaches for the mitigation of heavy metal pollution are urgently needed.
Human exposure to heavy metals is currently inevitable. Some lactic acid bacteria such as lactobacilli are known capable of binding and removing heavy metals in vitro 12. As such, heavy metal remediation via supplementation with lactobacillus has been studied. Previous researches have found that oral administration of Lactobacillus plantarum (L. plantarum) can protect mice against toxicity of Cd, Pb, copper, and aluminum by facilitating the excretion of heavy metals through feces and inhibiting absorption via intestinal barrier13,14,15,16. Further references on how L. plantarum remediate heavy metals in vivo are scarce. On the other hand, a previous study using germ-free mice has shown that intestinal microbiome plays an essential role in fending off heavy metals exposure17. Ingestion of Cd, Pb, and nickel can lead to alterations in the composition of gut microbiota18,19,20. Hence, whether oral supplementation with L. plantarum could protect against heavy metal toxicity by regulating the composition and function of intestinal microbiota deserves to be investigated.
Chromium (Cr) is a common toxic heavy metal that is both mutagenic and carcinogenic to humans21. In the present study, we examined the protective effects of L. plantarum TW1-1, a candidate probiotic strain derived from fermented dairy products with known Cr reduction capability, against Cr toxicity in mice, and demonstrated that TW1-1 can effectively attenuate Cr(VI) toxicity22. Furthermore, we evaluated the effectiveness of “gut remediation", a process that involves both direct and indirect remediation of heavy metal pollution by L. plantarum.
Results
Cr(VI)-reduction ability of Lactobacillus strains
Four lactobacillus strains capable of reducing Cr(VI) were identified from, including L. plantarum TW1-1, L. delbrueckii ATCC 11842, L. casei BL23, and L. paracasei LZU-D2. Phylogenetic analysis of these four strains was performed using the neighbor-joining method (Fig. S1). The Cr(VI)-reduction abilities of the four strains are shown in Fig. 1. Strain TW1-1 reduced 60% of 0.5 mM Cr(VI) within 48 h of incubation, whereas strains BL23, ATCC11842, and LZU-D2 reduced about 20% of Cr(VI) over the same time period. The precipitates were re-oxidized by Na2S2O3, with results indicating that Cr(VI) was reduced rather than absorbed (Fig. S2). Thus, strain TW1-1 was chosen for further experimental study in mice, and strain LZU-D2 was used as a control lactobacillus strain with less Cr-reducing ability.
Cr levels declined in tissues and increased in feces following TW1-1 treatment
The growth of mice from all five groups were monitored throughout the experiment and did not exhibit significant differences between groups (Fig. S3). Cr (Cr[VI] + Cr[III]) content in tissues and feces were measured after mice were sacrificed. After Cr(VI) treatment, Cr levels were found sharply increased (Fig. 2a–d). Concentrations of Cr(VI) in the kidneys and liver were twice that of control mice, and concentration in the small intestine increased from 0.1405 mg/g to 0.9602 mg/g. Concentrations of Cr(III) in the kidneys and liver were four-fold higher than in mice in the control group, and concentrations in the small intestine rose from 0.1703 mg/g to 0.7774 mg/g. However, both Cr(VI) and Cr(III) levels were lower in the liver, kidneys, and small intestines of mice treated with Cr(VI) + TW1-1 than in mice treated solely with Cr(VI). At the same time, Cr(III) content was markedly increased in feces with TW1-1 treatment, rising from 0.0325 mg/g to 3.5166 mg/g, whereas Cr(VI) was nearly absent (Fig. 2d). In addition, administration of LZU-D2 also resulted in reductions in Cr concentrations in tissue samples, albeit to a lesser degree. The Cr found in feces was in the form of Cr(III), suggesting that most of the Cr(VI) entering the body via oral intake was reduced to Cr(III) and then directly excreted in the feces. Therefore, it would appear that TW1-1 is capable of reducing Cr concentrations in body tissues.
(a) to (d) Determination of Cr content in tissues and feces by atomic absorption spectrophotometer. (□) Cr(VI) content in tissues and feces; ( ) Cr(III) content in tissues and feces. Cr(III) content was detected via sodium hydroxide precipitation. (e) TEM images of hepatic tissue of mice and EDX data analysis. Arrows show accumulations of Cr particles. The particles are located on the cell membrane in Cr(VI) group, and in the cytoplasmic matrix and near the cell nucleus in Cr(VI) + LZU-D2 mice. Scale bars (from left to right in each row) are 10 μm and 2 μm. All images were taken at 150 kV.
) Cr(III) content in tissues and feces. Cr(III) content was detected via sodium hydroxide precipitation. (e) TEM images of hepatic tissue of mice and EDX data analysis. Arrows show accumulations of Cr particles. The particles are located on the cell membrane in Cr(VI) group, and in the cytoplasmic matrix and near the cell nucleus in Cr(VI) + LZU-D2 mice. Scale bars (from left to right in each row) are 10 μm and 2 μm. All images were taken at 150 kV.
Rates of Cr accumulation in the hepatic cells were further shown by transmission electron microscopy (TEM) images in Fig. 2e. Dark-colored precipitates were observed on the cell membrane of hepatocytes of mice treated with Cr(VI), and lower levels of Cr accumulation were revealed in the Cr(VI) + TW1-1 and Cr(VI) + LZU-D2 groups, although the effect was more apparent in the former group.
Strain TW1-1 attenuated Cr(VI)-induced oxidative stress and inflammation
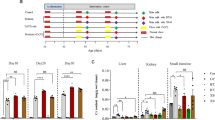
The levels of total superoxide dismutase (T-SOD), glutathione (GSH), malonaldehyde (MDA), catalase (CAT), glutathione disulfide (GSSG), and tumor necrosis factor α (TNF-α) were assayed in mouse tissues (Fig. 3 and Fig. S4). In the liver, MDA levels increased in mice treated solely with Cr(VI), and levels of GSH, CAT, and T-SOD decreased. With TW1-1 treatment, however, the levels of these markers were greatly restored (P < 0.01). In the kidney, similar restorative effects of TW1-1 were observed. Cr(VI) exposure also resulted in a marked increase of the inflammatory cytokine TNF-α in various tissues (P < 0.01), and TW1-1 reversed the changes in TNF-α to a great degree, especially in the kidneys (P < 0.05) (Fig. 3e). Considered together, our results suggested that TW1-1 was effective in mitigating oxidative stress and inflammation caused by exposure to Cr(VI).
Tissue damages induced by Cr(VI) were effectively alleviated after TW1-1 intervention
Hepatic tissues collected from mice in the control group appeared to be normal, whereas those of mice administered Cr exhibited a variety of pathologic alterations, including loss of intact liver plates, hepatocyte vacuolization, and chromatin condensation in the liver (Fig. 4). Co-treatment with TW1-1 resulted in restoration of a close-to-normal appearance of liver tissues. And moderate histological restorations were observed in mice treated with LZU-D2. There were no obvious histological alterations detected in the TW1-1-only group. Taken together, these results indicated that TW1-1 could be used to alleviate Cr (VI)-induced hepatic damages safely and effectively.
Representative photomicrographs of H&E staining of mouse liver tissues. (a),(f): Normal appearance of hepatic tissue in the control group. (b),(g): Clear Cr-induced alterations with loss of intact liver plates, hepatocyte vacuolization, and chromatin condensation. (c),(h): TW1-1 only group with no apparent histological alterations. (d),(i): Hepatocyte vacuolization and chromatin condensation were greatly alleviated in Cr(□) + TW1-1 group. (e),(j): Moderate histological restoration in Cr(VI) + LZU-D2 group. Magnification in panels (a) to (e) is 12.6×, and magnification in (f) to (j) is 25.2×. Scale bar in e represents 100 μm; scale bar inj represents 50 μm.
Colonization of TW1-1 increased the Cr(VI)-reduction ability of fecal microbes
One important trait of probiotics is their ability to survive transit through the upper gastrointestinal tract. As such, we investigated levels of TW1-1 in feces using real time (RT)-quantitative (q) PCR. The relative expression level (the expression level of TW1-1/the expression level of total bacteria) of TW1-1 was significantly higher in mice administered TW1-1 (P < 0.01) (Fig. 5a), indicating that TW1-1 colonized in the gastrointestinal tracts of treated mice.
Quantification of TW1-1 in culturable bacteria of feces inoculated in MRS medium (a) and Cr reduction ability of feces of four groups in vitro at the same time (b). The bars in b represent the reduction ability of 1010 fecal bacteria. Same letters indicate no significant differences (P > 0.05); different letters indicate a significant difference (P < 0.05).
To further investigate whether TW1-1 enhanced the reduction of Cr(VI) of gut microbiota, we compared the Cr(VI)-reduction ability of fecal microbes between the groups. As shown in Fig. 5b, the ability of fecal bacteria to reduce Cr(VI) doubled in mice given Cr diets supplemented with TW1-1 (P < 0.01), an indication that TW1-1 indeed enhances the ability of gut microbiota to reduce Cr(VI). Interestingly, the enhancement was only observed in Cr(VI) + TW1-1 group but not in the TW1-1-only group.
Structural changes of the gut microbiota in response to Cr(VI) treatment and probiotic intervention
20 fecal samples were collected after a 7-week treatment and subjected to MiSeq sequencing of the bacterial 16 S rRNA gene V4 region, with 4,794 reads generated from each sample. The majority of the phyla consisted of Bacteroidetes, Firmicutes, Acidobacteria, and Proteobacteria in all groups (Fig. 6a). Seven weeks of Cr(VI) treatment induced significant changes in gut microbial community structure, with the relative abundances of Bacteroidetes (45.62% to 66.64%, P = 0.0015) and Tenericutes (from 0.52% to 2.54%, P = 0.0285) increasing, and the abundance of Firmicutes significantly declined (39.07% to 16.97%, P = 0.0006). Cr(VI) supplemented with TW1-1, however, partially reversed the effects of Cr(VI) on the microbial abundance at phylum level, with decreased abundance of Bacteroidetes (66.64% to 52.63%, P = 0.0315) and increased abundance of Firmicutes (16.97% to 26.8%).
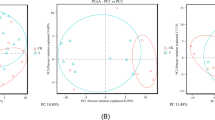
Analysis of MiSeq data by R. (a) Relative abundance (% of total reads) of bacterial 16S rRNA gene at phylum level. (b) NMDS analysis of OTUs. (c) and (d) 79 OTUs that were changed in abundance by Cr exposure according to redundancy analysis. Heatmap of the abundance of 46 OTUs enriched and 33 OTUs reduced by Cr(VI). The taxonomy of the OTUs (genus, family, and phylum) is depicted on the right. *OTUs in which abundance was changed by Cr(VI) and then reversed by TW1-1.
An NMDS was then performed to examine the relationships among the four groups. As shown in Fig. 6b, samples segregated into four distinct groups based on treatment type, suggesting that treatment had an influence on the composition of the bacterial community; the microbial communities of the two TW1-1 groups were determined to be more closely related to one another than to the control and Cr(VI)-only groups.
Treatment with Cr(VI) significantly altered the abundance of 79 OTUs, as compared to that in the control group, with 46 OTUs enhanced and 33 OTUs reduced (Fig. 6c–d). Most of the OTUs assigned to Bacteroidetes were enriched as a result of Cr(VI) treatment, whereas the majority of OTUs that decreased were associated with Firmicutes. Administration of TW1-1 partially reversed the abundances of 49 of the 79 OTUs altered by Cr(VI) treatment.
Modulation effects of TW1-1 intervention on gut microbiota
Sequences data were analyzed at the family, genus and species levels as well (Tables S2 and S3, Fig. S5). At the family level, Cr exposure led to increases in the proportion of Paraprevotellaceae (from 3.76% to 9.04%, P = 0.036) and S24-7 (from 16.38% to 25.09%), whereas the relative abundance of Lachnospiraceae declined (from 6.34% to 2.22%, P = 0.007). Co-treatment with TW1-1 significantly reversed the effects of Cr(VI) on the microbial abundance, with decreased abundance of Paraprevotellaceae (9.04% to 1.72%, P = 0.012) and increased Lachnospiraceae (2.22% to 6.51%, P = 0.00009) at the family level. Overall, TW1-1 mitigated many of the effects that Cr(VI) treatment had on microbial populations, with the exception of the genus Lactobacillus.
Supplementation with TW1-1 alone led to alterations in the proportion of Proteobacteria at the phylum level, and increased relative abundance of the Bacteroidaceae, Prevotellaceae, S24-7 and Paraprevotellaceae (from 3.76% to 9.04%, P = 0.036) at the family level, and changes in the proportion of genera Prevotella and Faecalibacterium. In general, both Cr(VI) and TW1-1 treatment respectively led to changes in the relative abundance of some bacterial species.
Discussion
In this study, we investigated the effects and the mechanisms of a Cr-reducing strain TW1-1 in remediating heavy metal Cr in mouse.
Some lactobacilli are Cr-resistant and would be useful for Cr detoxification and bioremediation, e.g. L. paracase CL1107 manifested in vitro Cr(VI) reduction ability, though not as competent as TW1-123,24. TW1-1 reduced tissue absorption of Cr by promoting its excretion through feces, implying that TW1-1 might exert its effects in two possible ways. First, inhabiting TW1-1 might quickly reduce Cr(VI) entering through the digestive tract to the less soluble Cr(III), thus reducing the amount of Cr absorbed via the intestinal tract and increasing the amount of Cr excreted with feces. This protective mechanism of TW1-1 against Cr accumulation in mice is similar to several reports focusing on L. plantarum strains25,26. Cr(VI) toxicity can be attributed to its high solubility, rapid permeability through cell membranes, and subsequent interaction with intracellular proteins and nucleic acids27, whereas Cr(III) is much less soluble and less able to permeate through cell membranes; as such, reducing Cr(VI) to Cr(III) would “trap” Cr in microbes and ultimately it would be expelled from the body along with these microbes. Second, live TW1-1 might improve bowel movement activity, which is suppressed by Cr, thus increasing fecal excretion of Cr25.
The effects of Cr(VI) exposure on oxidative stress and inflammation was evidenced by the decrease in the levels of such antioxidant markers as T-SOD, CAT, and GSH, and the increase in the peroxidation marker MDA, in body tissues. These results corroborate those of previous research in rats28. MDA is an end product and marker of the lipid peroxidation process, whereas SOD and GSH are thought to be important components of the host’s antioxidant defense system29. The exhaustion of SOD enzymes might be due to the sharp increase in ROS caused by Cr exposure, which may exceed the anti-oxidative capacities of both SOD and CAT30. Increasing levels of TNF-α suggested that host tissues might undergo further damage in response to Cr-induced stress31. Our work demonstrated that TW1-1 could partially reverse the changes in the biomarkers induced by Cr(VI) exposure.
To gain some insight on the mechanism of TW1-1-aided remediation, Cr(VI)-reduction ability of fecal microbes of Cr(VI) + TW1-1 mice was measured and found enhanced. Specifically, the capacity of 1010 fecal bacterial cells in Cr(VI) + TW1-1 mice to reduce Cr(VI) was approximately 18 μM higher than in mice treated solely with Cr(VI) (P < 0.01), which may be attributed to the Cr(VI)-reducing capacity of TW1-1 cells in the fecal bacteria or to functional changes of the gut microbiota as a whole. Based on the reducing ability of TW1-1 strain as shown in Fig. 1 (roughly 3 μM Cr(VI)/1010 TW1-1 cells after 48 h), we estimated that the 18 μM increase in the reducing ability of fecal bacteria in Cr(VI) + TW1-1 group was unlikely solely due to the reducing capacity of TW1-1; more likely, TW1-1 intake causes structural and functional changes in the gut microbiota, which in turn enhance rates of Cr(VI) reduction. The profiling changes of RNA and proteins in the gut microbial community in response to TW1-1 intervention deserve further investigation. In addition, enhanced of Cr(VI) reduction was not observed in group supplied with TW1-1 alone, suggesting that the intake of Cr(VI) was necessary for the induction of Cr resistance in the fecal microbes, consistent with a previous study showing that heavy metal remediation genes are upregulated by Cr(VI) treatment in soil microbiota32.
Given the phenotypic results, gut microbial populations were analyzed to further explore the protective mechanisms of candidate probiotic TW1-1. Gut microbiota is characterized by temporal stability and resilience, however many environmental perturbations that exceed the resilience capacity of gut microbiota can lead to dysbiosis33. In our case, mice were treated with 100 mg Cr/L water, a dosage that mimics acute Cr(VI) intoxication in human. The results showed that high doses of Cr(VI) led to dysbiosis of the gut microbiota, with significant shifts in the relative abundances of the Bacteroidetes and Firmicutes; similar transformations in microbial community composition have been observed in mice treated with Cd(II) and Pb(II)34. Moreover, Cr(VI) treatment increased the abundance of S24-735, Prevotella, and Clostridiales, but lowered Lachnospiraceae abundance. It was previously reported that Prevotella are involved in the transferable and colitogenic activity in a colitis mouse model36,37. Clostridiales are involved in hepatic metabolic activity and immune function, and affect the production of short chain fatty acids38,39. The Lachnospiraceae produce butyrate, which plays an important role in energy supply and the development of intestinal epithelial cells; lower abundances might thus correlate with the inflammation and oxidative stress observed in mice exposed to Cr(VI)40,41. Supplementation with TW1-1 restored the abundance of Bacteroidetes and Firmicutes to almost normal levels, decreased concentrations of both unclassified S24-7 and Prevotella, and increased the abundance of unclassified Lachnospiraceae to weaken the adverse effects of Cr(VI) exposure. Overall, our results suggested that TW1-1 could partially repair gut microbiota dysbiosis caused by Cr(VI) possibly by enriching the abundance of species beneficial to health and suppressing harmful species.
Ingestion of TW1-1 resulted in major structural changes of the gut microbiota, but did not lead to dysbiosis of gut microbiota in mice, a finding consistent with those previously reported42. Administration of TW1-1 led to increases in the abundance of Proteobacteria, which are involved in the metabolism of carbohydrates and maintenance of normal physiological functioning43, and the abundance of Faecalibacterium, which presumably play a protective role in the intestine44. Other species known for their anti-inflammatory effects increased in response to TW1-1 as well, including Faecalibacterium prausnitzii 45 and Prevotella copri 46. Generally, TW1-1 administration did not trigger inflammation, oxidative stress, or obvious dysbiosis of gut microbiota, suggesting that TW1-1 could be a potential candidate in remediating heavy metals with no obvious harmful side effects.
In conclusion, our study demonstrated the utility of L. plantarum TW1-1 for attenuating Cr(VI)-induced toxicity in mice. TW1-1 might remediate heavy metal toxicity by modifying the structure and functioning of the gut microbial community, a process we termed “gut remediation” (Fig. 7). Compared to current remediation technology, gut remediation appears to be direct and efficient at repairing tissue damages caused by heavy metal pollution, and might play a role in treating heavy metal toxicity in human.
Methods
Bacteria and media
The strains L. delbrueckii ATCC 1184247, L. casei BL2348, and L. plantarum TW1-122, originally derived fermented dairy producted, were provided by Dr. J. Kong (Shandong University, Jinan, China), and L. paracasei LZU-D249, isolated from fermented milk of yak from Tibetan plateau, was obtained from Dr. X. Guo (Lanzhou University, Lanzhou, China) (Table 1). The Genebank ID of 16S rRNA gene sequence of TW1-1 is KJ026561.1. A de Man, Rogosa, and Sharpe (MRS, Beijing Solarbio Science & Techinology, Beijing, China) growth medium was used to incubate the bacteria. A solid medium was prepared by adding 2% (w/w) agar to the MRS medium. Stock solutions of potassium dichromate (Shuangshuang Chemical Reagent Company, Yantai, China) were prepared at 1 mM in distilled water and 2 mM in MRS medium.
Determination of Cr(VI)-reducing abilities of Lactobacillus strains and fecal microbes in vitro
All strains were cultivated in MRS at 37 °C for 24 h under aerobic conditions, following which cultures were centrifuged and washed before being re-suspended in 0.9% NaCl solution. Re-suspended cells were inoculated in saline solution containing 500 µmol/L Cr(VI). After 48 h, cultured cells were centrifuged, with concentrations of Cr(VI) in the supernatant determined using the DPC technique50.
To determine Cr(VI) reduction in feces, approximately 0.1 g of feces pellets were collected from two mice in each group, and re-suspended in 5 ml of MRS medium and incubated overnight at 37 °C in anaerobic chamber. Next, 1 ml of the culture were inoculated in 100 ml of MRS medium containing 100 µM of Cr(VI) and incubated for additional 12 h at 37 °C under anaerobic conditions. The solution was then centrifuged to obtain cells and washed with PBS. Cells were then cultured in 100 ml of PBS containing 500 µM Cr(VI) anaerobically. After 48 h, samples were centrifuged; the supernatant was used to determine Cr(VI) concentrations, and the cells were used to obtain DNA for use in the qRT-PCR. The absolute numbers of total bacteria in each feces sample were calculated based on qRT-PCR results.
Animals, experimental design
Fifty adult female Kunming mice with an average weight of 26 g were purchased from the Animal Facility of the Medical School of Lanzhou University (Lanzhou, China), and maintained in a 12 h light/dark cycle. After 1 week of acclimatization and diet adaptation, the mice were randomly divided into five groups, with 10 mice per group (Table 2). Group 1 served as the control (no exposure to Cr[VI] or TW1-1), and Groups 2, 3, and 4 were administered 1 mM K2Cr2O7 in drinking water based on our pilot study. In addition to Cr(VI), mice in Groups 3 and 4 were administered TW1-1 and LZU-D2 via oral gavage, respectively, at the rate of 1 × 109 CFU/mouse/every second day for 7 weeks. Mice in Group 5 were administered TW1-1 only. Bacterial suspensions were prepared as previously described51. Solutions of Cr(VI) and water were replaced twice weekly with fresh preparations. All procedures and protocols followed in this study conform to the institutional guidelines and were approved by the Ethics Committee of Lanzhou University.
Determination of Cr(VI) concentrations in tissues and feces
Tissues and feces were harvested and stored in liquid nitrogen until use (Fig. S6). Frozen samples were thawed and air-dried in oven at 70 °C for 6 h until constant weight. The dessicated samples were then digested overnight in 10 ml of concentrated hydrochloric acid in enclosed caps. The digested solution was heated in a microwave until the solution was clear. The solution was then divided into two parts, one of which was treated with excess alkali, as Cr(III) can be precipitated to form insoluble oxides and hydroxides in water at neutral pH level52. The two parts were filtered through quantitative filter paper and diluted with distilled water to a volume of 25 ml, then analyzed using a graphite furnace atomic absorption spectrometer (ZEEnit®700 P Analytik Jena AG). The D-value between the two samples represented the Cr(VI) concentrations in tissues and feces.
Estimation of enzyme levels and enzymatic activity in tissues
The activities of CAT, T-SOD, MDA, GSH and GSSG levels in the liver and kidneys were measured using commercial assay kits (Jiancheng Bioenginnering Institute, Nanjing, China). An enzyme-linked immunosorbent assay kit (Shanghai Enzyme-linked Biotechnology, China) was used to determine the TNF-α level in the liver, kidney, and small intestine. All experimental procedures were performed in strict accordance with the manufacturer’s instructions.
Histological and TEM analysis
Histology of liver, kidney, and small intestine segments were carried out in standard procedures and sections were examined by light microscopy.
For TEM, tissues were immersed in 2.5% glutaraldehyde solution in PBS (Sigma, USA), and then embedded in epoxy resin and sliced with an ultramicrotome. TEM and energy-dispersive X-ray (EDX) spectroscopy analyses were performed by core facility of Lanzhou University.
Quantification of TW1-1 population in feces
Total genomic DNA of cells in feces samples was extracted using a TIANamp Stool DNA kit (TIANGEN Biotech, Shanghai, China). 16S rRNA of TW1-1 was amplified by PCR and cloned to T vector (Takara, Dalian, China), and serially diluted and used as quantitative PCR templates to make a standard curve. qPCR was performed with SYBR Premix ExTaTM II (TaKaRa, Dalian, China) on a real-time quantification PCR instrument (Bio-RAD CFX96, USA). The PCR program consisted of 95 °C for 30 s and 40 cycles at 95 °C for 5 s, 58 °C for 30 s, and 95 °C for 10 s. The copy numbers of the TW1-1 16S rRNA genes were calculated based on the standard curve and normalized against total 16S rRNA gene products amplified with universal 16 rRNA primers53. The primers are listed in Table 3.
Illumina MiSeq sequencing of fecal bacterial 16S rRNA gene V4 region and data analysis
Genomic DNA was extracted and checked by PCR with universal 16S rRNA primers 27 F/1492 R. After confirmation, the DNA was lyophilized and sent for Illumina MiSeq sequencing (Nuozhou Biotech Co., Chengdu, China)54,55. Briefly, universal primers 515F-806R plus a 10-nt barcode were used amplify the hypervariable V4 region of the bacterial 16SrRNA gene. Each PCR was performed in duplicate, with the products recovered from the agarose gel, and then prepared for sequencing. The sequence data were processed using QIIME Pipeline v. 1.7.0 (http://qiime.org/). All sequence reads were trimmed and assigned to each sample based on their barcodes. Sequences of high quality (>150 bp in length, no ambiguous base “N,” and an average base quality score >30) were used for downstream analyses. Sequences were clustered into operational taxonomic units (OTUs) at a 97% identity threshold. The aligned ITS gene sequences were used for a chimera check with the Uchime algorithm. All samples were randomly resampled to 3,760 reads. Alpha-diversity measurements (Shannon), species richness (observed, chao1), and rarefaction curves were generated from the observed species. Taxonomy was assigned using the Ribosomal Database Project (RDP) classifier. Sequencing data of the bacterial communities was transformed to a quantitative matrix, and further analyzed by non-metric multidimensional scaling (NMDS), as NMDS analysis is generally considered to be the most effective ordination method for ecological-community data. The NMDS were conducted using the ‘vegan’ package (v. 2.0–10) in R software (v. 3.1.1).
Statistical analysis
Data were expressed as the standard error of the mean (SEM) for each group. Analyses between groups were performed with one-way analysis of variance (ANOVA) tests using SPSS22. Differences were considered statistically significant at a P value < 0.05.
References
Chen, H., Zheng, C., Tu, C. & Shen, Z. Chemical methods and phytoremediation of soil contaminated with heavy metals. Chemosphere 41, 229–234 (2000).
Dong, W. Q. Y., Cui, Y. & Liu, X. Instances of soil and crop heavy metal contamination in China. Soil and Sediment Contamination 10, 497–510 (2001).
Van Liedekerke, M., Prokop, G., Rabl-Berger, S., Kibblewhite, M. & Louwagie, G. Progress in the management of contaminated sites in Europe. Reference Report by the Joint Research Centre of the European Commission (2014).
Meers, E. et al. The use of bio-energy crops (Zea mays) for ‘phytoattenuation’of heavy metals on moderately contaminated soils: a field experiment. Chemosphere 78, 35–41 (2010).
Ali, H., Khan, E. & Sajad, M. A. Phytoremediation of heavy metals—concepts and applications. Chemosphere 91, 869–881 (2013).
Aoshima, K. [Itai-itai disease: cadmium-induced renal tubular osteomalacia]. Nihon eiseigaku zasshi. Japanese journal of hygiene 67, 455–463 (2012).
Gavrilescu, M. Removal of heavy metals from the environment by biosorption. Engineering in Life Sciences 4, 219–232 (2004).
Kapoor, A., Viraraghavan, T. & Cullimore, D. R. Removal of heavy metals using the fungus Aspergillus niger. Bioresource technology 70, 95–104 (1999).
Boopathy, R. Factors limiting bioremediation technologies. Bioresource technology 74, 63–67 (2000).
Ghosh, M. & Singh, S. A review on phytoremediation of heavy metals and utilization of it’s by products. Asian J Energy Environ 6, 18 (2005).
Tangahu, B. V. et al. A review on heavy metals (As, Pb, and Hg) uptake by plants through phytoremediation. International Journal of Chemical Engineering 2011 (2011).
Monachese, M., Burton, J. P. & Reid, G. Bioremediation and tolerance of humans to heavy metals through microbial processes: a potential role for probiotics? Applied and environmental microbiology 78, 6397–6404 (2012).
Zhai, Q. et al. Protective effects of Lactobacillus plantarum CCFM8610 against chronic cadmium toxicity in mice: Intestinal sequestration is not the only route of protection. Applied and environmental microbiology, 00762–00714 (2014).
Tian, F. et al. Lactobacillus plantarum CCFM8661 alleviates lead toxicity in mice. Biological trace element research 150, 264–271, https://doi.org/10.1007/s12011-012-9462-1 (2012).
Tian, F. et al. Protective Effects of Lactobacillus plantarum CCFM8246 against Copper Toxicity in Mice. PloS one 10, e0143318, https://doi.org/10.1371/journal.pone.0143318 (2015).
Yu, L. et al. Lactobacillus plantarum CCFM639 alleviates aluminium toxicity. Applied microbiology and biotechnology 100, 1891–1900, https://doi.org/10.1007/s00253-015-7135-7 (2016).
Breton, J. et al. Gut microbiota limits heavy metals burden caused by chronic oral exposure. Toxicol Lett 222, 132–138, https://doi.org/10.1016/j.toxlet.2013.07.021S0378-4274(13)01249-6 (2013). [pii].
Liu, Y., Li, Y., Liu, K. & Shen, J. Exposing to cadmium stress cause profound toxic effect on microbiota of the mice intestinal tract. PloS one 9, e85323 (2014).
Wu, B. et al. Toxicological effects of dietary nickel chloride on intestinal microbiota. Ecotoxicology and environmental safety 109, 70–76 (2014).
Breton, J. et al. Ecotoxicology inside the gut: impact of heavy metals on the mouse microbiome. BMC Pharmacol Toxicol 14, 62, https://doi.org/10.1186/2050-6511-14-622050-6511-14-62 (2013). [pii].
Zhitkovich, A. Chromium in drinking water: sources, metabolism, and cancer risks. Chemical research in toxicology 24, 1617–1629 (2011).
Sun, Z., Kong, J. & Kong, W. Characterization of a cryptic plasmid pD403 from Lactobacillus plantarum and construction of shuttle vectors based on its replicon. Molecular biotechnology 45, 24–33, https://doi.org/10.1007/s12033-010-9242-0 (2010).
Mishra, R., Sinha, V., Kannan, A. & Upreti, R. K. Reduction of Chromium-VI by Chromium Resistant Lactobacilli: A Prospective Bacterium for Bioremediation. Toxicology international 19, 25–30, https://doi.org/10.4103/0971-6580.94512 (2012).
Huang, G., Wang, W. & Liu, G. Simultaneous chromate reduction and azo dye decolourization by Lactobacillus paracase CL1107 isolated from deep sea sediment. Journal of environmental management 157, 297–302, https://doi.org/10.1016/j.jenvman.2015.04.031 (2015).
Zhai, Q. et al. Protective effects of Lactobacillus plantarum CCFM8610 against acute cadmium toxicity in mice. Applied and environmental microbiology 79, 1508–1515 (2013).
Tian, F. et al. Lactobacillus plantarum CCFM8661 alleviates lead toxicity in mice. Biological trace element research 150, 264–271 (2012).
Cervantes, C. et al. Interactions of chromium with microorganisms and plants. FEMS Microbiology Reviews 25, 335–347 (2001).
Samuel, J. B. et al. Ameliorative effect of vitamin C on hexavalent chromium-induced delay in sexual maturation and oxidative stress in developing Wistar rat ovary and uterus. Toxicology and industrial health, 0748233711422728 (2011).
Bagchi, D., Bagchi, M., Hassoun, E. & Stohs, S. Cadmium-induced excretion of urinary lipid metabolites, DNA damage, glutathione depletion, and hepatic lipid peroxidation in Sprague-Dawley rats. Biological trace element research 52, 143–154 (1996).
Thijssen, S. et al. Low cadmium exposure triggers a biphasic oxidative stress response in mice kidneys. Toxicology 236, 29–41 (2007).
Soares, P. M. et al. Gastrointestinal dysmotility in 5-fluorouracil-induced intestinal mucositis outlasts inflammatory process resolution. Cancer chemotherapy and pharmacology 63, 91–98 (2008).
Yu, Z. et al. The shifts of sediment microbial community phylogenetic and functional structures during chromium (VI) reduction. Ecotoxicology, 1–12 (2016).
Pérez-Cobas, A. E. et al. Gut microbiota disturbance during antibiotic therapy: a multi-omic approach. Gut 62, 1591–1601 (2013).
Breton, J. et al. Ecotoxicology inside the gut: impact of heavy metals on the mouse microbiome. BMC Pharmacology and Toxicology 14, 1 (2013).
Nakajima, M. et al. Oral administration of P. gingivalis induces dysbiosis of gut microbiota and impaired barrier function leading to dissemination of enterobacteria to the liver. PloS one 10, e0134234 (2015).
Brinkman, B. M. et al. Gut microbiota affects sensitivity to acute DSS-induced colitis independently of host genotype. Inflammatory bowel diseases 19, 2560–2567 (2013).
De Filippis, F. et al. High-level adherence to a Mediterranean diet beneficially impacts the gut microbiota and associated metabolome. Gut. https://doi.org/10.1136/gutjnl-2015-309957 (2015).
Delzenne, N. M. & Cani, P. D. Interaction between obesity and the gut microbiota: relevance in nutrition. Annual review of nutrition 31, 15–31 (2011).
Imhann, F. et al. Interplay of host genetics and gut microbiota underlying the onset and clinical presentation of inflammatory bowel disease. Gut, gutjnl-2016-312135 (2016).
Sekirov, I., Russell, S. L., Antunes, L. C. M. & Finlay, B. B. Gut microbiota in health and disease. Physiological reviews 90, 859–904 (2010).
Harris, K., Kassis, A., Major, G. & Chou, C. J. Is the gut microbiota a new factor contributing to obesity and its metabolic disorders? Journal of obesity 2012 (2012).
Ohland, C. L. & MacNaughton, W. K. Probiotic bacteria and intestinal epithelial barrier function. American Journal of Physiology-Gastrointestinal and Liver Physiology 298, G807–G819 (2010).
Flint, H. J., Scott, K. P., Louis, P. & Duncan, S. H. The role of the gut microbiota in nutrition and health. Nature Reviews Gastroenterology and Hepatology 9, 577–589 (2012).
Miquel, S. et al. Faecalibacterium prausnitzii and human intestinal health. Current opinion in microbiology 16, 255–261 (2013).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. science 308, 1635–1638 (2005).
Cénit, M., Matzaraki, V., Tigchelaar, E. & Zhernakova, A. Rapidly expanding knowledge on the role of the gut microbiome in health and disease. Biochimica et Biophysica Acta (BBA)-Molecular Basis of Disease 1842, 1981–1992 (2014).
van de Guchte, M. et al. The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution. P Natl Acad Sci USA 103, 9274–9279, https://doi.org/10.1073/pnas.0603024103 (2006).
Hazebrouck, S. et al. Efficient production and secretion of bovine β-lactoglobulin by Lactobacillus casei. Microbial Cell Factories 6, 12, https://doi.org/10.1186/1475-2859-6-12 (2007).
Zhang, H. et al. [Effect of lactic acid bacteria isolated from Tibetan Plateau on silage fermentation quality of Elms nutans]. Wei sheng wu xue bao = Acta microbiologica Sinica 55, 1291–1297 (2015).
Pettine, M. & Capri, S. Removal of humic matter interference in the determination of Cr (VI) in soil extracts by the diphenylcarbazide method. Analytica chimica acta 540, 239–246 (2005).
Wang, J. et al. Modulation of gut microbiota during probiotic-mediated attenuation of metabolic syndrome in high fat diet-fed mice. The ISME journal 9, 1–15 (2015).
Pandey, S., Singh, N. K., Bansal, A. K., Arutchelvan, V. & Sarkar, S. Alleviation of toxic hexavalent chromium using indigenous aerobic bacteria isolated from contaminated sites of tannery industry. Preparative Biochemistry and Biotechnology (2015).
Castillo, M. et al. Quantification of total bacteria, enterobacteria and lactobacilli populations in pig digesta by real-time PCR. Veterinary microbiology 114, 165–170 (2006).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naïve Bayesian Classifier for Rapid Assignment of rRNA Sequences into the New Bacterial Taxonomy. Applied and Environmental Microbiology 73, 5261–5267, https://doi.org/10.1128/aem.00062-07 (2007).
Acknowledgements
This study was supported by National Natural Science Foundation grant (31470224); MOST international cooperation grant (2014DFA91340); Gansu provincial international cooperation grants (1604wkca013, 1504WKCA089-2); fundamental research funds for the central universities (2022013zr0090, lzujbky-2016-82). We thank Dr. J. Kong and Dr. X. Guo for providing the bacterial strains.
Author information
Authors and Affiliations
Contributions
G.W. and X.X. carried out the experiments. P.F. and P.L. drafted the manuscript. F.X. and Z.Y. analyzed the MiSeq sequencing data. W.Y. assisted in histological analysis. P.L. and X.L. designed the study and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, G., Xiao, X., Feng, P. et al. Gut remediation: a potential approach to reducing chromium accumulation using Lactobacillus plantarum TW1-1. Sci Rep 7, 15000 (2017). https://doi.org/10.1038/s41598-017-15216-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15216-9
This article is cited by
-
Potential of Synbiotics and Probiotics as Chemopreventive Agent
Probiotics and Antimicrobial Proteins (2024)
-
Guardians of the Gut: Harnessing the Power of Probiotic Microbiota and Their Exopolysaccharides to Mitigate Heavy Metal Toxicity in Human for Better Health
Probiotics and Antimicrobial Proteins (2024)
-
Oxidative, epigenetic changes and fermentation processes in the intestine of rats fed high-fat diets supplemented with various chromium forms
Scientific Reports (2022)
-
The influence of preferred habitat and daily range of the European hare on its contamination by heavy metals: a case study from the West Carpathians
Environmental Science and Pollution Research (2021)
-
Probiotics: a Promising Generation of Heavy Metal Detoxification
Biological Trace Element Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.