Abstract
Combinatorial design is an effective strategy to acquire the optimal solution in complex systems. In this study, the combined effects of pathway combination, promoters’ strength fine-tuning, copy numbers and integration locus variations caused by δ-integration were explored in Saccharomyces cerevisiae using geranylgeraniol (GGOH) production as an example. Two GGOH biosynthetic pathway branches were constructed. In branch 1, GGOH was converted from isopentenyl pyrophosphate (IPP) and farnesyl diphosphate (FPP). In branch 2, GGOH was derived directly from IPP and dimethylallyl pyrophosphate (DMAPP). Regulated by 10 combinations of 11 diverse promoters, a fusion gene BTS1-ERG20, a heterologous geranylgeranyl diphosphate synthase from Sulfolobus acidocaldarius (GGPPSsa) and an endogenous N-terminal truncated gene 3-hydroxyl-3-methylglutaryl-CoA reductase isoenzyme 1 (tHMGR), were incorporated into yeast by δ-integration, leading to a series of GGOH producing strains with yields ranging from 18.45 mg/L to 161.82 mg/L. The yield was further increased to 437.52 mg/L by optimizing the fermentation medium. Consequently, the GGOH yield reached 1315.44 mg/L in a 5-L fermenter under carbon restriction strategy. Our study not only opens large opportunities for downstream diterpenes overproductions, but also demonstrates that pathway optimization based on combinatorial design is a promising strategy to engineer microbes for overproducing natural products with complex structure.
Similar content being viewed by others
Introduction
(E, E, E)-Geranylgeraniol (GGOH), a kind of ingredient for perfumes, can also be used to synthesize vitamins A and E1,2. In Saccharomyces cerevisiae, GGOH is the dephosphorylated derivative of geranylgeranyl pyrophosphate (GGPP) by its endogenous phosphatases. In general, GGOH production is too low to be detected in the wild strains. Since GGPP is the direct substrate for diterpenoids synthesis in microorganisms, developing GGPP overproduction platform in yeast would open many opportunities for other high value compounds biosynthesis3. In the meanwhile, taken the difficulty of GGPP detection into consideration, GGOH can be utilized as another direct reporter for GGPP overproduction except lycopene and β-carotene. Recently, researchers have focused on the overproduction of GGOH in microbial hosts through metabolic engineering and synthetic biology strategies. Tokuhiro et al. achieved a titer of 228.8 mg/L GGOH in an engineered diploid prototrophic S. cerevisiae through overexpression of BTS1-DPP1 fusion gene, BTS1-ERG20 fusion gene, and the HMG1 genes4. Ohto et al. obtained a yield of 138.8 mg/L GGOH in a 5-L jar by combined effects of co-expression of HMG1 and the BTS1-ERG20 fusion and culture conditions optimization5. However, all above efforts just focused on fine-tuning endogenous GGPP biosynthesis pathway using IPP and FPP as precursors, in which the FPP is also an intermediate for ergosterol accumulation. It was also reported some heterologous GGPPS could directly catalyze IPP/DMAPP into GGPP, which can efficiently avoid the competition from ergosterol biosynthesis. Thus, co-overexpression and combinatorial fine-tuning of the two pathways would be crucial for GGPP overproduction as well as GGOH accumulation.
Combinatorial design has been widely used in synthetic biology and metabolic engineering areas for microbial production of diverse chemicals6,7,8. Firstly, multiple pathways with same or similar function can be combinatorially assembled to enhance target products output, Lv et al. succeeded in boosting isoprene synthesis in S. cerevisiae by dual metabolic engineering of cytoplasmic and mitochondrial acetyl-CoA utilization9. Secondly, as promoters are the key regulators of gene expression, different strength of promoters facilitated the fine-tuning of gene expression levels10,11,12,13. Kim et al. reported enhanced production of 2,3-butanediol was achieved in engineered S. cerevisiae through fine-tuning of pyruvate decarboxylase and NADH oxidase activities14. The manipulation of copy numbers and insertion sites of targeted genes are also an effective solution to realize diverse expression levels15,16. The δ-integration has been widely used for the integration of heterologous genes and the construction of biosynthetic pathways and can simultaneously achieve significant disturbances in both gene copy numbers and insertion locations17,18. The delta-sequences are yeast retrotransposon Ty1 long terminal repeats (LTR) sequences, and they have more than 400 copies distributed throughout the yeast genome. Unlike other single copy sites, δ-site integration has the potential to achieve multiple stable integrations at a time through yeast homologous recombination, and the integration efficiency is related to some factors, such as the size and the number of the integration fragments. Semkiv et al. constructed a vector for multicopy δ-integration in S. cerevisiae using a modified selective marker and a reporter gene PHO8. Their system provided integration of 3–10 copies of the module in the genome of S. cerevisiae 19. Therefore, combining all above aspects could efficiently improve the output of desired products in a more systematic way.
In the present study, three main modules involved in GGOH production, endogenous gene tHMGR, a fusion gene BTS1-ERG20, and a heterologous gene GGPPSsa from S. acidocaldarius under the control of 10 combinations of 11 various promoters were introduced into yeast by δ-integration, generating a series of GGOH producing strains, in which the yields varied from 18.45 mg/L to 161.82 mg/L. The effects of integration sites and copy numbers on GGOH production were also investigated. Finally, a titer of 1315.44 mg/L GGOH was achieved in a 5-L bioreactor based on carbon restriction strategy. Our research provides an efficient platform for GGOH and GGPP producing pool, which can largely promote downstream terpenes biosynthesis. Our study also sets a reference for microbial overproduction of desired chemicals and pharmaceuticals through combinatorial pathways design.
Results and Discussion
Pathway combination for GGOH biosynthesis
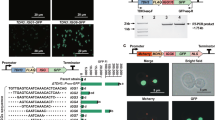
The pathway combination for the construction of GGOH biosynthetic pathway in yeast was described in Fig. 1a. To compete with hidden pathways that may inhibit the production of our desired product and to increase the flux from IPP to GGPP, both of the branch 1 and branch 2 for GGPP production were combinatorially constructed by co-overexpression of the fusion gene BTS1-ERG20 and the exogenous GGPPSsa gene (Fig. 1a). Another major regulatory control point of the MVA pathway is HMG-CoA reductase (HMGR). Early studies by Donald et al.20 and Ohto et al.5 showed that over-expression of the catalytic domain of HMGR could lead to improved productions of isoprenoids. We thus integrated the tHMGR cassette into the genome in this study as well.
Paradigm of combinatorial design for GGOH biosynthesis in S. cerevisiae. (a) Pathway combination for GGOH production. Key enzymes are tHMGR, fusion protein BTS1-ERG20 and GGPPSsa. (b) Combinatorial design of GGOH biosynthetic modules with different promoter strength. Each color and line type represented an independent combination. (c) Schematic diagram for one-step δ-integration.
Since YPRC15 locus was reported with higher gene expression level among 20 different integration sites in S. cerevisiae genome21, we transformed the three modules (GGPPSsa, BTS1-ERG20, tHMGR) into the YPRC15 locus in the genome of BY4742, obtaining strain SyBE_Sc01010369. As shown in Supplementary Fig. S1, after 72 h in shake flask cultivation, a distinct peak with retention time 13.44 min was detected in strain SyBE_Sc01010369, which was identified as GGOH based on retention time and mass spectrum (69, 93, and 119 m/z), indicating that the biosynthesis pathway of GGOH was successfully constructed in yeast. In addition, the titer of this strain we firstly constructed was only 24.93 mg/L.
The diversity of GGOH production obtained by promoters’ combination and δ-integration
Since there are two constructed pathway branches for co-producing GGOH (Fig. 1), the relative low yield of GGOH production at present may be due to unbalanced expression levels of the three modules. Systematic and combinatorial fine-tuning of the interactive modules would be an efficient solution to obtain balanced metabolic flux towards GGOH. This fine-tuning can be obtained through a selection of promoters, integration sites and copy numbers using δ-integration. According to the work of Zhao et al.11, 11 promoters with different strength were chosen for strains construction. The strength of all the promoters involved in this study were characterized by RFP as shown in Supplementary Fig. S2. In present study, as shown in Fig. 1b and Fig. 1c, one-step genomic δ-integration of the three modules with 10 combinations of 11 diverse promoters was performed in BY4742 strain. In the meanwhile, eight transformants of each constructed combination were chosen and cultivated, generating a wide dynamic library of GGOH-producing strains (obtaining strains SyBE_Sc00011201 to SyBE_Sc00011280) (Table 1) in which the yields varied from 18.43 mg/L to 161.82 mg/L (Fig. 2). Previous study by Yuan and Ching22 obtained a amorpha-4, 11-diene yield range approximately from 5 mg/L to 64 mg/L by the δ-integration of the MVA pathway genes together with carotenoid-based screening approach. The consistency in production library between our study and other group just support the efficiency of one-step δ-integration in manipulation of multiple modules. Based on this combinatorial design, a highest GGOH producing strain named SyBE_Sc00011224 was generated for further bioprocess optimization.
As described in Fig. 2, it was also found that the GGOH yields were significantly different when changing the promoters of the GGPPSsa (combinations 1, 2, 3 and 4), indicating that fine-tuning of the GGPPSsa gene might be more favorable for higher GGOH production. However, the promoter of the GGPPSsa with the highest GGOH production (TDHIp) was not the strongest one (TDH3p) among the four promoters tested. GGPPSsa could catalyze IPP/DMAPP, GPP and FPP to GGPP, but the enzyme activity for these substrates decreased sequentially23. We deduced that the fluctuations of the GGOH production caused by fine-tuning of the expression level of GGPPSsa were due to the complicate and non-linear catalytic characteristics and the wide compatibility with substrates as well as its synergistic effect with the other pathway branch in this study. This assumption was further proved by the modest but relatively apparent fluctuation of GGOH production caused by regulating BTS1-ERG20 in branch 1 (combinations 1, 5, 6 and 7). Similar results were found when promoters’ strength of tHMGR changed (combinations 1, 8, 9 and 10). We believed that fine-tuning the promoter strength of each single gene or pathway branch simultaneously led to balanced metabolic flux and thereby influenced the overall production of desired product in our study.
Furthermore, the gene expression levels of tHMGR, BTS1-ERG20, and GGPPSsa in strains with the highest GGOH production in each combination were also investigated by RT-PCR for better understanding of the underlying mechanisms that would affect the GGOH production (Fig. 3). It was shown that the diversity of expression levels for each module was realized by one-step δ-integration and the transcriptional levels of GGPPSsa were positively correlated with the GGOH yields, indicating that GGPPSsa played a dominant role in the GGOH production pathway. The GGPPSsa can directly use the IPP and DMAPP, which had higher priority and stronger competitiveness on the substrates supply compared to BTS1-ERG20, leading to more metabolic flux to GGPP than FPP. Furthermore, previous study showed GGPPSsa not only had IPP catalytic activity, but also possessed GPP and FPP catalytic activity24. In addition, the Km value for IPP and FPP of GGPPSsa was 0.361 μM and 1.8 μM, which was lower than the corresponding Km value of BTS1 as 0.8 μM and 3.2 μM25, which also indicated that GGPPSsa had a better affinity for these allylic diphosphates, and thus played a decisive role.
In addition, the δ-integration could lead to multiple copies integrating in different sites, and thus improve the integration efficiency. Regardless of the difference of viability between parent strains, the reason for the production variation between 8 transformants in each combination was thought to be the different copy numbers and integration sites caused by δ-integration. Firstly, in order to verify the effect of copy number on GGOH yield, we selected combinations 3, 4 with high-yield, and combinations 5, 7 with low-yield to verify the effects of the integration copy number on GGOH production in each transformant in these combinations by qPCR. In all the 32 transformants, 20, 3, 8 and 1 transformants possessed 1, 2, 3 and 5 copy integrations, respectively, which was consistent with the previous study that the use of auxotrophic selection marker often resulted in single integration into yeast chromosome26. Further confirmation of multiple-copy site integration and increasing the size of the screening library are likely to obtain strains with higher copies in future. Besides, the copy number could be further increased using recently developed antibiotic selection markers19,27, and CRISPR-mediated δ-integration28. As shown in Fig. 4, taken the yields into consideration, there was a nonlinear relationship between yields and copy numbers. Taking combination 7 as an example, the GGOH yield in strain with 2 copies was 54.54 mg/L, while the yield was only 47.57 mg/L in strain with 5 copies. In addition, there were also differences in yields among strains with the same copy number and promoter combination. For example, the yields of the 1 copy integration in combination 5 ranged from 31.95 mg/L to 55.32 mg/L. Recent study has also shown that position effects accounted for increased variability in gene expression levels16. Thus, expression level differences among these strains with one copy δ-integration may be due to the position effect, even though its inherent reason is not clear yet. Thus, we also have tried to figure out the differences in the integration sites, 10 specific strains were selected from combinations 3, 4, 9 and 10 (see Supplementary Table S1) with diverse GGOH production, and tested by PCR using one certain forward primer annealing to the end of the integrated module with reverse primers bonding to the downstream of the delta sequence on the chromosome (see Supplementary Fig. S3a). 25 δ-sites gained from NCBI Nucleotide BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) with more than 90% sequence similarity, were chosen for verification by PCR, and 6 δ-sites were confirmed with fragments integration. The result showed that the integration sites of delta-integration were diversified, but the correlation between integration sites and GGOH productions was not apparent (see Supplementary Fig. S3b and Supplementary Table S1). In summary, our results demonstrated that the variety by δ-integration would facilitate a large range for productions of target compounds. The δ-integration combined with fine-tuning of promoters’ strength provided a comprehensive and efficient solution for obtaining high-producing strains through synthetic biology and metabolic engineering. Nevertheless, the fact that different integration sites gave rise to different production levels does not mean that all production levels can be explained through copy numbers and integration sites, which needs further illustration in future work.
GGOH production promoted by medium optimization and fed batch fermentation
In order to determine the optimal fermentation medium for further bioreactor production of GGOH, systematic optimization was performed using the highest GGOH yield strain in the combination 3, named SyBE_Sc00011224. Firstly, the 2-level fractional factorial design was utilized to investigate the main effects and interactions of glucose, yeast extract, peptone, MgSO4 and KH2PO4 (see Supplementary Table S2). The result (see Supplementary Fig. S4) showed that the composition of glucose, yeast extract and peptone significantly influenced GGOH production. Then, the three critical factors were further determined by applying Response Surface Methodology (RSM) with a Box-Behnken design (see Supplementary Table S3) whereas concentrations of KH2PO4 and MgSO4 were fixed on the constant levels. The experimental results were analyzed statistically using ANOVA. The 3D surface plots of every two factors were shown in Fig. 5a,b,and c, and the final equation in terms of coded factors was presented in Equation S1. Based on the equation, optimal solution of the concentration was 52.9 g/L glucose, 16.4 g/L yeast extract and 32.8 g/L peptone, and the maximum titer of GGOH was predicted to reach 420.00 mg/L. The verification experiment was done with the predicted concentration of medium and the actual GGOH production was 437.52 ± 30.18 mg/L, which matched the prediction very well. The good agreement between the prediction and experimental results verified the validity of the model. This titer in the optimized fermentation medium was increased by 1.75-fold than that in the traditional YPD medium.
Medium optimization and fed-batch fermentation. (a) Response surface methodology for medium optimization. (b) 5-L Fed-batch fermentation. Line in dark, green, yellow, red, blue and purple represented glucose, ethanol, acetate, glycerol, OD600 and GGOH, respectively. All the data points were calculated from duplicates.
Statistical optimization is an important step for overproduction of the desired product and involves a number of physiochemistry parameters29,30. Microbes that are engineered to target metabolites always have different requirements for nutrients from the original ones due to the shift of metabolic flux caused by introduction of heterologous pathways. Through rational optimization, the most favorable and well-balanced compositions of nutrients can be easily obtained which facilitates the maximum yield of the desired product31,32. The enhancement of cell growth and GGOH production through statistical optimization in this study illustrated that there would be still huge potential for shifts in metabolism by regulating medium components. It could be speculated that combinatorial integrative design provided a high possibility for increasing the yield of target product by RSM.
In order to fully explore the GGOH producing ability in the engineered strains, fed-batch fermentation of the highest GGOH accumulating strain (437.52 ± 30.18 mg/L in strain SyBE_00011224) was carried out in a 5-L bioreactor (Fig. 5d) with liquid-liquid two-phase fermentation using 20% (v/v) dodecane as overlay. The organic solvent was introduced to prevent air stripping of secreted GGOH from the fermentation medium. In defined media with controlled glucose feeding, GGOH production of the strain SyBE_Sc00011224 increased to 1315.44 mg/L within 200 h. In addition, the biomass increased dramatically during the first 35 h when the yield of GGOH reached 238.49 mg/L. Then, the strain kept a relatively stable growth rate during the next 165 h, and the titer increased significantly from 238.49 mg/L to 1315.44 mg/L. During the whole process of fermentation, GGOH maintained a stable growth rate, which proved that there was great potential to further increase production. However, it was also noticed that the fermentation time was quite long. As recent study in process optimization has demonstrated great potential in isoprene overproduction (up to 24.0 g/L)33, we believe that GGOH and related other terpenoids production by our engineered strain would be further improved by continuous efforts in metabolic engineering, synthetic biology and fermentation optimization.
Conclusion
In this study, combinatorial design was carried out to optimize the GGOH titer in engineered S. cerevisiae. A GGOH overproducing S. cerevisiae with the titer of 1315.44 mg/L in the 5-L bioreactor was obtained by constructing two GGOH biosynthetic pathway branches, fine-tuning the promoters’ strength of key genes and optimizing the fermentation conditions. This study provides a possibility for obtaining high-production strains by combinatorial design. Thus, it would be a helpful strategy for constructing microbes by combinatorial design to overproduce terpenoid and other natural products. The improvement of the GGOH titer in S. cerevisiae chassis constructs a sufficient GGPP supply pool and would eventually offer a great opportunity for the biosynthesis of taxol precursors overproduction, such as taxadiene and oxygenated taxanes in microbes.
Methods
Strains and plasmids
All the plasmids were constructed in E. coli DH5a, and all the GGOH producing strains were constructed in the yeast strain BY4742, a derivative of S288C34, was obtained from EUROSCARF and used as the parent strain for all yeast strains. Plasmids extraction was purchased from BEIJING Biomed Co., Ltd. The yeast expression plasmid pRS425 was purchased from Addgene (American) and the ampicillin resistant gene of the plasmid was substituted by kanamycin resistant gene to construct pRS425K. The yeast strains constructed in this study were listed in Table 1.
Combinatorial design and plasmid construction
The GGOH biosynthetic pathway branches were constructed by combinatorial design (Fig. 1a). The GGOH producing strains were engineered followed the illustration shown in Fig. 1b and c. For the production of GGOH, yeast endogenous gene tHMGR, ERG20 and BTS1 were amplified by PCR from the genomic DNA of the strain BY4742. The fusion enzyme encoding gene BTS1-ERG20 was constructed using overlap extension PCR (OE-PCR) according to the method described by Zhou et al.35. The codon optimization and complete synthesis of GGPPS gene from S. acidocaldarius (GGPPSsa) were performed by AuGCT Company (China). Promoters with different strength used in this study were shown in Fig. 1b and Supplementary Table S4, and were all amplified by PCR from the genomic DNA of the strain BY4741. All the cassettes built in this study were listed in Supplementary Table S4, and primers used were listed in Supplementary Table S5. In order to integrate the pathway modules into the YPRC15 locus, four cassettes YPRC15up-ura3-CYC1t, CYC1t-TPI1p-GGPPSsa-TEF1t, TEF1t-GPM1p-BTS1-ERG20-TPI1t, and TPI1t-PDC1p-tHMGR-FBA1t-YPRC15dn were built in the vector pRS425K separately with their terminator acting as homologous fragment to each other. Specifically, for the pRS425K- YPRC15up-ura3-CYC1t cassette, the YPRC15 locus upstream 600 bp homologous fragment YPRC15up, the URA3 marker gene from Kluyveromyces lactis and the CYC1 terminator were constructed by OE-PCR. The resulting fragments were digested with NotI, and introduced into the corresponding sites of vector pRS425K to form pRS425K-YPRC15up-ura3-CYC1t. For pRS425K-CYC1t-TPI1p-GGPPSsa-TEF1t cassette, the empty cassette pRS425K-CYC1t-TPI1p-TEF1t was first constructed by OE-PCR, consisting of upstream homologous fragment CYC1 terminator, promoter TPI1p, and downstream terminator TEF1 into the plasmid pRS425K. Then, the gene GGPPSsa was introduced into the empty cassettes constructed above to create pRS425K-CYC1t-TPI1p-GGPPSsa-TEF1t cassettes using the Golden Gate method36. The pRS425K-TEF1t-GPM1p-BTS1-ERG20-TPI1t cassette was constructed following the similar steps as pRS425K-GGPPSsa cassette. For pRS425K-TPI1t-PDC1p-tHMGR-FBA1t-YPRC15dn cassette, YPRC15dn was introduced into the end of the second terminator as the downstream homologous region with yeast chromosome. Correct constructions of the cassettes were confirmed by diagnostic PCR using primers annealing to the end of the promoter and the beginning of the terminator. In each case, the correct insertion was confirmed by vector insert sequencing. Then, the upstream (YPRC15up) and downstream (YPRC15dn) homologous modules were replaced by delta-site upstream homologous fragment delta1 and downstream homologous fragment delta 2 for δ-integration. Besides, promoters with different strengths were introduced into corresponding cassettes following the same method above except for the TEF1p regulated cassettes. Since there was a BsaI restriction site in the nucleotide sequence of TEF1p, PmeI site was introduced into the TEF1p end instead of BsaI and yeast homologous recombination was applied to in vivo assembling the PmeI cleaved plasmid and the gene.
The expression levels of the 11 promoters used in this study were tested by RFP. Different promoters were digested by BglII /XbaI and were introduced into the corresponding site of vector pRS425K-KAX-GFP-Leu, which was integrated at the KAX site of the BY4742 strain.
Yeast transformation, verification and cultivation
The target pathways were constructed by DNA assembler37. The cassettes described above were digested with corresponding restriction enzymes, and the URA3 maker fragment and the gene fragments related to the GGOH pathway were co-transformed into yeast after gel purification as designed in Fig. 1c. Yeast was transformed using the LiAc/SS carrier DNA/PEG method followed by selection on SC-drop agar plates (0.2% amino acid mixture, 0.67% yeast nitrogen base without amino acid, 2% glucose, 1.8% agar) without supplementation of uracil. After incubating at 30 °C for 36 h, 12 colonies were selected to be cultivated in SC-drop medium at 220 rpm for 20 h before the genomes were extracted. The genomes of the 12 colonies were extracted and verified by PCR with primers listed in Supplementary Table S5 and the genome of the BY4742 chassis was used as negative control. For GGOH production, seed was firstly inoculated from SC-drop agar plate into 20 mL culture tubes containing 5 mL SD medium and cultivated at 30 °C, 220 rpm for 20 h to the exponential phase (OD600≈5.0). And then aliquots were diluted to an initial OD600 of 0.2 in 5 mL of SD medium and cultivated at 30 °C, 220 rpm for 12 h until OD600 reached about 5.0. Aliquots were diluted to an initial OD600 of 0.1 in 50 mL of YPD medium (1% yeast extract, 2% peptone, 2% glucose) in 250 mL flasks and were cultivated at 30 °C, 220 rpm. Dodecane was added aseptically to 5% (v/v) of the culture at 10 h to minimize the loss of GGOH. And the organic layer was harvested for GGOH analysis after 70 h by centrifugation of the fermentation broth at 11,000 rpm for 10 min.
Fed batch Fermentation for GGOH production
The experiment of fractional factorial designs and response surface methodology (RSM) were calculated using Design Expert Software, Stat-Ease, Inc. 1.5 g/L citric acid which served as chelating agent was introduced into the medium as MgSO4 and KH2PO4 were added. The fermentation processes at shake-flask level were performed as described above.
A 5-L bioreactor (BLBIO-5GJG-2, Shanghai, China) was used for this study. Firstly, cells from a single colony of SyBE_00011224 on the SC-drop agar plate was cultivated in a 100 mL flask containing 20 mL SC-drop medium at 30 °C, 200 rpm for 20 h, and then aliquots were diluted to an initial OD600 of 0.2 in 200 mL of SD medium and cultivated for 10 h until OD600 reached about 5.0. Secondly, 200 mL seed culture was transferred into the bioreactor containing 1.8 L fermentation medium (5% glucose, 1% yeast extract, 3% peptone, 0.8% KH2PO4, and 0.6% MgSO4). Oxygen was supplied by filtered air at 1 vvm. The agitation was 350 rpm. The temperature was maintained at 30 °C. The pH of the culture was controlled at 5.8 using 10 M ammonia water. 20% (v/v) dodecane was added aseptically into the culture at 6 h of the fermentation to start the biphasic liquid-liquid fermentation. The cell growth and the concentration of glucose were constantly monitored during the fermentation process. Glucose was feed into the culture continuously and the speed of the glucose feeding was controlled to keep its concentration in the culture between 0~1 g/L. GGOH (98% pure) was prepared to construct a standard curve for determining GGOH production.
GGOH analysis by GC-TOF/MS
The GGOH analysis was carried out by GC-TOF/MS according to the previous studies5,38. The dodecane layer was sampled and diluted in hexane. 1 μL sample was injected by Agilent 7683 autosampler into Agilent 6890 GC, which was equipped with a fused silica capillary column (30 m × 0.25 mm i.d., 0.25 mm DB-5MS, J&W Scientific, Folsom, CA). The injector temperature was set at 260 °C. The column effluent was introduced into the ion source (250 °C) of TOF/MS. And ions were generated by 40 mA ionization current of a 70 eV electron beam. The mass scan range was 50–800 m/z.
For GC-TOF/MS analysis of GGOH, the oven temperature was first kept constant at 70 °C for 1 min, then increased to 115 °C at a rate of 15 °C/min and kept for 1 min. Next, it increased to 250 °C at a rate of 20 °C/min, and finally increased to 300 °C at a rate of 25 °C/min, kept for 7 min. The total run time was 20.75 min. The GGOH was identified by the mass fragments 69, 93, and 119 m/z.
Real-time PCR for transcriptional analysis and copy numbers verification
Transcriptional expression levels of the genes BTS1-ERG20, GGPPSsa, and tHMGR in each constructed strain with different GGOH productions were evaluated by qPCR. The extraction of total RNA was performed using Trizol solution (Invitrogen). The PCR procedure was carried out on a CFX96 real time PCR system (Bio-Rad) in a total volume of 20 μL containing diluted cDNA (1 μL), 2 × SsoFast supermix (10 μL), 10 μM forward primer and reverse primer (0.8 μL for each) and 50 × ROX Reference Dye II (0.4 μL). The cycling condition used was 95 °C for 2 min, followed by 40 cycles of 95 °C for 10 s and 58 °C for 20 s.
δ-integration method can obtain a various range of copy numbers. Thus, it’s necessary to verify the copy numbers of each engineered strain by quantitative PCR, which can uncover the relationship between the copy numbers and yields. As the three genes BTS1-ERG20, GGPPSsa and tHMGR were integrated sequentially into the yeast chromosome, their copy numbers were the same theoretically except the endogenous ones. Thus, only the gene GGPPSsa of each constructed strain with different GGOH productions were evaluated by qPCR. ALG9 gene in the chromosome was chosen as the reference gene that was verified with one copy in the strain genome.
References
Gershenzon, J. & Dudareva, N. The function of terpene natural products in the natural world. Nat Chem Biol 3, 408–414, https://doi.org/10.1038/nchembio.2007.5 (2007).
You, L. F., Guo, L. Q., Lin, J. F., Ren, T. & Wang, J. R. Overproduction of geranylgeraniol in Coprinopsis cinerea by the expression of geranylgeranyl diphosphate synthase gene. J Basic Microbiol 54, 1387–1394, https://doi.org/10.1002/jobm.201400152 (2014).
Krivoruchko, A. & Nielsen, J. Production of natural products through metabolic engineering of Saccharomyces cerevisiae. Curr Opin Biotechnol 35, 7–15, https://doi.org/10.1016/j.copbio.2014.12.004 (2015).
Tokuhiro, K. et al. Overproduction of geranylgeraniol by metabolically engineered Saccharomyces cerevisiae. Appl Environ Microbiol 75, 5536–5543, https://doi.org/10.1128/AEM.00277-09 (2009).
Ohto, C., Muramatsu, M., Obata, S., Sakuradani, E. & Shimizu, S. Overexpression of the gene encoding HMG-CoA reductase in Saccharomyces cerevisiae for production of prenyl alcohols. Appl Microbiol Biotechnol 82, 837–845, https://doi.org/10.1007/s00253-008-1807-5 (2009).
Jin, Y. S. & Stephanopoulos, G. Multi-dimensional gene target search for improving lycopene biosynthesis in Escherichia coli. Metab Eng 9, 337–347, https://doi.org/10.1016/j.ymben.2007.03.003 (2007).
Lechner, A., Brunk, E. & Keasling, J. D. The need for integrated approaches in metabolic engineering. Cold Spring Harb Perspect Biol 8, https://doi.org/10.1101/cshperspect.a023903 (2016).
Essani, K., Glieder, A. & Geier, M. Combinatorial pathway assembly in yeast. AIMS Bioengineering 2, 423–436, https://doi.org/10.3934/bioeng.2015.4.423 (2015).
Lv, X. et al. Dual regulation of cytoplasmic and mitochondrial acetyl-CoA utilization for improved isoprene production in Saccharomyces cerevisiae. Nat Commun 7, 12851, https://doi.org/10.1038/ncomms12851 (2016).
Redden, H., Morse, N. & Alper, H. S. The synthetic biology toolbox for tuning gene expression in yeast. FEMS Yeast Res 15, 1–10, https://doi.org/10.1111/1567-1364.12188 (2015).
Sun, J. et al. Cloning and characterization of a panel of constitutive promoters for applications in pathway engineering in Saccharomyces cerevisiae. Biotechnol Bioeng 109, 2082–2092, https://doi.org/10.1002/bit.24481 (2012).
Yu, J. H. et al. Combinatorial optimization of CO2 transport and fixation to improve succinate production by promoter engineering. Biotechnol Bioeng 113, 1531–1541, https://doi.org/10.1002/bit.25927 (2016).
Freestone, T. S. & Zhao, H. Combinatorial pathway engineering for optimized production of the anti-malarial FR900098. Biotechnol Bioeng 113, 384–392, https://doi.org/10.1002/bit.25719 (2016).
Kim, J. W. et al. Enhanced production of 2,3-butanediol by engineered Saccharomyces cerevisiae through fine-tuning of pyruvate decarboxylase and NADH oxidase activities. Biotechnol Biofuels 9, 265, https://doi.org/10.1186/s13068-016-0677-9 (2016).
Lian, J., Jin, R. & Zhao, H. Construction of plasmids with tunable copy numbers in Saccharomyces cerevisiae and their applications in pathway optimization and multiplex genome integration. Biotechnol Bioeng 113, 2462–2473, https://doi.org/10.1002/bit.26004 (2016).
Chen, X. & Zhang, J. The Genomic landscape of position effects on protein expression level and noise in yeast. Cell Syst 2, 347–354, https://doi.org/10.1016/j.cels.2016.03.009 (2016).
Dujon, B. The yeast genome project: what did we learn? Trends in Genetics Tig 12, 263–270 (1996).
Lee, W. & Dasilva, N. A. Application of sequential integration for metabolic engineering of 1,2-propanediol production in yeast. Metab Eng 8, 58–65, https://doi.org/10.1016/j.ymben.2005.09.001 (2006).
Semkiv, M. V., Dmytruk, K. V. & Sibirny, A. A. Development of a system for multicopy gene integration in Saccharomyces cerevisiae. J Microbiol Methods 120, 44–49, https://doi.org/10.1016/j.mimet.2015.10.023 (2016).
Donald, K. A., Hampton, R. Y. & Fritz, I. B. Effects of overproduction of the catalytic domain of 3-hydroxy-3-methylglutaryl coenzyme A reductase on squalene synthesis in Saccharomyces cerevisiae. Appl Environ Microbiol 63, 3341–3344 (1997).
Flagfeldt, D. B., Siewers, V., Huang, L. & Nielsen, J. Characterization of chromosomal integration sites for heterologous gene expression in Saccharomyces cerevisiae. Yeast 26, 545–551, https://doi.org/10.1002/yea.1705 (2009).
Yuan, J. & Ching, C. B. Combinatorial engineering of mevalonate pathway for improved amorpha-4,11-diene production in budding yeast. Biotechnol Bioeng 111, 608–617, https://doi.org/10.1002/bit.25123 (2014).
Ohnuma, S., Suzuki, M. & Nishino, T. Archaebacterial ether-linked lipid biosynthetic gene. Expression cloning, sequencing, and characterization of geranylgeranyl-diphosphate synthase. J Biol Chem 269, 14792–14797 (1994).
Ohnuma, Si, Hemmi, H., Ohto, C., Nakane, H. & Nishino, T. Effects of random mutagenesis in a putative substrate-binding domain of geranylgeranyl diphosphate synthase upon intermediate formation and substrate specificity. J Biochem 121, 696–704, https://doi.org/10.1093/oxfordjournals.jbchem.a021642 (1997).
Lo, C. H. et al. Combined experimental and theoretical study of long-range interactions modulating dimerization and activity of yeast geranylgeranyl diphosphate synthase. J Am Chem Soc 131, 4051–4062, https://doi.org/10.1021/ja808699c (2009).
Lee, F. W. & Da Silva, N. A. Sequential delta-integration for the regulated insertion of cloned genes in Saccharomyces cerevisiae. Biotechnol Prog 13, 368–373, https://doi.org/10.1021/bp970055d (1997).
Yuan, J. & Ching, C. B. Combinatorial assembly of large biochemical pathways into yeast chromosomes for improved production of value-added compounds. ACS Synth Biol 4, 23–31, https://doi.org/10.1021/sb500079f (2015).
Shi, S., Liang, Y., Zhang, M. M., Ang, E. L. & Zhao, H. A highly efficient single-step, markerless strategy for multi-copy chromosomal integration of large biochemical pathways in Saccharomyces cerevisiae. Metab Eng 33, 19–27, https://doi.org/10.1016/j.ymben.2015.10.011 (2016).
Izmirlioglu, G. & Demirci, A. Enhanced bio-ethanol production from industrial potato waste bystatistical mediumoptimization. Int J Mol Sci 16, 24490–24505, https://doi.org/10.3390/ijms161024490 (2015).
Mandenius, C. F. & Brundin, A. Bioprocess optimization using design-of-experiments methodology. Biotechnol Prog 24, 1191–1203, https://doi.org/10.1002/btpr.67 (2008).
Choudhari, S. & Singhal, R. Media optimization for the production of beta-carotene by Blakeslea trispora: a statistical approach. Bioresour Technol 99, 722–730, https://doi.org/10.1016/j.biortech.2007.01.044 (2008).
Witek-Krowiak, A., Chojnacka, K., Podstawczyk, D., Dawiec, A. & Pokomeda, K. Application of response surface methodology and artificial neural network methods in modelling and optimization of biosorption process. Bioresour Technol 160, 150–160, https://doi.org/10.1016/j.biortech.2014.01.021 (2014).
Yang, C. et al. Synergy between methylerythritol phosphate pathway and mevalonate pathway for isoprene production in Escherichia coli. Metab Eng 37, 79–91, https://doi.org/10.1016/j.ymben.2016.05.003 (2016).
Baker Brachmann, C. et al. Designer deletion strains derived fromSaccharomyces cerevisiae S288C: A useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14, 115–132, https://doi.org/10.1002/(sici)1097-0061(19980130)14:2115::aid-yea2043.0.co;2-2 (1998).
Zhou, Y. J. et al. Modular pathway engineering of diterpenoid synthases and the mevalonic acid pathway for miltiradiene production. J Am Chem Soc 134, 3234–3241, https://doi.org/10.1021/ja2114486 (2012).
Engler, C., Gruetzner, R., Kandzia, R. & Marillonnet, S. Golden gate shuffling: a one-pot DNA shuffling method based on type IIs restriction enzymes. PLoS One 4, e5553, https://doi.org/10.1371/journal.pone.0005553 (2009).
Shao, Z., Zhao, H. & Zhao, H. DNA assembler, an in vivo genetic method for rapid construction of biochemical pathways. Nucleic Acids Res 37, e16, https://doi.org/10.1093/nar/gkn991 (2009).
Engels, B., Dahm, P. & Jennewein, S. Metabolic engineering of taxadiene biosynthesis in yeast as a first step towards Taxol (Paclitaxel) production. Metab Eng 10, 201–206, https://doi.org/10.1016/j.ymben.2008.03.001 (2008).
Acknowledgements
This work was funded by the National Natural Science Foundation of China (21621004; 21390203, 21676190, 21676192, 81502976); the Ministry of Science and Technology of China (“973” Program: 2014CB745100); Innovative Talents and Platform Program of Tianjin (16PTSYJC00050), and Foundation for the Author of National Excellent Doctoral Dissertation of PR China (201456); Tianjin Municipal Science and Technology Commission (13RCGFSY19800).
Author information
Authors and Affiliations
Contributions
W.H.X. and Y.J.Y. conceived and supervised the experiments; T.Q.S., M.Z.D. and F.Z. performed the experiments; M.Z.D. and D.L. provided technical assistances; H.L. and T.Q.S. identified GGOH and analyzed the data; T.Q.S., M.Z.D. and F.Z. drafted the manuscript and all author contributed to writing.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Song, TQ., Ding, MZ., Zhai, F. et al. Engineering Saccharomyces cerevisiae for geranylgeraniol overproduction by combinatorial design. Sci Rep 7, 14991 (2017). https://doi.org/10.1038/s41598-017-15005-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15005-4
This article is cited by
-
Metabolic engineering of Saccharomyces cerevisiae for enhanced taxadiene production
Microbial Cell Factories (2024)
-
Improved production of Taxol® precursors in S. cerevisiae using combinatorial in silico design and metabolic engineering
Microbial Cell Factories (2023)
-
De novo biosynthesis of 8-prenylnaringenin in Saccharomyces cerevisiae improved by screening and engineering of prenyltransferases and precursor pathway
Systems Microbiology and Biomanufacturing (2023)
-
Enhanced production of taxadiene in Saccharomyces cerevisiae
Microbial Cell Factories (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.