Abstract
Seasonal variation in temperature fluctuations may provide corals and their algal symbionts varying abilities to acclimate to changing temperatures. We hypothesized that different temperature ranges between seasons may promote temperature-tolerance of corals, which would increase stability of a bacterial community following thermal stress. Acropora muricata coral colonies were collected in summer and winter (water temperatures were 23.4–30.2 and 12.1–23.1 °C, respectively) from the Penghu Archipelago in Taiwan, then exposed to 6 temperature treatments (10–33 °C). Changes in coral-associated bacteria were determined after 12, 24, and 48 h. Based on 16S rRNA gene amplicons and Illumina sequencing, bacterial communities differed between seasons and treatments altered the dominant bacteria. Cold stress caused slower shifts in the bacterial community in winter than in summer, whereas a more rapid shift occurred under heat stress in both seasons. Results supported our hypothesis that bacterial community composition of corals in winter are more stable in cold temperatures but changed rapidly in hot temperatures, with opposite results for the bacterial communities in summer. We infer that the thermal tolerance ranges of coral-associated bacteria, with a stable community composition, are associated with their short-term (3 mo) seawater thermal history. Therefore, seasonal acclimation may increase tolerance of coral-associated bacteria to temperature fluctuations.
Similar content being viewed by others
Introduction
Coral-associated bacteria are critical to the health and function of coral ecosystems and coral holobiont1,2. Coral-associated bacteria are dynamic and often form stable homeostatic associations with their host3. Therefore, changes in these bacterial communities may be an indicator of health of coral holobionts4.
Increased temperature is a stressor for coral reefs and may adversely affect coral holobiont physiology5,6. Temperature stress from global climate change, often a critical factor in massive coral bleaching7,8, can directly or indirectly influence coral health, increasing susceptibility to disease or facilitating pathogen propagation9,10. Shifts in coral bacterial communities are common after heat stress9,11,12,13. Further to this, cold stress may also induce coral bleaching14,15,16 and cause mortality17,18. Cold stress may decrease growth, metabolic activity, respiration and chlorophyll a content in corals16,19,20. However, effects of cold stress on coral-associated bacterial communities have not been well characterized.
To reduce detrimental impacts of thermal stress corals may acclimatize21,22. Mechanisms responsible for physiological acclimation are not well understood, but the process is likely affected by thermal history23,24. Corals can resist temperature stress or acclimate to dramatic temperature fluctuations25,26,27,28. Interestingly, coral-associated bacterial communities also exhibit a similar phenomenon. After acclimatization, the bacterial community in corals may remain stable before and after heat stress29,30. Santos and co-workers (2014)29 preheated coral colonies prior to heat treatment; in the study, communities in heat-acclimatized samples were more stable than in those without pretreatment. Similarly, Ziegler and co-workers (2017)30 transplanted coral colonies to pools with higher temperature for acclimation before a heat-treatment experiment. After 1 y of acclimation, transplanted colonies had consistent bacterial communities before and after heat treatment, whereas colonies that had remained in original habitats had altered composition of bacterial communities after treatment.
Responses of coral-associated bacteria to acclimation, more specifically variation in responses to acute stress, have not been well characterized. Seasonal acclimation in coral-associated bacteria is unknown, as only between-season variations in the coral-associated bacterial community have been reported31,32,33,34,35. To better understand this phenomenon, we conducted a comprehensive study to determine effects of temperature stress on changes in bacteria with different seasonal thermal histories.
The objectives were to detect short-term dynamics in coral-associated bacterial communities under various temperatures and to compare this variation between summer and winter. Acropora muricata coral colonies were collected from the Penghu Archipelago, located in western Taiwan, proposed as a climate-change refuge for corals in a previous study36. It has a unique thermal regime; warm waters of the Kuroshio Current pass through the islands from the south in summer, whereas in winter, cold waters of the China Coastal Current move down the islands from the north. Therefore, this area has a wide range (18 °C) in temperatures annually (from 12.1 to 30.1 °C). Consequently, average monthly seawater temperature in Penghu ranges from 18.1 °C in winter to 27.7 °C in summer. Therefore, we conducted tank experiments with temperature treatments that compassing a range from 10 to 33 °C (2 °C lower than the lowest temperature in Penghu and 3 °C higher than the highest temperature), which were 10 °C, 15 °C, 20 °C, 25 °C /26 °C, 30 °C and 33 °C. This study is the first to characterize the impacts of cold and heat stressors on changes in coral-associated bacterial communities after seasonal acclimation (summer and winter). We concluded that seasonal thermal history may enhance stability of the bacterial community against thermal stress.
Results
Composition of bacterial communities between coral and seawater in 2 seasons
To detect short-term dynamics in coral-associated bacterial communities exposed to various temperatures and to compare this variation between summer and winter, Acropora muricata coral colonies were collected from the Penghu Archipelago, in western Taiwan (Fig. 1). Nubbins from Acropora muricata were exposed to 6 acute temperature treatments, 10 °C, 15 °C, 20 °C, 25 °C (in winter or 26 °C in summer), 30 °C, and 33 °C in both summer and winter. Changes in coral-associated bacteria were determined after 12, 24 and 48 h (Table 1).
Sampling locations in Wukan (23°32′38.9″N, 119°37′32.3″E). Acropora muricata were collected in June 2012 and February 2013. This map was generated using Generic Mapping Tools (version 5; http://gmt.soest.hawaii.edu).
Analysis of bacterial community composition was based on OTUs with 97% nucleotide sequence identity and matrices of Bray-Curtis similarity. Overall, bacterial communities differed between coral and seawater samples, both in summer (2-way crossed ANOSIM, R = 0.653, P < 0.001) and winter (2-way crossed ANOSIM, R = 0.769, P < 0.001; Fig. 2a). Community compositions also differed between samples collected in summer and winter (Fig. 2b). In coral samples, variation was more obvious between seasons (2-way nested ANOSIM, R = 0.904, P = 0.002), compared to that under various temperature treatments (2-way nested ANOSIM, R = 0.23, P = 0.003). Bacterial communities were also different between seasons in seawater samples (2-way crossed ANOSIM, R = 0.965, P < 0.001).
NMDS based on Bray-Curtis similarity matrix with complete linkage among samples. All OTUs abundance in samples were nomalised by numbers of sequences in each sample. The 3 biological replicates of coral sample from each treatment were combined into 1 representative sample. Similarities among samples > 10 are circled by grey lines. (a) Bacterial communities varied between coral and seawater samples in both summer (left) and winter (right). (b) In both coral (left) and seawater (right) samples, bacterial communities differed between samples collected in summer versus winter.
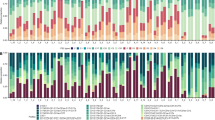
To facilitate detection of differences in bacterial community composition, OTUs assigned to the same bacterial class were combined and presented in a pie chart. Regarding class-level changes in bacterial composition, for samples before temperature treatment, Gammaproteobacteria was dominant (95.2 and 66.2% of relative abundance) in coral samples in both seasons, whereas Alphaproteobacteria was dominant (64.7 and 50.7%) in seawater samples (Fig. 3a). More Alphaproteobacteria and Epsilonproteobacteria (4.1 and 1.9%) were detected in winter than in summer samples. However, when coral samples were under thermal treatment (Fig. 3b), Epsilonproteobacteria and Deltaproteobacteria increased, particularly in winter samples with heat stress (10.3% after 48 h at 30 °C and 15.9% after 48 h at 33 °C), whereas after 48 h of heat stress, Gammaproteobacteria decreased in all coral samples from both seasons (minimum relative abundance = 26.5%).
Bacterial composition (class level) in samples. (a) Before treatment (0 h), Gammaproteobacteria were dominant in coral samples and dominant Alphaproteobacteria in seawater samples in both summer and winter. (b) Changes in the bacterial community under temperature stress from 12 to 48 h. For coral samples in winter, Epsilonproteobacteria and Deltaproteobacteria increased under heat stress.
Diversity indices of bacterial communities
Before treatment (0 h), coral samples nearly always had lower diversity, evenness or richness than seawater samples in both seasons (Supplementary Table S1a). Winter coral samples had greater richness and diversity than summer coral samples. However, after heat treatment, diversity in summer coral samples was highest among all samples (Fig. 4, Supplementary Fig. S1 and Table S1b).
Average of alpha-diversity in bacteria communities over time in 2 seasons. Squares and light grey represent summer samples, whereas circles and dark grey represent winter samples. Differences (P < 0.05) among samples are indicated by different letters in each panels. Averages and error bars of triplicate samples are in Supplementary Figure S1.
Diversity fluctuated among sampling times during treatment. Under heat treatment (30 and 33 °C), OTU number of summer coral samples decreased between 12 and 24 h, but rapidly increased after 24 h. Furthermore, diversity of samples under cold treatment (10 °C) had a different dynamic pattern, as it increased between 12 and 24 h of treatment, but decreased after 24 h. In general, dynamics of alpha-diversity in coral-associated bacterial communities had distinctively different patterns between heat and cold stresses.
Influence factors for the variation of bacterial composition in coral samples
Three factors were crucial in this study, namely season, temperature and duration of treatment. To determine which of these factors were associated with variation in bacterial composition, canonical correspondence analysis (CCA; Fig. 5) was performed. Eigen values of constrained axes for season, temperature and time were 0.499, 0.233 and 0.197, respectively. Therefore, season was most closely associated with bacterial composition of coral samples, with the majority of OTUs clearly separated into summer or winter groups.
CCA analysis for effects of 3 major factors (temperature, time and season) on distribution of bacteria in coral samples. All OTUs (crosses) in coral samples (circles) are included and distribution of OTUs represents differences between seasons. The scales of the 3 factors are shown in top and right axes. Light grey ellipses indicate most OTUs distributed in each season.
Transition of bacterial taxa in coral samples under thermal stress
To evaluate dominant OTUs, a heatmap was prepared to display relative abundance of the top 20 OTUs in all coral samples and their taxonomic affiliations, based on Greengenes database and the blast result of NCBI (Fig. 6). Our Bray-Curtis similarity results showed the presence of 3 clusters, which corresponded to samples with a similar bacterial community as control samples, samples under heat stress (“Heat” group in Fig. 6), and samples under cold stress (“Cold” group in Fig. 6), respectively. Samples within a cluster had Bray-Curtis similarity >20, except the 3 samples under cold stress. Moreover, the cluster, including control samples, was further separated into 2 sub-clusters, each with Bray-Curtis similarity >40, that corresponded to samples collected in summer and in winter (“Summer” and “Winter” groups in Fig. 6). Therefore, coral samples were divided into 4 groups: “Summer”, “Winter”, “Cold”, and “Heat”.
Heatmap of top 20 OTUs in coral samples with Bray-Curtis similarity clustering. The top 20 OTUs in the y-axis had >1% accumulative relative abundance in all coral samples. Coral samples in x-axis were clustered based on Bray-Curtis similarity between all OTUs. Coral samples were separated into 4 groups: “Winter”, “Summer”, “Cold”, and “Heat”. Relative abundance of the top 20 OTUs in samples was presented (after transformation with a z-score). Red represents higher abundance, whereas green indicates relative rare abundance in each sample. Sample names in x-axis with different colors indicate temperature of treatment, and duration of treatments in each sample are presented below sample names.
For the “Summer” and “Winter” clusters, composition of bacterial community was dominated by OTU1, affiliated with the bacteria Candidatus Endozoicomonaceae in the Greengenes database and belonging to the genus Endozoicomonas, Family Hahellaceae in SILVA and RDP databases. However, there were more dominant OTUs detected in the winter cluster, OTU3 and OTU6, affiliated with Endozoicomonaceae and Xanthomonadales, respectively.
The “Cold” cluster was comprised of 2 summer samples (S410 and S415) and 1 winter sample (W410). The 2 summer samples were dominated by OTU7 (Pseudoalteromonas), whereas the winter sample was dominated by 2 OTUs, OTU10 (Idiomarinaceae) and OTU28 (Vibrionaceae). Unlike other clusters, this cluster was not fully supported by Bray-Curtis similarity, and no common OTU was detected between summer and winter samples. OTU1 and OTU3 (Endozoicomonaceae) shifted to OTU7 (Pseudoalteromonas) in summer samples. A clear transition of bacterial communities was observed after 24 h treatment at either 15 or 10 °C. Samples S215 and S210 were co-dominated by OTU1 and OTU7. With more prolonged incubation (48 h), OTU1 decreased and summer samples were solely dominated by OTU7. A similar dynamic pattern was also present in winter samples incubated at 10 °C for 24 or 48 h. Co-occurrence of the dominant OTU1, OTU3 (Endozoicomonaceae) and OTU28 (Idiomarinanceae) in sample W210 (at 10 °C for 24 h) shifted to OTU28 and OTU10 (Vibronaceae) that were dominant in sample W410 (10 °C for 48 h).
The last “Heat” cluster was composed of 6 winter samples and only 2 summer samples. The latter were under heat stress at 30 or 33 °C for 48 h (S430, S433). In contrast, the bacterial community of the winter samples in this cluster changed quickly after 12 h at 33 °C (W133). In the beginning of the shift, the dominant bacterial profile changed from OTU1, OTU3 and OTU6 to OTU4 (Arcobacteraceae), followed by OTU13 (Peptostreptococcaceae) that predominated at 24 h, whereas OTU11 (Arcobacteraceae) became dominant after 48 h of incubation at 30 or 33 °C.
Temporal changes in 4 key bacteria at various temperatures
Four dominant bacterial taxa were highly correlated with various temperature treatments (Fig. 6), including Endozoicomonaceae (OTU1, 3, 799, 686), Alteromonadales (OTU7, 8, 10, 17), Vibrionaceae (OTU28, 29), and Arcobacteraceae (OTU4, 11, 20). To characterize dynamics of these 4 bacterial taxa during treatment, xyplots were generated (Supplementary Fig. S3).
Relative abundance of Endozoicomonaceae in summer samples exceeded 80% (Supplementary Fig. S3 and Fig. 7b), and was higher than that in winter samples (only 40% in samples before treatment). Abundance in summer decreased to nearly 0% under heat or cold stress (30 and 33 °C for heat stress and 10 and 15 °C for cold stress) for 48 h, whereas those abundances were >60% at 20 and 25 °C. The rate of decrease was faster under cold treatment than heat treatment in summer samples. In winter samples, the highest relative abundance of Endozoicomonaceae was in samples at 15 °C (68% at 12 h and 26% at 48 h) rather than in samples at 20 or 26 °C. For winter samples under heat treatment >30 °C, relative abundances rapidly decreased to 0% with 24 h incubation, which was faster than winter samples under cold treatment (near 0% at 10 °C for 48 h). This pattern was opposite to summer samples under temperature treatment.
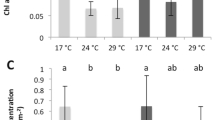
Xyplot for (a) photosynthetic efficiency of Symbiodinium and (b) relative abundance of Endozoicomonaceae in samples under various temperature treatments over time. Average and standard error of the mean of 3 biological replicates are in the y axis, whereas the x axis represents time after treatment.
Compared to Endozoicomonaceae, abundance of Alteromonadales, Arcobacteraceae, and Vibrionaceae was near 0% in samples before treatment (summer or winter; Supplementary Fig. S3). However, Alteromonadales in summer samples increased to >60% under cold stress (10 or 15 °C) at 48 h, whereas Endozoicomonaceae decreased to near 0% after the same incubation. In winter samples, abundance of Alteromonadales did not increase as high as those in summer, but still reached 15% relative abundance under 10 °C with 48 h incubation.
For Vibrionaceae, abundances were very low (<3%) in all summer samples, but increased quickly (up to 35%) within 12 h in winter samples under heat treatment at 30 °C. In addition, under cold stress (10 °C), abundance of Vibrionaceae was maintained at 15% after 24 h (Supplementary Fig. S3).
Relative abundance of Arcobacteraceae that belonged to Epsilonproteobacteria in winter samples, like Vibrionaceae, greatly increased (up to 50%) within 12 h under heat stress (33 °C). After 48 h treatment in winter samples, relative abundance was maintained at >15% in all samples except those at 25 °C, whereas those of Vibrionaceae under heat stress in winter samples were low and decreased after 24 h. However, Arcobacteraceae were low in summer samples (averaged < 3%) except for samples under 30 °C at 48 h (average = 22%). In summary, dynamics of bacterial community composition under temperature stresses with various time treatments were more complex in winter versus summer samples.
Short-term dynamics of Symbiodinium and coral-associated microbes under heat and cold stress
To compare responses of bacteria and Symbiodinium in coral under stress, photosynthetic efficiency (Fv/Fm) and density of Symbiodinium were determined (Fig. 7a and Supplementary Fig. S4). Under heat treatment, both density and photosynthetic efficiency of Symbiodinium were low compared to control (Fv/Fm < 0.2, density < 1000 cells) after 48 h. Photosynthetic efficiency of winter samples was decreased at 24 h under 30 °C; this was sooner than those of the summer samples. On the contrary, under cold treatment, both parameters of Symbiodinium in samples at 10 and 15 °C treatments remained stable, except photosynthetic efficiency of Symbiodinium decreased in summer samples at 10 °C with 48 h incubation (Fig. 6a).
Discussion
This study compared changes between the 2 seasons in Acropora muricata-associated bacterial communities under an array of temperature stresses. Bacterial communities collected in the 2 seasons had distinct patterns. Shifts in the bacterial community under cold stress were slower in winter than in summer, whereas a more rapid shift occurred under heat stress in both seasons. Compared to previous research, the present study had the lowest similarity of bacterial communities between samples of winter and summer (<10 Bray-Curtis similarity using cluster with complete linkage in Fig. 2, or <40 using cluster with average linkage in Fig. 6). To our knowledge, there are only 5 published studies comparing coral-associated bacterial communities between seasons. Two reports had results similar to ours33,37, whereas the other 3 detected limited differences between seasons31,34,38. Apparent inconsistencies among reports for compositional shifts in bacterial community between seasons were likely due to multiple factors, including differences in environments, time, coral species, or methods. Of those factors, environmental variation may have had the biggest impact. In this study, the waters around Penghu are dominated by varying currents between seasons, resulting in a much larger seasonal range in temperature than reefs in other reports, including those in the Mediterranean Sea, the Great Barrier Reef and South Taiwan (tropical region).
According to the meteorological database, cold waters of the China Coastal Current coming from high latitudes cause low temperatures at the Penghu sampling site in winter (the lowest historical temperature was 12.1 °C; Supplementary Fig. S5) compared to Cheng-Kung Township on the east coast of Taiwan (the lowest recorded temperature was 19.9 °C in December of 2009). Based on this study, the low temperature likely affected the bacterial community in A. muricata, although more evidence should be collected. The bacterial group Xanthomonadales predominated in winter samples (Fig. 6), but not in summer samples, and contributed to major variations between seasons. This bacterial group was also abundant in the region of a deep coral, Lophelia pertusa at a high-latitude area, the SE Rockall Bank in the NE Atlantic (55°28′55.2″N 15°48′19.8″W), with a water temperature <10 °C at the sampling site39 (van Bleijswijk et al., 2015).
Our study is 1 of only 2 recent studies to determine dynamics of bacterial community in corals under heat stress <1 d in duration. Ziegler and coworkers (2017)30 used Acropora hyacinthus samples from back-reef pools of Ofu Island of the U.S. National Park of American Samoa, and demonstrated the coral-associated bacterial community responded rapidly (within 20 h) to heat stress. In the present study, the response of bacterial community in Acropora muricata samples was also rapid and varied between seasons, within 12 h in winter and 48 h in summer under acute heat stress (Fig. 6 and Supplementary Fig. S3).
There were 4 dominant and sensitive bacterial groups for early responses to temperature stresses (Supplementary Fig. S3), namely Endozoicomonaceae, Alteromonadales, Vibrionaceae, and Arcobacteracea, in terms of changes in relative abundance. Those 4 bacterial groups were likely associated with changes in coral holobiont status under stress.
Endozoicomonaceae is a prominent group, likely associated with coral health4,40,41,42, and also a common marine invertebrate-associated bacterium, present in various reefs worldwide43. In our study, coral-associated Endozoicimonaceae were sensitive to thermal stress. The relative abundance of these bacteria significantly decreased under either heat (30 and 33 °C) or cold stress (10 and 15 °C), consistent with previous research. For instance, relative abundance of Endozoicomonas has been reported to largely decrease in response to abiotic stresses, e.g. temperature increases44, ocean acidification45, or anthropogenic impacts (sedimentation and sewage)46. Furthermore, bleaching of Acropora corals in the Great Barrier Reef caused substantial disappearance of Endozoicomonas 11. Collectively, based on present results and previous reports, we inferred that rapid changes in Endozoicomonas could be a robust bioindicator for changes in the coral holobiont.
Alteromonadales was associated with stressed or diseased corals47 and was regarded as an indicator of aged mucus of coral or mucus in disturbed coral4. Interestingly, Pseudoalteromonas, belonging to Alteromonadales, inhibited Vibrio 3,48 and had antibacterial activity in the coral holobiont49. There was a high relative abundance of Pseudoalteromonas in summer samples under cold stress (OTU7 in S410, S415, S210 and S215 samples; Fig. 6) and concurrently, low relative abundance of Vibrio (OTU28 and 29) in these 4 summer samples.
Vibrio bacteria are highly associated with corals under stress50. Moreover, increased Vibrio was often reported when Acropora spp. were under stress46,51, especially heat stress11,44. In the present study, relative abundance of Vibrionaceae in winter increased not only under heat stress but also under cold stress (Supplementary Fig. S3). Although Vibrio have been widely reported to be highly associated with coral bleaching or temperature stress11, their relative abundance in this study was not as high as other pathogens (i.e., Arcobacteraceae; Supplementary Fig. S3).
All 3 OTUs that belonged to Arcobacteraceae in the top 20 dominant OTUs (OTU4, 11, and 20) were all assigned to the genus Arcobacter in the NCBI database. These 3 OTUs were detected in the winter coral samples with a low relative abundance at the beginning, but they increased at subsequent sampling times. In the present study, Arcobacter was a stress-associated bacterial group and quickly grew in coral under thermal stress. Arcobacter have been reported in association with coral diseases, including White Syndrome and Brown Band Disease in Acropora muricata in the Great Barrier Reef47 and White Plague Disease in Montastraea faveolata in the Caribbean Sea52. Perhaps Arcobacter is an opportunistic bacterium in coral rather than a specific pathogen.
The link between stability of a bacterial community and adjustable heat tolerance after acclimation was recently proposed29,30. The present study also supports a similar link between seasonal acclimation and thermal tolerance range in bacterial community. As bacteria were under cold stress (15 and 10 °C), shifting of dominant OTUs occurred at 10 °C in winter, lower than that in summer (15 °C) (Fig. 6). With 15 °C treatment, the relative abundance of Endozoicomonaceae was highest among all other treatments in winter, but these bacteria decreased to near undetectable levels in summer (Fig. 7b). Therefore, bacteria in winter samples more readily acclimated themselves to a colder temperature than those in summer samples. Similarly, seasonal acclimation was also detected under heat stress. Shifting of dominant bacteria was more quickly in winter samples than in the summer samples under heat stress (Fig. 6). In winter, the bacterial community changed quickly into the “Heat” group (heat stress) when exposed to 33 °C for just 12 h, whereas summer samples were grouped into the “Heat” group only after treatment for 48 h. In addition, increased relative abundance of Arcobacter (stress-associated bacteria) was slower in summer than winter (48 versus 12 h; Supplementary Fig. S3).
Notably, there was a high relative abundance of Endozoicomonaceae between 15 and 30 °C (Fig. 7b and Supplementary Fig. S3). This temperature range was different from the optimal growth temperature of Endozoicomonas spp., reported as 25 to 30 °C53,54,55,56. Above 30 °C, relative abundance of Endozoicomonaceae rapidly decreased in this study (Fig. 7b). However, these bacteria in winter samples were still largely detected at a temperature (15 °C for 48 h) colder than the lowest optimal temperature (25 °C). Therefore, we hypothesize that there was some cold acclimation in this bacterial group (OTU1 and OTU3 in Fig. 6). The inconsistency of the temperature ranges might have been caused by species variation and perhaps other factors, including cold acclimation. It was noteworthy that Endozoicomonaceae were also widely detected in cold habitats. Although there is no information about these bacteria in hexacorals under cold stress, Endozoicomonaceae dominates in azooxanthellate octocorals in cold-water environments31,57. For example, Endozoicomonaceae were detected in a deep-sea coral gorgonian Anthothela sp., collected at 400-600 m, where seawater temperatures were 5.5 to 7.1 °C58. High relative abundance of Endozoicomonaceae through various seasons was also reported in another cold-water gorgonian Eunicella verrucosa collected at 15-27 m along the south-west coast of England with a seawater temperature 13.9 to 17.0 °C34. Collectively, it was apparent that Endozoicomonaceae have potential to acclimate to a cold environment.
In this study, Acropora muricata-associated bacteria in Penghu had different tolerance ranges to thermal stress between summer and winter after seasonal acclimation (Supplementary Fig. S5). We further inferred that the response and tolerance ranges of coral-associated bacteria were associated with short-term “memory” in thermal history. Tolerance ranges in the 2 seasons were estimated to be >15 °C to <30 °C in summer, and >10 °C to <30 °C in winter. To better define the tolerance ranges for Acropora-associated bacterial community in Penghu, finer scale of temperature treatments is needed. Regardless, we concluded that the tolerance range may cover 15 to 25 °C in winter and 20 to 26 °C in summer, as the bacterial community remained similar to the control sample in the same cluster (Fig. 6 and Supplementary Fig. S5a).
This estimated temperature range based on community stability differed from average temperatures in summer and winter in the thermal history of the sampling site (Supplementary Fig. S5a)59. The upper limit of the estimated tolerance range for coral under heat stress in summer was 26 °C, which was lower than the average temperature in summer in Penghu (27.1 °C from the averages of June, July, and August). On the contrary, the lower limit of the estimated tolerance range for coral under cold stress was 15 °C, lower than the average temperature in winter (February, March, and April), 18.8 °C. Therefore, both ranges did not match to one another (Supplementary Fig. S5a).
However, the estimated temperature ranges of bacterial community aligned well with the seawater temperature profile 3 mo before the sampling was done, in either winter or summer (orange or blue square in Supplementary Fig. S5b), suggesting that their response was more likely to associate with the temperature history within the previous 3 mo, rather than the entire thermal history (i.e. from 12.1 to 30.2 °C). This apparent short-term acclimation was also supported by another study30, where the ability of thermal tolerance disappeared 1 y after corals were transplanted from a high-temperature region to lower temperature region. We inferred that the short-term “memory” of temperature in a coral holobiont was important, at least for stabilization of the bacterial community.
We modified our hypothesis that the tolerance temperature range for the bacterial community is adjustable in different seasons, based on acclimation to thermal history within the short-term (previous 3 mo in this study). In future studies, smaller differences in temperature and interval, and more prolonged stress treatments should provide a clearer picture of the thermal tolerance range for coral-associated bacteria.
Conclusions
In this study, variations of microbial communities associated with corals under both heat and cold stress for up to 2 d were characterized in 2 seasons, summer and winter. This was apparently the first study to document responses of the coral-associated bacterial community to cold stress. In this study, the bacterial community in Acropora from Penghu varied according to season. Variations affected the response and tolerance range of stable homeostasis in coral-associated bacteria under temperature stress. The tolerance range of coral-associated bacteria under various temperatures was related to short-term thermal history (3 mo before stress). Finally, observed shifts in the abundance and diversity of bacterial communities were further considered in the context of known interactions between the coral holobiont and associated bacteria.
Materials and Methods
Study site, sample collection and experimental design
Summer and winter experiments were conducted in June 2012 and February 2013, respectively, in sub-tropical, Penghu, Taiwan (Fig. 1 and Table 1). The map in Fig. 1 was generated using the Generic Mapping Tools (GMT v.5; http://gmt.soest.hawaii.edu). Branches from 5 colonies of Acropora muricata were collected from Wukan (23°32′38.9″N, 119°37′32.3″East). Branches from each colony were used to make 2–3 cm nubbins that were attached (with epoxy) to 2-ml Eppendorf tubes. All nubbins were maintained in a 26 °C (summer) or 20 °C (winter) acclimation tank for 2–3 d prior to the onset of experiment period. After acclimation, 3 coral nubbins each from 5 colonies and 50 mL seawater were sampled, whereas remaining nubbins were distributed into 6 experimental seawater tanks (15 × 15 × 15 cm, ~3 l capacity) maintained at temperatures of 10, 15, 20, 26, 30 or 33 °C respectively. Each tank was placed in a bigger tank (that was used as a water bath). Circulation inside experimental tanks was a closed system, with 2/3 of the water replaced with filtered (0.2 μm) seawater every 24 h. Temperature of the water bath was maintained by Hailea 150 A aquarium chillers (set to 26 and 20 °C in summer and winter, respectively). Each of the 6 experimental tanks contained small aquarium heater rods to maintain temperatures. Each tank had at least 60 nubbins (12 nubbins from each of 5 colonies), with 3 nubbins from different colonies randomly sampled after 12, 24, and 48 h, respectively. A total of 57 nubbins each during summer and winter experiment were sampled and fixed in absolute ethanol and 19 seawater samples each during summer and winter experiment were collected and filtered immediately through cellulose acetate membranes with 0.2-μm pores (Adventec, Tokyo, Japan), transported at 4 °C and stored at −20 °C prior to DNA extraction.
Symbiodinium density and photosynthetic efficiency
To calculate density of Symbiodinium per unit coral surface area, coral surface areas were measured using an aluminum foil method60. Symbiodinium in hospite were removed from coral branches using an airbrush with filtered seawater (0.2-µm mesh filter) and fixed in 4% formaldehyde. The solution of Symbiodinium and coral tissue was centrifuged (4000 × g for 10 min) to separate coral tissue from Symbiodinium cells. Fresh-filtered seawater was added to the resulting pellet, mixed thoroughly and centrifuged (4000 × g for 10 min). For counting Symbiodinium cells, 3 aliquots (1 ml each) were used and cells were counted (hemocytometer and light microscope) 3 times for each aliquot. The average number of cells was converted to Symbiodinium density normalized to coral surface area (cm2).
The dark-adapted maximum quantum yield of photosystem II (Fv/Fm) was measured using a Walz® Junior-pulse amplitude-modulated (PAM) flurometer with a 0.8 s saturating pulse of >4500 µmol photons m−2s−1 and gain value of 12. Dark-adapted yield values (Fv/Fm) were determined from 5 coral nubbins from each colony, collected 2 h after sunset. Analysis of variance (ANOVA) was used to detect variations among coral samples.
Total DNA extraction, amplification of V6-V8 region in bacterial 16S rRNA gene, and tagging PCR
Coral nubbins were washed with TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8), frozen in liquid nitrogen, and homogenized using a sterile mortar and pestle. Coral powder and seawater samples were transferred into TE buffer for total DNA extraction using a modified CTAB method61,62.
Amplification of the 16S ribosomal RNA gene was performed by PCR using a pair of universal bacterial primers: 968 F (5′-AACGCGAAGAACCTTAC-3′) and Uni1391R (5′-ACGGGCGGTGWGTRC-3′) specifically designed for the bacterial V6-V8 hypervariable region42,63,64. The PCR was performed in 50-μL reaction volumes, consisting of 1.5 U TaKaRa Ex Taq (Takara Bio, Otsu, Japan), 1X TaKaRa Ex Taq buffer, 0.2 mM deoxynucleotide triphosphate mixture (dNTP), 0.2 mM of each primer, and 50 to 150 ng purified total DNA. The thermocycler was set to an initial step of 94 °C for 5 min, 30 cycles of 94 °C for 30 s, 52 °C for 20 s and 72 °C for 45 s, and a final extension at 72 °C for 10 min. Target DNA bands (~420 bp) were electrophoresed on a 1.5% agarose gel, and specific bands were eluted using a QIAEX II Gel Extraction Kit (Qiagen, Valencia, CA, USA).
To tag each bacterial V6-V8 amplicon with a unique barcode sequence, tag primers were designed with 4 overhanging nucleotides at 5′ ends of common primers. The tagging reaction was performed with a 5-cycle PCR, with a reaction program of 94 °C for 30 s, 52 °C for 20 s, and 72 °C for 45 s with the modified primers. Amplicons were purified by the same gel elution method described above. Concentration of DNA was determined with a Qubit dsDNA HS assay (Invitrogen, Carlsbad, CA, USA).
Illumina MiSeq pair-end sequencing and data processing
Summer and winter V6-V8 amplicons were pooled into 2 independent libraries for 2 × 250 pair-end reads in Illumina MiSeq sequencing (Yourgene Bioscience, Taipei, Taiwan). Overall, 581,860 and 567,874 raw reads were obtained after pair-end reads were merged. After sorting and trimming, high-quality reads were extracted using MOTHUR65 with the following criteria: (1) read lengths between 380 and 450 bps; (2) average quality score >27; (3) homopolymer length <8 bp; and (4) removal of reads with any ambiguous base (N). Thereafter, the 4 nucleotide tags and primer sequences were removed. Chimeric reads were inspected and eliminated by UCHIME66 with USEARCH v7.0.1090 (parameters: reference mode, rdp_gold database, and mindiv of 5). Qualified sequences were retained for subsequent analyses.
For operational taxonomic units (OTU) analysis, qualified reads were pooled and assigned OTUs with a cutoff value at 97% identity by the UPARSE pipeline67. Each OTU was classified with a bootstrap value set to 0.8 using RDP classifier68 implemented in MOTHUR. The alignment template was the SILVA reference database (release 102). Sequences were annotated with taxonomies in the Greengenes database. There were 114 OTUs that belonged to unclassified bacteria, according to the database of Greengenes taxonomy. These OTUs were further examined and blasted with the RDP Seqmatch tool. In subsequent analyses, 27 OTUs affiliated to bacteria with >0.8 ab_score were retained, whereas the remaining 87 OTUs were removed.
Data analyses
Diversity indices of microbial communities were presented without singleton OTU after the UPARSE pipeline, except for calculation of richness. Diversity indices of Shannon-Weaver and Gini-Simpson, values of richness and evenness were calculated using MOTHUR software. Average and standard error (3 biological replicates) of alpha-diversity were calculated. Two-way analysis of variance and Tukey’s HSD comparisons were conducted to detect significant differences in diversity among sampling times and between seasons.
Three biological replicates from each treatment were combined into one representative sample and relative abundances of each classified bacterial OTU in individual samples were calculated by dividing the sum of sequence numbers in the sample37. For composition of bacterial community in each sample, OTUs assigned to the same class were combined, and relative abundances of each class in an individual sample were presented in a pie chart. It is noteworthy that primer bias from different regions of the 16S rRNA gene may cause variation in analysis of relative abundance69.
To analyze beta-diversity and determine relationships of bacterial communities between samples, relative abundances of OTUs in individual samples were incorporated into a matrix to estimate a distance matrix (Bray-Curtis distance). Then, the result was presented in a non-Metric multi-Dimensional Scaling (nMDS) analysis with complete linkage using Primer 6 software (PRIMER-E, Lutton, Ivybridge, UK), a plot of Canonical Correspondence Assay (CCA), and a heatmap with hierarchical clustering (CLUSTER) by average linkage using an R program (https://www.r-project.org/).
In the nMDS analysis, differences in bacterial community composition among samples were tested by ANOSIM analysis. Differences among bacterial communities in coral samples from various temperatures were tested within seasons, using 2-way nested ANOSIM analysis, whereas differences among bacterial communities in seawater samples from different temperatures and seasons were determined using 2-way crossed ANOSIM.
The 20 most abundant OTUs were selected for both CCA and heatmap analyses. In the heatmap, relative abundance data of the 20 OTUs were presented after transformation to z-scores, and their taxonomies identified and listed. Clustering of coral samples along the x-axis was based on Bray-Curtis similarity matrix with average linkage using relative abundance of all OTUs in each coral sample, although only the relative abundances of the top 20 OTUs were shown.
Xyplot for Endozoicomonaceae, Alteromonadels, Vibrionaceae, and Arcobacteraceae
Average and standard error (3 biological replicates) of relative abundances of sequences of Endozoicomonaceae, Alteromonadels, Vibrionaceae, and Arcobacteraceae were calculated. Analysis of variance (ANOVA) was used to determine significance of variation among coral samples.
Data Accessibility
Multiplex sequenced reads (bacterial 16S V6-V8 region) were deposited in the NCBI Sequence Read Archive (accession number PRJNA379752).
References
Rohwer, F., Seguritan, V., Azam, F. & Knowlton, N. Diversity and distribution of coral-associated bacteria. Mar. Ecol. Prog. Ser. 243, 1–10 (2002).
Rosenberg, E., Koren, O., Reshef, L., Efrony, R. & Zilber-Rosenberg, I. The role of microorganisms in coral health, disease and evolution. Nat. Rev. Microbiol. 5, 355–362 (2007).
Thompson, J. R., Rivera, H. E., Closek, C. J. & Medina, M. Microbes in the coral holobiont: partners through evolution, development, and ecological interactions. Front. Cell Infect. Microbiol. 4, 176, https://doi.org/10.3389/fcimb.2014.00176 (2015).
Glasl, B., Herndl, G. J. & Frade, P. R. The microbiome of coral surface mucus has a key role in mediating holobiont health and survival upon disturbance. ISME J. 10, 2280–2292 (2016).
Brown, B. E. Coral bleaching: causes and consequences. Coral Reefs 16, S129–S138 (1997).
Kleypas, J. A., McManus, J. W. & Menez, L. A. B. Environmental limits to coral reef development: Where do we draw the line? Am. Zool. 39, 146–159 (1999).
Baird, A. H., Bhagooli, R., Ralph, P. J. & Takahashi, S. Coral bleaching: the role of the host. Trends Ecol. Evol. 24, 16–20 (2009).
Hoegh-Guldberg, O. Climate change, coral bleaching and the future of the world’s coral reefs. Mar. Freshwater. Res. 50, 839–866 (1999).
Littman, R., Willis, B. L. & Bourne, D. G. Metagenomic analysis of the coral holobiont during a natural bleaching event on the Great Barrier Reef. Environ. Microbiol. Rep. 3, 651–660 (2011).
Thurber, R. V. et al. Metagenomic analysis of stressed coral holobionts. Environ. Microbiol. 11, 2148–2163 (2009).
Bourne, D., Iida, Y., Uthicke, S. & Smith-Keune, C. Changes in coral-associated microbial communities during a bleaching event. ISME J. 2, 350–363 (2008).
Littman, R. A., Bourne, D. G. & Willis, B. L. Responses of coral-associated bacterial communities to heat stress differ with Symbiodinium type on the same coral host. Mol. Ecol. 19, 1978–1990 (2010).
Sunagawa, S., Woodley, C. M. & Medina, M. Threatened corals provide underexplored microbial habitats. PLoS ONE 5, e9554, https://doi.org/10.1371/journal.pone.0009554 (2010).
Hoegh-Guldberg, O. & Fine, M. Low temperatures cause coral bleaching. Coral Reefs 23, 444–444 (2004).
Hoegh-Guldberg, O. et al. Coral bleaching following wintry weather. Limnol. Oceanogr. 50, 265–271 (2005).
Saxby, T., Dennison, W. C. & Hoegh-Guldberg, O. Photosynthetic responses of the coral Montipora digitata to cold temperature stress. Mar. Ecol. Prog. Ser. 248, 85–97 (2003).
Kavousi, J., Parkinson, J. E. & Nakamura, T. Combined ocean acidification and low temperature stressors cause coral mortality. Coral Reefs 35, 903–907 (2016).
Kemp, D. W. et al. Catastrophic mortality on inshore coral reefs of the Florida Keys due to severe low-temperature stress. Glob. Chang. Biol. 17, 3468–3477 (2011).
Roth, M. S., Goericke, R. & Deheyn, D. D. Cold induces acute stress but heat is ultimately more deleterious for the reef-building coral Acropora yongei. Sci. Rep. 2, 240, https://doi.org/10.1038/srep00240 (2012).
Roth, M. S. & Deheyn, D. D. Effects of cold stress and heat stress on coral fluorescence in reef-building corals. Sci. Rep. 3, 1421, https://doi.org/10.1038/srep01421 (2013).
Brown, B. E., Clarke, K. R. & Warwick, R. M. Serial patterns of biodiversity change in corals across shallow reef flats in Ko Phuket, Thailand, due to the effects of local (sedimentation) and regional (climatic) perturbations. Mar. Biol. 141, 21–29 (2002).
Fang, L. S., Huang, S. P. & Lin, K. L. High temperature induces the synthesis of heat-shock proteins and the elevation of intracellular calcium in the coral Acropora grandis. Coral Reefs 16, 127–131 (1997).
Bellantuono, A. J., Hoegh-Guldberg, O. & Rodriguez-Lanetty, M. Resistance to thermal stress in corals without changes in symbiont composition. Proc. Biol. Sci. 279, 1100–1107 (2012).
Middlebrook, R., Hoegh-Guldberg, O. & Leggat, W. The effect of thermal history on the susceptibility of reef-building corals to thermal stress. J. Exp. Biol. 211, 1050–1056 (2008).
Brown, B. E., Downs, C. A., Dunne, R. P. & Gibb, S. W. Exploring the basis of thermotolerance in the reef coral Goniastrea aspera. Mar. Ecol. Prog. Ser. 242, 119–129 (2002).
Palumbi, S. R., Barshis, D. J., Traylor-Knowles, N. & Bay, R. A. Mechanisms of reef coral resistance to future climate change. Science 344, 895–898 (2014).
Schoepf, V., Stat, M., Falter, J. L. & McCulloch, M. T. Limits to the thermal tolerance of corals adapted to a highly fluctuating, naturally extreme temperature environment. Sci. Rep. 5, 17639, https://doi.org/10.1038/srep17639 (2015).
Tang, C. H. et al. Cellular membrane accommodation to thermal oscillations in the coral Seriatopora caliendrum. PLoS ONE 9, e105345, https://doi.org/10.1371/journal.pone.0105345 (2014).
Santos, H. F. et al. Climate change affects key nitrogen-fixing bacterial populations on coral reefs. ISME J. 8, 2272–2279 (2014).
Ziegler, M., Seneca, F. O., Yum, L. K., Palumbi, S. R. & Voolstra, C. R. Bacterial community dynamics are linked to patterns of coral heat tolerance. Nat. Commun. 8, 14213, https://doi.org/10.1038/ncomms14213 (2017).
La Riviére, M., Roumagnac, M., Garrabou, J. & Bally, M. Transient shifts in bacterial communities associated with the temperate gorgonian Paramuricea clavata in the Northwestern Mediterranean Sea. PLoS ONE 8, e57385 (2013).
Lema, K. A., Bourne, D. G. & Willis, B. L. Onset and establishment of diazotrophs and other bacterial associates in the early life history stages of the coral Acropora millepora. Mol. Ecol. 23, 4682–4695 (2014).
Li, J. et al. Bacterial dynamics within the mucus, tissue and skeleton of the coral Porites lutea during different seasons. Sci. Rep. 4, 7320, https://doi.org/10.1038/srep07320 (2014).
Ransome, E., Rowley, S. J., Thomas, S., Tait, K. & Munn, C. B. Disturbance to conserved bacterial communities in the cold-water gorgonian coral Eunicella verrucosa. FEMS Microbiol. Ecol. 90, 404–416 (2014).
Roder, C., Bayer, T., Aranda, M., Kruse, M. & Voolstra, C. R. Microbiome structure of the fungid coral Ctenactis echinata aligns with environmental differences. Mol. Ecol. 24, 3501–3511 (2015).
Cacciapaglia, C. & van Woesik, R. Climate-change refugia: shading reef corals by turbidity. Glob. Chang. Biol. 22, 1145–1154 (2016).
Chen, C. P., Tseng, C. H., Chen, C. A. & Tang, S. L. The dynamics of microbial partnerships in the coral Isopora palifera. ISME J. 5, 728–740 (2011).
Lema, K. A., Willis, B. L. & Bourne, D. G. Amplicon pyrosequencing reveals spatial and temporal consistency in diazotroph assemblages of the Acropora millepora microbiome. Environ. Microbiol. 16, 3345–3359 (2014).
van Bleijswijk, J. D. L. et al. Microbial assemblages on a cold-water coral mound at the SE Rockall Bank (NE Atlantic): interactions with hydrography and topography. Biogeosciences 12, 4483–4496 (2015).
Bourne, D. G., Morrow, K. M. & Webster, N. S. Insights into the coral microbiome: underpinning the health and resilience of reef ecosystems. Annu. Rev. Microbiol. 70, 317–340 (2016).
Ding, J. Y., Shiu, J. H., Chen, W. M., Chiang, Y. R. & Tang, S. L. Genomic Insight into the host-endosymbiont relationship of Endozoicomonas montiporae CL-33(T) with its coral host. Front. Microbiol. 7, 251, https://doi.org/10.3389/fmicb.2016.00251 (2016).
Lee, S. T. M., Davy, S. K., Tang, S. L., Fan, T. Y. & Kench, P. S. Successive shifts in the microbial community of the surface mucus layer and tissues of the coral Acropora muricata under thermal stress. FEMS Microbiol. Ecol. 91, 12, https://doi.org/10.1093/femsec/fiv142 (2015).
Neave, M. J., Apprill, A., Ferrier-Pages, C. & Voolstra, C. R. Diversity and function of prevalent symbiotic marine bacteria in the genus Endozoicomonas. Appl. Microbiol. Biotechnol. 100, 8315–8324 (2016).
Tout, J. et al. Increased seawater temperature increases the abundance and alters the structure of natural Vibrio populations associated with the coral Pocillopora damicomis. Front. Microbiol. 6, 432, https://doi.org/10.3389/fmicb.2015.00432 (2015).
Webster, N. S. et al. Host-associated coral reef microbes respond to the cumulative pressures of ocean warming and ocean acidification. Sci. Rep. 6, 19324, https://doi.org/10.1038/srep19324 (2016).
Ziegler, M. et al. Coral microbial community dynamics in response to anthropogenic impacts near a major city in the central Red Sea. Mar. Pollut. Bull. 105, 629–640 (2016).
Sunagawa, S. et al. Bacterial diversity and White Plague Disease-associated community changes in the Caribbean coral Montastraea faveolata. ISME J. 3, 512–521 (2009).
Rypien, K. L., Ward, J. R. & Azam, F. Antagonistic interactions among coral-associated bacteria. Environ. Microbiol. 12, 28–39 (2010).
Shnit-Orland, M., Sivan, A. & Kushmaro, A. Antibacterial activity of Pseudoalteromonas in the coral holobiont. Microb. Ecol. 64, 851–859 (2012).
O’Brien, P. A., Morrow, K. M., Willis, B. & Bourne, D. Implications of ocean acidification for marine microorganisms from the free-Living to the host-associated. Front. Mar. Sci. 3, 1–14 (2016).
Kvennefors, E. C. E., Sampayo, E. M., Ridgway, T., Barnes, A. C. & Hoegh-Guldberg, O. Bacterial communities of two ubiquitous great barrier reef corals reveals both site- and species-specificity of common bacterial associates. PLoS ONE 5, e10401, https://doi.org/10.1371/journal.pone.0010401 (2010).
Sweet, M. & Bythell, J. Ciliate and bacterial communities associated with white syndrome and brown band disease in reef-building corals. Environ. Microbiol. 14, 2184–2199 (2012).
Kurahashi, M. & Yokota, A. Endozoicomonas elysicola gen. nov., sp nov., a gamma-proteobacterium isolated from the sea slug Elysia ornata. Syst. Appl. Microbiol. 30, 202–206 (2007).
Nishijima, M., Adachi, K., Katsuta, A., Shizuri, Y. & Yamasato, K. Endozoicomonas numazuensis sp. nov., a gammaproteobacterium isolated from marine sponges, and emended description of the genus Endozoicomonas Kurahashi and Yokota 2007. Int. J. Syst. Evol. Microbiol. 63, 709–714 (2013).
Pike, R. E., Haltli, B. & Kerr, R. G. Description of Endozoicomonas euniceicola sp. nov. and Endozoicomonas gorgoniicola sp. nov., bacteria isolated from the octocorals Eunicea fusca and Plexaura sp., and an emended description of the genus. Endozoicomonas. Int. J. Syst. Evol. Microbiol. 63, 4294–4302 (2013).
Yang, C. S. et al. Endozoicomonas montiporae sp. nov., isolated from the encrusting pore coral Montipora aequituberculata. Int. J. Syst. Evol. Microbiol. 60, 1158–1162 (2010).
Vezzulli, L., Pezzati, E., Huete-Stauffer, C., Pruzzo, C. & Cerrano, C. 16SrDNA pyrosequencing of the Mediterranean gorgonian Paramuricea clavata reveals a link among alterations in bacterial holobiont members, anthropogenic influence and disease outbreaks. PLoS ONE 8, e67745, https://doi.org/10.1371/journal.pone.0067745 (2013).
Lawler, S. N. et al. Coral-associated bacterial diversity is conserved across two deep-sea Anthothela species. Front. Microbiol. 7, 458, https://doi.org/10.3389/fmicb.2016.00458 (2016).
Central Weather Bureau. Report on typhoons in 2000. (ed. Ministry of Transportation and Communications) 130–162. (Taiwan, 2000).
Marsh, J. A. Primary productivity of reef-building calcareous red algae. Ecology 51, 254–263 (1970).
Hong, M. J., Yu, Y. T., Chen, C. A., Chiang, P. W. & Tang, S. L. Influence of species specificity and other factors on bacteria associated with the coral Stylophora pistillata in Taiwan. Appl. Environ. Microbiol. 75, 7797–7806 (2009).
Wilson, K. Preparation of genomic DNA from bacteria in Current Protocols in Molecular Biology (ed. John Wiley & Sons, Inc.) Unit2.4; doi:https://doi.org/10.1002/0471142727.mb0204s56 (2001).
Jorgensen, S. L. et al. Correlating microbial community profiles with geochemical data in highly stratified sediments from the Arctic Mid-Ocean Ridge. Proc. Natl. Acad. Sci. USA 109, E2846–2855 (2012).
Nubel, U. et al. Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J. Bacteriol. 178, 5636–5643 (1996).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, vommunity-dupported doftware for fescribing and vomparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1 (2013).
Acknowledgements
This work was financially supported by a grant from the Ministry of Science and Technology of Taiwan (105–2611-M-001–004) and Biodiversity Research Center, Academia Sinica, Taiwan. S.K. thanks Nien-Yun Sophie Cheng and Laura del Can~o Gonza ́lezfor their help in performing temperature stress experiments.
Author information
Authors and Affiliations
Contributions
S.K. and C.C. generated the research concept and S.K. designed the experiments. S.K., H.C. and H.H. collected samples. S.L., P.C., H.C. and S.K. performed the experiments. J.S. and P.C. analyzed the data. J.S. elaborated the figures and tables. J.S. and S.T. contributed to writing the manuscript. S.T. and S.K. critically revised this article. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shiu, JH., Keshavmurthy, S., Chiang, PW. et al. Dynamics of coral-associated bacterial communities acclimated to temperature stress based on recent thermal history. Sci Rep 7, 14933 (2017). https://doi.org/10.1038/s41598-017-14927-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14927-3
This article is cited by
-
Comparative genomics: Dominant coral-bacterium Endozoicomonas acroporae metabolizes dimethylsulfoniopropionate (DMSP)
The ISME Journal (2020)
-
The coral Platygyra verweyi exhibits local adaptation to long-term thermal stress through host-specific physiological and enzymatic response
Scientific Reports (2019)
-
Characterization of coral-associated microbial aggregates (CAMAs) within tissues of the coral Acropora hyacinthus
Scientific Reports (2019)
-
Comparison of microbiomes of cold-water corals Primnoa pacifica and Primnoa resedaeformis, with possible link between microbiome composition and host genotype
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.