Abstract
Paladin (Pald1, mKIAA1274 or x99384) was identified in screens for vascular-specific genes and is a putative phosphatase. Paladin has also been proposed to be involved in various biological processes such as insulin signaling, innate immunity and neural crest migration. To determine the role of paladin we have now characterized the Pald1 knock-out mouse in a broad array of behavioral, physiological and biochemical tests. Here, we show that female, but not male, Pald1 heterozygous and homozygous knock-out mice display an emphysema-like histology with increased alveolar air spaces and impaired lung function with an obstructive phenotype. In contrast to many other tissues where Pald1 is restricted to the vascular compartment, Pald1 is expressed in both the epithelial and mesenchymal compartments of the postnatal lung. However, in Pald1 knock-out females, there is a specific increase in apoptosis and proliferation of endothelial cells, but not in non-endothelial cells. This results in a transient reduction of endothelial cells in the maturing lung. Our data suggests that Pald1 is required during lung vascular development and for normal function of the developing and adult lung in a sex-specific manner. To our knowledge, this is the first report of a sex-specific effect on endothelial cell apoptosis.
Similar content being viewed by others
Introduction
Paladin or Pald1 is a phosphatase-domain containing protein and we have identified it in two independent screens for novel regulators of angiogenesis and vascular function1,2. Pald1 expression is prominent in developing endothelial cells during early embryonic development and, in certain vascular beds, Pald1 expression shifts to mural cells as the vasculature matures, e.g. in the CNS3. Furthermore, Pald1 was also found in hematopoietic cells and other non-vascular cells, such as neural crest cells3,4. A role of Pald1 as a regulator for neural crest cell formation and migration in the chick embryo has been shown by morpholino knock-down experiments. It was suggested that Pald1 would not require any catalytic activity for its role in neural crest migration since mutation of the putative catalytic cysteine to serine still affected neural crest migration after over expression4.
Pald1 has also been shown to negatively regulate expression and phosphorylation of the insulin receptor, as well as phosphorylation of its downstream target kinase Akt in cell culture, even though no phosphatase activity could be detected in vitro 5. Pald1 has therefore been proposed to be a pseudo-phosphatase, and as such been implicated to indirectly regulate cell signaling5,6,7. However, others have predicted that Pald1 can possess catalytic activity, even though this have not been shown experimentally yet8. Furthermore, Pald1 has been suggested to affect another type of receptor signaling, acting as a negative regulator of Toll-like receptor 99.
While there is a strong expression of Pald1 in both the prenatal and adult lung3, its role in lung development or function is unknown. A central role for angiogenesis has been proposed for lung diseases like chronic obstructive pulmonary disease (COPD) and emphysema10. COPD is a chronic progressive disease characterized by clinical symptoms such as dyspnea and coughing, due to obstruction of the airways. COPD is a major cause of death worldwide and it is estimated that COPD will become the third leading cause of death by 202011,12. COPD is frequently accompanied by emphysema, i.e. the destruction of the alveolar walls and the consequential dilation of the distal airways10. Alterations of mRNA and protein expression of vascular endothelial growth factor A (VEGF-A) and its receptor VEGF receptor 2 (VEGFR2) are associated with patients exhibiting COPD and emphysema13. Experimental disruption of VEGF-A/VEGFR2 signaling has also been shown to be sufficient to induce emphysema and COPD-like changes in mice and rats through apoptosis14,15,16. There are sex-specific differences in COPD and emphysema in humans where subsets of women develop more severe emphysema than men. Female smokers with early onset COPD or severe emphysema have smoked significantly less than their male counterparts17. It has also been shown that female mice develop earlier and more severe emphysema than male mice in response to cigarette smoke18. However, the mechanisms responsible for this apparent sex-linkage remain unclear.
Here we describe the phenotype of Pald1 +/− and Pald1 −/− mice19,20 and characterize lung defects linking Pald1 to sex-specific endothelial cell apoptosis, development of emphysema and COPD-like changes in females. To our knowledge, this is the first report of a sex-specific effect on endothelial cell apoptosis.
Results
Pald1 −/− mice exhibit an obstructive lung phenotype
In order to characterize the Pald1 −/− mouse, we performed a wide array of behavioral, physiological and biochemical tests at the German Mouse Clinic (GMC). Although most of the results turned out normal for Pald1 −/− mice (Table S1 and Figure S1), it was noteworthy that female, but not male, Pald1 −/− mice exhibited increased lung volumes and compliance and decreased resistance, which are changes also observed in elastase-induced emphysema mouse model21 (Table 1).
Paladin is expressed broadly in the adult lung
We have previously shown that Pald1 protein is abundant in the mesenchymal compartment of embryonic lungs and that Pald1 is also highly expressed in the adult lung3. Single cell sorting and sequencing from mouse embryonic lung strongly supports the conclusion that Pald1 is exclusively expressed in endothelial cells during embryonic development22,23.
To begin to understand the role of Pald1 in postnatal lung development and function, we mapped Pald1 expression in detail. We analyzed the expression at three distinct time points during postnatal development and in the adult, postnatal day 5 (P5), 4 and 19 weeks of age as it covers main aspects of postnatal development of the major gas exchange unit of the lungs – the alveoli. The tremendous increase in gas exchange surface that occurs postnatally by formation of alveoli is accelerated from P5 until 2 weeks of age and peaks at 5–6 weeks. It is then stable until 40 weeks of age, and subsequently there is a gradual loss of alveoli. The decrease in alveolar septal thickness, that also facilitates gas exchange, continues from P5 until 4 weeks of age. Therefore, we decided to map the expression of Pald1 and assess the phenotype of Pald1 −/− animals at P5, 4 and 19 weeks of age, to cover the dynamic process of postnatal lung development24. First we isolated single endothelial cells and pneumocytes type II from lungs using mT/mG reporter mice expressing Cre-recombinase under the endothelial Tie2 promoter and under the pneumocyte type II-specific surfactant protein C (SPC) promoter, respectively25,26. mRNA isolation and quantitative PCR for Pald1 showed that Pald1 is expressed in endothelial cells, but also in type II pneumocytes (Fig. 1a). To map the expression of Pald1 at cellular resolution, we took advantage of the LacZ reporter of Pald1 knock-out construct3. Murine Pald1 heterozygous (Pald1 LacZ/+) and homozygous mutant (Pald1 LacZ/LacZ) lungs were harvested at postnatal day (P) 5, week 4 and week 19. The LacZ reporter indicated a broad expression of Pald1 in the murine lung in both males and females across all ages analyzed apart from bronchiolar epithelium. LacZ expression pattern was the same in Pald1 LacZ/+ and Pald1 LacZ/LacZ lungs (Fig. 1b and Figure S2). Combining LacZ staining with antibodies specific for each of the main cell types in the alveoli revealed that Pald1 can be detected at all stages in the endothelial cells forming the capillaries around the alveoli but is less frequently detected in the endothelium of larger vessels (Fig. 2, and Figure S3). Paladin was also detected in vascular smooth muscle cells, macrophages, pneumocytes type I and II and Platelet-derived Growth Factor Receptor α (PDGFRα)-positive mesenchymal cells (Fig. 2). However, the surfactant protein positive type II pneumocytes were not LacZ positive at P5.
Paladin has a broad expression in the postnatal lung. (a) Endothelial cells and pneumocytes type II were isolated from lungs of mT/mG mice expressing endothelial-specific Tie2-Cre (left) or pneumocyte type II-specific SPC-Cre (right). Q-PCR of sorted single cells indicates Pald1 mRNA expression in both endothelial cells and non-endothelial cells, including pneumocytes type II, both at 3 weeks and 3 months of age. Expression was normalized to 18 S RNA. (b) Pald1 LacZ reporter activity (blue) is detected in Pald1 LacZ/LacZ mice, 5 days, 4 weeks and 19 weeks after birth. LacZ is broadly expressed in the lung tissue, except for the bronchial epithelium, which shows no reporter activity (arrow). Scale bar = 200 µm.
Paladin is expressed both in the epithelial and mesenchymal compartment of the postnatal lung (4 weeks). (a–e) Combined X-gal (black) and immunofluorescence staining of Pald1 LacZ/LacZ mice show Pald1 LacZ expression in the vasculature, i.e. endothelial cells in capillaries (a) but to a lesser extent in endothelial cells of larger blood vessels (b) as indicated by Erg staining (endothelial cell nuclei, green). In large blood vessels LacZ expression can be detected in vascular smooth muscle cells (c, α-smooth muscle actin, red). Paladin LacZ reporter is also active in pneumocytes type II (d, SPC, green) and pneumocytes type I/II (e, cytokeratin, red). Scale bar = 20 µm.
Taken together, Pald1 mRNA and protein are strongly expressed in endothelial cells as well as in other cell types in the postnatal lung, except for bronchial epithelial cells.
Lack of Pald1 leads to enlarged distal airspaces in female mice
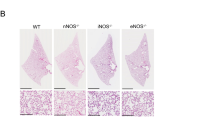
COPD in humans is often accompanied by emphysema, characterized by destruction of alveolar walls and distal airspace enlargement27. Histological analysis of Pald1 wild type and knock-out lungs revealed enlarged distal airspaces in female mice, but not in males, assessed by both blinded lung pathologist and quantification of mean linear intercept (MLI) (Fig. 3a–c). Increase in MLI, i.e. increase in distance between airway walls, was seen already at P5, at the end of the saccular stage and before alveolarization28. The distal airway dilation was still present at 4 and 19 weeks of age. Interestingly, we observed that female Pald1 +/− mice also exhibited a significant increase in distal airspace enlargement as quantified by MLI (Fig. 3d). However, no difference in alveolar septal thickness was detected (Pald1 +/+ 2,67+/− 0,14 vs. Pald1 −/− 2,48+/−0,07 µm3/µm2, P = 0,13).
Pald1 −/− mice show reduced complexity of the lung tissue. (a,b) Dark field images of distal airspace from 4 weeks old female Pald1 wild type (a) and knock-out animals (b), suggest a general airspace enlargement in the Pald1 knock-out lungs. Scale bars = 200 µm. (c) Quantification of interseptal alveolar distance using Mean Linear Intercept (MLI) of hematoxylin-eosin stained lung tissue at P5 (n = 3), 4 (n = 5) and 19 weeks (n = 4) shows increased air spaces in female knock-out animals at all stages, but was particularly increased at 4 weeks. ANOVA per age group: P5 p < 0.0001; 4 weeks p = 0.0002; 19 weeks p = 0.0004. Pald1 −/− was compared to Pald1 +/+ of the same sex within each age group. (d) MLI of Pald1 wild type, heterozygous and knock-out female mice at the indicated ages. ANOVA per age group: 4 weeks p = 0.0001, 19 weeks p = 0.0007. Pald1 −/− and Pald1 +/− were compared to Pald1 +/+ within each age group. Error bars: SD, *p ≤ 0.05, ***p ≤ 0.001, ****p ≤ 0.0001.
Reduced number, increased apoptosis and proliferation of endothelial cells in female Pald1 knock-out lungs
To further dissect the lung phenotype of female Pald1 knock-out mice, we used cell type-specific markers to assess the relative contribution in the lung. Cell type-specific markers for pneumocytes type I or II, macrophages or PDGFRα-positive cells did not reveal any differences of cell number between wild type and Pald1 knock-out mouse lungs over time (Fig. 4c–e and Figure S4). However, using the endothelial cell nuclei-specific marker Erg showed a reduction of endothelial cell number in the vascular compartment in females at 4 weeks of age by 14% (Fig. 4a and b).
Pald1 −/− show a decrease in the endothelial cell population at 4 weeks. (a–e) Quantification of the relative proportion of endothelial cells (a,b, Erg, n = 3–5), pneumocytes type II (c, SPC, n = 3), pneumocytes type I/II (d, cytokeratin, n = 3), and macrophages (e, CD68, n = 3–7). The specific cell type contribution was unaltered in Pald1 −/− mice except for a 15% decrease of the endothelial cell population at 4 weeks in Pald1 −/− mice. Scale bar = 20 µm, Error bars: SD, t-test between genotypes of each age group. ***p ≤ 0.001.
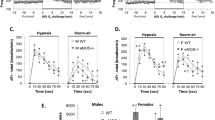
Given the reduced number of endothelial cells at 4 weeks, we determined the frequency of apoptotic cells by cleaved caspase-3 (CC3) staining in both endothelial and non-endothelial cells in these lungs (Figure S5). There was a more than six-fold increase in apoptotic cells (0.75% ± 0.41% (SD) vs. 4.90% ± 0.76% CC3+ cells, p < 0.0001) in the endothelial compartment in female Pald1 −/− compared to Pald1 +/+ mice at 4 weeks of age. However, there were no differences in non-endothelial cell apoptosis or apoptosis comparing male wild type and knock-out mice. The increase in cleaved caspase-3 was still more than two-fold at both P5 and 19 weeks of age comparing wild type and knock-out Pald1 females (Fig. 5a and b). The enhanced endothelial apoptosis in female Pald1 −/− was also accompanied by an increase of proliferation, as assessed by Ki-67 staining (Fig. 5c and Figure S5).
Increased apoptosis and proliferation of endothelial cells in Pald1 knock-out female lungs. (a,b) Quantification of cleaved caspase-3 positive (CC3+) cells in 4 and 19-week old lungs revealed a significant increase in the number of cleaved caspase-3 positive endothelial cells (Erg positive) at 4 weeks of age and at 19 weeks in female (a, n = 3–4), but not in male Pald1 −/− mice (b, n = 3–4). There was no significant increase in non-endothelial cells (Erg negative) at both 4 and 19 weeks. (c,d) Quantification of Ki67 positive cells in 4 and 19-week old lungs revealed an increased number of Ki67-positive endothelial cells (Erg positive) in female (c, n = 3–4), but not in male Pald1 −/− mice (d, n = 3–4). There was no significant increase in non-endothelial cells (Erg negative) at both 4 and 19 weeks. Error bars: SD, t-test between genotypes within each age group. * = p ≤ 0.05, **** = p ≤ 0.0001.
Sex-dependent protein expression differences in lung tissue
To begin to understand the mechanism of sex-specific phenotype in Pald1 knock-out lungs, we employed lung proteome analysis using mass spectrometry based proteomics29. We identified in total 10099 proteins at 1% protein false discovery rate, and 8,635 proteins were quantified in all 16 lung samples (n = 4 per sex and genotype) from 4 weeks old mice. One female Pald1 knock-out sample was excluded from further analyses because of contamination as described in Materials and Methods. Hierarchical clustering based on the expression of all proteins showed that the major factor for clustering is sex, and not genotype (Fig. 6a). Three proteins including Pald1 showed significantly altered expression between wild type and knock-out mice in a sex-independent manner. Hpgd (hydroxyprostaglandin dehydrogenase 15 (NAD)), showed higher expression in knock-out lungs than wild type, which was confirmed by Western blot (Fig. 6b and Figure S6). In addition, Mycbp2, MYC binding protein 2, as well as Pald1 was significantly down regulated (Figure S7). The most differentially expressed protein between wild type male and female lungs was HSD17B7, a protein essential for cholesterol biosynthesis and with the capacity to catalyze conversion of estrone to estradiol30. There were no statistically significant protein expression changes that were specific to female knock-out mice, after correction for multiple testing.
Proteomics data overview and significantly differentially expressed proteins. (a) Proteomics reveals sex-dependent protein expression differences in lung tissue. The heatmap shows proteome data overview of 8635 proteins with overlapping quantification in all 4 weeks of age lung samples using hierarchical clustering. Columns and rows represent lung samples and proteins, respectively. The samples are labels as sex_genotype_ID. The major factor for clustering is sex, and not genotype. (b) Boxplot of the three significantly differentially expressed proteins. The y-axis shows the ratios to the pool. For each gene, the ratios to the pool in the wild type group (WT) and knock-out group (KO) are plotted in the boxplot, respectively.
Discussion
Several independent groups have identified Pald1 as a vascular enriched gene1,2,31 and we have previously shown that Pald1 is expressed in both endothelial and mural cells in the vasculature3.
Extensive phenotypic screening revealed morphological and functional lung defects in female Pald1 −/− mice. This was accompanied by increased distal air spaces and elevated apoptosis and proliferation exclusively in the endothelial compartment of female mice. It is tempting to speculate, that Pald1 has a unique role in the vasculature of the lung, as this is where we observe the strongest phenotype, even though the expression of Pald1 is broader and we cannot exclude that the increased endothelial turnover is secondary to alteration in other cell types.
A central role for endothelial cell apoptosis in the development of emphysema has been proposed. This is based on both the correlation of increased endothelial cell apoptosis and down regulation of VEGF in human patients exhibiting emphysema13, and experimental evidence in animal models. Genetic ablation of VEGF-A in the lung using adenoviral Cre led to endothelial and non-endothelial cell apoptosis, without a compensatory increase in proliferation16. Inhibition of VEGFRs using either low molecular weight kinase inhibitors or specific VEGFR2 (but not VEGFR1) blocking antibodies was also sufficient to trigger alveolar apoptosis and development of emphysema in mice and rats14,15. In addition, direct induction of lung endothelial cell apoptosis using an endothelial-homing peptide triggers development of emphysema32, suggesting that alveolar structures cannot be maintained without endothelial cells, as well as that endothelial cell apoptosis in the lung culminates in emphysema. Comparison of MLI measurements in the above-mentioned VEGF-A/VEGFR2 targeted mice and rats shows that the emphysema development in Pald1 −/− female mice is comparable to what is seen in those models. However, alterations of VEGF and VEGFR levels were not the reason to emphysema in the Pald1 knock-out mice as no differences in VEGFA or VEGFR2 protein levels were detected in the proteomic analysis, or by western blot.
The increase in distal airspace in female Pald1 knock-out mice is stable from early postnatal stage to 19 weeks of age. The mechanism of airspace enlargement must be distinct from emphysematous development in adult humans as it occurs already before alveolar septation and is non-progressive. Despite this, further studies into this phenotype, might provide additional information on what type of cellular changes and biochemical pathways that can lead to the end stage phenotype that we refer to as emphysema. Further studies will also be necessary to pinpoint the mechanism whereby Pald1 regulates endothelial cell survival and proliferation in a sex-specific fashion. The proteomics data showed that sex, but not genotype, was the most significant factor for differential protein expression. This is consistent with the accumulating evidence regarding sex differences and lung biology33. Somewhat surprisingly, no specific protein expression differences were detected between female wild type and knock-out lungs. However, Hpgd, the major enzyme for degradation of prostaglandins, showed higher expression in all knock-out lungs compared to wild type, irrespective of sex. Given that prostaglandin signaling is abundant and important for lung and vascular function as well as angiogenesis34, it could be speculated that prostaglandin signaling is associated with the emphysema phenotype, but that the Pald1 −/− males are somehow protected. The lack of significant protein changes between female wild type and knock-out lungs, despite of the morphological differences and endothelial apoptosis observed at the time point analyzed, could be due to several reasons including sensitivity of the proteomics screen, use of complex tissue containing several cell types and/or due to time point of analysis. Even though we observe the greatest morphological changes at 4 weeks of age, the potential protein expression differences causing those changes might have occurred earlier.
Paladin was previously identified as a negative regulator of insulin signaling by in vitro screening for FOXO1A-driven reporter gene expression using a human cDNA library5. We detected a minor reduction in the ability to clear glucose from the blood stream in male mice as assessed by an intraperitoneal glucose tolerance test (Table S1). However, this is in contrast to the reported negative effect of Pald1 on insulin receptor signaling in cells. Further studies will be needed to determine the significance of this finding.
It was reported that Pald1 modulated the expression of key regulatory genes in neural crest development, and plasmid mediated over expression and morpholino-based knock-down of Pald1 delayed neural crest migration in the chick embryo4. Even though we have noted a prominent expression of Pald1 in migrating neural crest cells during embryonic development3, we have not noted any differences in neural crest derived tissues such as cardiac outflow tract, melanocytes, cranial bones or myelination in adult Pald1 −/− mice. However, a transient role of Pald1 in neural crest migration during development has not been assessed.
Taken together, our comprehensive description of the Pald1 −/− mouse revealed that the putative phosphatase Pald1 plays a role in the development and function of the lung, and specifically in female pulmonary endothelial cell survival and proliferation. Further studies are necessary to address how the lack of Pald1 leads to endothelial cell apoptosis and proliferation, and how that is related to the emphysema phenotype in a sex-specific manner.
Materials and Methods
Paladin nomenclature and mouse model
Paladin is encoded by Pald1 (phosphatase domain containing, paladin 1, also known as x99384 or mKIAA1274) in mice. C57BL/6 mice with constitutive deletion for Pald1 (Exon 1–18 replaced by a LacZ reporter cassette) have been generated3 and backcrossed for 10 generations. ROSA mT/mG × Tie2-Cre mice35 were generated as previously described25. ROSA mT/mG mice (Jackson Stock 007576) were crossed to Sftpc-Cre26 to generate ROSA mT/mG × Sftpc-Cre. All animal experiments were performed in accordance with the relevant laws and institutional guidelines and were approved by the Uppsala University board of animal experimentation. At the GMC mice were maintained in IVC cages with water and standard mouse chow according to the GMC housing conditions and German laws. All tests performed at the GMC were approved by the responsible authority of the district government of Upper Bavaria, Germany.
FACS and quantitative PCR
Isolation of single cells from ROSA mT/mG × Tie2-Cre and ROSA mT/mG × Sftpc-Cre lungs by FACS, mRNA extraction and quantitative PCR was done as previously described25.
Western blot analysis
Snap frozen lungs from 2, 3 and 22-week old mice were lysed in 20 mM HEPES, 150 mM NaCl, 1% NP40, 1x protease inhibitor cocktail (Roche), homogenized with Tissue Tearor (BioSpec Products) and sonicated six times for 5 sec at 200 W (Bioruptor, diagenode). Tissue lysates were incubated for one hour at 4 °C with rotation, and centrifuged at 21’100 g for 20 min at 4 °C. Protein concentration was measured with the BCA protein detection kit (Thermo Fisher Scientific). Lung lysates were denatured in sample buffer (Life Technologies) and proteins were separated on a 4–12% BisTris polyacrylamide gel (Novex by Life Technologies). Proteins were transferred to an Immobilon-P PVDF membrane (Millipore) using the XCell II™ Blot Module (Novex by LifeTechnologies). The membrane was blocked with 5% skimmed milk in TBS 0.1% Tween and incubated with rabbit anti-Pald1 (1:1000; Atlas Antibodies, HPA017343) or goat anti-actin (1:1000, Santa Cruz, sc-1615) antibodies overnight at 4 °C. Membranes were washed in TBS 0.1% Tween and incubated with horseradish peroxidase (HRP) conjugated secondary anti-rabbit (1:10’000, GE Healthcare) and anti-goat antibodies (1:10’000, Invitrogen), respectively. Membranes were washed in TBS 0.1% Tween and developed using ECL prime (GE Healthcare). Luminescence signal was detected by the ChemiDoc MP system (BioRad).
X-gal staining and immunohistochemistry of lung sections
Lungs were inflated and fixed intratracheally with 4% PFA. The trachea was tied under pressure, and the lung was fixed for 2 h in 4% PFA at 4 °C. For histochemical analysis, paraffin sections of fixed lungs were dehydrated gradually in a series of 70% ethanol to xylene and soaked in paraffin (for 6 h in total) prior to embedding. Paraffin sections (6 µm) were deparaffinized using xylene, rehydrated in graded alcohol series (99.6% to 70% ethanol) and rinsed in distilled water. For H&E staining, sections were immersed sequentially in hematoxylin and eosin solutions (Histolab). Stained sections were dehydrated (70% ethanol to 99.6% ethanol, xylene) and mounted in PERTEX mounting media (Histolab). Interalveolar septal wall distance was measured by the mean linear intercept (MLI). Images from H&E stained lung sections were acquired on a Nikon light microscope (Nikon Eclipse 80i, Nikon digital camera DXM 1200) at a 400x magnification. The MLI was obtained by dividing the length of a line drawn across the lung section by the total number of intercepts encountered in 10 lines per picture. Twenty-five images per section, and two sections per lung, were analyzed. One section was from the inferior lobe and another one from the superior lobe.
For frozen lung sections, lungs were collected and fixed as described. After overnight incubation in 30% sucrose in PBS, lung sections of 5–10 µm were cut, blocked (3% BSA, 0.1% Triton x-100, 5% Normal Donkey serum [Jackson Immunoresearch], 5% Normal Mouse serum [Invitrogen] in PBS) and stained with rabbit anti-ERG (1:100, Abcam, ab92513), mouse anti-ERG (1:100, Abcam, ab140520), mouse anti-αSMA (1:100, Sigma, C6198), rabbit anti-prosurfactant Protein C (1:100, Abcam, ab40879), mouse anti-cytokeratin (1:100, Sigma, P2871), rat anti-CD68 (1:100, AbD serotec, MCA1957), rabbit anti-cleaved caspase 3 Ab-5 (1:3000, Neomarkers, RB-1611-P1), and mouse anti-ki67 (1:100, Dako, M7240) in combination with appropriate fluorophore-coupled secondary antibodies. Images were obtained with the Zeiss LSM700 confocal microscope, 63x objective and analyzed by ImageJ. For x-gal staining, frozen lung sections were post-fixed in 0.2% PFA for 10 min on ice, rinsed in PBS with 2 mM MgCl2 and permeabilized in detergent rinse (2 mM MgCl2, 0.01% sodium deoxycholate, 0.02% Nonidet P-40, PBS) for 10 min on ice, prior to overnight staining at 37 °C with 1 mg/ml x-gal (Promega) diluted in staining solution (detergent rinse containing 5 mM potassium ferricyanide, 5 mM potassium ferrocyanide). Sections were washed twice for 10 min in detergent rinse, followed by PBS.
Statistical analysis of data sets of two groups was done by Student’s t-test and of three or more groups by one-way ANOVA using GraphPad Prism6.
Lung function analysis
Lung function analysis were performed as previously reported36. Briefly, lung function was assessed by a forced maneuver system and a Fine-Pointe RC system (Buxco Research Systems; Wilmington, NC, USA).
Statistical analyses were performed using R-scripts (version 3.0.2, Foundation of Statistical Computing, Vienna, Austria) implemented in the database (MausDB). Differences between genotypes were evaluated by Wilcoxon test. Statistical significance was assumed at p < 0.05. Data are presented as median values ± interquartile range.
Proteome analysis
Mouse lung samples were lysed by SDS and prepared for mass spectrometry analysis using a modified version of the FASP protocol29. Peptides were labelled with TMT10plex reagent according to the manufacturer’s protocol (Thermo Scientific) and separated by immobilized pH gradient - isoelectric focusing (IPG-IEF) on 3–10 strips as described previously29. Extracted peptide fractions from the IPG-IEF were separated using an online 3000 RSLCnano system coupled to a Thermo Scientific Q Exactive. MSGF + Percolator in the Galaxy platform was used to match MS spectra to the Ensembl 82 mouse protein database37.
One of the samples in Pald1 knock-out females was excluded from further analyses because it is highly likely that squamous epithelium could have contaminated the sample; protein expression pattern was completely different from others and some proteins specific to squamous epithelial cells had been detected. The remaining quantified proteomics data were processed in R software. To identify differentially expressed genes between different groups (knock-out group versus controls, male versus female), student’s t test was used and multiple test correction was implemented using the false discovery rate method38. Heatmap analysis was performed using the pheatmap packages in R software. The genes were clustered using Pearson correlation distance and the samples were clustered using Euclidean distance. The average linkage cluster method was used to build the cluster dendrogram.
Data Availability
Mass spectrometry proteomics data is deposited to jPOSTrepo39 with the dataset identifier JPST000225 & PXD005625.
All other data generated or analyzed during this study are included in this published article (and its Supplementary Information files).
References
Kalen, M. et al. Combination of reverse and chemical genetic screens reveals angiogenesis inhibitors and targets. Chem Biol 16, 432–441, https://doi.org/10.1016/j.chembiol.2009.02.010 (2009).
Wallgard, E. et al. Identification of a core set of 58 gene transcripts with broad and specific expression in the microvasculature. Arterioscler Thromb Vasc Biol 28, 1469–1476, https://doi.org/10.1161/ATVBAHA.108.165738 (2008).
Wallgard, E. et al. Paladin (X99384) is expressed in the vasculature and shifts from endothelial to vascular smooth muscle cells during mouse development. Dev Dyn 241, 770–786, https://doi.org/10.1002/dvdy.23753 (2012).
Roffers-Agarwal, J., Hutt, K. J. & Gammill, L. S. Paladin is an antiphosphatase that regulates neural crest cell formation and migration. Dev Biol 371, 180–190, https://doi.org/10.1016/j.ydbio.2012.08.007 (2012).
Huang, S. M., Hancock, M. K., Pitman, J. L., Orth, A. P. & Gekakis, N. Negative regulators of insulin signaling revealed in a genome-wide functional screen. PloS one 4, e6871, https://doi.org/10.1371/journal.pone.0006871 (2009).
Reiterer, V., Eyers, P. A. & Farhan, H. Day of the dead: pseudokinases and pseudophosphatases in physiology and disease. Trends in cell biology 24, 489–505, https://doi.org/10.1016/j.tcb.2014.03.008 (2014).
Roffers-Agarwal, J., Hutt, K. J. & Gammill, L. S. Paladin is an antiphosphatase that regulates neural crest cell formation and migration. Dev Biol 371, 180–190, https://doi.org/10.1016/j.ydbio.2012.08.007 (2012).
Alonso, A. & Pulido, R. The extended human PTPome: a growing tyrosine phosphatase family. FEBS J 283, 2197–2201, https://doi.org/10.1111/febs.13748 (2016).
Li, S., Wang, L., Berman, M., Kong, Y. Y. & Dorf, M. E. Mapping a dynamic innate immunity protein interaction network regulating type I interferon production. Immunity 35, 426–440, https://doi.org/10.1016/j.immuni.2011.06.014 (2011).
Taraseviciene-Stewart, L. & Voelkel, N. F. Molecular pathogenesis of emphysema. The Journal of clinical investigation 118, 394–402, https://doi.org/10.1172/JCI31811 (2008).
Lopez, A. D. & Murray, C. C. The global burden of disease, 1990–2020. Nat Med 4, 1241–1243, https://doi.org/10.1038/3218 (1998).
Murray, C. J. & Lopez, A. D. Alternative projections of mortality and disability by cause 1990-2020: Global Burden of Disease Study. Lancet 349, 1498–1504, https://doi.org/10.1016/S0140-6736(96)07492-2 (1997).
Kasahara, Y. et al. Endothelial cell death and decreased expression of vascular endothelial growth factor and vascular endothelial growth factor receptor 2 in emphysema. American journal of respiratory and critical care medicine 163, 737–744, https://doi.org/10.1164/ajrccm.163.3.2002117 (2001).
Kasahara, Y. et al. Inhibition of VEGF receptors causes lung cell apoptosis and emphysema. The Journal of clinical investigation 106, 1311–1319, https://doi.org/10.1172/JCI10259 (2000).
McGrath-Morrow, S. A. et al. Vascular endothelial growth factor receptor 2 blockade disrupts postnatal lung development. American journal of respiratory cell and molecular biology 32, 420–427, https://doi.org/10.1165/rcmb.2004-0287OC (2005).
Tang, K., Rossiter, H. B., Wagner, P. D. & Breen, E. C. Lung-targeted VEGF inactivation leads to an emphysema phenotype in mice. Journal of applied physiology 97, 1559–1566; discussion 1549, https://doi.org/10.1152/japplphysiol.00221.2004 (2004).
Hardin, M. et al. Sex-specific features of emphysema among current and former smokers with COPD. Eur Respir J 47, 104–112, https://doi.org/10.1183/13993003.00996-2015 (2016).
March, T. H. et al. Modulators of cigarette smoke-induced pulmonary emphysema in A/J mice. Toxicol Sci 92, 545–559, https://doi.org/10.1093/toxsci/kfl016 (2006).
Fuchs, H. et al. Mouse phenotyping. Methods 53, 120–135, https://doi.org/10.1016/j.ymeth.2010.08.006 (2011).
Gailus-Durner, V. et al. Introducing the German Mouse Clinic: open access platform for standardized phenotyping. Nat Methods 2, 403–404, https://doi.org/10.1038/nmeth0605-403 (2005).
Vanoirbeek, J. A. et al. Noninvasive and invasive pulmonary function in mouse models of obstructive and restrictive respiratory diseases. American journal of respiratory cell and molecular biology 42, 96–104, https://doi.org/10.1165/rcmb.2008-0487OC (2010).
Du, Y., Guo, M., Whitsett, J. A. & Xu, Y. ‘LungGENS’: a web-based tool for mapping single-cell gene expression in the developing lung. Thorax 70, 1092–1094, https://doi.org/10.1136/thoraxjnl-2015-207035 (2015).
Guo, M., Wang, H., Potter, S. S., Whitsett, J. A. & Xu, Y. SINCERA: A Pipeline for Single-Cell RNA-Seq Profiling Analysis. PLoS Comput Biol 11, e1004575, https://doi.org/10.1371/journal.pcbi.1004575 (2015).
Pozarska, A. et al. Stereological monitoring of mouse lung alveolarization from the early postnatal period to adulthood. American journal of physiology. Lung cellular and molecular physiology 312, L882–L895, https://doi.org/10.1152/ajplung.00492.2016 (2017).
Niaudet, C. et al. Gpr116 Receptor Regulates Distinctive Functions in Pneumocytes and Vascular Endothelium. PloS one 10, e0137949, https://doi.org/10.1371/journal.pone.0137949 (2015).
Okubo, T., Knoepfler, P. S., Eisenman, R. N. & Hogan, B. L. Nmyc plays an essential role during lung development as a dosage-sensitive regulator of progenitor cell proliferation and differentiation. Development 132, 1363–1374, https://doi.org/10.1242/dev.01678 (2005).
O’Donnell, D. E., Laveneziana, P., Webb, K. & Neder, J. A. Chronic obstructive pulmonary disease: clinical integrative physiology. Clin Chest Med 35, 51–69, https://doi.org/10.1016/j.ccm.2013.09.008 (2014).
Ten Have-Opbroek, A. A. The development of the lung in mammals: an analysis of concepts and findings. Am J Anat 162, 201–219, https://doi.org/10.1002/aja.1001620303 (1981).
Branca, R. M. et al. HiRIEF LC-MS enables deep proteome coverage and unbiased proteogenomics. Nat Methods 11, 59–62, https://doi.org/10.1038/nmeth.2732 (2014).
Jokela, H. et al. Hydroxysteroid (17beta) dehydrogenase 7 activity is essential for fetal de novo cholesterol synthesis and for neuroectodermal survival and cardiovascular differentiation in early mouse embryos. Endocrinology 151, 1884–1892, https://doi.org/10.1210/en.2009-0928 (2010).
Bhasin, M. et al. Bioinformatic identification and characterization of human endothelial cell-restricted genes. BMC genomics 11, 342, https://doi.org/10.1186/1471-2164-11-342 (2010).
Giordano, R. J. et al. Targeted induction of lung endothelial cell apoptosis causes emphysema-like changes in the mouse. J Biol Chem 283, 29447–29460, https://doi.org/10.1074/jbc.M804595200 (2008).
Carey, M. A. et al. It’s all about sex: gender, lung development and lung disease. Trends Endocrinol Metab 18, 308–313, https://doi.org/10.1016/j.tem.2007.08.003 (2007).
Nakanishi, M. et al. Prostaglandin E2 stimulates the production of vascular endothelial growth factor through the E-prostanoid-2 receptor in cultured human lung fibroblasts. American journal of respiratory cell and molecular biology 46, 217–223, https://doi.org/10.1165/rcmb.2010-0115OC (2012).
Kisanuki, Y. Y. et al. Tie2-Cre transgenic mice: a new model for endothelial cell-lineage analysis in vivo. Dev Biol 230, 230–242, https://doi.org/10.1006/dbio.2000.0106 (2001).
Sarker, R. S. et al. Coactivator-Associated Arginine Methyltransferase-1 Function in Alveolar Epithelial Senescence and Elastase-Induced Emphysema Susceptibility. American journal of respiratory cell and molecular biology 53, 769–781, https://doi.org/10.1165/rcmb.2014-0216OC (2015).
Boekel, J. et al. Multi-omic data analysis using Galaxy. Nat Biotechnol 33, 137–139, https://doi.org/10.1038/nbt.3134 (2015).
Hochberg, Y. B. A. Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society. Series B (Methodological) 57, 289–300 (1995).
Okuda, S. et al. jPOSTrepo: an international standard data repository for proteomes. Nucleic Acids Research 45 (D1):D1107–D1111 (2017).
Acknowledgements
This work was supported by the Swedish Cancer foundation, Beijer foundation, Åke Wiberg’s foundation, Magnus Bergwall’s foundation and Gustav Adolf Johansson foundation (IE, HK). In part this work has been funded by the German Federal Ministry of Education and Research to the GMC (Infrafrontier grant 01KX1012) and to the German Center for Vertigo and Balance Disorders (grant 01 EO 0901). We thank biostatistician Johan Westerberg Uppsala Clinical Research Center, Uppsala, Sweden, for advise regarding statistical methods. Dr Patrick Micke, pathologist at Akademiska University Hospital, Sweden, for blinded assessment of histological sections of Pald1 −/− mice. We thank Hisamitsu Hayashi, Department of Surgical Sciences, Uppsala University, for help with cochleae dissections, and Ross Smith, Department of Immunology, Genetics and Pathology, Uppsala University, for help with setting-up the GMC screens. The SciLife Lab BioVis platform at Rudbeck laboratory, Uppsala University, Sweden was used for acquiring confocal images and sorting of cells. Mass spectrometry analysis was performed by the Clinical Proteomics Mass Spectrometry facility, Karolinska Institutet/ Karolinska University Hospital/ Science for Life Laboratory. We are grateful for support from Hillevi Andersson Sand & Jessie Thorslund from the facility.
Author information
Authors and Affiliations
Author notes
A comprehensive list of consortium members appears at the end of the paper.
Consortia
Contributions
Experimental design and analysis of data: I.E., H.K., A.N., L.B., C.B.L., C.N., M.P., W.L., M.V., A.V., H.R.A., H.J.J., L.H., A.Ö.Y., G.M.C. Assembling manuscript figures: I.E., A.N., H.K. Supervision of project: H.F., V.G.D., T.K., A.Ö.Y., M.H.A., H.R.A., J.L., M.H. Manuscript writing: I.E., H.K., A.N., A.Ö.Y., M.H. All authors reviewed manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Egaña, I., Kaito, H., Nitzsche, A. et al. Female mice lacking Pald1 exhibit endothelial cell apoptosis and emphysema. Sci Rep 7, 15453 (2017). https://doi.org/10.1038/s41598-017-14894-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14894-9
This article is cited by
-
The proteomic landscape of in vitro cultured endothelial cells across vascular beds
Communications Biology (2024)
-
Paladin, overexpressed in colon cancer, is required for actin polymerisation and liver metastasis dissemination
Oncogenesis (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.