Abstract
Populating transient and partially unfolded species is a crucial step in the formation and accumulation of amyloid fibrils formed from pathogenic variants of human lysozyme linked with a rare but fatal hereditary systemic amyloidosis. The partially unfolded species possess an unstructured β-domain and C-helix with the rest of the α-domain remaining native-like. Here we use paramagnetic relaxation enhancement (PRE) measured by NMR spectroscopy to study the transient intermolecular interactions between such intermediate species. Nitroxide spin labels, introduced specifically at three individual lysine residues, generate distinct PRE profiles, indicating the presence of intermolecular interactions between residues within the unfolded β-domain. This study describes the applicability to PRE NMR measurements of selective lysine labeling, at different sites within a protein, as an alternative to the introduction of spin labels via engineered cysteine residues. These results reveal the importance of the β-sheet region of lysozyme for initiating self-assembly into amyloid fibrils.
Similar content being viewed by others
Introduction
Globular proteins with cooperative and persistent folds under native conditions can form amyloid fibrils and cause protein deposition diseases as a result of global or partial unfolding of the structure1,2. Human lysozyme is a well-known glycosidase containing 130 amino acid residues3. In 1993, two mutational variants of human lysozyme were directly associated with a rare form of systemic amyloidosis4. Since then it has been found that a reduction in native state stability and global cooperativity of the pathogenic variants give rise to the increased population of the transient intermediate species which initiates protein aggregation into amyloid fibrils5,6,7. Under destabilizing conditions (high temperatures and low pH) we reported that the intermediate species appear to be an ensemble of interconverting conformers with varying degrees of denaturation8, and that the rate of seeded fibril formation by human lysozyme is directly proportional to the population of this state9. However, despite the importance of the transient intermediate species in fibril formation, it is highly challenging to investigate the initial events of the aggregation process due to the heterogeneous nature and low population of the intermediate species, making them recalcitrant to most biophysical techniques.
NMR spectroscopy has proved to be one of the most powerful techniques for investigating the structure of transient protein molecules present at low population10,11. A number of NMR methods with the ability to monitor such states have been developed, but two relaxation based methods stand out: paramagnetic relaxation enhancement (PRE) and relaxation dispersion (RD) NMR12,13. Whilst RD provides kinetic (exchange rates) and thermodynamic (population) information on systems that undergo chemical exchange on millisecond to microsecond timescales, PRE provides structural information through the measurements of distances between a paramagnetic center and the NMR active nuclei of interest. The last decade has seen significant advances in the application of PRE to a variety of biological systems12 including visualization of ultra-weak protein self-association14, and use of PRE to determine intermolecular protein interactions contributing to amyloid fibril formation15,16.
To investigate the intermolecular interactions between transiently populated human lysozyme molecules by PREs, we first need to incorporate appropriate probes. With the exception of metalloproteins that contain intrinsic paramagnetic groups, most proteins must be labeled with an extrinsic paramagnetic group in order to conduct PRE measurements and a range of spin labeling techniques have been applied to PRE measurements of proteins12,17. Introduction of paramagnetic groups into proteins includes the use of nitroxide spin labels (MTSL18, PROXYL19) and metal chelators (EDTA20, ATCUN21). Most commonly, MTSL is introduced via a thiol group, as it is often relatively straightforward to introduce a single cysteine residue into the protein of interest using site-directed mutagenesis. In the case of human lysozyme, however, this strategy is not practical due to the presence of eight cysteine residues which generate four disulfide bonds that are crucial for the native-state structural integrity3,22. Although in WT lysozyme, a spin label has previously been incorporated via its single histidine residue, the information obtained remains limited without further labeling sites23,24.
In this study we have aimed to incorporate nitroxide spin labels using our selective lysine labeling strategy25 in order to investigate transient intermolecular interactions between monomeric species present during the early events of lysozyme fibril formation. Intramolecular PRE experiments of spin-labeled I59T lysozyme variants, under native conditions, were used to confirm the functionality and applicability of PRE NMR experiments (Fig. 1). These spin-labeled proteins were further studied, under fibril forming conditions, and intermolecular PREs between transiently populated species were observed, demonstrating that the earliest interactions occur between the unfolded β-domain region of the non-native species.
Human lysozyme is site-specifically labeled with a nitroxide group via a lysine residue. Intra- and intermolecular paramagnetic relaxation enhancement (PRE) was measured by NMR to report on native-state structure, but more importantly on transient intermolecular interactions between partially unfolded protein species. Specific lysine labeling provides an alternative strategy for systems that are not amenable to the use of cysteine-targeting methods.
Results
WT human lysozyme is spin-labeled at K33
Previously, we reported selective biotin labeling of human lysozyme at the most reactive of its five lysine residues (K33) using an active ester (N-hydroxysuccinimide, NHS) at relatively low pH (pH 5). Here we have extended our labeling method by introducing a nitroxide to K33 of WT lysozyme for PRE measurements. At a molar ratio of 1:100 (lysozyme-to-SpinNHS (1-oxyl-2,2,5,5-tetramethylpyrroline-3-carboxylate N-hydroxysuccinimide ester)), the majority of WT lysozyme was labeled with a single nitroxide as shown in Fig. 2a. The single, double and non-labeled proteins could be separated by cation exchange chromatography on a MonoS® column, as the protein becomes less positively charged when the ε-amino group is labeled with the nitroxide moiety (Fig. 2b). Analysis by mass spectrometry of the samples collected from each elution peak confirms the identity of the labeled species (Fig. 2c).
Site-specific nitroxide spin labeling of WT human lysozyme at K33. (a) MALDI MS analysis of the effects of SpinNHS at different concentrations on the labeling reaction. The black circle, red star and blue square represent WT and spin-labeled WT human lysozyme with single and double nitroxide labeling (molecular weights of 14,694, 14,860 and 15,026 Da, respectively). Samples were incubated with 7 μM WT human lysozyme and different SpinNHS concentrations (1:0, 1:5, 1:25 and 1:100 lysozyme-to-SpinNHS molar ratio) in MES buffer (0.1 M, pH 5) at 20 °C for 20 hrs. (b) Elution profile of the 1:100 labeling reaction from MonoS cation exchange chromatography. (c) MALDI MS analysis of the samples in (b). The black circle, red star and blue square represent the same proteins as in (a). (d) Overlaid HSQC spectra of sample B (red, reduced by sodium ascorbate) and C (black) from (c) recorded at pH 5.0, 37 °C and 500 MHz. Amide protons of residues with noticeable chemical shift perturbations are labeled.
HSQC spectra of WT human lysozyme and the WT containing a single nitroxide (SpinHuL) were recorded and are shown in Fig. 2d. The assignment of the SpinHuL protein was confirmed by a HNCA experiment (see Supplementary Fig. S1). As with our previously reported selective biotinylation of WT lysozyme at K3325, the SpinHuL protein shows only marginal chemical shift perturbation of amide peaks (Δδ) in a few residues that are close to K33, either in amino acid sequence or in space, confirming that the overall native fold of the WT protein is retained (Supplementary Fig. S1).
Intramolecular PREs confirm the location and functionality of the nitroxide spin label on the lysine side chains
To demonstrate that the nitroxide labeling is amenable to measuring PREs we recorded intramolecular PREs on the native state of WT lysozyme at pH 5 and 37 °C (Supplementary Fig. S2). The HSQC spectrum of the paramagnetic sample was recorded, after which the sample was reduced by sodium ascorbate and used directly to record a subsequent HSQC spectrum of the diamagnetic sample (see Supplementary Fig. S2). The reduced sample was preferred over non-labeled protein as the diamagnetic sample, as observing the same protein after reduction avoids introducing any unwanted structural and biophysical differences between the spin-labeled and non-labeled protein samples. The effect of the addition of sodium ascorbate to the total volume of the sample was minimal (<0.5%) and taken into account when calculating the intensity ratios (Ipara/Idia). The Ipara/Idia of the SpinHuL was calculated from the HSQC spectra and mapped onto the structure of the protein (Supplementary Fig. S2). A significant number of residues experience PRE effects due to close proximity to the K33 spin label, either in amino acid sequence (residues 30–40) or in the native fold (N-terminus and the region near residue 125) (Supplementary Fig. S2), confirming the location of the spin label at K33 and its utility for PRE experiments.
In order to enhance the chances of detecting intermolecular interactions between intermediate species that are produced only at low concentrations, we used the well-characterized I59T human lysozyme for the PRE experiments as this variant is able to populate the intermediate species more readily than the WT8,26. A nitroxide spin label was introduced to K33 of I59T lysozyme and to obtain more information from the PRE experiments spin labels were included at two additional sites. The labeling of other lysine residues was achieved using an I59T variant containing a Lys33–to-Arg33 mutation (I59T-K33R) which removes the most reactive lysine residue. The I59T-K33R variant shows effectively identical biophysical characteristics to the I59T variant (see Supplementary Fig. S3 and Supplementary Table S1), and can be labeled with a single nitroxide spin label at either K13 (K13spin) or K97 (K97spin), as the reactivity of these two lysine residues appear similar. Despite the fact that K13spin and K97spin would be expected to have similar pIs, we observed two discernable peaks when the reaction mixture was separated using a MonoS cation exchange column (see Supplementary Fig. S4), indicating that the location of the spin label influences the affinity of the protein for this ion exchange resin. Although K13spin and K97spin eluted at 24 and 26 min respectively, there was significant overlap in the chromatogram (Supplementary Fig. S4); fractions were, however, carefully collected to ensure sample purity (Supplementary Fig. S4) and the HSQC spectra of the paramagnetic and diamagnetic samples of these proteins were recorded and are shown in Supplementary Fig. S4. The native fold of I59T was retained in both proteins, as in I59T with a nitroxide label at K33, and each protein shows a high level of purity (>90% and >95% for K13spin and K97spin, respectively).
The location and the functionality of the spin label at the three individual lysine residues were confirmed by measuring intramolecular PREs within the native states (pH 5) (Fig. 3), which showed the decrease in the intensity ratios of a number of residues located in distinct positions in the sequence relative to the position of each spin label.
Intramolecular PRE effects induced by nitroxide-spin labels at three individual lysine residues (K33 in I59T, K13 or K97 in the I59T-K33R) at pH 5, 37 °C. Intensity ratios of the correlation peaks in the paramagnetic and diamagnetic samples were measured and used for mapping the structures. The intensity ratios that appear higher than 1.0, due to experimental error, are mapped as 1.0. Low and high PRE effects are shown in blue and red, respectively. Residues where PRE values could not be measured are shown in dark grey and the yellow arrows indicates the location of the nitroxide spin labels. The intramolecular PREs of natively folded I59T with the nitroxide label at K33 show an effectively identical PRE profile to the K33 spin labeled WT protein (see Supplementary Fig. S2). The PRE effects from spin labels located at K97 and K13 relate residues which are close to the spin label either in the amino acid sequence (residues 85–105 for K97; residues 5–30 for K13) or in spatial terms (residues 10–25 for K97; residues 125–130 for K13).
Intermolecular PREs are observed from non-native forms of the I59T variant present at the thermal denaturation temperatures
To detect potential intermolecular interactions between the transient intermediate states in the early stages of lysozyme fibril formation, intermolecular PREs were first measured using SpinHuL and 15N-labeled WT lysozyme. The 1HN-Γ2 values of the WT protein were calculated using the difference between 1HN-R2 values of the paramagnetic and diamagnetic samples (Fig. 4) in order to circumvent potential limitations of the intensity ratio method, which can also be affected by other factors such as 1HN-Γ1 and coherence transfer delay12,27.
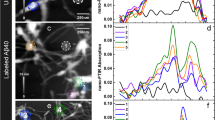
Intermolecular PRE relaxation experiments for WT lysozyme (40 and 50 °C, 500 MHz) at pH 1.2. Spin-labeled human lysozyme and 15N-isotopically labeled protein were mixed to give a 1:1 molar ratio and R2 relaxation experiments were performed for both paramagnetic and diamagnetic samples. Eight relaxation delay times (8, 13, 18, 23, 28, 38, 48 and 68 ms) were used to collect 1HN-R2 data. (a–c) HSQC spectrum of WT (a), 1HN-R2 (b) and 1HN-Γ2 (c) of the native state peaks at 40 °C. (d–f) HSQC spectrum of WT (d), 1HN-R2 (e) and 1HN-Γ2 (f) of the unfolded state peaks at 50 °C. (g) Structure of WT colored by 1HN-R2 values measured from the unfolded state peaks in (e). Red and green data points in (b,e) indicate 1HN-R2 of the paramagnetic and diamagnetic samples. Red coloring on the lysozyme structure in (g) indicates the regions of the protein that are flexible and predominantly unfolded as indicated by low 1HN-R2 values.
Overall, no significant intermolecular PREs were detected for the native or the unfolded states at 40 and 50 °C (Fig. 4c and f). This indicates that the non-native protein species which are postulated to experience intermolecular interactions, thus resulting in PRE effects, are either too scarce to give rise to detectable Γ2, or the value of Γ2 in the complex is itself very small. One interesting feature is present in the R2 values of the unfolded states at 50 °C (Fig. 4e), in that some residues, such as 1–3 and 41–52, show noticeably reduced 1HN-R2 values (<10 s−1), as expected for an unstructured or denatured protein. In contrast, other regions of the unfolded state show faster R2 rates in a similar trend to that of the R2 rates of HEWL28,29, where the unstructured parts of the protein show slow R2 rates in contrast to the other regions that have faster R2 rates as a result of residual hydrophobic structures and long range intramolecular interactions mediated by the presence of disulfide bonds under oxidized conditions. Likewise, our data suggest that the residues with slower R2 rates are likely to be more flexible in the unfolded state due to the partial unfolding of this region, whereas other regions show faster R2 rates potentially because of the presence of the disulfide bonds that are maintained in the unfolded state. Indeed the residues with slower R2 rates correspond to those in the S1 and S2 strands of the β-sheet (41–52), which unfold first as the temperature increases at the same pH in the I56T variant8. Also, the residues that display fast R2 rates are near the eight cysteine residues that form disulfide bonds within the native state. Therefore, despite the absence of detectable PREs under these conditions in the WT human lysozyme, relaxation experiments allow structural insights, indicating that the unfolded state is not fully unstructured, but maintains some residual structures likely due to the presence of the four disulfide bridges.
In order to increase the population of the intermediate state species, thereby increasing the chances of detecting any intermolecular interactions, intermolecular PREs were measured for the I59T variant. At pH 1.2, I59T shows a larger intermediate state population than the WT protein due to lower native stability and a loss of global cooperativity8. To minimize the chances of inducing unwanted denaturation of the protein via acid-hydrolysis, we chose to use two relaxation delays instead of eight, thereby reducing the duration of the experiment, as it is possible to measure relaxation rates with high accuracy by using two relaxation delays20. Intermolecular PREs were measured on the 15N-labeled I59T human lysozyme in the presence of the same protein containing a K33 spin label using pH 1.2 at different temperatures (35, 40, 45 and 50 °C) (Fig. 5).
Temperature evolution of the intermolecular PREs for I59T lysozyme (pH 1.2, at 35, 40, 45 and 50 °C) with a K33 nitroxide spin label. HSQC spectra of the diamagnetic sample at each temperature are shown on the left. The spectra at 25 and 45 °C with assignments are shown in Supplementary Figs S5 and S6, respectively. 1HN-Γ2 values (defined as the difference between 1HN-R2, para and 1HN-R2, dia) were calculated from 1HN-R2 values measured using two relaxation delay times (7, 37 ms) at 700 MHz. Red shading indicates regions of the N-terminus and the small loop at the domain interface where noticeable PREs are observed. Grey squares and black circles represent 1HN-Γ2 values measured from native and non-native peaks, respectively.
The native peaks give rise to no significant 1HN-Γ2 values at low temperatures; however, a large number of non-native state residues were found to have measurable 1HN-Γ2 values at higher temperatures with their peak intensities increasing when the protein progressively unfolds. Moreover, two regions of the protein (the N-terminus and the domain interface) show noticeable PREs that were not observed within the more stable WT protein (Fig. 4). The magnitudes of the 1HN-Γ2 values reach a maximum at 40 °C, then decrease with further temperature increases as the population of the conformational species giving rise to these PRE decreases. Intramolecular PRE measurements on K33 spin labeled WT and I59T (see Supplementary Figs S7 and S8) under similar conditions suggest that the non-native protein species possess a significant degree of residual structure. Analysis of the thermal unfolding of I59T monitored by far- and near-UV circular dichroism (CD) spectroscopy predicts that the maximum population of the intermediate state should exist at ca. 40 °C, based on a three-state unfolding model8. Overall the data suggest that the ultra-weak protein interactions14, observed in the intermolecular PREs, have resulted from lysozyme monomers that are distinct from the native-state, but still retain residual compact structures similar to the partially unfolded protein species that are crucial in lysozyme fibril formation.
Intermolecular PREs from three different spin labels (K13, K33 and K97) report on interactions involving the unfolded β-domain
In order to obtain additional structural information on the intermolecular interactions, PREs induced by K13spin and K97spin were measured at the optimized temperature of 40 °C (Fig. 6a). The results clearly show that the spin labels at different locations generate distinct PRE profiles; the K97spin shows PRE values only in the loop region at the domain interface (residues 55–57), whereas the K13spin shows no significant PREs. The effects are not due to non-specific solvent PREs, as the same molar ratio of free nitroxide molecules added to I59T lysozyme does not show any PRE effects (data not shown). It is interesting to note that analysis of 1HN-R2 values for the K33spin protein shows that the regions adjacent to those displaying intermolecular PREs have reduced relaxation rates, suggesting a potential interplay of these two neighboring regions (Fig. 6b).
(a) Intermolecular PREs for I59T lysozyme (pH 1.2, 40 °C) with spin labels at three different locations: K33, K97 and K13. 1HN-Γ2 values were calculated from the 1HN-R2 values measured using two relaxation delay times (7, 37 ms) at 700 MHz and used for mapping the structures. Red shading indicates regions of the N-terminus and the small loop at the domain interface where noticeable PREs are observed as in Fig. 3. (b) 1HN-R2 of I59T with spin label at K33 at 40 °C and the structure colored by 1HN-R2 of the diamagnetic sample. Filled squares and open circles indicate 1HN-R2 of the paramagnetic and diamagnetic samples, respectively. Data points from diamagnetic samples are used for mapping the structure. Blue shading indicates regions of the N-terminus and the β-hairpin regions where low 1HN-R2 values are observed (<15 s−1).
Discussion
Central to specific spin labeling of protein molecules for PRE measurement by NMR has been the use of site-directed mutagenesis to substitute a desired amino acid with a single cysteine residue, which can subsequently be reacted with thiol-reactive reagents, such as MTSL18. This strategy is advantageous as solvent-exposed cysteines can be readily labeled without inducing side reactions. It has some limitations, however, in cases where the naturally existing cysteines have critical structural (i.e. disulfide bridges) or functional (i.e. acting as a ligand or a catalytic center in various reactions) roles and cannot be mutated into serine or alanine30. For such systems our approach of selectively introducing nitroxide labels at lysine residues via a nucleophilic acyl substitution reaction is facile and offers an alternative labeling strategy avoiding interference with naturally existing cysteine residues. As this selective lysine-targeting method hinges on different reactivities of lysine based on their pKa values, it also has limitations as it is not uncommon to find a protein that contains multiple lysine residues with similar mobility and accessibility to the solvent. In such situations, however, a number of other labeling strategies can be applied, such as total-synthesis17, tag-promoted complexation31 or direct biosynthesis32. These methodologies require a greater degree of protein engineering; however, enable highly specific introduction of the desired probes.
When the native state (N) of lysozyme unfolds, it populates an ensemble of partially denatured species (D) that are transient and heterogeneous8. Under such conditions there can be three types of intermolecular interactions between these species: N-N, N-D and D-D. The first and second type of interactions do not contribute to the observed PRE effects as no intermolecular PREs are detected for native state residues; therefore, the observed intermolecular PREs arise solely from the interactions between the species that form the denatured ensemble. Given that the magnitude of the intermolecular PREs from protein conformers is small and decreases with increasing temperature, it is probable that the PREs are detected from a specific type of partially unfolded species that is only marginally and transiently populated near the melting temperature.
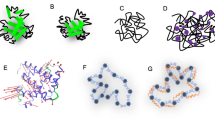
The gradual appearance of the unfolded state peaks in the HSQC spectra of human lysozyme with increasing temperature reveals that at 40 °C, residues at the N-terminus (residues 1–8), in the β-hairpin of the native state (residues 41–52) and the small loop at the domain interface (residues 55–59) are predominantly unstructured, whilst the rest of the protein retains highly native-like structure. In addition, the residues at the N-terminus and β-hairpin show reduced 1HN-R2 values (<10 s−1) in both the WT and I59T proteins, representative of flexibility that is close to that of a completely random coil structure (Fig. 3). Under these conditions the spin labels that are close to the β-sheet region (K33 and K97) display distinct PRE signals, whilst the spin label that is spatially distant from this region (K13) shows little or no effects. Using the results obtained from our experiments, we illustrate the interactions in a schematic diagram shown in Fig. 7. Based on this information we conclude that in the partially unfolded lysozyme state, initial intermolecular contacts are made through interactions involving the unfolded region of the structure that, in its native state, forms the β-domain, and these interactions are consistent with the proposed aggregation model of human lysozyme, which identifies the population of the partially unfolded intermediate species as crucial to amyloid fibril formation6,33.
Schematic diagram of the proposed unfolded β-domain region involved in early intermolecular interactions of I59T lysozyme. The structural elements shown in blue represent the regions that are predominantly unstructured and flexible (1HN-R2 < 10 s−1) (Fig. 3(b)), whereas the regions experiencing high PREs (1HN-Γ2 > 10 s−1) are shown in red. Green circles mark the nitroxide spin labels that are located at the three lysine residues (K13, K33 and K97, Fig. 3(a)). The black lines indicate the four disulfide bonds. Dashed lines indicate the potential intermolecular contacts giving rise to significant 1HN-Γ2 values.
Conclusion
We have demonstrated that transient protein interactions can be determined by PRE NMR measurements using the selective lysine labeling strategy discussed in this study. This strategy could be widely applied to other systems, including those where more conventional site-directed engineering to introduce single cysteine residues for labeling is not practical, as modification of lysine residues does not usually alter the chemical nature of a protein due to the high solvent-accessibility and mobility of these residues34,35. By utilizing a strategy to attach spin-labels specifically to individual lysine residues in human lysozyme, we have been able to capture the early intermolecular interactions between non-native I59T lysozyme molecules that are likely to trigger the subsequent aggregation process which leads to amyloid deposition in patients possessing pathological mutations in their lysozyme gene.
Methods
All chemicals and reagents were purchased from Sigma Aldrich Ltd. (Gillingham, UK) unless otherwise stated.
Preparation of I59T-K33R plasmid
Site-directed mutagenesis to introduce K33R was performed on pPIC9 containing the I59T human lysozyme gene using the Quick Change Site-Directed Mutagenesis protocol (Agilent Technologies, Oxford, UK). The K33R mutation was confirmed by DNA sequencing, performed at the Sequencing Facility in the Department of Biochemistry, University of Cambridge.
Site-specific modification of lysine side-chains of human lysozyme with nitroxide label
The WT, I59T and I59T-K33R human lysozymes were expressed in Pichia pastoris and purified as previously described36,37. In order to produce isotopically labeled samples for NMR experiments, 15N ammonium sulfate was used to label the protein produced in P. pastoris as previously described37. The level of isotope labeling was assessed by MALDI mass spectrometry and found to be essentially complete (>99%).
K33 of WT and I59T human lysozymes was labeled with nitroxide by incubating the protein with 1-oxyl-2,2,5,5-tetramethylpyrroline-3-carboxylate N-hydroxysuccinimide ester (SpinNHS) at a lysozyme-to-SpinNHS molar ratio of 1:100 (WT) and 1:300 (I59T) using the same reaction conditions as those reported for biotinylation of WT lysozyme25. Mass spectrometry of the samples at the endpoint of the reactions confirmed that the predominant species (80–90%) was singly-labeled lysozyme. Dialyzed samples were flash frozen in liquid nitrogen and lyophilized. Nitroxide spin-labeled WT and I59T samples were dissolved in Tris buffer (50 mM, pH 8) and purified using a MonoS® ion exchange column (GE Healthcare, Amersham, UK) on an ÄKTA Pure purification system (GE Healthcare). The protein was eluted using a sodium chloride gradient (0–1 M) and multiple fractions were collected. The fractions containing protein were dialyzed against deionized water (dH2O) (2 × 4 h and 24 h) or centrifuged (4 × 10 min, 15,000 g, dH2O was replenished to the filter unit after each repetition) in an Amicon® Ultra-0.5 centrifugal filter unit (3,000 nominal molecular weight limit, 500 μL, Millipore, Watford, UK) to remove the remaining sodium chloride. Samples in dH2O were flash frozen in liquid nitrogen, lyophilized and used for further analysis by dissolving in the respective buffers. Mass spectrometry of the single spin-labeled sample after the purification indicated that the sample was pure and homogeneous (>99%, Figure S1(b)). K97 and K13 of I59T-K33R were spin-labeled in the same way as described above using a higher lysozyme-to-SpinNHS molar ratio (1:500). The sample after incubation was purified in the same way on the MonoS® ion exchange column. Interestingly, this reaction yielded two populations that were resolved as separate peaks on the chromatogram (retention times at 24 and 26 min, respectively) for spin labeled K13 and K97; with the latter being the predominant species. Collection of the individual peaks resulted in the expected mass for each species, and high purity of each species (>95 and >90% for K97 and K13 spin-labeled lysozyme, respectively) was confirmed by HSQC NMR spectra.
Circular dichroism spectroscopy
CD spectroscopy experiments were performed using a Jasco J-810 spectropolarimeter (JASCO Ltd, Great Dunmow, UK) equipped with a Peltier temperature controller. Protein samples (20 μM) were dissolved in sodium citrate buffer (10 mM, pH 5) and analyzed in a 0.1 cm cuvette. Thermal denaturation was monitored at 222 or 270 nm with temperatures increasing from 5 to 95 °C (1 °C min−1). Ellipticity values were normalized to the fraction of unfolded protein (Fu) using equation 1:
where θ is the observed ellipticity, and θN and θU are the ellipticities of the native and the unfolded states, respectively. θN and θU were extrapolated from pre- and post-transition baselines. Experimental data were fitted to a two-state unfolding model assuming linear baselines for both native and unfolded states using equation 2:
where SN, SU are the signals for native and unfolded states at T = 0, whilst mN and mU are the slopes of the baseline for native and unfolded states, respectively. ΔG represents the change of the free energy for the transition. The fitting was performed by OriginPro 8.0 (OriginLab Corporation, Northampton, MA, USA) and the mid-point Tm values are defined as the temperatures where Fu is 0.5.
Thermal denaturation monitored by 8-anilino-1-napthalenesulfonic acid (ANS) fluorescence emission
Thermal denaturation monitored by ANS fluorescence emission was recorded on a Cary Eclipse spectrofluorimeter (Agilent Ltd., Oxford, UK) using excitation and emission wavelengths of 350 and 475 nm, respectively (slit-widths 5 nm) with increasing temperatures from 20 to 95 °C (0.5 °C min−1). Samples contained 2 μM protein (0.1 M sodium citrate (pH 5.0), 360 μM ANS). Transition curves of ANS alone (360 µM) were collected and subtracted from all samples. The fluorescence data were normalized with respect to the ANS emission spectrum in the presence of the I56T variant. Experimental data were fitted to a Gaussian expression using OriginPro 8.0
Hydrogen/deuterium exchange (HDX) monitored by mass spectrometry
EX1 HD exchange of I59T and I59T-K33R variants were monitored as previously described38.
Lysozyme aggregation monitored by thioflavin-T binding
Aggregation studies were performed in triplicate with lysozyme variants (6.8 μM, 0.1 M sodium citrate buffer, pH 5.0, 62.5 μM thioflavin-T (ThT)) incubated with stirring at 60 °C in a Cary Eclipse spectrofluorimeter. ThT fluorescence was measured with excitation and emission wavelengths of 440 nm and 480 nm (slit-widths 5 nm) respectively.
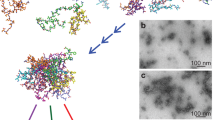
Transmission Electron Microscopy (TEM)
Samples for TEM were prepared on carbon support film, 400 mesh, 3 mm copper grids (EM Resolutions Ltd., Saffron Walden, UK) and stained with 2% uranyl acetate (w/v). The samples were imaged on a FEI Tecnai G2 transmission electron microscope in the Cambridge Advanced Imaging Centre (CAIC, University of Cambridge, Cambridge, UK). Images were analyzed using the SIS Megaview II Image Capture system (Olympus, Tokyo, Japan).
NMR spectroscopy
The backbone resonances of the isotopically double-labeled (13C, 15N) nitroxide spin-labeled lysozyme (SpinHuL) were assigned at 37 °C using HNCA experiments39. Double-labeled SpinHuL (100 μM) was dissolved in sodium acetate buffer (20 mM, pH 5.0) containing a 90% H2O/10% D2O mixture. The experiment had 1792 × 48 × 128 complex points (1H × 15N × 13C) with spectral widths of 14 ppm (1H), 31 ppm (15N) and 40 ppm (13C). For reference, HSQC experiments were run with NS = 8 scans in this series of experiments. Spectra were recorded on a Bruker Avance 700 MHz NMR spectrometer (Bruker, Coventry, UK) and processed with NMRPipe40 and Sparky41. The peak intensities were quantified by FuDA42.
Spin-labeled WT human lysozyme (200 μM) was dissolved in sodium acetate buffer (20 mM, pH 5.0) with 90% H2O/10% D2O and HSQC spectra were recorded at 37 °C on a Bruker Avance 500 MHz NMR spectrometer. Paramagnetic samples were reduced by adding sodium ascorbate (1 mM) and used as the diamagnetic counterparts. Reduction of the paramagnetic samples was monitored by multiple 1D proton and 2D HSQC spectra over time. After full reduction (when there were no further increases in the peak intensities of both 1D and 2D spectra) multiple HSQC spectra with the same parameters for paramagnetic samples were recorded on the diamagnetic samples. The peak heights of the amide protons were measured and the intensity ratios (para/dia) were calculated, these are colored on the crystal structure of the protein (Fig. 1).
14N-SpinHuL (250 μM) and 15N-WTHuL (200 μM) were mixed in phosphate buffer (50 mM, pH 1.2) with 90% H2O/10% D2O. 1HN-R2 was measured by relaxation experiments at 40 °C and 50 °C based on published pulse sequences20,21 using eight (8, 13, 18, 23, 28, 38, 48 and 68 ms) relaxation delays on a Bruker Avance 500 MHz NMR spectrometer. 1HN-R2 was calculated by fitting a single exponential decay curve to the data. 1HN-Γ2 was calculated by the difference between 1HN-R2, para and 1HN-R2, dia. The experiment was performed using two relaxation delays (7, 27/37ms) under the same conditions and no noticeable differences were observed in either 1HN-R2 or 1HN-Γ2 values as described and investigated previously20. For I59T measurements two relaxation delays were included to reduce the duration of the experiment and the occurrence of unwanted denaturation of the protein via acid-hydrolysis. The 1HN-Γ2 values were measured by performing relaxation-based measurements at 30, 35, 40 and 45 °C on a Bruker Avance 700 MHz NMR spectrometer and using the intensities of the paramagnetic and diamagnetic samples (Ipara and Idia, respectively) with equation 3:
where Ta and Tb are the two relaxation times20. The errors in 1HN-Γ2 (σ) are calculated by the equation 4:
where σdia and σpara are the standard deviations of the background noise in the spectra for diamagnetic and paramagnetic samples20.
Solvent PRE effects that are caused by diffusion and random elastic collision14,15 were tested by measuring 1HN-Γ2 values on I59T variant in the presence of equimolar SpinNHS at 40 and 45 °C. No noticeable difference between the 1HN-R2 values of the paramagnetic and diamagnetic sample (after reduction) was observed and all measured 1HN-Γ2 values were effectively zero. 1D (1H) and 2D (1H-15N HSQC or HMQC) experiments were performed before and after all intermolecular PRE measurements and confirmed that there was no degradation or aggregation of the protein. The concentration of the protein samples measured by UV absorbance at 280 nm was also identical before and after the measurements, indicating that no detectable increase in light scattering occurred due to oligomerization or aggregation.
References
Dobson, C. M. Protein folding and misfolding. Nature 426, 884–890 (2003).
Chiti, F. & Dobson, C. M. Amyloid formation by globular proteins under native conditions. Nat. Chem. Biol. 5, 15–22 (2008).
Dumoulin, M., Kumita, J. R. & Dobson, C. M. Normal and aberrant biological self-assembly: Insights from studies of human lysozyme and its amyloidogenic variants. Acc. Chem. Res. 39, 603–610 (2006).
Pepys, M. et al. Human lysozyme gene mutations cause hereditary systemic amyloidosis. Nature 362, 553–557 (1993).
Booth, D. R. et al. Instability, unfolding and aggregation of human lysozyme variants underlying amyloid fibrillogenesis. Nature 385, 787–793 (1997).
Canet, D. et al. Local cooperativity in the unfolding of an amyloidogenic variant of human lysozyme. Nat. Struct. Biol. 9, 308–315 (2002).
Dumoulin, M. et al. Reduced global cooperativity is a common feature underlying the amyloidogenicity of pathogenic lysozyme mutations. J. Mol. Biol. 346, 773–788 (2005).
Dhulesia, A. et al. Local cooperativity in an amyloidogenic state of human lysozyme observed at atomic resolution. J. Am. Chem. Soc. 132, 15580–15588 (2010).
Buell, A. K. et al. Population of nonnative states of lysozyme variants drives amyloid fibril formation. J. Am. Chem. Soc. 133, 7737–7743 (2011).
Clore, G. M. Exploring sparsely populated states of macromolecules by diamagnetic and paramagnetic NMR relaxation. Protein Sci. 20, 229–246 (2011).
Sekhar, A. & Kay, L. E. NMR paves the way for atomic level descriptions of sparsely populated, transiently formed biomolecular conformers. Proc. Natl. Acad. Sci. USA 110, 12867–12874 (2013).
Clore, G. M. & Iwahara, J. Theory, practice, and applications of paramagnetic relaxation enhancement for the characterization of transient low-population states of biological macromolecules and their complexes. Chem. Rev. 109, 4108–4139 (2009).
Korzhnev, D. M. & Kay, L. E. Probing invisible, low-populated states of protein molecules by relaxation dispersion NMR spectroscopy: an application to protein folding. Acc. Chem. Res. 41, 442–451 (2008).
Tang, C., Ghirlando, R. & Clore, G. M. Visualization of transient ultra-weak protein self-association in solution using paramagnetic relaxation enhancement. J. Am. Chem. Soc. 130, 4048–4056 (2008).
Wu, K.-P. & Baum, J. Detection of transient interchain interactions in the intrinsically disordered protein α-synuclein by NMR paramagnetic relaxation enhancement. J. Am. Chem. Soc. 132, 5546–5547 (2010).
Karamanos, T. K., Kalverda, A. P., Thompson, G. S. & Radford, S. E. Visualization of transient protein-protein interactions that promote or inhibit amyloid assembly. Mol. Cell 55, 214–226 (2014).
Klare, J. P. & Steinhoff, H.-J. Spin labeling EPR. Photosynth. Res. 102, 377–390 (2009).
Berliner, L. J., Grunwald, J., Hankovszky, H. O. & Hideg, K. A novel reversible thiol-specific spin label: Papain active site labeling and inhibition. Anal. Biochem. 119, 450–455 (1982).
Gillespie, J. R. & Shortle, D. Characterization of long-range structure in the denatured state of staphylococcal nuclease. I. Paramagnetic relaxation enhancement by nitroxide spin labels. J. Mol. Biol. 268, 158–169 (1997).
Iwahara, J., Tang, C. & Marius Clore, G. Practical aspects of 1H transverse paramagnetic relaxation enhancement measurements on macromolecules. J. Magn. Reson. 184, 185–195 (2007).
Donaldson, L. W. et al. Structural characterization of proteins with an attached ATCUN motif by paramagnetic relaxation enhancement NMR spectroscopy. J. Am. Chem. Soc. 123, 9843–9847 (2001).
Artymiuk, P. & Blake, C. Refinement of human lysozyme at 1.5 Å resolution analysis of non-bonded and hydrogen-bond interactions. J. Mol. Biol. 152, 737–762 (1981).
Mildvan, A. & Cohn, M. Aspects of enzyme mechanisms studies by nuclear spin relaxation induced by paramagnetic probes. Adv. Enzymol. Relat. Areas Mol. Biol. 33, 1–70 (1970).
Wien, R. W., Morrisett, J. D. & McConnell, H. M. Spin-label-induced nuclear relaxation. Distances between bound saccharides, histidine-15, and tryptophan-123 on lysozyme in solution. Biochemistry 11, 3707–3716 (1972).
Ahn, M. et al. Analysis of the native structure, stability and aggregation of biotinylated human lysozyme. PLoS One 7, e50192 (2012).
Hagan, C. L. et al. A non-natural variant of human lysozyme (I59T) mimics the in vitro behaviour of the I56T variant that is responsible for a form of familial amyloidosis. Prot. Eng. Design Select. 23, 499–506 (2010).
Xue, Y. et al. Paramagnetic relaxation enhancements in unfolded proteins: Theory and application to drkN SH3 domain. Protein Sci. 18, 1401–1424 (2009).
Schwalbe, H. et al. Structural and dynamical properties of a denatured protein. Heteronuclear 3D NMR experiments and theoretical simulations of lysozyme in 8 M urea. Biochemistry 36, 8977–8991 (1997).
Klein-Seetharaman, J. et al. Long-range interactions within a nonnative protein. Science 295, 1719–1722 (2002).
Schmidt, M. J. et al. EPR distance measurements in native proteins with genetically encoded spin labels. ACS Chem. Biol. 10, 2764–2771 (2015).
Baldauf, C., Schulze, K., Lueders, P., Bordignon, E. & Tampé, R. In‐situ spin labeling of His‐tagged proteins: distance measurements under in‐cell conditions. Chem. Eur. J. 19, 13714–13719 (2013).
Schmidt, M. J., Borbas, J., Drescher, M. & Summerer, D. A genetically encoded spin label for electron paramagnetic resonance distance measurements. J. Am. Chem. Soc. 136, 1238–1241 (2014).
Kumita, J. R. et al. The extracellular chaperone clusterin potently inhibits human lysozyme amyloid formation by interacting with prefibrillar species. J. Mol. Biol. 369, 157–167 (2007).
Brown, E. M., Pfeffer, P. E., Kumosinski, T. F. & Greenberg, R. Accessibility and mobility of lysine residues in β-Lactoglobulin. Biochemistry 27, 5601–5610 (1988).
Andersson, L. K., Caspersson, M. & Baltzer, L. Control of lysine reactivity in four-helix bundle proteins by site-selective pKa depression: expanding the versatility of proteins by postsynthetic functionalisation. Chem. Eur. J. 8, 3687–3697 (2002).
Spencer, A. et al. Expression, purification, and characterization of the recombinant calcium-binding equine lysozyme secreted by the filamentous fungus Aspergillus niger: Comparisons with the production of hen and human lysozymes. Prot. Express. Purif. 16, 171–180 (1999).
Johnson, R. J. et al. Rationalising lysozyme amyloidosis: Insights from the structure and solution dynamics of T70N lysozyme. J. Mol. Biol. 352, 823–836 (2005).
Ahn, M. et al. The significance of the location of mutations for the native-state dynamics of human lysozyme. Biophys. J. 111, 2358–2367 (2016).
Sattler, M., Schleucher, J. & Griesinger, C. Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog. Nucl. Magn. Reson. Spectrosc. 34, 93–158 (1999).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Sparky 3, University of California, San Francisco (2008).
Function and Data Analysis, Hansen, D. F., http://www.biochem.ucl.ac.uk/hansen/fuda.
Acknowledgements
No competing financial interests have been declared. This research was supported by the Cambridge Centre for Misfolding Diseases and funded, in part, by the Biotechnology and Biological Sciences Research Council (BB/E019927/1; JRK and CMD), the Medical Research Council (G1002272; EDG and CMD), the Wellcome Trust (094425/Z/10/Z; JRK and CMD) and the Wellcome Trust Investigator Award (097806/Z/11/Z; CAW and JC). MA was supported by a Korean Government Scholarship for Overseas Studies. The NMR facility (Department of Chemistry, University of Cambridge) is supported, in part, by an EPSRC Core Capability grant (EP/K039520/1). The authors wish to thank Mrs Ewa Klimont for her help with the purification of the lysozyme variants and Dr. Dijana Matak-Vinkovic for assistance with the HDX mass spectrometry. We acknowledge the use of the UCL NMR Centre and the MRC Biomedical NMR Centre at the Francis Crick Institute, London, and thank the staff for their support.
Author information
Authors and Affiliations
Contributions
M.A., C.A.W., C.M.D. and J.R.K. designed the research. M.A., C.A.W., A.B.-G., E.D.G., J.C. and J.R.K. performed experiments. M.A., C.A.W., A.B.-G., E.D.G., A.D., X.S., J.C., C.M.D. and J.R.K. analyzed the data. M.A., C.A.W., A.B.-G., E.D.G., A.D., X.S., J.C., C.M.D. and J.R.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahn, M., Waudby, C.A., Bernardo-Gancedo, A. et al. Application of Lysine-specific Labeling to Detect Transient Interactions Present During Human Lysozyme Amyloid Fibril Formation. Sci Rep 7, 15018 (2017). https://doi.org/10.1038/s41598-017-14739-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14739-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.