Abstract
While yeast is one of the most studied organisms, its intricate biology remains to be fully mapped and understood. This is especially the case when it comes to capture rapid, in vivo fluctuations of metabolite levels. Secondary electrospray ionization-high resolution mass spectrometry SESI-HRMS is introduced here as a sensitive and noninvasive analytical technique for online monitoring of microbial metabolic activity. The power of this technique is exemplarily shown for baker’s yeast fermentation, for which the time-resolved abundance of about 300 metabolites is demonstrated. The results suggest that a large number of metabolites produced by yeast from glucose neither are reported in the literature nor are their biochemical origins deciphered. With the technique demonstrated here, researchers interested in distant disciplines such as yeast physiology and food quality will gain new insights into the biochemical capability of this simple eukaryote.
Similar content being viewed by others
Introduction
Metabolites are small molecules that provide a direct readout of organisms’ phenotypes and cellular activity1. During metabolic processes2, a subset of the metabolites, the so-called volatile organic compounds (VOCs) are released to the ambient at normal pressure and temperature. VOCs are organic compounds typically C5-C20 3 with molecular weights up to 500 Da4, boiling point up to 250 °C5 and high vapor pressure. Many living organisms including humans, animals, plants, and even microorganisms, produce large varieties of VOCs6. The sum of all VOCs produced by an organism has been termed the volatilome7,8,9. While some VOCs have dedicated pathways, many are intermediates of anabolic pathways or are synthesized by moonlighting activities of enzymes10, hence reflect promiscuous enzymatic activities in the underlying metabolic network11,12. Although generally produced in small concentrations (parts-per-million to parts-per-trillion range)13, VOCs reflect the metabolic state of a cell and, thus, can be used to understand many biological processes such as, e.g., oxidative stress14,15,16. VOCs are also at the core of the flavor and fragrance manufacturing, an industry with a focus on plant-derived VOCs. Volatile metabolites are also of importance in the food industry as taste and smell is informing the consumer on the quality of a product. A volatile metabolite, industrially produced in large quantities, is alcohol (i.e., ethanol), synthesized by yeast from glucose during fermentation. Interestingly, since ethanol is neutral-flavored, the taste of fermented drinks originates from minor compounds, such as higher alcohols, aldehydes, esters, and acids. Most of these metabolites are volatile and intensely flavored. Indeed, monitoring VOCs (i.e., diacetyl, 2,3-pentanedione) abundance in beer is industrial standard, since it has a strong effect on the product quality17. The economic impact of monitoring such compounds is thus vast18.
Gas chromatography-mass spectrometry (GC-MS)-based methods have been the workhorse to analyze VOC profiles. The main limitation of such methods is that it requires sample manipulation, resulting in laborious procedures19,20. A more recently deployed approach is ion mobility spectrometry, which provides near real-time analysis, but its poor resolution compared to mass spectrometry compromises metabolite coverage and compound identification capabilities21,22,23,24. Real-time mass spectrometric methods used for on-line quantification of a handful of VOCs include proton transfer reaction (PTR)13 and selected ion flow tube (SIFT)25. A recent mass spectrometric method based on direct flow injection, has shown to monitor metabolite dynamics in the 15–30 second range and was used to gain insights into cellular responses to environmental changes26.
In line with such efforts to develop instrumentation capable of monitoring time-resolved metabolic information, we show here how secondary electrospray ionization (SESI)27,28,29,30,31,32,33,34,35,36,37,38,39 coupled to high resolution mass spectrometry (HRMS) captures on-line an unprecedented wealth of volatile analytes emitted in vivo by growing baker’s yeast (Saccharomyces cerevisiae).
Results and Discussion
Volatiles emitted by wild type Saccharomyces cerevisiae during growth in13C6-glucose
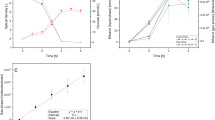
Initially, to confirm that SESI-HRMS is indeed suitable to track growth-related metabolites in real-time, we grew the yeast in glucose minimal medium and monitored well-known end products40 like ethanol or acetic acid. As expected, we observed the production of ethanol despite full aeration because baker’s yeast is Crabtree positive. Figure 1a shows such an example, in which ethanol production kicked off shortly after the start of the growth experiment. The concentration increased exponentially to then level off, presumably concomitant with glucose depletion. Eventually, ethanol started to decrease again. Such a diauxic growth is expected for yeast fermentation on glucose. The depletion kinetic, however, seems too fast to be solely explained by metabolization. Evaporation of this volatile metabolite might have been enforced by the vigorous aeration (25 gas volume flow per unit of liquid volume per minute; vvm) of the bioreactor. When ethanol was almost completely depleted, we spiked glucose into the medium (~27 h; see Methods section for experimental details). The cells immediately re-started glucose consumption as evidenced by the instantaneous production of ethanol and acetic acid.
Real-time monitoring of volatile metabolites during yeast growth; (a) raw time profiles of ethanol (detected as the dimer) and acetic acid during growth on 12C6-glucose and re-injection of glucose after ~27 h; (b) normalized raw time profiles of ethanol (detected as the dimer), acetic acid and the image signal captured by a time-lapse camera during growth on 13C6-glucose; c) heatmap showing 263 time-dependent signals during yeast growth. All the signals in the heatmap were labeled with 13C. For reference, the ethanol signal is shown on the top.
One of the advantages of SESI coupled to high-resolution mass spectrometry (HRMS) is that it can detect a wide range of analytes, including species of low vapor pressure at concentrations in the low parts-per-trillion range41. Indeed, along with the well-known volatile metabolites ethanol and acetic acid, numerous other volatile metabolites were detected during yeast growth. Thus, we replicated the experiment independently; however, we administered glucose labelled with 13C in all six carbon atoms. 13C label incorporation into metabolites synthesized from glucose allowed differentiating analytes of biological origin from environmental contaminants or media components. Figure 1b shows the resulting time profiles of ethanol, acetic acid, and yeast biomass estimated from image grayscale levels in a time-lapse video. The observed correlation between increase of ethanol and production of biomass confirmed that the analytical setup allowed capturing biological events soundly. 13C-label incorporation confirmed that 263 signals observed in the previous experiment originated from the yeast’s metabolism. In addition, we conducted a negative control experiment whereby a bioreactor was run under the very same conditions but without inoculation with yeast (monitored for ~17 hours). As expected, neither ethanol nor acetic acid were produced (Figure S1).
The time-resolved dataset of the yeast fermentation enabled a comprehensive overview of the volatilome dynamics (Fig. 1c). The heatmap is ordered by a cluster analysis, easing the visualization of the different time-profiles. Note that these metabolites were captured simultaneously and in real-time without any sample manipulation. Table S1 lists the 263 signals detected in both experiments (i.e., labeled and non-labeled glucose) as they appear in the heatmap of Fig. 1c. Interestingly, homologous series of compounds tend to cluster together, suggesting similar kinetics for closely related metabolites. For example, one cluster comprised the homologous series C6H10, C7H12, C8H14 and C9H16. This bouquet of volatile metabolites remains to be annotated, but the chemical formula suggests that these compounds could well be alkadienes. Several olefinic compounds have been reported for diverse organisms42,43,44,45,46. 1,3-octadiene (C8H14), for example, is produced by several fungi and predicted to be a degradation product of linoleic acid, but the responsible enzymes and encoding genes are yet to be identified47. None of the potential alkadienes detected here were previously identified in the headspace of yeast fermentations.
Volatiles emitted by wild type and mutants of Saccharomyces cerevisiae during growth in 13C1-glucose
To demonstrate that the detected series of hydrocarbons were produced by the baker’s yeast and not by a biological contamination of the bioreactor, we repeated the measurements. Specifically, we performed a comparison of two baker’s yeast mutants (i) devoid of the oxidative pentose phosphate pathway (zwf1), which is associated with oxidative stress response or (ii) limited in glycolysis (pfk1); and compared them with a wild type (WT) strain. To discriminate in vivo pathway usage, we fed WT, zwf1, and pfk1 with 13C1-glucose. Figure 2a shows a zoom of the m/z range 200–215 of a typical mass spectrum acquired during such measurements for WT during the stationary phase. It shows two distinct bell-shaped isotopic distributions. The first one assigned to a series of Cx(13C)yH25O2 with x ranging 11 to 7 and y 1 to 5 (i.e., molecular formula C12H25O2). Similarly, another more abundant distribution of C15H25 (13C ranging from 2 to 9) was clearly observed. Apart from high resolution (~ 30,000) enabling the discrimination of complete 13C isotopic envelopes, MS/MS capability is just another advantage of our proposed technique over other common on-line technologies used to monitor volatiles in industrial processes48. Thus, further insights in this respect were gained by identifying some of the detected compounds. For example, the two compounds in Fig. 2a were assigned to ethyl decanoate and farnesene based on the MS/MS spectrum (Figs 2b and S2). Ethyl esters have fruity and floral flavor and constitute important aroma compounds in wine or beer49. It is hypothesized that fatty acid ethyl esters are formed to prevent accumulation of medium chain fatty acid under anaerobic conditions during which fatty acid synthesis is inhibited and these fatty acids are prematurely released from the fatty acid synthase50. The sesquiterpene farnesene is synthesized from farnesyl diphosphate, an intermediate of the mevalonate pathway.
Growth of WT and mutants on 13C1-glucose led to complex mass spectra that revealed a unique in vivo metabolic response; (a) Centroided mass spectrum of the region m/z 200–215 during the stationary phase of WT yeast growth upon injection of 13C1-glucose into the system. Two isotopic envelopes for C12H25O2 and C15H25 were clearly resolved. The incorporation of 13C into ethyl decanoate and farnesene reflects the metabolism of the yeast; (b) Fragmentation (SESI MS/MS) spectrum produced using m/z 201 as the precursor ion from a yeast sample (top) and ethyl decanoate standard (bottom); (c) Set of odd-numbered carbon molecules built up during growth of the zwf1 mutant but absent in WT and pfk1; (d) Heatmap for the 636 signals detected for the three strains. For reference, ethanol profiles are shown on the top.
Figure 2c shows an example of some representative time profiles for a set of metabolites present in the zwf1 mutant but absent in WT and pfk1. Notably, this set of metabolites is a homologous series of odd-numbered carbons ranging from C7 to C15. This observation was replicated in an independent experiment (Figure S3). We hypothesize that these compounds are methyl ketones, which can be derived from β-oxidation of fatty acids, but their identity remains to be fully elucidated51. Likewise, some analytes were characteristic for pfk1 (see example in Figure S4).
Figure 2d (top) shows the time traces for ethanol. We found that the WT produced approximately twice as much ethanol as the mutants under the same experimental conditions. The same results were obtained in a replicate experiment (Figure S3). This is consistent with previous work, where similar, reduced glucose uptake and growth rate ―and as a consequence, a low ethanol formation rate― was found for the zwf1 mutant52. As for Fig. 1c, we subjected the time traces for all detected signals of the three strains to a cluster analysis and visualized them in heatmaps. Figure 2d (bottom) shows the results, providing an overview of the differences in volatile profiles between the strains. While the general picture seems relatively similar for WT and pfk1, the zwf1 mutant showed marked differences for some compounds. For example, at the bottom of the heatmap one can observe the cluster of molecules from Fig. 2b. Table S2 lists all the peaks detected in this experiment. Overall, this initial assessment illustrates the potential of SESI-HRMS to provide a comprehensive overview of VOC production by baker’s yeast in real-time.
Although the pfk1 cells showed a different metabolic profile than WT and zwf1 cells, our study showed an unexpected result. The pfk1 strain was able to undergo glycolysis (apparent from lower than expected loss of 13C) after an extended time of lag-phase growth. It is not entirely clear to us, how the pfk1 strain was able to accomplish this. However, S. cerevisiae’s phosphofructokinase is a heterooctameric enzyme with an alpha subunit encoded by PFK1 and a beta subunit encoded by PFK2. Thus, a single gene deletion mutant deficient in either of these genes might retain phosphofructokinase activity, which may explain the observed labeling pattern53,54.
Another common approach to visualize multivariate data is principal component analysis (PCA). The score plot of the mass spectra during the exponential phase (Fig. 3a) clearly shows a separation between the three strains, confirming the existence of a distinct volatile profile for each strain. Please note that we have excluded ethanol and acetic acid from the analysis in order to show that even without these major end-products, the less abundant volatile compounds suggest a very distinct profile. As expected, a similar result was obtained when we included ethanol and acetic acid in the PCA matrix (Figure S5). To understand which metabolites contributed mostly to the separation shown in the score plot, Fig. 3b displays the loadings for the first PC, which separate WT from the mutants. Some other highly discriminating molecules were identified via MS/MS and by comparison with standards (Figure S2). The esters ethyl octanoate and ethyl decanoate were major contributors, along with farnesene and octanoic acid. Tables S3 and S4 list the major contributions for PC1 and PC2, respectively. When ethanol was included in the analysis, it was by far the major contributor to the separation shown in the score plot (Figure S5).
Distinct volatile metabolic profiles and production kinetics for yeast WT and mutants; (a) PCA score plot of average spectra in the exponential phase for WT and mutants suggests a clear distinction based on the volatile metabolic profile; (b) the corresponding loading plot for score 1 shows that esters, acids, and sesquiterpenes are major contributors to the separation between WT and mutants. Identified compounds are highlighted in red. Note that ethanol and acetic acid were excluded from the PCA analysis; (c) Kinetic profiles of the food-relevant metabolite ethyl octanoate for WT, pfk1 and zwf1 illustrates the potential to monitor industrial processes; (d) series of fatty acids detected in negative ion mode during WT growth in 13C1-glucose; (e) isotopic distribution for farnesene obtained during the stationary phase. As expected, it shows a greater accumulation of 13C for zwf1; (f) Time profiles of 13C/12C ratios for ethanol dimer (i.e., m/z 95/93; black) and acetic acid (red) for the three strains investigated. Note the different kinetic profiles. The average spectra during the stationary phase for the ethanol dimer is shown in the insets.
Some of the identified compounds are not only relevant biologically, but also industrially. For example, ethyl octanoate is a yeast-based food product with sour apple aroma55. Figure 3c shows the time traces for ethyl octanoate for the three strains investigated. It clearly shows that the levels produced by WT were about four times as much as that of the mutants, reinforcing the potential to monitor industrial processes. Interestingly, ethyl hexanoate showed a very similar trend (this and additional relevant examples are shown in Figure S6). Fatty acids such as hexanoic, octanoic, and decanoic acids are just some other examples of relevant metabolites for the food and cosmetic industry. To demonstrate that these products can also be conveniently monitored, we switched the polarity to negative ion mode. SESI-HRMS spectra in negative ion mode are dominated by deprotonated fatty acids56,57. Therefore, isomeric interferences such as esters could be excluded. Figure 3d shows the time traces of a complete series of 11 fatty acids detected in WT after injecting 13C1-glucose. Acetic, octanoic, and decanoic acids were the major products. The complete list of all 43 compounds detected in this experiment is given in supplementary Table S5.
Thus, in total, 8 compounds were identified with a high degree of confidence by real-time MS/MS, although isomeric structures cannot be excluded. 11 fatty acids were also confirmed in negative ion mode with a high degree of confidence based on prior studies56,57. Additional hints of potential compounds were sought by querying publicly available databases. Of the 263 peaks associated to glucose metabolism in positive ion mode, 111 formulae matched at least one compound present in the yeast metabolome database (Table S1)58.
By injecting 13C1-glucose, we also aimed to confirm the validity of our approach to capture well-known biological processes. For example, while the contribution of the oxidative pentose phosphate pathway is rather small in baker’s yeast grown on glucose, this little contribution should be detectable59. This is indeed suggested in Fig. 3c, which shows the normalized isotopic distributions for farnesene for the three strains. The maximum isotope accumulation for all three strains was found at 13C4, but clearly, the relative abundance of C5 to C9 is greater in the culture of the zwf1 knockout strain as no 13CO2 is released during breakdown of glucose to acetyl-CoA. The same trend was observed for acetic acid, ethanol, ethyl octanoate, ethyl decanoate, and octanoic acid. All these observations were reproduced in a replicate experiment (Figure S7).
As metabolic systems are highly plastic, the ability to monitor metabolite dynamics is crucial for understanding cellular processes. Here, our time-resolved data was pivotal for understanding the metabolic pathways involved in the metabolism of glucose that are favored by each yeast strain. Figure 3f shows the 13C/12C ratio for ethanol and acetic acid. Note that, in the case of ethanol, we monitored the dimer, which lead to a triplet-like isotopic distribution (i.e., m/z 93 for C2H6O-C2H6O; m/z 94 for C(13C)H6O-C2H6O and m/z 95 for C(13C)H6O-C(13C)H6O; inset in Fig. 3f). Both, the kinetic and isotopic ratios were different for the three strains. Mutant zwf1 incorporated 13C into ethanol steeply to reach a plateau of 13C/12C ratio of ~1.2 after 5 h. The label incorporation in ethanol formed by the WT was similar but reached lower final fractional labeling. In contrast, no labeled ethanol was observed for pfk1 in the first 4 h after the 13C tracer injection and did not reach a steady labeling within the 25 h of the experiment. This is explained by a slowed-down glucose metabolism and the delayed ethanol formation in this mutant. Interestingly, the 13C in acetic acid built up immediately upon glucose injection in all strains. Other compounds showed different kinetics (Figure S8). The time resolution capability allowed to exquisitely capture fine details of kinetic profiles, which is crucial to adjust industrial parameters to tune in real-time the desired volatile profile.
In conclusion, we deployed a sensitive and selective, yet, real-time mass spectrometric technique to investigate the production of volatile metabolites during yeast growth. The technique gently tracks biological processes at the metabolic level in vivo with a time resolution of less than one minute. Thus, we benchmarked the technique by observing well-known processes such as production of ethanol from glucose. However, despite being one of the most widely studied organisms, the rich volatile profiles (~300 metabolites combining positive and negative mode) of S. cerevisiae detected in these analyses, including non-reported analytes suggests that much work remains to be accomplished to fully map the metabolism of yeast. Such comprehensive metabolic coverage may also have potential to tune industrial processes where yeast fermentation is involved.
Methods
Secondary Electrospray Ionization-High Resolution Mass Spectrometry (SESI-HRMS)
SESI-HRMS experiments were carried out using a commercial ion source (SEADM S.L.)60 plugged onto a Thermo-Fisher LTQ Orbitrap mass spectrometer. The SESI solvent was 0.1% formic acid in water infused at ~100 nL/min through a 20 μm ID silica capillary. The electrospray voltage was set to 5.4 kV in positive mode (i.e., focusing electrode 2.59 kV and impact electrode 1.6 kV) and to 5 kV in negative mode. The sweep gas used for cleaning the electrospray region was set to 2 a.u. All other internal Orbitrap parameters were optimized during calibration. Orbitrap scan parameters: scan type: FTMS full MS [50–500 Da]; source fragmentation: off; resolution: 30.000; polarity: positive and negative; Typical mass accuracies were within 2 ppm by using common chemical noise encountered in SESI-HRMS background as lock masses (i.e. m/z 149.0233, 279.1591 and 445.1200)61,62. AGC target: 30.000; Maximum injection time: 150 ms; micro scans: 30, which led to acquire a profile mass spectrum every ~49 seconds to prevent the generation of intractable large files during several hours of volatiles monitoring. The system was properly calibrated in the respective polarities prior to the analysis. MS/MS fragmentation spectra for both, yeast and standards, were obtained under identical conditions: Same collision energies; 5 microscans; Isolation width: 1 Da; Activation Q: 0.25 and Activation time: 30 ms.
Yeast cell cultures
The bioreactor consisted of an autoclaved 100 mL three-neck flask filled with 20 mL of medium, stirred at 800 rpm with a Teflon magnet and uniformly heated at 40 °C in a water bath. The metabolites produced during yeast growth were dragged downstream towards the SESI source by a continuous flow of compressed air at 0.5 L/min. The compressed air was filtered and humidified upstream the bioreactor. The bioreactor was connected to the ion source through a stainless-steel tube (OD 6 mm) heated at 130 °C to minimize metabolite adsorption onto the tube walls.
All the described experiments in this study were performed with Saccharomyces cerevisiae, prototrophic, YSBN.6 (wild-type) strain. Cells were grown in minimal defined medium: BD (DIFCO) yeast nitrogen base (#233520); and 2% glucose (as only carbon-source). For each experiment, pre-cultures were inoculated from SD plates and grown at 30 °C while shaking with 300 rpm for 8 to 10 hours in 1 mL pre-culture tubes. Then, the cells were inoculated (starting optical density; OD ~0.1) in 500 mL Erlenmeyer flasks with 50 mL of growth medium. Cultivation was performed at 30 °C with stirrer bar at 300 rpm until an OD of 1.2. The YSBN.6 strains zwf1Δ (Δzwf::Kan) and pfk1Δ (ΔPFK1::Kan), were grown in a similar minimal defined medium with the addition of 0.02% (v/v) of kanamycin and under the same time, temperature, and mixing conditions. To start the experiment, 1 mL of inoculum with an OD (between 0.8 to 1.0) was injected into the bioreactor for obtaining a starting OD of 0.1. When the ethanol signal decayed, 0.8 mL of 50% glucose was injected to prolong the exponential growth phase.
During the experiment, a time-lapse camera took a picture of the bioreactor every minute. Using ImageJ, an open platform for scientific image analysis63, yeast concentrations of relative values were obtained by averaging grayscale level in a region of interest of the image in a video. Absolute OD values were also obtained at the beginning and at the end of each experiment with a Thermo electron corporation GENESYS 10 UV spectrophotometer.
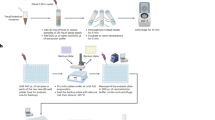
Figure 4 shows a schematic of the experimental set-up (photograph in Figure S9).
Experimental set-up: 0.5 L/min of compressed air flowed constantly through an active carbon filter (1); it was humidified in a gas washing bottle (2) before entering the bioreactor (3), a three-neck bottle with rubber stoppers filled with 50 mL of medium, heated to 30 °C and stirred at 800 rpm. The gas-phase metabolites were dragged to the SESI source (4) to be analyzed in the mass spectrometer.
Data analysis
The raw mass spectra were transformed into mzXML format via MSConvert (Proteowizard)64 and imported to MATLAB (R2016b). Each sample file was interpolated linearly (106 points in the range 50–500 Da). These interpolated profile mass spectra were then centroided (intensity threshold of 50 a.u.), resulting in ~4,400 mass spectral features (number of peaks dependent of course on the experiment performed). Subsequently, we computed the time traces for each of these peaks by adding the signal intensities ± 2, 5 mDa around the peaks. The resulting time traces (i.e., signal intensity for each m/z value as a function of time) were subjected to an agglomerative hierarchical cluster tree (Ward method; Euclidean distance; Figs 1c and 2d). To ease visualization, the time traces were smoothed (moving mean; span = 25). The 263 features finally considered (Table S1) were retained as they raised upon the 12C-glucose and 13C-glucose injection but did not experience any temporal change in a blank experiment were glucose was spiked in sterile fermentation medium. The signal to noise ratios were determined as the ratio of the variances of the signals of the experiment with yeast and the negative control where we spiked glucose into sterile medium (Figure S1).
The generation of molecular formulae from accurate mass was performed assuming protonated ions in the positive ion mode and deprotonated ions in negative mode. The following rules65 were taken into account: Masses up to 500 Da; considered elements were restricted to C, H, N and O; restricted maximum number of each of these elements, bounded H/C ratio and heteroatom/C ratio and limited number of double-bond equivalent.
Principal component analysis (PCA) was also used for dimensionality reduction and visualization of the mass spectra (Fig. 3a,b). The matrix subjected for PCA consisted of 30 rows (i.e., ten time points during the exponential growth for each of the three strains investigated) and 636 variables (i.e., peaks listed in Table S2). The data was mean-centered prior PCA. No further transformation of the matrix was applied.
References
Patti, G. J., Yanes, O. & Siuzdak, G. Metabolomics: the apogee of the omics trilogy. Nat. Rev. Mol. Cell Biol. 13, 263–269, https://doi.org/10.1038/nrm3314 (2012).
Skogerson, K., Wohlgemuth, G., Barupal, D. K. & Fiehn, O. The volatile compound BinBase mass spectral database. BMC Bioinformatics 12, https://doi.org/10.1186/1471-2105-12-321 (2011).
Bergström, G., Rothschild, M., Groth, I. & Crighton, C. Oviposition by butterflies on young leaves: Investigation of leaf volatiles. CHEMOECOLOGY 5, 147–158, https://doi.org/10.1007/bf01240599 (1994).
Lemfack, M. C., Nickel, J., Dunkel, M., Preissner, R. & Piechulla, B. mVOC: a database of microbial volatiles. Nucleic Acids Res 42, D744–748, https://doi.org/10.1093/nar/gkt1250 (2014).
Indoor air quality: organic pollutants. Report on a WHO meeting. EURO Rep Stud, 1–70 (1989).
Kesselmeier, J. & Staudt, M. Biogenic volatile organic compounds (VOC): An overview on emission, physiology and ecology. J. Atmos. Chem. 33, 23–88, https://doi.org/10.1023/A:1006127516791 (1999).
Ebert, B. E., Halbfeld, C. & Blank, L. M. Exploration and Exploitation of the Yeast Volatilome. Curr. Metabolomics 4, 1–17, https://doi.org/10.2174/2213235X04666160818151119 (2016).
Amann, A. et al. The human volatilome: Volatile organic compounds (VOCs) in exhaled breath, skin emanations, urine, feces and saliva. J. Breath Res. 8 (2014).
Bicchi, C. & Maffei, M. The Plant Volatilome: Methods of Analysis. Methods Mol. Biol. 918, https://doi.org/10.1007/978-1-61779-995-2_15 (2012).
Boukouris, A. E., Zervopoulos, S. D. & Michelakis, E. D. Metabolic Enzymes Moonlighting in the Nucleus: Metabolic Regulation of Gene Transcription. Trends Biochem. Sci. 41, 712–730, https://doi.org/10.1016/j.tibs.2016.05.013 (2016).
Copley, S. D. Enzymes with extra talents: moonlighting functions and catalytic promiscuity. Curr. Opin. Chem. Biol. 7, 265–272, https://doi.org/10.1016/S1367-5931(03)00032-2 (2003).
Schwab, W. Role of metabolome diversity in fruit and vegetable quality: multifunctional enzymes and volatiles. Woodhead Publ Food S, 272–286, https://doi.org/10.1533/9781845694296.4.272 (2008).
Mallette, N. D., Knighton, W. B., Strobel, G. A., Carlson, R. P. & Peyton, B. M. Resolution of volatile fuel compound profiles from Ascocoryne sarcoides: a comparison by proton transfer reaction-mass spectrometry and solid phase microextraction gas chromatography-mass spectrometry. AMB Express 2, 23, https://doi.org/10.1186/2191-0855-2-23 (2012).
Calenic, B. et al. Oxidative stress and volatile organic compounds: interplay in pulmonary, cardio-vascular, digestive tract systems and cancer. Open Chem 13, 1020–1030, https://doi.org/10.1515/chem-2015-0105 (2015).
Sun, D. D. et al. Effects of Growth Parameters on the Analysis of Aspergillus flavus VolatileMetabolites. Separations 3, ARTN 13.10.3390/separations3020013 (2016).
Caldeira, M. et al. Profiling allergic asthma volatile metabolic patterns using a headspace-solid phase microextraction/gas chromatography based methodology. J. Chromatogr. 1218, 3771–3780, https://doi.org/10.1016/j.chroma.2011.04.026 (2011).
Krogerus, K. & Gibson, B. R. Influence of valine and other amino acids on total diacetyl and 2, 3-pentanedione levels during fermentation of brewer’s wort. Appl. Microbiol. Biotechnol. 97, 6919–6930, https://doi.org/10.1007/s00253-013-4955-1 (2013).
Capozzi, V. et al. PTR-MS Characterization of VOCs Associated with Commercial Aromatic Bakery Yeasts of Wine and Beer Origin. Molecules 21, 483, https://doi.org/10.3390/molecules21040483 (2016).
Vas, G. & Vékey, K. Solid-phase microextraction: a powerful sample preparation tool prior to mass spectrometric analysis. Journal of Mass Spectrometry 39, 233–254, https://doi.org/10.1002/jms.606 (2004).
Arthur, C. L. & Pawliszyn, J. Solid phase microextraction with thermal desorption using fused silica optical fibers. Analytical Chemistry 62, 2145–2148, https://doi.org/10.1021/ac00218a019 (1990).
Krebs, M. D. et al. Detection of biological and chemical agents using differential mobility spectrometry (DMS) technology. IEEE Sensors Journal 5, 696–703, https://doi.org/10.1109/JSEN.2005.845515 (2005).
Lambertus, G. R. et al. Silicon Microfabricated Column with Microfabricated Differential Mobility Spectrometer for GC Analysis of Volatile Organic Compounds. Analytical Chemistry 77, 7563–7571, https://doi.org/10.1021/ac051216s (2005).
Vautz, W. & Baumbach, J. I. Analysis of bio-processes using ion mobility spectrometry. Eng. Life Sci. 8, 19–25, https://doi.org/10.1002/elsc.200720221 (2008).
Vautz, W. & Baumbach, J. I. Exemplar application of multi-capillary column ion mobility spectrometry for biological and medical purpose. Int. J. Ion Mobil. Spectrom. 11, 35–41, https://doi.org/10.1007/s12127-008-0007-4 (2008).
Scotter, J. M., Langford, V. S., Wilson, P. F., McEwan, M. J. & Chambers, S. T. Real-time detection of common microbial volatile organic compounds from medically important fungi by Selected Ion Flow Tube-Mass Spectrometry (SIFT-MS). Journal of microbiological methods 63, 127–134, https://doi.org/10.1016/j.mimet.2005.02.022 (2005).
Link, H., Fuhrer, T., Gerosa, L., Zamboni, N. & Sauer, U. Real-time metabolome profiling of the metabolic switch between starvation and growth. Nat. Methods 12, 1091–1097, https://doi.org/10.1038/Nmeth.3584 (2015).
Kiselev, P. & Fenn, J. B. In ESIMS Analysis of Vapors at Trace Levels, ASMS Conference on Mass Spectrometry and Allied Topics, Chicago, Illinois, May 27–31, 2001.
Fuerstenau, S., Kiselev, P. & Fenn, J. B. In ESIMS in the Analysis of Trace Species in Gases, ASMS Conference on Mass Spectrometry and Allied Topics, Dallas, Texas, 1999.
Whitehouse, C. M., Levin, F., Meng, C. K. & Fenn, J. B. In Further Adventures With An Electrospray Ion Source, ASMS Conference on Mass Spectrometry and Allied Topics, Cincinnati, Ohio, 1986.
Wu, C., Siems, W. F. & Hill, H. H. Secondary Electrospray Ionization Ion Mobility Spectrometry/Mass Spectrometry of Illicit Drugs. Anal. Chem. 72, 396–403 (2000).
Lee, C. Y. & Shiea, J. Gas chromatography connected to multiple channel electrospray ionization mass spectrometry for the detection of volatile organic compounds. Anal. Chem. 70, 2757–2761, https://doi.org/10.1021/ac971325+ (1998).
Zhu, J., Bean, H. D., Kuo, Y. M. & Hill, J. E. Fast detection of volatile organic compounds from bacterial cultures by secondary electrospray ionization-mass spectrometry. J. Clin. Microbiol. 48, 4426–4431, https://doi.org/10.1128/JCM.00392-10 (2010).
Bregy, L. et al. Differentiation of oral bacteria in in vitro cultures and human saliva by secondary electrospray ionization – mass spectrometry. Sci. Rep. 5, 15163, https://doi.org/10.1038/srep15163 http://www.nature.com/articles/srep15163#supplementary-information (2015).
Berchtold, C., Meier, L., Steinhoff, R. & Zenobi, R. A new strategy based on real-time secondary electrospray ionization and high-resolution mass spectrometry to discriminate endogenous and exogenous compounds in exhaled breath. Metabolomics 10, 1–11, https://doi.org/10.1007/s11306-013-0568-z (2013).
Li, X., Huang, L., Zhu, H. & Zhou, Z. Direct human breath analysis by secondary nano-electrospray ionization ultrahigh-resolution mass spectrometry: Importance of high mass resolution and mass accuracy. Rapid Commun. Mass Spectrom. 31, 301–308, https://doi.org/10.1002/rcm.7794 (2017).
Huang, L., Li, X., Xu, M., Huang, Z. & Zhou, Z. Identification of Relatively High Molecular Weight Compounds in Human Breath Using Secondary Nano Electrospray Ionization Ultrahigh Resolution Mass Spectrometry. Gaodeng Xuexiao Huaxue Xuebao/Chemical Journal of Chinese Universities 38, 752–757, https://doi.org/10.7503/cjcu20160821 (2017).
Gaugg, M. T. et al. Expanding metabolite coverage of real-time breath analysis by coupling a universal secondary electrospray ionization source and high resolution mass spectrometry – a pilot study on tobacco smokers. J. Breath Res. 10, 016010 (2016).
Barrios-Collado, C. et al. Capturing in Vivo Plant Metabolism by Real-Time Analysis of Low to High Molecular Weight Volatiles. Anal. Chem. 88, 2406–2412, https://doi.org/10.1021/acs.analchem.5b04452 (2016).
Li, X. et al. Drug Pharmacokinetics Determined by Real-Time Analysis of Mouse Breath. Angew. Chem. Int. Ed. 54, 7815–7818, https://doi.org/10.1002/anie.201503312 (2015).
Saad, S., Peter, M. & Dechant, R. In scarcity and abundance: metabolic signals regulating cell growth. Physiology (Bethesda) 28, 298–309, https://doi.org/10.1152/physiol.00005.2013 (2013).
Martínez-Lozano, P. & Rus, J. Fernández de la Mora, G., Hernández, M. & Fernández de la Mora, J. Secondary Electrospray Ionization (SESI) of Ambient Vapors for Explosive Detection at Concentrations Below Parts Per Trillion. J. Am. Soc. Mass Spectrom. 20, 287–294, https://doi.org/10.1016/j.jasms.2008.10.006 (2009).
Bos, L. D. J., Sterk, P. J. & Schultz, M. J. Volatile Metabolites of Pathogens: A Systematic Review. PLoS Pathog. 9, e1003311, https://doi.org/10.1371/journal.ppat.1003311 (2013).
Joblin, Y. et al. Detection of moulds by volatile organic compounds: Application to heritage conservation. Int. Biodeterior. Biodegrad. 64, 210–217, https://doi.org/10.1016/j.ibiod.2010.01.006 (2010).
Strobel, G. A. Bioprospecting-fuels from fungi. Biotechnol. Lett. 37, 973–982, https://doi.org/10.1007/s10529-015-1773-9 (2015).
Korpi, A., Jarnberg, J. & Pasanen, A. L. Microbial Volatile Organic Compounds. Crit. Rev. Toxicol. 39, 139–193, Pii 908581964 https://doi.org/10.1080/10408440802291497 (2009).
Jelen, H. H. & Grabarkiewicz-Szczesna, J. Volatile compounds of Aspergillus strains with different abilities to produce ochratoxin A. J. Agric. Food Chem. 53, 1678–1683, https://doi.org/10.1021/jf0487396 (2005).
Gianoulis, T. A. et al. Genomic Analysis of the Hydrocarbon-Producing, Cellulolytic, Endophytic Fungus Ascocoryne sarcoides. PLoS Genet. 8, doi:ARTN e1002558 https://doi.org/10.1371/journal.pgen.1002558 (2012).
Makhoul, S. et al. Proton-transfer-reaction mass spectrometry for the study of the production of volatile compounds by bakery yeast starters. J. Mass Spectrom. 49, 850–859, https://doi.org/10.1002/jms.3421 (2014).
Saerens, S. M. G., Delvaux, F. R., Verstrepen, K. J. & Thevelein, J. M. Production and biological function of volatile esters in Saccharomyces cerevisiae. Microb Biotechnol 3, 165–177, https://doi.org/10.1111/j.1751-7915.2009.00106.x (2010).
Nordstrom, K. Formation of Esters from Acids by Brewer’s Yeast: Formation from Unsaturated Acids. Nature 210, 99–100, https://doi.org/10.1038/210099a0 (1966).
Molimard, P. & Spinnler, H. E. Review: Compounds involved in the flavor of surface mold-ripened cheeses: Origins and properties. J. Dairy Sci. 79, 169–184 (1996).
Blank, L. M., Kuepfer, L. & Sauer, U. Large-scale 13C-flux analysis reveals mechanistic principles of metabolic network robustness to null mutations in yeast. Genome Biol. 6 (2005).
Arvanitidis, A. & Heinisch, J. J. Studies on the function of yeast phosphofructokinase subunits by in vitro mutagenesis. J. Biol. Chem. 269, 8911–8918 (1994).
Klinder, A., Kirchberger, J., Edelmann, A. & Kopperschlager, G. Assembly of phosphofructokinase-1 from Saccharomyces cerevisiae in extracts of single-deletion mutants. Yeast 14, 323–334, https://doi.org/10.1002/(SICI)1097-0061(19980315)14:4<323::AID-YEA223>3.0.CO;2-W (1998).
Saerens, S. M. G. et al. Parameters affecting ethyl ester production by Saccharomyces cerevisiae during fermentation. Appl. Environ. Microbiol. 74, 454–461, https://doi.org/10.1128/Aem.01616-07 (2008).
Martínez-Lozano, P. & de la Mora, F. J. On-line Detection of Human Skin Vapors. J. Am. Soc. Mass Spectrom. 20, 1060–1063, https://doi.org/10.1016/j.jasms.2009.01.012 (2009).
Martinez-Lozano, P. & de la Mora, F. J. Direct analysis of fatty acid vapors in breath by electrospray ionization and atmospheric pressure ionization-mass spectrometry. Anal. Chem. 80, 8210–8215, https://doi.org/10.1021/ac801185e (2008).
Ramirez-Gaona, M. et al. YMDB 2.0: a significantly expanded version of the yeast metabolome database. Nucleic Acids Res. 45, D440–D445, https://doi.org/10.1093/nar/gkw1058 (2017).
Blank, L. M. & Sauer, U. TCA cycle activity in Saccharomyces cerevisiae is a function of the environmentally determined specific growth and glucose uptake rates. Microbiol.-Sgm 150, 1085–1093, https://doi.org/10.1099/mic.0.26845-0 (2004).
Barrios-Collado, C., Vidal-de-Miguel, G. & Martinez-Lozano Sinues, P. Numerical Modeling and Experimental Validation of a Universal Secondary Electrospray Ionization Source for Mass Spectrometric Gas Analysis in Real-Time. Sensors and Actuators B: Chemical 223, 217–225, https://doi.org/10.1016/j.snb.2015.09.073 (2016).
Keller, B. O., Sui, J., Young, A. B. & Whittal, R. M. Interferences and contaminants encountered in modern mass spectrometry. Anal. Chim. Acta 627, 71–81, https://doi.org/10.1016/j.aca.2008.04.043 (2008).
Guo, X., Bruins, A. P. & Covey, T. R. Characterization of typical chemical background interferences in atmospheric pressure ionization liquid chromatography-mass spectrometry. Rapid Commun. Mass Spectrom. 20, 3145–3150, https://doi.org/10.1002/rcm.2715 (2006).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. ProteoWizard: Open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536, https://doi.org/10.1093/bioinformatics/btn323 (2008).
Kind, T. & Fiehn, O. Seven Golden Rules for heuristic filtering of molecular formulas obtained by accurate mass spectrometry. BMC Bioinformatics 8, 105, Artn 105 https://doi.org/10.1186/1471-2105-8-105 (2007).
Acknowledgements
We are thankful to Prof. Renato Zenobi for hosting the ACID project at ETH Zurich. We gratefully acknowledge Dr. Juan Zhang (Novartis AG) for the donation of the LTQ Orbitrap instrument used in this study and Martin Thomas Gaugg (ETH) for his support with some of the Matlab scripts. The research leading to these results has received funding from the European Community’s Seventh Framework Programme (FP7–2013-IAPP) within the project “Analytical Chemistry Instrumentation Development” (609691). Lars M. Blank and Birgitta E. Ebert acknowledge the German Federal Ministry for Education and Research (BMBF) for funding (Az. 031A301A). This work is dedicated to Prof. John B. Fenn (1917–2010) on the occasion of the centennial celebration of his birth.
Author information
Authors and Affiliations
Contributions
D.G.G., A.J.I. and P.M.L.S. designed the research; A.T.R., D.G.G., A.J.I. and P.M.L.S. performed the experiments; P.M.L.S. analyzed the data; all authors interpreted the data; A.T.R., B.E.E., L.M.B., A.J.I. and P.M.L.S. wrote the manuscript; all authors contributed to editing the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
Alberto Tejero Rioseras and Pablo Martinez-Lozano Sinues work in the framework of a project (Analytical Chemistry Instrumentation Development, FP7-2013-IAPP, 609691), one of whose main objectives is to develop the commercial ion source used in this study.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tejero Rioseras, A., Garcia Gomez, D., Ebert, B.E. et al. Comprehensive Real-Time Analysis of the Yeast Volatilome. Sci Rep 7, 14236 (2017). https://doi.org/10.1038/s41598-017-14554-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14554-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.