Abstract
The adenylate cyclase toxin (CyaA) of the whooping cough agent Bordetella pertussis subverts immune functions of host myeloid cells expressing the αMβ2 integrin (CD11b/CD18, CR3 or Mac-1). CyaA delivers into cytosol of cells an extremely catalytically active adenylyl cyclase enzyme, which disrupts the innate and adaptive immune functions of phagocytes through unregulated production of the key signaling molecule cAMP. We have used phosphoproteomics to analyze cAMP signaling of CyaA in murine bone marrow-derived dendritic cells. CyaA action resulted in alterations of phosphorylation state of a number of proteins that regulate actin cytoskeleton homeostasis, including Mena, Talin-1 and VASP. CyaA action repressed mTOR signaling through activation of mTORC1 inhibitors TSC2 and PRAS40 and altered phosphorylation of multiple chromatin remodelers, including the class II histone deacetylase HDAC5. CyaA toxin action further elicited inhibitory phosphorylation of SIK family kinases involved in modulation of immune response and provoked dephosphorylation of the transcriptional coactivator CRTC3, indicating that CyaA-promoted nuclear translocation of CRTC3 may account for CyaA-induced IL-10 production. These findings document the complexity of subversive physiological manipulation of myeloid phagocytes by the CyaA toxin, serving in immune evasion of the pertussis agent.
Similar content being viewed by others
Introduction
The Gram-negative coccobacillus Bordetella pertussis excels in sophistication of its immunomodulatory action. The bacterium causes the respiratory infectious disease called whooping cough, or pertussis, which can be lethal to unvaccinated infants1 and still accounts for an estimated 15 to 50 million cases and ~150,000–300,000 deaths annually world-wide2. Among the first cells of the immune system that respond to B. pertussis infection are the myeloid phagocytic cells that bear the complement receptor 3 (CR3, the αMβ2 integrin CD11b/CD18 or Mac-1). This includes macrophages, neutrophils and dendritic cells (DCs)3. B. pertussis employs several mechanisms to subvert their functions. A prominent role in paralysis of these sentinel cells is played by the CR3-binding adenylate cyclase (AC) toxin-hemolysin (CyaA, ACT, or AC-Hly). CyaA is a member of the Repeat In ToXin (RTX) family of leukotoxins4 and consists of a cell-invasive adenylyl cyclase (AC) enzyme fused to a pore-forming RTX cytolysin (Hly) moiety5. Upon binding to CR3 on cell surface, the toxin translocates its AC domain directly across the plasma membrane into cytosol of phagocytes. There, the AC enzyme is activated by calmodulin and catalyzes unregulated production of a key signaling molecule, the 3′,5′-cyclic adenosine monophosphate (cAMP). Supraphysiological concentrations of cAMP then signal through protein kinase A (PKA) and Exchange Protein directly Activated by cAMP (Epac) pathways6 and rapidly annihilate the bactericidal capacities of phagocytes. Signaling of CyaA-produced cAMP provokes massive but unproductive cell ruffling, inhibits opsonophagocytic uptake of bacteria, blocks induction of nitric oxide (NO) production, inhibits NADPH assembly and oxidative burst and induces macrophage apoptosis6,7,8,9. The molecular details of how CyaA-triggered cAMP signaling interferes with phagocyte functions remain, however, poorly defined. The high specific activity of the CyaA-delivered adenylyl cyclase (AC) enzyme represents, hence, a unique tool for analysis of the impact of cAMP signaling on myeloid cell function in general.
We have used stable isotope labelling by amino acids in cell culture (SILAC)10 for quantitative shotgun phosphoproteomic analysis of cAMP signaling resulting from CyaA toxin action on primary mouse bone marrow derived dendritic cells (BMDC). The results reveal that CyaA action causes alteration of phosphorylation of a number of proteins involved in regulation of actin cytoskeleton homeostasis, phagocytosis, translation, chromatin remodeling, IL-10 secretion and tolerogenic DC shaping.
Materials and Methods
CyaA toxin preparation
CyaA toxin and its enzymatically inactive CyaA-AC- toxoid were produced in Escherichia coli XL-1 Blue cells and purified as previously described11, including 60% isopropanol washes of the chromatography resin with bound CyaA, which reduced the endotoxin content of eluted CyaA below 300 IU/mg protein (QCL-1000 Limulus amebocyte lysate assay, Cambrex, East Rutherford, NJ).
Preparation and SILAC labelling of bone marrow-derived DCs (BMDCs)
The handling of animals was approved by the ethical committees of the Faculty of Military Health Sciences of the University of Defence and of the Institute of Microbiology of the Czech Academy of Sciences. Handling of animals and all experiments were performed in accordance with relevant guidelines and regulations, according to Guidelines for the Care and Use of Laboratory Animals, the Act of the Czech National Assembly, Collection of Laws No. 149/2004, inclusive of the amendments, on the Protection of Animals against Cruelty, and Public Notice of the Ministry of Agriculture of the Czech Republic, Collection of Laws No. 207/2004, on care and use of experimental animals.
The generation and SILAC labelling of C57BL/6 murine BMDCs was carried out as previously described12 and outlined in detail in the Supplementary methods section. BMDCs were generated from bone marrow progenitors isolated from femurs and tibias of 6- to 8-week-old female C57BL/6 mice. GM-CSF induces catabolism of 13C/15N-labelled arginine, yielding heavy arginine-derived proline and 15N isotope incorporation, thus skewing estimation of SILAC ratios. Therefore an optimized SILAC-labeling medium was used to suppress these effects, as described earlier12.
Toxin treatment
SILAC-labelled BMDCs were first incubated in D-MEM (1.9 mM Ca2+) for 2 hours at 37 °C. ‘Light’ isotope-labelled cells (12C6-arginine/12C6-lysine) were next treated for 10 or 30 minutes at 37 °C with 100 ng/ml of either CyaA toxin or CyaA-AC- toxoid dissolved in TUC buffer (50 mM Tris-HCl, 8 M urea, 2 mM CaCl2, pH 8). Corresponding ‘heavy’ isotope-labelled cells (13C6-arginine/13C6-lysine) were treated by TUC buffer alone and served as controls for both CyaA and CyaA-AC--treated BMDCs (Supplementary Fig. S1). The whole experiment was performed in biological triplicate and SILAC groups were swapped in one replicate. Cells were next washed in ice-cold PBS and lysed in Lysis buffer (50 mM NH4HCO3, 1% (w/v) sodium deoxycholate (SDC)) containing a cocktail of phosphatase inhibitors (cocktail set II; Merck). Lysis was accomplished by placing the cell suspension into a boiling water bath for 5 min13. After cooling of the lysed suspension to room temperature, the samples were subjected to benzonase treatment (Sigma) for 1 h and cell debris was removed by centrifugation (14,000 g, 10 min and 4 °C). Protein concentrations in supernatants were measured using the Micro BCA kit (Thermo Pierce) and the corresponding ‘light’ and ‘heavy’ isotope-labelled lysates were mixed at a 1:1 ratio based on their protein content.
Phosphoproteomic analysis
Details of sample preparation and data processing are provided in the Supplementary methods. Briefly, proteins in SILAC-labelled BMDC lysates were digested with trypsin and the peptides were fractionated by hydrophilic interaction liquid chromatography (HILIC)14. The phosphopeptides were enriched on TiO2 resin (GL Sciences), separated by reversed-phase nano-scale LC and analyzed by mass spectrometry on a Q-Exactive MS instrument (Thermo Scientific) operating in data-dependent acquisition (DDA) mode. Data processing, identification and quantitation was performed with MaxQuant software15,16 and the MS data were deposited via the PRIDE partner repository17 in the ProteomeXchange Consortium database (http://proteomecentral.proteomexchange.org) under the dataset identifier PXD004733. Both WT CyaA and CyaA-AC–treated samples were first compared to buffer-treated controls in order to determine the changes in phosphorylation status of each phosphosite. The resulting ratios for individual phosphosites were derived from normalized ratios of the least modified phosphopeptides in a given replicate. Ratio values > 1 then represent dephosphorylation and values < 1 represent increase in phosphorylation. The statistical significance of differences in the phosphorylation status at individual sites was determined by the Global Mean Rank Test18. Significantly enriched phosphorylation motifs were assembled using the web-based motif-x algorithm19. Assignment to known kinase motifs was performed by Motif Matcher, a web-based online tool from the PHOSIDA posttranslational modification database20,21. Alternatively, Fisher’s exact test analysis of kinase motif enrichment was performed for significantly regulated phosphosites by using the Perseus software22 with built-in Human Protein Reference Database23 as the source of kinase motifs. Functional association between significantly regulated phosphoproteins was visualized using the online-based STRING database (v10; http://string-db.org/; confidence values 0.900 and 0.400, respectively)24. Gene Ontology (GO)25 enrichment analysis was performed in Cytoscape26 using the plugin ClueGO27.
Results
CyaA triggers PKA and CaMK2-dependent phosphorylation of regulatory proteins
CR3-expressing intraepithelial DCs of myeloid origin (CD11b+) are a likely cellular target of the CyaA toxin in the course of B. pertussis infection of host airways. We thus assessed the impact of the subversive signaling of CyaA-produced cAMP in DCs by a quantitative phosphoproteomic analysis. Since human monocyte-derived dendritic cells exhibit high inter-donor variation, BMDCs from inbred C57BL/6 mice were used. The cells were labelled ex vivo with stable isotopes (SILAC)12 and advantage was taken of the use of a non-enzymatic CyaA-AC- toxoid as negative control. CyaA-AC- is unable to elevate cAMP in cells28,29 but retains the full capacity to bind the cell surface receptor CR3, to permeabilize cellular membrane and to elicit Ca2+ influx into cells and K+ efflux from cells, like intact CyaA. This control thus enabled exclusion of phosphorylation changes that were unrelated to cAMP signaling elicited by the CyaA toxin and were due to cell handling and/or cell permeabilization by the pore-forming activity of CyaA. To capture the early cAMP-triggered signaling events and to mimic the physiological concentrations of CyaA encountered by the phagocytes on B. pertussis-infected airway mucosa30,31, the SILAC-labelled BMDCs (106/ml) were exposed to 100 ng/ml of CyaA or of CyaA-AC- for 10 or 30 minutes.
SILAC-based phosphoproteomic analysis of tryptic digests of 1:1 mixtures of lyzates form ‘heavy’ and ‘light’ isotope-labelled cells was then accomplished as schematically outlined in Supplementary Fig. S1, using biological triplicates and including one label swap condition. This yielded a total of 19,310 identified phosphosites that could be confirmed by MS/MS analysis, including two phosphosites on CyaA itself. From all identified phosphosites, 14,389 were found to be class I sites having a localization probability of >0.7532. The observed global phosphorylation pattern of 12,562 serine (87.3%), 1,700 threonine (11.81%) and 127 tyrosine (0.9%) phosphosites (Supplementary Fig. S2B) resembled well the classical radioisotope-based estimates of phosphosite distribution in cells33. In total, 6,931 phosphosites were quantified for all three replicates in at least one experimental condition, including the serine residue 393 of CyaA. Compared to buffer control, the treatment of cells with CyaA for 10 or 30 min yielded a significant alteration of the phosphorylation state of the proteome for a total of 313 and 275 sites, respectively. In contrast, upon treatment with CyaA-AC- for 10 min, the alteration of phosphorylation state of only 3 sites passed the significance test. However, phosphorylation status of 56 sites was significantly modulated when BMDCs were exposed to the cell-permeabilizing CyaA-AC- toxoid for 30 min (Supplementary Fig. S2B-C, Supplementary Fig. S3 and Supplementary Table S1).
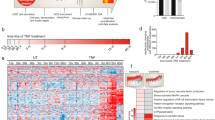
PKA isoforms are the central cAMP-regulated protein kinases that are likely to mediate most of the immunomodulatory effects of CyaA toxin action6,7,34,35. We therefore searched the phosphoproteome data specifically for cAMP-dependent alterations of the phosphorylation state of proteins (Supplementary Table S1) and the overrepresented phosphorylation patterns were extracted using motif-x19,36. As expected, the cAMP-activated PKA and Ca2+/calmodulin-dependent protein kinase II (CaMK2) were found to be the major kinases mediating cAMP signaling elicited by CyaA (Fig. 1A,B and E). Indeed, the involvement of CaMK2 goes well with the intrinsic capacity of CyaA to bind calmodulin and mediate influx of extracellular Ca2+ ions across the plasma membrane of cells37,38. Moreover, activation of CaMK2 by the other cAMP effector Epac was previously reported39, which suggested a potential indirect role of Epac in phosphoproteome alterations in CyaA-treated cells. The obtained dataset revealed a significant modulation of phosphorylation at sites recognized by the Aurora A kinase40 (Fig. 1A,B and E), in line with the previously reported inhibitory impact of CyaA toxin action on mitotic progression and G1/S transition of macrophage cells41,42. As further highlighted in Fig. 1C and D, action of CyaA led to the inhibition of cyclin-dependent (CDKs) and extracellular-signal-regulated (ERKs) kinases, yielding an overrepresentation of dephosphorylated proline-directed sites in the dataset.
Kinase enrichment analysis of significantly regulated phosphosites. Significantly regulated phosphosites in all replicates for the given experimental condition were identified by Global Mean Rank Test as described in Supplementary Methods. The kinase motifs were next extracted with the web-based motif-x (http://motif-x.med.harvard.edu/) application (Chou & Schwartz, Curr Protoc Bioinformatics. 2011) and assigned to characterized kinases using the PHOSIDA database (http://141.61.102.18/phosida/home.aspx) with the IPI mouse proteome data background (ftp://ftp.ebi.ac.uk/pub/databases/IPI), using a significance threshold of 0.000001 and a minimal occurrence value of the motif set to 20. (A) Overrepresented phosphosite motifs extracted as upregulated (phosphorylated) phosphosites by motif-x for samples incubated with CyaA for 10 or (B) 30 minutes. Occurrence value of the “..R..S..” motif was 19, hence, just below the set threshold of 20. (C) Overrepresented motifs extracted as downregulated (dephosphorylated) phosphosites by motif-x for samples incubated with CyaA for 10 or (D) 30 minutes. The “..SP…R..” motif found in the samples treated with CyaA for 30 min exhibited an occurrence number of 19, hence, below the threshold of 20. (E) Fold increase of extracted motifs shown in (A) and (B). Fold increase was calculated as the ratio (foreground matches divided by foreground size) divided by (background matches divided by background size), where the background is the predefined organism background from IPI Mouse Proteome. (F) Prediction of kinase-specific phosphorylation sites found among the significantly differentially regulated (both phosphorylated and dephosphorylated) phosphosites in samples treated with CyaA (with localization probability > 0.75 in all replicates in 10 and 30 min samples), compared to buffer-treated controls, by Fisher exact test (Benjamini-Hochberg FDR with the threshold value 0.05). Results are expressed as enrichment factors that are counted using the following formula: (number of hits for particular kinase divided by the number of significantly regulated phosphosites) divided by (number of hits of the particular kinase in the whole given cluster divided by the total size of the dataset). Only kinase enrichment factors > 1 are depicted. Perseus software package for shotgun proteomics data analyzes, version 1.5.1.6, was used for the analysis, employing the Human Protein Reference Database (Release 9).
To corroborate this analysis by an alternative approach, individual phosphosites were annotated for potential kinase motifs according to Human Protein Reference Database23 and the enrichment level for regulated sites was next evaluated by Fisher’s exact test using the Perseus software. Following CyaA treatment for 10 or 30 minutes, the PKA, CAMK2 and Aurora A kinase motifs were found to be overrepresented among the increasingly phosphorylated sites, while the Erk1/2 kinase motifs were overrepresented among significantly dephosphorylated sites (Fig. 1C–E). However, BMDC exposure to CyaA for 30 minutes had no effect on the phosphorylation status of the threonine 203 and tyrosine 205 residues in the activation loop of Erk kinases (Supplementary Fig. S4A and B). This indicates that CyaA-produced cAMP signaling activated some phosphatase(s) dephosphorylating the Erk1/2 target motifs.
CyaA is itself phosphorylated upon translocation into target cells
The 384 N-terminal amino residues of the AC enzyme domain of CyaA are fused to the ‘AC to Hly-linking segment’ (residues 385–485)43 that plays an essential role in AC domain translocation across the plasma membrane of cells44. The phosphoproteomic analysis revealed a phosphorylation signal of the serine 393 residue of CyaA (Supplementary Table S1, for processed MS/MS spectrum see Supplementary Fig. S5), showing that the N-proximal end of the ‘AC to Hly-linking segment’ penetrates into cell cytosol.
Adenylyl cyclase enzyme and pore-forming activities of CyaA exert counteracting effects on PI3K-Akt pathway
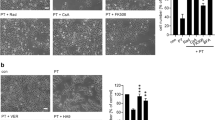
The impact of toxin and toxoid treatments on the phosphoproteome of BMDCs is summarized in Supplementary Fig. S3A-D. Only a small number of significantly regulated phosphosites was observed upon treatment of cells with the CyaA-AC- toxoid for 10 minutes. Therefore, only samples from the 30 minutes time point for the toxoid were analyzed in more detail. When the significantly regulated phosphosites of the toxoid-treated sample were plotted against the matching phosphosites from the CyaA-treated condition, a prominent alteration of the phosphorylation state of the threonine residue 1465 (1462 in man) of tuberin (product of the Tsc2 gene) was observed (Fig. 2). Tuberin, a well described mTOR-signaling suppressor45, was dephosphorylated in CyaA-treated cells and was phosphorylated in toxoid-treated cells (Fig. 2A and B). cAMP signaling of CyaA thus inhibited the activity of the PI3K-Akt pathway accounting for tuberin phosphorylation46, while the permeabilization of cells by the toxoid activated this pathway. Besides, Western blot analysis revealed that phosphorylation of the PRAS40 protein, yet another mTOR signaling inhibitor47, was deregulated. PRAS40 exhibited diminished phosphorylation of the threonine residue 24748 in CyaA-treated cells (Fig. 3A,B, Supplementary Figs S6A and S7A-C). For mTOR itself, only a mild decrease in phosphorylation of serine residue 2448 was observed after 30 min of CyaA action on cells (Fig. 3B, Supplementary Figs S6B and S7D-E). To test if these changes affected the downstream targets of mTOR, the phosphorylation status of serine residue 64 of 4E-BP1 was probed by a specific antibody. Dephosphorylation at this site was detected both by SILAC phosphoproteomics and Western blotting (Fig. 3, Supplementary Figs S6C and D7F-H and Supplementary Table S1).
Phosphorylation of tuberin (Tsc2) is oppositely manipulated by the cAMP signaling and pore-forming activities of CyaA. (A) Scatter plot of all significantly regulated phosphosites (based on signal ratios of toxin or toxoid-treated samples to buffer-treated samples). Values for the samples treated by CyaA-AC- toxoid for 30 minutes are plotted against the corresponding values for the samples treated by WT CyaA for 30 minutes. Tsc2, showing an inverse correlation between the two conditions, is highlighted. (B) Fold change of phosphorylation status (toxin- or toxoid-treated sample versus buffer-treated control) of the threonine residue 1465 of Tsc2 for three individual replicates (values > 0 represent phosphorylation, values < 0 represent dephosphorylation).
Changes in phosphorylation status of selected phosphoproteins. Members of mTOR signaling pathway after the toxin/toxoid treatment for (A) 10 minutes or (B) 30 minutes. (C) Phosphorylation status of serine 235 residue of cytoskeletal regulator VASP. In general, 106 BMDCs in 1 mL were exposed for 10 or 30 minutes to 100 ng of CyaA or of the CyaA-AC- toxoid, cellular lyzates were separated by SDS-PAGE and phosphorylation of indicated proteins was probed by Western blotting with specific antibodies. Images representative of at least three biological replicates are shown and quantifications of the Western blots of mTOR signaling pathway members are shown in Supplementary Fig. S6. Cropped images are shown and full-length blots are presented in Supplementary Fig. S7. “Buffer” stands for TUC-treated cells (TUC buffer −50 mM Tris-HCl, 8 M urea, 2 mM CaCl2, pH 8; used for toxin/toxoid dilution).
CyaA-elicited phosphoproteome alterations modulate small GTPase signaling and actin cytoskeleton homeostasis
When Gene Ontology (GO)25 terms enrichment analysis was applied to the phosphoproteome dataset, the action of CyaA on BMDCs resulted in the over-representation of terms related to GTPase signaling pathways and regulation of actin cytoskeleton (Fig. 4). This was further corroborated by construction of protein-protein interaction networks, using the STRING v10 database software24 to decode the signaling processes triggered by toxin action in BMDCs (Fig. 5A–C). The network capturing of CyaA action in the first 10 minutes consisted of 43 nodes of phosphoproteins (Fig. 5A), with the most highly interconnected nodes comprising Mena (Enah), Talin-1 (Tln1), VASP (Fig. 3C, Supplementary Fig. S7I-J) or Abl2 proteins, which are all involved in the regulation of cytoskeleton homeostasis49,50,51,52. This goes well with the previous observation that CyaA-elicited cAMP signaling provokes transient inhibition of RhoA and triggers massive actin cytoskeleton rearrangements and membrane ruffling9. CyaA-triggered signaling remained similar at 30 min of BMDC treatment with CyaA, yielding a network of 57 nodes (Fig. 5B). The interconnected cluster of the largest subnetwork then again contained components of the small GTPase signaling pathways, such as the small GTPase-activators Gmip53 and Arhgap11a54, which are involved in cytoskeleton homeostasis regulation. Altogether, these results show that the early effects of CyaA-elicited cAMP signaling largely affect small GTPases that regulate actin remodeling and cytoskeletal rearrangements.
Functional analysis of CyaA-induced phosphoproteome changes. Analysis was performed using Gene Ontology Term enrichment and the top 20 terms (p ≤ 0.05) enriched in „Biological process“ library were ranked according to their p-values (corrected with Benjamini-Hochberg).The enrichment was counted as a number of genes belonging to each GO term divided by the total number of genes in each input gene set.
Functional association between significantly regulated phosphoproteins. Visualization of functional association of significantly regulated phosphoproteins was performed using STRING database v10 at the highest confidence level of 0.900 for (A) the sample treated with 100 ng/ml of WT CyaA for 10 min and (B) for 30 min. (C) Association at medium confidence 0.400 for significantly regulated phosphoproteins in the sample treated with 100 ng/ml CyaA-AC- toxoid. C). The software-assigned coloring of nodes has no particular meaning and many of the indicated proteins are involved in several signaling pathways in parallel. Confidence view with stronger associations is represented by thicker lines.
For the sample treated with CyaA-AC- (Fig. 5C), the STRING-based interaction network of affected phosphoproteins at the 30 min time point could only be constructed upon lowering of the stringency conditions (medium confidence 0.400). The resulting network then comprised, among others, the Heterochromatin protein 1 homolog alpha (CBX5), the E3 ubiquitin-protein ligase MDM2, the Rho guanine nucleotide exchange factor 7 (Arhgef7), PML and the SAFB-like transcription modulator (SLTM). Alterations of phosphorylation of these proteins might thus reflect the cell-permeabilizing activity of the CyaA-AC- toxoid.
CyaA-elicited cAMP signaling impacts on phosphorylation status of SIK kinases involved in immune response
Based on their annotation25,55,56 (selected GO terms are listed in Supplementary Table S2) we identified 32 kinases that contain at least one phosphosite having a significantly altered phosphorylation state upon BMDCs treatment with CyaA (Supplementary Table S3). Among them, the phosphorylation of regulatory serine residues of several salt-inducible kinases (SIK), known for their involvement in modulation of immune responses57,58,59,60, was importantly altered (Fig. 6).
Changes in phosphorylation status of SIK family kinases and CRTC3/TORC3. Changes in the phosphorylation status of selected phosphosites are depicted here as fold change, representing the ratios of signals from toxin- or toxoid-treated samples to the corresponding buffer-treated control samples. Each data point represents one ratio value for one of the three replicates. The values > 0 represent phosphorylation, values < 0 represent dephosphorylation.
Discussion
We report here the first phosphoproteome-wide analysis of the signaling impact of the cAMP-elevating activity of B. pertussis adenylate cyclase toxin in primary myeloid cells. The AC domain of the CyaA toxin binds calmodulin in the cytosol of cells and acquires an extremely high specific enzymatic activity of about 2,000 molecules of ATP converted to cAMP per second per AC enzyme molecule61. This yields a steep and uncontrolled elevation of intracellular cAMP concentration in cells. The data reported here thus provides important insight into the pleiotropic effects of cAMP signaling in myeloid cells and point to two main areas of CyaA toxin interference within normal DC physiology. These comprise cAMP signaling towards inhibition of mTOR activity, signaling promoting cytoskeletal rearrangements and signaling regulating cytokine production by dendritic cells, as discussed below.
Pro-survival signaling of mTOR is negatively regulated by the protein tuberin, the activity of which is inhibited by phosphorylation of the threonine residue 1465 by Akt45,46. Tuberin is also known to be inhibited by Erk62. In contrast to previous reports using mouse or human macrophage or neutrophil cells7,63, we observed here that the Thr203 and Tyr205 residues in the activation loop of Erk1/2 were not dephosphorylated and the Erk kinases likely remained active even after 30 minutes of BMDC incubation with both the active CyaA toxin and its CyaA-AC- toxoid (c.f. Supplementary Fig. S4). Nevertheless, we found by phosphoproteomics that in CyaA-treated cells the threonine 1465 residue of tuberin was dephosphorylated. However, we were unable to detect tuberin by immunoblotting with antibodies from various suppliers, which may suggest that only very low levels of tuberin are produced in BMDCs. Alternatively, these cells may express an isoform of tuberin that is not detected by the commercially available antibodies. Nevertheless, the dephosphorylation of threonine 1465 of tuberin, as detected by SILAC phosphoproteomics, would yield activated tuberin46 capable of inhibiting mTOR and its pro-survival signaling. In line with this, WT CyaA action caused inhibition of Akt, as observed by dephosphorylation of serine 475 and threonine 308 residues64 (see Supplementary Fig. S8 and the work of Ahmad et al.7). Deactivation of Akt kinase also yielded dephosphorylation of the threonine 247 residue of PRAS40, promoting mTOR inhibition47 (Figs 3 and 7). As a result, the downstream effectors of mTOR involved in the synthesis of cellular proteins, such as 4E-BP1 (serine 64)65 (c.f. Fig. 3 and Supplementary Fig. S7F-H) and p70S6K (serine 427)66, were found to be dephosphorylated in CyaA-treated cells (Supplementary Table S1), where hypophosphorylated 4E-BP1 strongly binds to eukaryotic translation initiation factor and prevents translation67. The proposed model of mTOR signaling regulation by CyaA in BMDCs is then shown in Fig. 7. mTOR acts as an important regulator of the antibacterial inflammatory response in monocytes, macrophages and primary dendritic cells68,69. Inhibition of mTOR also leads to the activation and nuclear translocation of NF-κB and CyaA-provoked inhibition of mTOR would go well with our previous observation that NF-κB is activated upon CyaA-produced elevation of cAMP in macrophages6,68. The CyaA-provoked inhibition of mTOR is likely to have rather pleiotropic effects and these would vary in different DC subtypes70,71. For example, inhibition of mTOR was reported to result in the inhibition of IL-4-dependent maturation of BMDCs and in lowering of their T cell stimulatory capacity72,73. This would go well with our observations that CyaA treatment skews TLR-induced maturation of dendritic cells, inhibits their capacity to present antigens to both CD4+ and CD8+ T cells and enables the toxin-treated DCs to expand regulatory T cells in vitro 74.
Schematic depiction of CyaA-induced signaling that would lead to induction of IL-10 production, inhibition of protein translation and deregulation of actin cytoskeleton homeostasis. Elevation of intracellular cAMP concentration due to the catalytic action of internalized AC domain of CyaA provokes activation of the protein kinase A, which performs an inhibitory phosphorylation of the SIK family proteins. This would promote nuclear localization of the transcriptional co-activator CRTC3. In parallel, PKA phosphorylates CREB and the phospho-CREB-CRTC3 complex would activate expression of the IL-10 gene. cAMP-dependent phosphorylation of VASP, presumably by PKA, affects actin cytoskeleton homeostasis. CyaA action further yields inhibition of Akt signaling, thus provoking inhibition of mTOR signaling. Pointed arrowheads indicate an activating effect and flat arrowheads indicate an inhibitory effect under normal physiological conditions. The red color of the arrowhead indicates an inhibitory effect, or interference, resulting from cAMP signaling elicited by the CyaA toxin and the green color indicates an enhancing effect of CyaA/cAMP signaling action. For the broader context of these signaling processes the reader is referred to dedicated reviews on immunity to B. pertussis infections and on CyaA toxin action3,97.
CyaA action elicited phosphorylation of the hub-like Crk adaptor protein on the conserved serine 125 residue that is located between the SH2 and SH3 domains (c.f. Supplementary Table S1). This is likely to be of relevance during Bordetella infections, as hijacking of Crk appears to be implicated in bacterial pathogenesis through diverse mechanisms, including subversion of cellular ruffle formation (reviewed in Martinez-Quiles et al.75). The Crk-associated proteins retained in the interaction network (c.f. Fig. 5B) then comprised Talin-1, Filamin-A and the APBB1-interacting protein 1/RIAM, which are involved in cellular actin remodeling and phagocytosis. It is of note that CyaA action was previously reported to manipulate RhoA signaling and induce unproductive membrane ruffling9. Phosphorylation of the vasodilator-stimulated phosphoprotein (VASP) by the cAMP-activated PKA was then shown to play an important role in membrane ruffle formation, causing a significant reduction of the retraction of membrane ruffles76. In line with that, increased phosphorylation of the known PKA/PKG target VASP on serine residue 235 was observed in WT CyaA-treated BMDCs samples (c.f. Fig. 3C and Supplementary Table S1). This VASP site lies within the EVH2 domain that has been reported to account for VASP oligomerization and F-actin binding77. Phosphorylation of VASP then prevents actin polymerization by inhibiting the formation of profilin-actin complexes in myogenic cells50. This modification may thus complement the impact exerted on cytoskeleton rearrangements by CyaA-provoked RhoA inhibition. However, what causes this particular RhoA inhibition remains unknown.
Indeed, three possible mechanisms of small GTPase inhibition due to cAMP signaling can plausibly be invoked. The activities of small GTPases like RhoA, Cdc42 or Rac1 involve cycling between GDP-bound (inactive) and GTP-bound (active) states and are regulated by the interplay of activating GEFs (activating guanine nucleotide exchange factor) and inhibitory GAPs (GTPase-activating proteins)78. Besides, RhoA, as other small GTPases, can also be kept in an inactive state by guanine nucleotide dissociation inhibitors (GDIs)79. We observed that the action of CyaA provoked alterations of the phosphorylation state of several members of the GEF protein family, some of which were previously shown to directly promote RhoA activity80. For example, the protein known as Solo (or ARHGEF40) was shown to specifically induce RhoA activity in vascular endothelial cells81. Here, the treatment of BMDCs with CyaA resulted in dephosphorylation of the serine residue located in the region closest to the GEF domain of Solo. Another RhoA activator that is regulated by CyaA action is PDZRhoGEF (PRG; also known as ARHGEF11). This protein was found to be phosphorylated on the serine residue (c.f. Supplementary Table S1) that is localized in the center of the proposed key autoinhibitory element82, which may affect the conformation of the surrounding regions and may eventually lead to changes in PRG and affect its affinity for RhoA.
Among the GAPs known to be involved in RhoA regulation, three were found to be phosphorylated upon CyaA action on BMDCs (c.f. Supplementary Table S1), including the unconventional myosin-IXb (Myo9b), whose depletion was shown to impair the migratory capacity of dendritic cells and alter their interaction with T cells83. The other GAPs found to be regulated upon CyaA action was the ARHGAP18 protein, which is required for cellular polarization involved in cell migration84 and the ARHGAP11A protein, a key regulator of RhoA during cell division (also called MP-GAP or M Phase GAP)85.
The prolonged course of the whooping cough disease, often lasting for up to 12 weeks or longer, raises the hypothesis that B. pertussis infection may exert long-lasting effects on the cells of host mucosa and these would persist long after the bacteria have been cleared from the airway86,87. Although this effect is presumably due to the changes in epithelial cells, we hypothesize, that also airway intraepithelial DC might be affected similarly. Therefore, we examined the phosphorylation state of epigenetic effectors and chromatin remodeling proteins88 in CyaA-treated DC. Indeed, the class II histone deacetylase, HDAC5, was found to be dephosphorylated on serine 270 (serine 279 in human orthologue) in the nuclear localization signal. This would go well with the report of Taniguchi et al.89 that cAMP-stimulated nuclear import of HDAC5 requires dephosphorylation of Ser 270 by protein phosphatase 2 A (PP2A). Furthermore, CyaA action on DC provoked alterations of phosphorylation state of additional proteins that are involved in chromatin remodeling and epigenetic regulation. Among them, the Cbx5 gene-encoded HP1α protein, involved in propagation and maintenance of pericentric heterochromatin90, was found to be phosphorylated on serine 93. Another Chromo domain-containing protein that was significantly phosphorylated (on serine 305) upon CyaA treatment of BMDC was the transcriptional repressor CBX8, also known as PC391. In addition, significant phosphorylation of the replication-dependent histone H1.4 (Hist1h1e) on threonine 35 and serine 36 was observed. This goes well with the observation that H1.4 can be phosphorylated by PKA during mitosis and when it dissociates from mitotic chromatin92. CyaA-triggered changes in chromatin state and histone modifications could then also result in alteration of pre-mRNA splicing93 and numerous other effects on gene expression and cell cycle progression.
An important effect of CyaA toxin action, with relevance for host defense capacities, was observed for the phosphorylation state of the regulatory serine residues of the salt-inducible kinases (SIK). These are known for their involvement in modulation of immune responses57,58,59,60. Indeed, inhibition of SIKs by cAMP-activated and PKA-mediated phosphorylation was shown to suppress secretion of the pro-inflammatory cytokines IL-6, IL-12, and TNF-α by macrophages and appears to promote formation of regulatory (M2b) macrophages59. One of the SIK targets is the CREB-regulated transcription co-activator 3 (CRTC3) that can be phosphorylated at several serine residues. SIK-mediated phosphorylation was then shown to promote sequestering of CRTC3 into cytosolic complexes with the 14-3-3 protein, inhibiting CRTC3-assisted transcription of CREB-dependent genes, such as the anti-inflammatory cytokine IL-1059. In turn, phosphorylation of CRTC3 is inhibited by the cAMP-activated PKA and this enables translocation of the non-phosphorylated CRTC3 into the nucleus, where interaction of CRTC3 with CREB, activated by CyaA-produced cAMP94, may upregulate IL-10 gene transcription. In line with that we observed significant dephosphorylation of the serine 329 residue of CRTC3 and increased phosphorylation of the upstream regulatory kinases SIK1 (SIK), SIK2 (QIK) and SIK3 (QSK) in CyaA-treated BMDCs (c.f. Fig. 6). There was also enhanced phosphorylation of SIK1 on serine 577, which would promote the export of SIK1 from cell nucleus into the cytoplasm95, rescuing CRTC3-dependent transcription. Similarly, in CyaA-treated cells an increased phosphorylation of serine 343 at the critical regulatory site of SIK2/QIK was observed, where this modification was shown to inhibit the ability of SIK2/QIK to phosphorylate CRTC360. Moreover, we also observed the enhanced phosphorylation of serine 358 of SIK2. This modification is known to promote cytoplasmic localization of SIK2 into complexes that contain the 14-3-3 proteins96. Last but not least, SIK3 phosphorylation was also observed. All of these results are thus in line with previous reports that TLR-activated DCs exposed to CyaA secrete increased amounts of the IL-10 cytokine and promote IL-10 secretion by regulatory T cells74. The sum of the data then allows proposing a model of immunosubversive signaling activated by CyaA in myeloid cells, as depicted in Fig. 7. It predicts that activation of PKA isoforms by the CyaA-produced cAMP yields inhibitory phosphorylation of the SIK family kinases and CRTC3-activated transcription of the CREB-dependent IL-10 gene. Altogether these results document the pleiotropic immunosubversive effects of the massive cAMP signaling that is elicited in primary myeloid cells by the AC enzyme action of the CyaA toxin.
References
Rocha, G., Soares, P., Soares, H., Pissarra, S. & Guimarães, H. Pertussis in the newborn: certainties and uncertainties in 2014. Paediatr Respir Rev 16, 112–118, https://doi.org/10.1016/j.prrv.2014.01.004 (2015).
Snyder, J. & Fisher, D. Pertussis in childhood. Pediatr Rev 33, 412-420; quiz 420-411, https://doi.org/10.1542/pir.33-9-412 (2012).
Higgs, R., Higgins, S. C., Ross, P. J. & Mills, K. H. Immunity to the respiratory pathogen Bordetella pertussis. Mucosal Immunol 5, 485–500, https://doi.org/10.1038/mi.2012.54 (2012).
Linhartová, I. et al. RTX proteins: a highly diverse family secreted by a common mechanism. FEMS Microbiol Rev 34, 1076–1112, https://doi.org/10.1111/j.1574-6976.2010.00231.x (2010).
Ladant, D. & Ullmann, A. Bordetella pertussis adenylate cyclase: a toxin with multiple talents. Trends Microbiol 7, 172–176 (1999).
Cerny, O. et al. Bordetella pertussis Adenylate Cyclase Toxin Blocks Induction of Bactericidal Nitric Oxide in Macrophages through cAMP-Dependent Activation of the SHP-1 Phosphatase. J Immunol 194, 4901–4913, https://doi.org/10.4049/jimmunol.1402941 (2015).
Ahmad, J. N. et al. cAMP signalling of Bordetella adenylate cyclase toxin through the SHP-1 phosphatase activates the BimEL-Bax pro-apoptotic cascade in phagocytes. Cell Microbiol, https://doi.org/10.1111/cmi.12519 (2015).
Eby, J. C., Gray, M. C. & Hewlett, E. L. Cyclic AMP-mediated suppression of neutrophil extracellular trap formation and apoptosis by the Bordetella pertussis adenylate cyclase toxin. Infect Immun 82, 5256–5269, https://doi.org/10.1128/IAI.02487-14 (2014).
Kamanova, J. et al. Adenylate cyclase toxin subverts phagocyte function by RhoA inhibition and unproductive ruffling. J Immunol 181, 5587–5597, https://doi.org/10.4049/jimmunol.181.8.5587 (2008).
Mann, M. Functional and quantitative proteomics using SILAC. Nat Rev Mol Cell Biol 7, 952–958, https://doi.org/10.1038/nrm2067 (2006).
Karimova, G. et al. Charge-dependent translocation of Bordetella pertussis adenylate cyclase toxin into eukaryotic cells: implication for the in vivo delivery of CD8(+) T cell epitopes into antigen-presenting cells. Proc Natl Acad Sci USA 95, 12532–12537 (1998).
Fabrik, I. et al. Application of SILAC labeling to primary bone marrow-derived dendritic cells reveals extensive GM-CSF-dependent arginine metabolism. J Proteome Res 13, 752–762, https://doi.org/10.1021/pr4007798 (2014).
Rogers, L. D., Fang, Y. & Foster, L. J. An integrated global strategy for cell lysis, fractionation, enrichment and mass spectrometric analysis of phosphorylated peptides. Mol Biosyst 6, 822–829, https://doi.org/10.1039/b915986j (2010).
McNulty, D. E. & Annan, R. S. Hydrophilic interaction chromatography reduces the complexity of the phosphoproteome and improves global phosphopeptide isolation and detection. Mol Cell Proteomics 7, 971–980, https://doi.org/10.1074/mcp.M700543-MCP200 (2008).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26, 1367–1372, https://doi.org/10.1038/nbt.1511 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J Proteome Res 10, 1794–1805, https://doi.org/10.1021/pr101065j (2011).
Vizcaíno, J. A. et al. The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res 41, D1063–1069, https://doi.org/10.1093/nar/gks1262 (2013).
Klammer, M., Dybowski, J. N., Hoffmann, D. & Schaab, C. Identification of significant features by the Global Mean Rank test. PLoS One 9, e104504, https://doi.org/10.1371/journal.pone.0104504 (2014).
Chou, M. F. & Schwartz, D. Biological sequence motif discovery using motif-x. Curr Protoc Bioinformatics Chapter 13, Unit13.15–24, https://doi.org/10.1002/0471250953.bi1315s35 (2011).
Gnad, F. et al. PHOSIDA (phosphorylation site database): management, structural and evolutionary investigation, and prediction of phosphosites. Genome Biol 8, R250, https://doi.org/10.1186/gb-2007-8-11-r250 (2007).
Gnad, F., Gunawardena, J. & Mann, M. PHOSIDA 2011: the posttranslational modification database. Nucleic Acids Res 39, D253–260, https://doi.org/10.1093/nar/gkq1159 (2011).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods, https://doi.org/10.1038/nmeth.3901 (2016).
Keshava Prasad, T. S. et al. Human Protein Reference Database–2009 update. Nucleic Acids Res 37, D767–772, https://doi.org/10.1093/nar/gkn892 (2009).
Szklarczyk, D. et al. STRINGv10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43, D447–452, https://doi.org/10.1093/nar/gku1003 (2015).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25, 25–29, https://doi.org/10.1038/75556 (2000).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–2504, https://doi.org/10.1101/gr.1239303 (2003).
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093, https://doi.org/10.1093/bioinformatics/btp101 (2009).
Ladant, D., Glaser, P. & Ullmann, A. Insertional mutagenesis of Bordetella pertussis adenylate cyclase. J Biol Chem 267, 2244–2250 (1992).
Fiser, R. et al. Calcium influx rescues adenylate cyclase-hemolysin from rapid cell membrane removal and enables phagocyte permeabilization by toxin pores. PLoS Pathog 8, e1002580, https://doi.org/10.1371/journal.ppat.1002580 (2012).
Gonyar, L. A., Gray, M. C., Christianson, G. J., Mehrad, B. & Hewlett, E. L. Albumin, in the Presence of Calcium, Elicits a Massive Increase in Extracellular Bordetella Adenylate Cyclase Toxin. Infect Immun 85, https://doi.org/10.1128/IAI.00198-17 (2017).
Eby, J. C. et al. Quantification of the adenylate cyclase toxin of Bordetella pertussis in vitro and during respiratory infection. Infect Immun 81, 1390–1398, https://doi.org/10.1128/IAI.00110-13 (2013).
Olsen, J. V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648, https://doi.org/10.1016/j.cell.2006.09.026 (2006).
Hunter, T. & Sefton, B. M. Transforming gene product of Rous sarcoma virus phosphorylates tyrosine. Proc Natl Acad Sci USA 77, 1311–1315 (1980).
Serezani, C. H., Ballinger, M. N., Aronoff, D. M. & Peters-Golden, M. Cyclic AMP: master regulator of innate immune cell function. Am J Respir Cell Mol Biol 39, 127–132, https://doi.org/10.1165/rcmb.2008-0091TR (2008).
Aronoff, D. M., Canetti, C., Serezani, C. H., Luo, M. & Peters-Golden, M. Cutting edge: macrophage inhibition by cyclic AMP (cAMP): differential roles of protein kinase A and exchange protein directly activated by cAMP-1. J Immunol 174, 595–599, https://doi.org/10.4049/jimmunol.174.2.595 (2005).
Schwartz, D. & Gygi, S. P. An iterative statistical approach to the identification of protein phosphorylation motifs from large-scale data sets. Nat Biotechnol 23, 1391–1398, https://doi.org/10.1038/nbt1146 (2005).
Wolff, J., Cook, G. H., Goldhammer, A. R. & Berkowitz, S. A. Calmodulin activates prokaryotic adenylate cyclase. Proc Natl Acad Sci USA 77, 3841–3844 (1980).
Fiser, R. et al. Third activity of Bordetella adenylate cyclase (AC) toxin-hemolysin. Membrane translocation of AC domain polypeptide promotes calcium influx into CD11b + monocytes independently of the catalytic and hemolytic activities. J Biol Chem 282, 2808–2820, https://doi.org/10.1074/jbc.M609979200 (2007).
Pereira, L. et al. The cAMP binding protein Epac modulates Ca2 + sparks by a Ca2 + /calmodulin kinase signalling pathway in rat cardiac myocytes. J Physiol 583, 685–694, https://doi.org/10.1113/jphysiol.2007.133066 (2007).
Marumoto, T., Zhang, D. & Saya, H. Aurora-A - a guardian of poles. Nat Rev Cancer 5, 42–50, https://doi.org/10.1038/nrc1526 (2005).
Jantscher, F., Pirker, C., Mayer, C. E., Berger, W. & Sutterluety, H. Overexpression of Aurora-A in primary cells interferes with S-phase entry by diminishing Cyclin D1 dependent activities. Mol Cancer 10, 28, https://doi.org/10.1186/1476-4598-10-28 (2011).
Gray, M. C. & Hewlett, E. L. Cell cycle arrest induced by the bacterial adenylate cyclase toxins from Bacillus anthracis and Bordetella pertussis. Cell Microbiol 13, 123–134, https://doi.org/10.1111/j.1462-5822.2010.01525.x (2011).
Masin, J. et al. Negatively charged residues of the segment linking the enzyme and cytolysin moieties restrict the membrane-permeabilizing capacity of adenylate cyclase toxin. Sci Rep 6, 29137, https://doi.org/10.1038/srep29137 (2016).
Karst, J. C. et al. Identification of a region that assists membrane insertion and translocation of the catalytic domain of Bordetella pertussis CyaA toxin. J Biol Chem 287, 9200–9212, https://doi.org/10.1074/jbc.M111.316166 (2012).
Inoki, K., Li, Y., Zhu, T., Wu, J. & Guan, K. L. TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signalling. Nat Cell Biol 4, 648–657, https://doi.org/10.1038/ncb839 (2002).
Manning, B. D., Tee, A. R., Logsdon, M. N., Blenis, J. & Cantley, L. C. Identification of the tuberous sclerosis complex-2 tumor suppressor gene product tuberin as a target of the phosphoinositide 3-kinase/akt pathway. Mol Cell 10, 151–162 (2002).
Sancak, Y. et al. PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase. Mol Cell 25, 903–915, https://doi.org/10.1016/j.molcel.2007.03.003 (2007).
Kovacina, K. S. et al. Identification of a proline-rich Akt substrate as a 14-3-3 binding partner. J Biol Chem 278, 10189–10194, https://doi.org/10.1074/jbc.M210837200 (2003).
Courtemanche, N., Gifford, S. M., Simpson, M. A., Pollard, T. D. & Koleske, A. J. Abl2/Abl-related gene stabilizes actin filaments, stimulates actin branching by actin-related protein 2/3 complex, and promotes actin filament severing by cofilin. J Biol Chem 290, 4038–4046, https://doi.org/10.1074/jbc.M114.608117 (2015).
Harbeck, B., Hüttelmaier, S., Schluter, K., Jockusch, B. M. & Illenberger, S. Phosphorylation of the vasodilator-stimulated phosphoprotein regulates its interaction with actin. J Biol Chem 275, 30817–30825, https://doi.org/10.1074/jbc.M005066200 (2000).
Haining, A. W., Lieberthal, T. J. & Del Río Hernández, A. Talin: a mechanosensitive molecule in health and disease. FASEB J 30, 2073–2085, https://doi.org/10.1096/fj.201500080R (2016).
Vasioukhin, V., Bauer, C., Yin, M. & Fuchs, E. Directed actin polymerization is the driving force for epithelial cell-cell adhesion. Cell 100, 209–219 (2000).
Aresta, S., de Tand-Heim, M. F., Béranger, F. & de Gunzburg, J. A novel Rho GTPase-activating-protein interacts with Gem, a member of the Ras superfamily of GTPases. Biochem J 367, 57–65, https://doi.org/10.1042/BJ20020829 (2002).
Kagawa, Y. et al. Cell cycle-dependent Rho GTPase activity dynamically regulates cancer cell motility and invasion in vivo. PLoS One 8, e83629, https://doi.org/10.1371/journal.pone.0083629 (2013).
Consortium, G. O. Gene Ontology Consortium: going forward. Nucleic Acids Res 43, D1049–1056, https://doi.org/10.1093/nar/gku1179 (2015).
Huntley, R. P. et al. The GOA database: gene Ontology annotation updates for 2015. Nucleic Acids Res 43, D1057–1063, https://doi.org/10.1093/nar/gku1113 (2015).
Yong Kim, S. et al. Salt-inducible kinases 1 and 3 negatively regulate Toll-like receptor 4-mediated signal. Mol Endocrinol 27, 1958–1968, https://doi.org/10.1210/me.2013-1240 (2013).
Sundberg, T. B. et al. Small-molecule screening identifies inhibition of salt-inducible kinases as a therapeutic strategy to enhance immunoregulatory functions of dendritic cells. Proc Natl Acad Sci USA 111, 12468–12473, https://doi.org/10.1073/pnas.1412308111 (2014).
Clark, K. et al. Phosphorylation of CRTC3 by the salt-inducible kinases controls the interconversion of classically activated and regulatory macrophages. Proc Natl Acad Sci USA 109, 16986–16991, https://doi.org/10.1073/pnas.1215450109 (2012).
MacKenzie, K. F. et al. PGE(2) induces macrophage IL-10 production and a regulatory-like phenotype via a protein kinase A-SIK-CRTC3 pathway. J Immunol 190, 565–577, https://doi.org/10.4049/jimmunol.1202462 (2013).
Shen, Y. et al. Physiological calcium concentrations regulate calmodulin binding and catalysis of adenylyl cyclase exotoxins. EMBO J 21, 6721–6732 (2002).
Ma, L., Chen, Z., Erdjument-Bromage, H., Tempst, P. & Pandolfi, P. P. Phosphorylation and functional inactivation of TSC2 by Erk implications for tuberous sclerosis and cancer pathogenesis. Cell 121, 179–193, https://doi.org/10.1016/j.cell.2005.02.031 (2005).
Cerny, O., Anderson, K. E., Stephens, L. R., Hawkins, P. T. & Sebo, P. cAMP Signaling of Adenylate Cyclase Toxin Blocks the Oxidative Burst of Neutrophils through Epac-Mediated Inhibition of Phospholipase C Activity. J Immunol 198, 1285–1296, https://doi.org/10.4049/jimmunol.1601309 (2017).
Alessi, D. R. et al. Mechanism of activation of protein kinase B by insulin and IGF-1. EMBO J 15, 6541–6551 (1996).
Tee, A. R. et al. Tuberous sclerosis complex-1 and -2 gene products function together to inhibit mammalian target of rapamycin (mTOR)-mediated downstream signaling. Proc Natl Acad Sci USA 99, 13571–13576, https://doi.org/10.1073/pnas.202476899 (2002).
Han, J. W., Pearson, R. B., Dennis, P. B. & Thomas, G. Rapamycin, wortmannin, and the methylxanthine SQ20006 inactivate p70s6k by inducing dephosphorylation of the same subset of sites. J Biol Chem 270, 21396–21403 (1995).
Gingras, A. C. et al. Regulation of 4E-BP1 phosphorylation: a novel two-step mechanism. Genes Dev 13, 1422–1437 (1999).
Weichhart, T. et al. The TSC-mTOR signaling pathway regulates the innate inflammatory response. Immunity 29, 565–577, https://doi.org/10.1016/j.immuni.2008.08.012 (2008).
Schmitz, F. et al. Mammalian target of rapamycin (mTOR) orchestrates the defense program of innate immune cells. Eur J Immunol 38, 2981–2992, https://doi.org/10.1002/eji.200838761 (2008).
Thomson, A. W., Turnquist, H. R. & Raimondi, G. Immunoregulatory functions of mTOR inhibition. Nat Rev Immunol 9, 324–337, https://doi.org/10.1038/nri2546 (2009).
Haidinger, M. et al. A versatile role of mammalian target of rapamycin in human dendritic cell function and differentiation. J Immunol 185, 3919–3931, https://doi.org/10.4049/jimmunol.1000296 (2010).
Hackstein, H. et al. Rapamycin inhibits IL-4–induced dendritic cell maturation in vitro and dendritic cell mobilization and function in vivo. Blood 101, 4457–4463, https://doi.org/10.1182/blood-2002-11-3370 (2003).
Taner, T., Hackstein, H., Wang, Z., Morelli, A. E. & Thomson, A. W. Rapamycin-treated, alloantigen-pulsed host dendritic cells induce ag-specific T cell regulation and prolong graft survival. Am J Transplant 5, 228–236, https://doi.org/10.1046/j.1600-6143.2004.00673.x (2005).
Adkins, I. et al. Bordetella adenylate cyclase toxin differentially modulates toll-like receptor-stimulated activation, migration and T cell stimulatory capacity of dendritic cells. PLoS One 9, e104064, https://doi.org/10.1371/journal.pone.0104064 (2014).
Martinez-Quiles, N., Feuerbacher, L. A., Benito-León, M. & Hardwidge, P. R. Contribution of Crk adaptor proteins to host cell and bacteria interactions. Biomed Res Int 2014, 372901, https://doi.org/10.1155/2014/372901 (2014).
Lee, S. & Chung, C. Y. Role of VASP phosphorylation for the regulation of microglia chemotaxis via the regulation of focal adhesion formation/maturation. Mol Cell Neurosci 42, 382–390, https://doi.org/10.1016/j.mcn.2009.08.010 (2009).
Bachmann, C., Fischer, L., Walter, U. & Reinhard, M. The EVH2 domain of the vasodilator-stimulated phosphoprotein mediates tetramerization, F-actin binding, and actin bundle formation. J Biol Chem 274, 23549–23557 (1999).
Bos, J. L., Rehmann, H. & Wittinghofer, A. GEFs and GAPs: critical elements in the control of small G proteins. Cell 129, 865–877, https://doi.org/10.1016/j.cell.2007.05.018 (2007).
Cherfils, J. & Zeghouf, M. Regulation of small GTPases by GEFs, GAPs, and GDIs. Physiol Rev 93, 269–309, https://doi.org/10.1152/physrev.00003.2012 (2013).
Patel, M. & Karginov, A. V. Phosphorylation-mediated regulation of GEFs for RhoA. Cell Adh Migr 8, 11–18, https://doi.org/10.4161/cam.28058 (2014).
Abiko, H. et al. Rho guanine nucleotide exchange factors involved in cyclic-stretch-induced reorientation of vascular endothelial cells. J Cell Sci 128, 1683–1695, https://doi.org/10.1242/jcs.157503 (2015).
Bielnicki, J. A. et al. Insights into the molecular activation mechanism of the RhoA-specific guanine nucleotide exchange factor, PDZRhoGEF. J Biol Chem 286, 35163–35175, https://doi.org/10.1074/jbc.M111.270918 (2011).
Xu, Y. et al. Dendritic cell motility and T cell activation requires regulation of Rho-cofilin signaling by the Rho-GTPase activating protein myosin IXb. J Immunol 192, 3559–3568, https://doi.org/10.4049/jimmunol.1300695 (2014).
Maeda, M. et al. ARHGAP18, a GTPase-activating protein for RhoA, controls cell shape, spreading, and motility. Mol Biol Cell 22, 3840–3852, https://doi.org/10.1091/mbc.E11-04-0364 (2011).
Zanin, E. et al. A conserved RhoGAP limits M phase contractility and coordinates with microtubule asters to confine RhoA during cytokinesis. Dev Cell 26, 496–510, https://doi.org/10.1016/j.devcel.2013.08.005 (2013).
Melvin, J. A., Scheller, E. V., Miller, J. F. & Cotter, P. A. Bordetella pertussis pathogenesis: current and future challenges. Nat Rev Microbiol 12, 274–288, https://doi.org/10.1038/nrmicro3235 (2014).
Mattoo, S. & Cherry, J. D. Molecular pathogenesis, epidemiology, and clinical manifestations of respiratory infections due to Bordetella pertussis and other Bordetella subspecies. Clin Microbiol Rev 18, 326–382, https://doi.org/10.1128/CMR.18.2.326-382.2005 (2005).
Suárez-Álvarez, B., Baragaño Raneros, A., Ortega, F. & López-Larrea, C. Epigenetic modulation of the immune function: a potential target for tolerance. Epigenetics 8, 694–702, https://doi.org/10.4161/epi.25201 (2013).
Taniguchi, M. et al. Histone deacetylase 5 limits cocaine reward through cAMP-induced nuclear import. Neuron 73, 108–120, https://doi.org/10.1016/j.neuron.2011.10.032 (2012).
Maison, C. & Almouzni, G. HP1 and the dynamics of heterochromatin maintenance. Nat Rev Mol Cell Biol 5, 296–304, https://doi.org/10.1038/nrm1355 (2004).
Bárdos, J. I., Saurin, A. J., Tissot, C., Duprez, E. & Freemont, P. S. HPC3 is a new human polycomb orthologue that interacts and associates with RING1 and Bmi1 and has transcriptional repression properties. J Biol Chem 275, 28785–28792, https://doi.org/10.1074/jbc.M001835200 (2000).
Chu, C. S. et al. Protein kinase A-mediated serine 35 phosphorylation dissociates histone H1.4 from mitotic chromosome. J Biol Chem 286, 35843–35851, https://doi.org/10.1074/jbc.M111.228064 (2011).
Luco, R. F., Allo, M., Schor, I. E., Kornblihtt, A. R. & Misteli, T. Epigenetics in alternative pre-mRNA splicing. Cell 144, 16–26, https://doi.org/10.1016/j.cell.2010.11.056 (2011).
Perkins, D. J., Gray, M. C., Hewlett, E. L. & Vogel, S. N. Bordetella pertussis adenylate cyclase toxin (ACT) induces cyclooxygenase-2 (COX-2) in murine macrophages and is facilitated by ACT interaction with CD11b/CD18 (Mac-1). Mol Microbiol 66, 1003–1015, https://doi.org/10.1111/j.1365-2958.2007.05972.x (2007).
Katoh, Y. et al. Salt-inducible kinase-1 represses cAMP response element-binding protein activity both in the nucleus and in the cytoplasm. Eur J Biochem 271, 4307–4319, https://doi.org/10.1111/j.1432-1033.2004.04372.x (2004).
Henriksson, E. et al. The AMPK-related kinase SIK2 is regulated by cAMP via phosphorylation at Ser358 in adipocytes. Biochem J 444, 503–514, https://doi.org/10.1042/BJ20111932 (2012).
Sebo, P., Osicka, R. & Masin, J. Adenylate cyclase toxin-hemolysin relevance for pertussis vaccines. Expert Rev Vaccines 13, 1215–1227, https://doi.org/10.1586/14760584.2014.944900 (2014).
Acknowledgements
This work was supported by the grants NV16-28126A, GA13-145475 (P.S.), the GAUK grant No. 228216 (J.N.), the EATRIS-CZ project CZ.02.1.01/0.0/0.0/16_013/0001818 funded by the Operational Programme Research, Development and Education, and The long-term organization development plan ‘Medical Aspects of Weapons of Mass Destruction’ of the Faculty of Military Health Sciences, University of Defence of the Czech Ministry of Defence. J.N. is PhD student of the Charles University, Faculty of Science, Czech Republic. I.F. is a PhD student of the University of Defence in Brno. We thank Jasper E. Manning from Institute of Molecular Genetics of the CAS, v. v. i. for careful reading of the manuscript and editorial assistance.
Author information
Authors and Affiliations
Contributions
J.N. performed research, analyzed the data and wrote the paper; I.F. performed research, prepared, collected and processed the samples and wrote the paper; I.L. contributed to study design and performed research; M.L. measured the samples; O.Č. prepared samples and analyzed data; J.S. contributed to the study design and P.Š. contributed study design, data interpretation and wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
P.Š. is co-inventor on patents protecting use of CyaA as pertussis vaccine antigen and is a founder and shareholder of Revabiotech SE, which develops a next generation of whole cell pertussis vaccines.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Novák, J., Fabrik, I., Linhartová, I. et al. Phosphoproteomics of cAMP signaling of Bordetella adenylate cyclase toxin in mouse dendritic cells. Sci Rep 7, 16298 (2017). https://doi.org/10.1038/s41598-017-14501-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14501-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.