Abstract
Undifferentiated embryonic cell transcription factor 1 (Utf1) is expressed in pluripotent embryonic stem cells (ESCs) and primordial germ cells (PGCs). Utf1 expression is directly controlled by pluripotency factors Oct4 and Sox2, which form a ternary complex with the Utf1 enhancer. The Utf1 protein plays a role in chromatin organization and epigenetic control of bivalent gene expression in ESCs in vitro, where it promotes effective cell differentiation during exit from pluripotency. The function of Utf1 in PGCs in vivo, however, is not known. Here, we report that proper development of Utf1 null embryos almost entirely depends on the presence of functional Utf1 alleles in the parental germline. This indicates that Utf1’s proposed epigenetic role in ESC pluripotency in vitro may be linked to intergenerational epigenetic inheritance in vivo. One component - or at least facilitator - of the relevant epigenetic mark appears to be Utf1 itself, since Utf1-driven tomato reporter and Utf1 are detected in mature germ cells. We also provide initial evidence for a reduced adult testis size in Utf1 null mice. Our findings thus point at unexpected functional links between the core ESC pluripotency factor network and epigenetic inheritance of pluripotency.
Similar content being viewed by others
Introduction
Undifferentiated embryonic cell transcription factor 1 (Utf1) is primarily expressed in pluripotent embryonic stem cells (ESCs) and primordial germline cells (PGCs)1,2,3. General interest in Utf1 resulted from the demonstration that Utf1 expression in mouse and human cells is directly regulated by core pluripotency factors Oct4, Sox2, and most likely Nanog, which form a ternary complex with the Utf1 enhancer4,5,6. Furthermore, Utf1 expression is a reliable early marker for and potent facilitator of ex vivo and in vivo cell reprogramming to full pluripotency6,7,8,9,10,11.
A number of molecular functions have been assigned to Utf1 in ESCs, such as transcription factor1, chromatin organizer12, and epigenetic factor controlling H3K27me3 deposition at bivalent genes via binding to thousands of loci around transcriptional start sites13. The latter is particularly interesting because it connects the pluripotency core to the polycomb-repressive complex 2 (PRC2) network, and the deposition of epigenetic chromatin marks in ESCs. These data, in conjunction with Utf1 knock-down (kd) and knock-out (ko) mouse ESCs studies, pointed at an important regulatory role for Utf1 during exit from ESC pluripotency en route to effective cell differentiation12,13.
A subsequent Utf1 ko (Utf1(−/−)) mouse study, however, revealed that Utf1 is not critical for pluripotency, because viable and fertile Utf1(−/−) mice can be obtained14. A noticeable Utf1(−/−) phenotype reported in this study was developmental delay during embryogenesis that often caused smaller pups and neonatal death, which was attributed to reduced placental growth. However, the delay can be resolved during neonatal growth, thus resulting in phenotypically normal adult Utf1(−/−) mice14. The in vivo relevance of Utf1-controlled maintenance of pluripotency, as proposed for ESCs in vitro, therefore remains an unresolved issue13,15.
Here, we report important insights into this topic by employing novel Utf1-tomato pluripotency reporter mice that carry either Utf1(−/+) or Utf1(−/−) ko alleles due to the tdTomato replacement of Utf1 exon 1. Strikingly, by genotyping of embryos at 16 days post coitum (dpc), we provide evidence that a regulatory role of Utf1 in pluripotency maintenance is linked to an intergenerational epigenetic mark in parental germ cells. Based on our data and in conjunction with information available in the literature, we propose a model in which the Utf1 protein and/or Utf1 mRNA is a key component or a facilitator of such an epigenetic mark. We also made an interesting initial finding that hinted at a negative correlation between adult testis vascularization and size with the Utf1(−/−) genotype.
Results
Utf1-tomato reporter mice strain for pluripotency
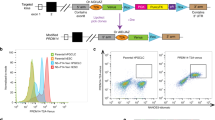
In order to probe into the in vivo relevance of the proposed Utf1-controlled maintenance of pluripotency13,15, we generated Utf1-tomato pluripotency reporter mice in which the tdTomato gene replaced a large segment of Utf1 exon 1 on chromosome 7. The inserted tdTomato gene carries two translational stop codons (Figure S1). Furthermore, the knock-in (ki) generated a +2 frameshift with respect to the unaltered part of the Utf1 coding sequence. Hence, the ki resulted in a Utf1 null allele, and we refer to the corresponding hetero- or homozygous transgenic animals as either Utf1(−/+)-tomato or Utf1(−/−)-tomato mice, respectively, and to the non-transgenic wild-type animals as Utf1(+/+).
First, we tested the functionality of the pluripotency reporter in vitro and derived fibroblasts from tail-tips of Utf1(−/+)-tomato mice. We subjected them to our established reprogramming protocol10,16 and found that clear fluorescent signals appeared in induced pluripotent stem cell (iPSC) colonies two weeks after “Yamanaka” factors transduction. The intensity of tdTomato expression correlated with iPSC/ESC colony morphology; i.e. colonies not resembling pluripotent stem cells showed weak or no detectable reporter expression (Fig. 1A, white arrows). Seven days later, iPSC colonies with sustained and strong reporter signals became visible (Fig. 1B).
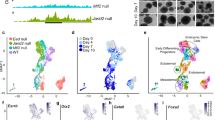
Characterization of Utf1-tomato pluripotency reporter mice. (A) iPSC colonies with ESC-like morphologies generated from Utf1-tomato mice begin to express the reporter around 14 days after the first “Yamanaka” factors transduction. (B) Expression of the tomato reporter in established iPSC colonies. Around 21 days after the “four factors” transduction, iPSC colonies show sustained and strong tdTomato expression. (C) An individual iPSC colony was followed during 12 days of DMSO-induced differentiation. (D) Utf1-tomato expression in mouse embryonic testis around 15 dpc, left: Utf1(−/+)-tomato embryonic testis; right: Utf1(−/−)-tomato embryonic testis. (E) Expression of Utf1-tomato in Utf1(−/+)-tomato (left) and Utf1(−/−)-tomato (right) ovaries at 15 dpc. (F) Utf1 protein expression in embryonic testes determined by Western Blotting (WB). A total protein lysate from mouse ESCs was used as control (left lane). The left panel shows that Utf1 is detectable in Utf1(+/+) and Utf1(−/+)-tomato embryonic testis, but not in lysates from Utf1(−/−)-tomato testis extracted from the same pregnant female that carried the Utf1(−/+)-tomato embryo. Bottom panel: WB showing equal amounts of histone H3 as loading control. Right panel: Ponceau S-staining of PVDF membrane as second loading control. The signals from core histones are indicated on the right.
We further tested our Utf1 reporter by DMSO-induced iPSC differentiation. While Tomato expression was maintained in the centre of iPSC colonies for several days, the outgrowing differentiated cells with fibroblast-like morphology had completely lost reporter signals (Fig. 1C). This had been observed earlier with different Utf1 reporter constructs2,6. Interestingly, the size of the fluorescent section in the centre of each colony did not change during 12 days of induced differentiation, indicating that cell proliferation and DNA replication at the edges of the central section triggers iPSC differentiation and concomitant down-regulation of Utf1. These results are in line with recent evidence for a functional link between the cell cycle and control of cell fate decisions in pluripotent stem cells17.
It has previously been shown that Utf1 is strongly expressed in gonocytes of rat testes during the embryonic and neonatal stage18. Furthermore, the highest expression level of Utf1 was observed in epiblast stage embryos (E6.5) and in PGCs (E12.5) in mice2. In order to verify that our reporter faithfully recapitulated known in vivo Utf1 expression patterns, we isolated testes from embryos at 15.5 dpc and found tdTomato expression in the cords of Utf1(−/+)-tomato and, at the expected higher levels, in Utf1(−/−)-tomato embryonic testes (Fig. 1D). In agreement with an earlier report2, we also detected reporter expression in PGCs of ovaries isolated at the same embryonic stage, although the signal intensities were clearly weaker (Fig. 1E).
We next determined Utf1 expression in embryonic testes by Western blotting and included mouse ESCs as positive control. It should be noted that the 36 kDa Utf1 protein is known to exhibit unusual mobility patterns in Western blots due to post-translational modification, such as phosphorylation19. We found that the vast majority of the protein was modified in embryonic Utf1(+/+) and in Utf1(−/+)-tomato testes, and that the protein was absent in all lysates prepared from testes obtained from different Utf1(−/−)-tomato embryos (Fig. 1F; Figure S2). The protein was also not detectable in embryonic mouse kidneys, irrespective of the genotype (Figure S2A). The latter result is in line with the known highly tissue-restricted Utf1 expression pattern during embryogenesis2,14. Taken together, we conclude that our Utf1-tomato reporter is a reliable indicator of Utf1 expression, and, as predicted, Utf1(−/−)-tomato mice do not express the Utf1 protein.
A functional parental Utf1 allele is important for Utf1(−/−) embryonic development
Utf1 is expressed in mouse and human PGCs, during oogenesis, and in the spermatogonial stem cell niche in testes3,18,20,21. The latter maintains the ability to self-renew and differentiate into functional spermatozoa22. However, functions for Utf1 in these important cell types are not known. Based on this well-established Utf1 expression pattern and the proposed role of Utf1 in epigenetic regulation of bivalent genes and maintenance of pluripotency in ESCs in vitro13,15, we became interested in interrogating whether Utf1 genotype frequencies of progenies might be affected by the parental Utf1 genotype. In order to prevent confounding effects due to neonatal developmental delays and associated postnatal lethality that we and others observed with Utf1(−/−)-tomato pups14, we decided to genotype progenies at embryonic stage 16 dpc.
Breeding Utf1(−/+)-tomato males with Utf1(−/+)-tomato females resulted in a 1:2:1 Mendelian Utf1 ratio with an average of 8.23 embryos/cage, which is in the normal litter size range for wild-type C57BL/6 J mice (Table 1). This result is in agreement with Utf1(−/+) breeding data from a previous study14 and indicated that Utf1(−) sperm cells produced by Utf1(−/+)-tomato males are functionally equivalent to Utf1(+) sperms, i.e. they produced the expected number of Utf1(−/+)-tomato and Utf1(−/−)-tomato embryos. However, breeding Utf1(−/−)-tomato males with Utf1(−/+)-tomato females produced embryos in a Utf1(−/+)-tomato:Utf1(−/−)-tomato ratio of about 2:1, instead of the expected 1:1 Mendelian ratio. We found that the reduced number of Utf1(−/−)-tomato embryos was statistically highly significant (p = 0.0012) (Table 1). As expected, the average number of embryos/cage was also reduced to 7.6. This reduction was not caused by a decrease in the reproductive potential of Utf1(−/−)-tomato reporter males, since, on average, they produced 4.9 Utf1(−/+)-tomato embryos/cage, which is even higher than the average 4.4 Utf1(−/+)-tomato embryos/cage produced by Utf1(−/+)-tomato males (Table 1). Instead, we frequently observed embryos which were arrested at various developmental stages (Fig. 2A). Hence, developmental arrest, probably in combination with uterine resorption, accounts for the observed reduction in the expected number Utf1(−/−)-tomato embryos.
Early embryonic developmental arrest. (A) Embryos arrested at different developmental stages from breeding Utf1(−/+)-tomato females and Utf1(−/−)-tomato males at 16 dpc. Embryos marked by red arrows in the uterus could not be isolated (upper panel). (B) Embryos from breeding Utf1(-/-)-tomato females and Utf1(−/−)-tomato males were developmentally arrested at different stages, which can be seen in the uterus (at 20dpc) (upper panels) and with isolated embryos (lower panels).
Together these data indicated that the 16 dpc embryonic development of Utf1(−/−)-tomato mice critically depended on the presence of at least one functional Utf1 allele in the paternal germ line. In order to test whether this also applied to the maternal germ line, we next examined progenies from breeding Utf1(−/−)-tomato males with Utf1(−/−)-tomato females. We found that out of fifty six breeding cages, only eight produced litters and a total of 14 viable pups, i.e. on average only 1.75 pups per litter. Examination of several pregnant females and their Utf1(−/−)-tomato embryos from this breeding revealed that they were developmentally arrested between 8 to 16 dpc, or earlier (Fig. 2B). This arrest was not caused by an uterine phenotype associated with Utf1(−/−)-tomato females, because breeding them with Utf1(−/+)-tomato males produced, on average, 7.1 viable pups/cage, which is in the range of the 6 to 7.6 pups/cage obtained with heterozygous Utf1(−/+)-tomato females (Table S1).
Utf1 is expressed during parental gametogenesis
Together our data showed that the full development potential of Utf1(−/−)-tomato embryos at 16 dpc critically depended on the parental Utf1 genotype, i.e. the presence of at least one functional maternal and paternal Utf1 allele. This can be explained by a role of Utf1 in intergenerational epigenetic inheritance23,24. Since it is known that Utf1 is expressed in the spermatogonial stem cell niche21,25, we hypothesized that a component or a facilitator of relevant intergenerational epigenetic mark(s) deposited in paternal germ cells is the Utf1 protein and/or mRNA. We therefore tested whether Utf1 promoter activity leads to the presence of fluorescent reporter in mature Utf1(−/−)-tomato sperm cells. Compared to an almost undetectable background in sperm cells isolated from Utf1(+/+) males, very strong tdTomato signals were detectable in the head and mid-piece regions of all examined sperm cells derived from two different Utf1(−/−)-tomato males at the age of 2.5 months (Fig. 3A; Figure S3A). We also analysed whether the reporter expression is maintained in older Utf1(−/−)-tomato males. Interestingly, we found that signal intensities were reduced and nearly lost in a number of sperm cells from males at the age of six to eight months (Fig. 3A; Figure S3A).
Tomato and Utf1 protein expression in sperm cells. (A) Expression of Tomato protein in sperm cells extracted from Utf1(−/−)-tomato males of different ages. Compared with control (sperm cells from age-matched Utf1+/+ mice), sperms from 2.5 months old mice showed strong tdTomato signals in the head and mid-piece regions. The bottom panels show that signal intensity became much more varied in 6 to 8 months old mice (see also Figure S3A). Magnification: 63X. (B) Utf1 protein can be detected in sperm cells of Utf1(+/+) and Utf1(−/+)-tomato mice. A lysate from mouse ESCs was used as positive control. The upper panel shows the WB result, and the lower panel depicts the corresponding membrane after Ponceau S-staining as loading control. Note that a faint non-specific band migrates above the specific Utf1 signal (see also Figure S3B).
In order to detect the Utf1 protein in sperm cells, we performed Western blotting and detected the Utf1 protein migrating slightly below a non-specific background signal in sperm lysates prepared from two different pairs of Utf1(+/+) and Utf1(+/-) males, but not in lysates from Utf1(−/−)-tomato males (Fig. 3B; Figure S3B). Interestingly, Utf1 has recently also been detected in various age groups of mouse metaphase II oocytes by both transcriptome and proteome analyses20. Combined these data reveal that Utf1 protein and mRNA are present in maternal and paternal wild-type Utf1 mouse germ cells.
Utf1 expression in PGCs and a potential role in testis development
Based on the high Utf1-tomato reporter activity observed in PGCs of Utf1(+/−) and Utf1(−/−)-tomato testes (Fig. 1D), we decided to examine this organ more carefully. Utf1(−/+)-tomato reporter mice were bred, and a pregnant female sacrificed at 15.5 dpc for the examination of testes from four Utf1(+/+) embryos. As expected, we found fully developed surface blood vessels on all eight organs (Fig. 4A). However, the four embryonic testes from Utf1(−/−)-tomato embryos that could also be obtained from the same pregnant female showed a marked reduction in visible blood vessels, which appeared much thinner and resulted in rather pale organs. This phenotype was also present, albeit less pronounced, with the two Utf1(−/+)-tomato embryonic testes that we could examine from the same pregnant female (Fig. 4A). Such genotype-specific differences in vasculature were not observed with embryonic ovaries from embryos that we extracted from another pregnant Utf1(−/+)-tomato female (Figure S4A).
Testicular vasculature and size in Utf1(−/−)-tomato testis. (A) Light microscopic images of embryonic testes. Compared with Utf1(+/+) and Utf1(−/+)-tomato testes, Utf1(−/−)-tomato testes exhibit fewer fully developed capillaries on the surface. (B) Immunofluorescence staining of mouse adult testis sections with IB4 (green) revealed that capillaries (yellow arrows) of Utf1(+/+) and Utf1(−/+)-tomato testes are better developed than those in Utf1(−/−)-tomato testes. (C) and (D) MRI analysis of adult testes of different genotypes obtained from the same males analysed in (B). Gradient echo images (FLASH, TR/TE = 1900/14 ms, resolution = 27 × 55 µm2, slice thickness = 0.4 mm, NA = 5, flip angles = 30°). 3 s and 13 s vertical sections of 2D MRI movies showed Utf1(+/+) and Utf1(−/+)-tomato testes have more pronounced and thicker blood vessels (red arrows) than Utf1(−/−)-tomato testes. The central images revealed that Utf1(+/+) and Utf1(−/−)-tomato testes have similar seminiferous tubules morphologies (purple arrows in (D). (See also SI movies for full organ examination). (E) Utf1(+/+), Utf1(−/+)-tomato and Utf1(−/−)-tomato pluripotency reporter mice from the same litter at 6 months of age are indistinguishable in appearance and size. (F) Testes from the adult Utf1(−/−)-tomato mice shown in (E) are much smaller than those from Utf1(+/+) and Utf1(−/+)-tomato mice, while the kidneys exhibited no size difference (see also Figure S6).
To probe further into the possibility of compromised testicular vasculature development in the Utf1(−/−)-tomato embryos, we examined the hemoglobin α contents in embryonic testes. Using SDS-PAGE combined with mass spectrometry, we found that the protein was present in high amounts in Utf1(+/+) and Utf1(−/+)-tomato testes, but clearly reached the detection limit of this method in Utf1(−/−)-tomato organs. These genotype-specific differences in blood content were not obvious in other embryonic organs, such as kidney, heart or ovaries (Figure S4B and S4C; Figure S5).
We next examined whether differences in vasculature also manifested in adult organs. Histological sections of adult testes obtained from three litter mates were analyzed by immunofluorescence using Isolectin IB4 Alexa Fluor 488, which is a marker for seminiferous tubules basement membranes and capillaries in the interstitial tissue26. Compared to the large and clearly visible capillaries in testes of Utf1(+/+) mice, those in Utf1(−/−)-tomato organs were much thinner and their appearance less frequent (Fig. 4B).
In order to confirm these results by a different method, we employed magnetic resonance imaging (MRI) to examine these testes. It confirmed that major capillaries of Utf1(−/−)-tomato adult testis are much thinner and less pronounced than those found in Utf1(−/+)-tomato and Utf1(+/+) organs (Fig. 4C; SI Movies 1–3). Based on our MRI analysis, we did not detect obvious genotype-specific differences in seminiferous tubules morphology (Fig. 4D; SI Movies 1–3). It appears, therefore, that lack of Utf1 expression in PGCs could affect testicular vasculature development during embryogenesis, which may also influence adult organ vasculature.
An impaired organ vascularization during development is known to have the potential to affect adult organ size27. We therefore examined the size of testes of adult Utf1(−/−)-tomato reporter mice. While some of these mice showed an overall reduced body size at birth, as reported earlier in a different genetic background14, their appearance became indistinguishable from wild-type and heterozygous littermates during postnatal development (Fig. 4E, Figure S6A,B). We found that the size and weight of testes was substantially reduced compared with testes of their Utf1(+/+) and Utf1(−/+)-tomato littermates (Fig. 4F, Figure S6A–D). We observed this with all ten testes from five Utf1(−/−)-tomato males that we were able to obtain together with the corresponding male littermate genotypes. We also examined kidneys and found no size or shape differences (Fig. 4F, Figure S6A,B). Hence, the correlation between reduced testis size and Utf1(−/−)-tomato genotype appears to be organ-specific.
We next examined the size of Utf1(−/−)-tomato embryonic testes and found differences in conjunction with the already described vascularization impairment at 18.5 dpc (Figure S7). However, since Utf1(−/−)-tomato embryos occasionally appear to be smaller than Utf1(−/+)-tomato and Utf1(+/+) embryos14, we cannot exclude that a general growth delay contributed to smaller testes at this stage. Interestingly, however, we observed that Utf1(−/−)-tomato embryos frequently exhibited a very pale appearance, indicating that organ vasculature development problems in the Utf1 null background could reach beyond testes (Figure S8). Together our data provide initial evidence for a potential role of Utf1 expression in PGSs in connection with testis development.
Discussion
The main result of our study on the in vivo function of Utf1 is that depending on the presence or absence of functional parental Utf1 alleles, Utf1(−/−)-tomato embryos in the next generation either develop normally or arrest at various stages of development, respectively. Such phenomenon can best be explained by intergenerational epigenetic inheritance rather than a classical paramutation23,24,28. In the latter case, we expect that the embryonic development phenotype should become detectable in F1 homozygous wild-type Utf1(+/+) embryos generated by Utf1(−/+)-tomato mice29. This is clearly not the case (Table 1). Furthermore, we demonstrated that Utf1 promoter-driven Tomato and the Utf1 protein is present in sperm cells of Utf1(−/−)-tomato and Utf1 wild-type males, respectively, and Schwarzer et al.20 identified the Utf1 protein and mRNA in stage II mouse oocytes.
Based on our data and information in the literature, we propose a model in which Utf1(−)-tomato sperm cells produced by Utf1(−/−)-tomato males lack a relevant epigenetic mark that is present in Utf1(−)-tomato sperms, which are produced in Utf1(−/+)-tomato testes (Fig. 5). It is clear from our data that such a mark also plays a role in the female germ line, because the development of the vast majority of embryos became arrested when Utf1(−/−)-tomato males were bred with homozygous Utf1(−/−)-tomato females, instead of Utf1(−/+)-tomato females (Fig. 5). Some of the earliest arrests may occur when the pluripotent primitive ectoderm begins to convert into the three primary germ layers30. Since the tomato reporter as well as the endogenous Utf1 protein is detectable in both mature sperm cells (this study) and oocytes20, we propose that a critical component of this epigenetic mark could be the Utf1 protein itself, most likely bound to parental DNA via its Myb domain2,13,19,31. The fact that the core pluripotency factors Oct4, Sox2, and Nanog, which control Utf1 expression in ESCs2,5, are also expressed in PGCs23 implies that the pluripotency core is directly connected to the proposed intergenerational epigenetic inheritance mediated by Utf1.
Model of an epigenetic role for Utf1 in intergenerational inheritance of pluripotency. Utf1(−)-tomato sperm cells produced by Utf1(−/−)-tomato males lack a relevant epigenetic mark that is present in Utf1(−)-tomato sperm cells, which are produced in Utf1(−/+)-tomato testes. Such a mark also plays a role in the female germ line, because the development of the vast majority of embryos became arrested when Utf1(−/−)-tomato males were bred with homozygous Utf1(−/−)-tomato females, instead of Utf1(−/+)-tomato females.
Jia et al.13 recently reported that Utf1 regulates PRC2 loading and H3K27me3 levels in mouse ESCs, thereby contributing to the control of bivalent gene expression. Furthermore, the study provided evidence that Utf1 is involved in the control of the stability of messenger RNAs transcribed from incompletely silenced bivalent genes in ESCs. Functionally, this dual gene regulatory role of Utf1 in mouse ESCs appears to be critical for the induction of proper ESC differentiation pathways in vitro13. Interestingly, recent studies indicated that H3K27me3 epigenetic marks are also present at bivalent domains of developmental regulatory genes in mouse PGCs, which are highly similar to the domains marked in ESCs32,33. In Drosophila melanogaster, H3K27-trimethylated nucleosomes remain heritably associated with silenced hox genes and can carry this epigenetic memory through multiple rounds of DNA replication34,35. Furthermore, this type of histone modification is also found at important gene regulatory regions in human sperm cells36,37,38. Since Utf1 is present in PGCs, spermatogonial stem cells, and functional sperm cells (Fig. 3A,B), we think it is possible that the protein contributes there to H3K27me3 deposition in a manner similar to its role in ESCs13. The same reasoning applies to oocytes, where Utf1 is also present20. It is also worth noting that the Utf1 gene itself is controlled by H3K27me3 epigenetic modification8, thus potentially indicating an epigenetic regulatory feedback mechanism.
Other, mutually not exclusive possibilities are that the parental Utf1 protein delivered by functional germ cells engages in interactions with target genes, components of the mRNA-decapping complex13 or, perhaps, other RNA modifying proteins, and thereby contributes to transcriptional or post-transcriptional regulation of gene expression during, for example, zygote-to-embryo transition39. In either case, such a contribution would ultimately become critical in the context of exit from pluripotency and efficient initiation of cell differentiation programs in the next generation embryos. Finally, it is also possible that only the Utf1 mRNA present in germ cells might play a functional role as an epigenetic mark.
Our study also presented initial findings which hinted at a contribution of Utf1 to the development of testicular vasculature and adult testes size. Clearly, this needs to be confirmed in other Utf1 ko genetic backgrounds. Given the high expression levels of Utf1 specifically in PGCs in male embryos, we think an interesting question to answer in the future is whether Utf1 is directly involved in cell signaling during testicular development.
In summary, because Utf1 is directly regulated by Oct4, Sox2, and, most likely, Nanog6, our data established the first example for intergenerational epigenetic inheritance of pluripotency mediated, at least in part, by a factor belonging to the core ESC pluripotency network. It will be very interesting to dissect in the future the molecular function(s) of the proposed Utf1-mediated epigenetic mark(s).
Materials and Methods
Generation of Utf1-tomato reporter mice
We generated an Utf1 gene target vector that carries a short arm of Utf1 homology, a tandem dimeric Tomato (tdTomato) cassette (Clonetech) with two stop codons, an excisable selection cassette, a long arm of homology, and a thymidine kinase screening cassette (Figure S1A). Homologous recombination in ESCs inserted Tomato into Utf1 exon 1, replacing the coding sequence for the first 173 amino acids which harbor its conserved DNA binding domain (Fukushima et al.19; Kooistra et al.12). The Tomato reporter is translated from the Utf1 start codon. Hence, with two stop codons at the end of the Tomato cassette, expression of Utf1 from a successfully targeted allele is not possible. The target vector was introduced into 129 ES cells and screened by PCR and Southern blotting for successful integration (data not shown). The genomic selection marker was subsequently excised, verified by PCR, and modified ES cells were injected into blastocysts. Resulting chimera were crossed with C57BL/6 J mice to generate germ-line-competent heterozygous Utf1-tomato mice. Heterozygous and homozygous animals were genotyped by PCR (Figure S1B), and sequences between arms of homology were verified by PCR/sequencing (Figure S1C; data not shown).
Genotyping
Utf1-tomato animals were genotyped by PCR, using tail-tip DNA as template, using primers utf-F: 5′ TGTCCCGGTGACTACGTCTGATGCC 3′; utf-R: 5′ ATCGTCCCCCAATAGCCCCATGAG 3′; tmt-F: 5′ TCAGGGCTCCGCCCCTCC CCAGGAG 3′; tmt-R: 5′ ACTTGGATCCAAGCTGGACATCACCTCCCACAACG 3′. The PCR was run with following settings: 95 °C 2 min, 35 cycles (95 °C 30 s, 55 °C 30 s, 72 °C 1 min), 72 °C 5 min. To verify the nucleotide sequence between the arms of homology, we used primers UTF1f: 5′ TCTCTGGTGAGGCCACGCCTTG 3′; UTF1r: 5′ TCGGGGTA GACTGGG AGTCG 3′, and amplified with the following setting: 95 °C 2 min, 35 cycles (95 °C 30 s, 56 °C 30 s, 72 °C 2 min), 72 °C 5 min.
IPSC reprogramming
Tail tips from Utf1(−/+)-tomato mice were used to generate fibroblasts in vitro for further induction to iPSCs. Viral supernatant from PLAT-E (Ectropic packaging cell line) cells, transfected with pMYc-based retroviral vectors16, was passed through 0.45 µm pore sized filters and supplemented with 4 μg/ml polybrene (Millipore). 6-well plates coated with 10 µg/ml fibronectin (r-fibronectin CH-296 from TAKARA) were centrifuged with the viral supernatant (no cells) for 60 min at 4000 rpm (4oC). 1 × 105 cells were added to each well containing the viral supernatant. After transduction for two consecutive days, cells were cultured in general MEF medium until day 5. Cells were then detached and seeded into a 6-well plate coated with approximately 2 × 104 inactivated MEF cells for each well. On day 7, the medium was replaced with mouse ESC medium (containing LIF). Culture medium was changed every day until iPSC colonies appeared. In order to differentiate generated iPSCs, 1% DMSO was added daily to differentiation medium (DMEM containing 20% FBS, without LIF) for the period of 12 days.
Animal experiments
The culture and experiments using C57BL/6 J reporter mice were performed according to protocols approved by the Biological Resource Centre (IACUC: #161125). For breeding, 2 to 8 months old transgenic mice were mated as single pairs or initially in trios (1 male and 2 females) and, subsequently, pregnant females were moved to a new cage before females dropped their litter. For isolation of embryonic testes and embryos, pregnant females were euthanized by CO2 and embryos were genotyped by PCR. To analyze adult mouse testis, male mice (2–6 months) were sacrificed.
Western Blotting
Proteins were extracted from embryonic testes or kidneys using a lysis buffer containing 10 mM Tris-HCl, pH7.4, 150 mM NaCl, 1 mM EDTA, 1% SDS in 1X PBS and 0.5 mM PMSF. The extracted samples were loaded on BoltTM 4–12% Bis-Tris Plus gradient gel (Invitrogen) and electrophoretically transferred to PVDF membrane. The protein loading was analyzed by Panceau S staining. After blocking with Super Block T20 (TBS) Blocking buffer (Thermo), the membrane was probed with primary antibody anti-Utf1 from rabbit (1:500 dilution, a24273 ABCAM). The antibody was detected with anti-rabbit HRP-conjugated secondary antibody (Dako). WB for mouse histone H3 as loading control employed polyclonal rabbit anti mouse histone H3 antibodies (1:5000 dilution; ab1971 ABCAM).
Mass spectrometry
Protein samples were loaded into BoltTM 4–12% Bis-Tris Plus gradient gel (Invitrogen), and the target protein gel band was excised and digested with trypsin as described40. After cleanup steps using C18 ZipTips (Millipore), the samples were mixed with an equal amount of matrix solution containing 10 mg/ml R-cyano-4-hydroxycinnamic acid in 0.1% TFA/50% ACN and spotted onto a 384-well stainless steel MALDI target plate (Applied Biosystems, Foster City, CA). An ABI 4800 Proteomics Analyzer MALDI TOF/TOF mass spectrometer (Applied Biosystems) was used to analyze the samples. For protein ID, combined MS and MS/MS data were submitted via GPS Explorer (version 3.6, Applied Biosystems) to Mascot server (version 2.0, Matrix Science). The swissprot database (including 545536 sequences, 194023197 residues) was utilized for the search.
Immunofluorescence staining
Testes were fixed in 4% paraformaldehyde at 4 degree for 6 hours and dehydrated in 30% sucrose overnight. OCT-embedded testes were cross sectioned at 10 µm thickness and stored in −80 degree. The slides were dried at room temperature overnight, and then washed with PBS three times. After blocking with 10% FBS in PBST, slides were incubated with isolectin IB4-Alexa Fluor488 conjugate (1:200 dilution, Invitrogen) at room temperature for 2 hours in the dark. They were subsequently washed with PBS three times, and sections were mounted and viewed under a fluorescence microscope.
MRI analysis
Testes of mice with different genotypes from the same litter were fixed with 4% formaldehyde, soaked in PBS overnight before MRI analysis. All MRI experiments were using a 14 Tesla Bruker Ascend 600WB vertical bore magnet, equipped with a MicWB40 micro-imaging probe in combination with a Micro2.5 gradient system. Images were obtained using Paravision 6.0.1 software. Excised testis samples were bedded on top of 1% agarose gel in a 10mm diameter glass tube, and a quadrature coil with an inner diameter of 25mm was used to transmit/receive the MR signals. Fast Low Angle Shot (FLASH) pulse sequence with gradient echo was used to acquire 2D images. Susceptibility weighted images (SWI) were reconstructed to obtain suitable contrast of blood vessels versus seminiferous tubules.
Sperm collection and protein extraction
Mature male mice (2 to 4 months) were sacrificed, then their cauda epididymides were removed and placed on sterile filter paper to blot away any blood and fluid. The removed cauda epididymides were put in a sperm dish containing 100 μL HTF media, then cut and dissected, and sperms were released. 2 μL of sperms were fixed with 4% formaldehyde, mounted and viewed under a confocal microscope. The sperm proteins were extracted as described41 and separated on a 10% SDS-PAGE gel for the Western Blot.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Change history
23 January 2018
A correction to this article has been published and is linked from the HTML version of this paper. The error has been fixed in the paper.
References
Okuda, A. et al. UTF1, a novel transcriptional coactivator expressed in pluripotent embryonic stem cells and extra-embryonic cells. EMBO J. 17, 2019–2032 (1998).
Galonska, C., Smith, Z. D. & Meissner, A. In Vivo and in vitro dynamics of undifferentiated embryonic cell transcription factor 1. Stem Cell Reports 2, 245–252 (2014).
Tang, W. W. et al. A Unique Gene Regulatory Network Resets the Human Germline Epigenome for Development. Cell 161, 1453–1467 (2015).
Nishimoto, M., Fukushima, A., Okuda, A. & Muramatsu, M. The gene for the embryonic stem cell coactivator UTF1 carries a regulatory element which selectively interacts with a complex composed of Oct-3/4 and Sox-2. Mol Cell Biol. 19, 5453–5465 (1999).
Reményi, A. et al. Crystal structure of a POU/HMG/DNA ternary complex suggests differential assembly of Oct4 and Sox2 on two enhancers. Genes Dev. 17, 2048–2059 (2003).
Tan, S. M., Wang, S. T., Hentze, H. & Dröge, P. A UTF1-based selection system for stable homogeneously pluripotent human embryonic stem cell cultures. Nucleic Acids Res. 35, e118 (2007).
Zhao, Y. et al. Two Supporting Factors Greatly Improve the Efficiency of Human iPSC Generation. Cell Stem Cell 3, 475–479 (2008).
Mansour, A. A. et al. The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature 488, 409–413 (2012).
Abad, M. et al. Reprogramming in vivo produces teratomas and iPS cells with totipotency features. Nature 502, 340–345 (2013).
Morshedi, A., Soroush Noghabi, M. & Dröge, P. Use of UTF1 genetic control elements as iPSC reporter. Stem Cell Rev. 9, 523–530 (2013a).
Yang, C. S., Chang, K. Y. & Rana, T. M. Genome-wide functional analysis reveals factors needed at the transition steps of induced reprogramming. Cell Rep. 8, 327–337 (2014).
Kooistra, S. M., Thummer, R. P. & Eggen, B. J. Characterization of human UTF1, a chromatin-associated protein with repressor activity expressed in pluripotent cells. Stem Cell Res. 2, 211–218 (2009).
Jia, J. et al. Regulation of pluripotency and self- renewal of ESCs through epigenetic-threshold modulation and mRNA pruning. Cell 151, 576–589 (2012).
Nishimoto, M. et al. In vivo function and evolution of the eutherian-specific pluripotency marker UTF1. PLoS ONE 8, e68119 (2013).
Laskowski, A. I. & Knoepfler, P. S. Utf1: Goldilocks for ESC bivalency. Cell Stem Cell 11, 732–734 (2012).
Morshedi, A., Ren, Z., Li, J. & Dröge, P. Probing into the Biological Processes Influenced by ESC Factor and Oncoprotein HMGA2 Using iPSCs. Stem Cell Rev. 9, 514–522 (2013b).
Boward, B., Wu, T. & Dalton, S. Control of Cell Fate Through Cell Cycle and Pluripotency Networks. Stem Cells 34, 1427–1436 (2016).
van Bragt, M. P. et al. Expression of the pluripotency marker UTF1 is restricted to a subpopulation of early A spermatogonia in rat testis. Reproduction 136, 33–40 (2008).
Fukushima, A. et al. Characterization of functional domains of an embryonic stem cell coactivator UTF1 which are conserved and essential for potentiation of ATF-2 activity. J Biol Chem. 273, 25840–25849 (1998).
Schwarzer, C. et al. Maternal age effect on mouse oocytes: new biological insight from proteomic analysis. Reproduction 148, 55–72 (2014).
Neuhaus, N. et al. Single-cell gene expression analysis reveals diversity among human spermatogonia. Mol Hum Rep. 23, 79–90 (2017).
Phillips, B. T., Gassei, K. & Orwig, K. E. Spermatogonial stem cell regulation and spermatogenesis. Philos. Trans. R. Soc. Lond. B Biol. Sci. 365, 1663–1678 (2010).
Tang, W. W., Kobayashi, T., Irie, N., Dietmann, S. & Surani, M. A. Specification and epigenetic programming of the human germ line. Nat Rev Genet. 17, 586–600 (2016).
van Otterdijk, S. D. & Michels, K. B. Transgenerational epigenetic inheritance in mammals: how good is the evidence? FASEB J. 30, 2457–2465 (2016).
Kristensen, D. M. et al. Presumed pluripotency markers UTF-1 and REX-1 are expressed in human adult testes and germ cell neoplasms. Hum Reprod. 23, 775–782 (2008).
Peters, B. P. & Goldstein, I. J. The use of fluorescein-conjugated Bandeiraea simplicifolia B4-isolectin as a histochemical reagent for the detection of [alpha]–galactopyranosyl groups: Their occurrence in basement membranes. Exp Cell Res. 120, 321–334 (1979).
Chung, A. S. & Ferrara, N. Developmental and pathological angiogenesis. Annu Rev Cell Dev Biol. 27, 563–584 (2011).
Hollick, J. B. Paramutation and related phenomena on diverse species. Nat Rev Gen. 18, 5–23 (2017).
Rassoulzadegan, M. et al. RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature 441, 469–474 (2006).
Surani, M. A., Durcova-Hills, G., Hajkova, P., Hayashi, K. & Tee, W. W. Germ line, stem cells, and epigenetic reprogramming. Cold Spring Harb Symp Quant Biol. 73, 9–15 (2008).
Kooistra, S. M. et al. Undifferentiated embryonic cell transcription factor 1 regulates ESC chromatin organization and gene expression. Stem Cells 28, 1703–1714 (2010).
Ng, J. H. et al. In vivo epigenomic profiling of germ cells reveals germ cell molecular signatures. Dev Cell 24, 324–333 (2013).
Sachs, M. et al. Bivalent chromatin marks developmental regulatory genes in the mouse embryonic germline in vivo. Cell Rep. 3, 1777–1784 (2013).
Coleman, R. T. & Struhl, G. Causal role for inheritance of H3K27me3 in maintaining the OFF state of a Drosophila HOX gene. Science 356, 6333, https://doi.org/10.1126/science.aai8236 (2017).
Laprell, F., Finkl, K. & Müller, J. Propagation of Polycomb-repressed chromatin requires sequence-specific recruitment to DNA. Science 356, 6333, https://doi.org/10.1126/science.aai8266 (2017).
Hammoud, S. S. et al. Distinctive chromatin in human sperm packages genes for embryo development. Nature 460, 473–478 (2009).
Brykczynska, U. et al. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat. Struct. Mol. Biol. 17, 679–687 (2010).
Erkek, S. et al. Molecular determinants of nucleosome retention at CpG-rich sequences in mouse spermatozoa. Nat. Struct. Mol. Biol. 20, 868–875 (2013).
Clit, D. & Schuh, M. Restarting life: fertilization and the transition from meiosis to mitosis. Nat Rev Mol Cell Bio. 14, 549–562 (2013).
Meng, W. et al. One-step procedure for peptide extraction from in-gel digestion sample for mass spectrometricanalysis. Anal Chem. 80, 9797–9805 (2008).
Kichine, E., Di Falco, M., Hales, B. F., Robaire, B. & Chan, P. Analysis of the sperm head protein profiles in fertile men: consistency across time in the levels of expression of heat shock proteins and peroxiredoxins. PLoS One 8, e77471 (2013).
Acknowledgements
This work was supported by the Singapore Ministry of Education through a Tier 2 grant (MOE2014-T2–005) and the National Medical Research Council, Singapore (NMRC1114/2007) (P.D.). MRI studies were supported by the Nanyang Institute of Structural Biology. W-P.Y. is supported by the Biomedical Research Council (BMRC), Singapore. We thank members of the P.D. laboratory and Xia Yun for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
Q.B. performed animal studies and molecular biology experiments. A.M. performed in vitro reprogramming. F.W., W-P.Y. and P.D. generated the Utf1-tomato reporter mouse strain. S.B., Q.B. and K.P. performed MRI analysis. Q.B. and P.D. designed the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Change History: A correction to this article has been published and is linked from the HTML version of this paper. The error has been fixed in the paper.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41598-018-20336-x.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bao, Q., Morshedi, A., Wang, F. et al. Utf1 contributes to intergenerational epigenetic inheritance of pluripotency. Sci Rep 7, 14612 (2017). https://doi.org/10.1038/s41598-017-14426-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14426-5
This article is cited by
-
Effects of early geometric confinement on the transcriptomic profile of human cerebral organoids
BMC Biotechnology (2021)
-
An Insight into the Role of UTF1 in Development, Stem Cells, and Cancer
Stem Cell Reviews and Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.