Abstract
The transforming growth factor-β (TGF-β) signaling pathway and its relevant genes have been correlated with an increased risk of developing various hallmarks of metabolic syndrome (MetS). In this study, we assessed whether the TGF-β signaling pathway-associated genes of SMAD family member 2 (SMAD2), SMAD3, SMAD4, transforming growth factor beta 1 (TGFB1), TGFB2, TGFB3, transforming growth factor beta receptor 1 (TGFBR1), and TGFBR2 are associated with MetS and its individual components independently, through complex interactions, or both in a Taiwanese population. A total of 3,000 Taiwanese subjects from the Taiwan Biobank were assessed. Metabolic traits such as waist circumference, triglyceride, high-density lipoprotein cholesterol, systolic and diastolic blood pressure, and fasting glucose were measured. Our results showed a significant association of MetS with the two single nucleotide polymorphisms (SNPs) of SMAD2 rs11082639 and TGFBR2 rs3773651. The association of MetS with these SNPs remained significant after performing Bonferroni correction. Moreover, we identified the effect of SMAD2 rs11082639 on high waist circumference. We also found that an interaction between the SMAD2 rs11082639 and TGFBR2 rs3773651 SNPs influenced MetS. Our findings indicated that the TGF-β signaling pathway-associated genes of SMAD2 and TGFBR2 may contribute to the risk of MetS independently and through gene–gene interactions.
Similar content being viewed by others
Introduction
The transforming growth factor-β (TGF-β) signaling pathway, a major intercellular signaling pathway in animal cells, plays a key role in regulating many aspects of cellular processes such as cell proliferation, cell differentiation, extracellular matrix production, embryonic development, cell adhesion, and cell apoptosis1,2. Given its widespread functionality, abnormalities in the TGF-β signaling pathway have been found to lead to diverse human diseases such as hypertension, hyperlipidemia, atherosclerosis, renal and cardiac fibrosis, and cancer, which are in turn associated with metabolic syndrome (MetS)3,4,5,6. The TGF-β family members include TGF-β, TGF-β type I receptor, TGF-β type II receptor, SMAD family member 2 (SMAD2), SMAD3, SMAD4, and other relevant proteins1,2. TGF-β ligands first bind to type II receptors, which then form a ligand–receptor complex with type I receptors1,2. The TGF-β type I receptor directly activates intracellular SMAD2 and SMAD3 proteins through their phosphorylation, which then mediate TGF-β signals. SMAD2 and SMAD3 proteins also target a common mediator, the SMAD4 protein, to form a heteromeric SMAD complex, which is subsequently translocated to the cell nucleus. The genes involved in the TGF-β signaling pathway include the SMAD2, SMAD3, SMAD4, transforming growth factor beta 1 (TGFB1), TGFB2, TGFB3, transforming growth factor beta receptor 1 (TGFBR1), and TGFBR2 genes.
The SMAD2, SMAD3, and SMAD4 genes are located on chromosome 18q21.1, 15q22.33, and 18q21.2, respectively. The proteins encoded by the SMAD2, SMAD3, and SMAD4 genes belong to the SMAD protein family, which mediates TGF-β signals1,2. By using imputation based on the 1000 Genomes Project, a genome-wide association study (GWAS) meta-analysis of the data from the CARDIoGRAMplusC4D Consortium indicated that the rs56062135 single nucleotide polymorphism (SNP) in the intron region of the SMAD3 gene is associated with coronary artery disease (CAD)7. A subsequent study by Turner et al. indicated that the SMAD3 rs17293632 SNP, which is in strong linkage disequilibrium (LD) with the SMAD3 rs56062135 SNP, may contribute to susceptibility to CAD8. In several animal studies, the TGF-β/SMAD signaling pathway has been correlated with the hallmarks of MetS, including insulin resistance, obesity, diabetes, and lipid metabolism9,10,11,12.
The TGFB1, TGFB2, and TGFB3 genes are located on chromosome 19q13.2, 1q41, and 14q24.3, respectively. The three structurally similar isoforms (TGF-β1, TGF-β2, and TGF-β3) encoded by the TGFB1, TGFB2, and TGFB3 genes are secreted ligands that belong to the TGF-β protein superfamily; these ligands bind to various TGF-β receptors to recruit and activate the SMAD protein family1,2. Studies have shown that TGFB1 SNPs are associated with various diseases such as CAD, stroke, chronic kidney diseases, and inflammatory diseases13. Evidence also supports the association of TGFB2 SNPs with end-stage renal disease14 and conotruncal heart defects15. Moreover, research has indicated that the TGFB3 rs11466414 SNP increases the risk of pregnancy-induced hypertension in a Hispanic population16.
The TGFBR1 and TGFBR2 genes are located on chromosome 9q22.33 and 3p24.1, respectively. The proteins encoded by the TGFBR1 and TGFBR2 genes belong to the TGF-β receptor subfamily; these proteins form a heterodimeric complex that binds to TGF-β proteins1,2. A previous study demonstrated that the TGFBR2 rs9838682 SNP is likely to influence the risk of sudden cardiac arrest in the setting of CAD in a Caucasian population17. In addition, the TGFBR2 rs6785358 SNP has been linked to a predisposition to congenital heart defects in a Chinese population18,19. Moreover, it has been observed that the overexpression of the TGF-β type I receptor and TGF-β type II receptor may be associated with autophagy and fibrogenesis in human heart diseases20. An animal study suggested that the deletion of the Tgfbr2 gene in hepatocytes influences systemic insulin resistance and body weight gain in mice during the development of nonalcoholic fatty liver disease, which is the hepatic manifestation of MetS12. Furthermore, several meta-analyses have suggested that TGFBR1 SNPs are associated with increased risks of breast, ovarian, and colorectal cancer21,22.
Based on the aforementioned findings, it is speculated that the TGF-β signaling pathway and its relevant genes play a key role in the development of MetS. Thus, we hypothesized that TGF-β signaling pathway-associated genes, namely SMAD2, SMAD3, SMAD4, TGFB1, TGFB2, TGFB3, TGFBR1, and TGFBR2 genes, may be linked to MetS. To the best of our knowledge, scant human studies have investigated the influence of these genes on MetS. Therefore, we conducted an association study to determine the relationships between susceptibility to MetS and SNPs in the SMAD2, SMAD3, SMAD4, TGFB1, TGFB2, TGFB3, TGFBR1, and TGFBR2 genes. We also assessed the effects of potential gene–gene interactions on MetS.
Results
Table 1 describes the demographic and clinical characteristics of the study population. First, we investigated the association between MetS and 8 TGF-β signaling pathway-associated genes, namely SMAD2, SMAD3, SMAD4, TGFB1, TGFB2, TGFB3, TGFBR1, and TGFBR2 genes. Based on LD, we filtered SNPs and selected 141 tag SNPs (Supplementary Table S2). Among the 141 tag SNPs assessed in this study (Supplementary Table S3), there were 20 tag SNPs, among those SNPs present in the SMAD2, SMAD3, SMAD4, TGFB2, TGFB3, and TGFBR2 genes, which showed evidence of an association (P < 0.05) with MetS (Table 2).
Furthermore, as shown in Table 2, the association of two key SNPs, namely SMAD2 rs11082639 and TGFBR2 rs3773651, with MetS remained significant after applying Bonferroni correction (P < 0.05/(141 × 3) = 0.0001). As demonstrated in Table 2, for the SMAD2 rs11082639 SNP, an increased risk of MetS was observed among subjects with MetS and those without MetS after adjustment for covariates such as age and sex for genetic models, including the additive model (odds ratio [OR] = 1.66; 95% confidence interval [CI] = 1.32–2.08; P = 1.4 × 10−5) and recessive model (OR = 2.82; 95% CI = 1.80–4.43; P = 6.6 × 10−6). Similarly, for the TGFBR2 rs3773651 SNP, an increased risk of MetS was observed among the subjects after adjustment for covariates for genetic models, including the dominant model (OR = 1.59; 95% CI = 1.28–1.98; P = 3.1 × 10−5). In addition, the rs11082639, rs1981, rs10853560, and rs948603 SNPs of the SMAD2 gene were found to be in strong LD (r2 > 0.8) to each other (Supplementary Table S2).
Table 3 shows the OR analysis of the association of two key SNPs, namely SMAD2 rs11082639 and TGFBR2 rs3773651, with the individual components of MetS: (a) high waist circumference vs. normal waist circumference; (b) high triglyceride vs. normal triglyceride; (c) low high-density lipoprotein (HDL) vs. normal HDL; (d) high blood pressure vs. normal blood pressure; and (e) high fasting glucose vs. normal fasting glucose. As shown in Table 3, for the SMAD2 rs11082639 SNP, an increased risk of high waist circumference was observed among the subjects after adjustment for covariates for genetic models, including the additive model (OR = 1.47; 95% CI = 1.17–1.84; P = 0.0008) and recessive model (OR = 2.19; 95% CI = 1.40–3.43; P = 0.0006). The effect of SMAD2 rs11082639 on high waist circumference remained significant after Bonferroni correction (P < 0.05/30 = 0.0017). In addition, we examined the association of the two implicated SNPs with MetS traits considered as continuous variables, including waist circumference, triglyceride, HDL, systolic blood pressure, diastolic blood pressure, and fasting glucose. As shown in Supplementary Table S4, the results suggest an association between SMAD2 rs11082639 and MetS traits such as waist circumference (P = 0.0048) or fasting glucose (P = 0.0027).
In addition, the generalized multifactor dimensionality reduction (GMDR) analysis was used to assess the effects of interaction between two key SNPs, namely SMAD2 rs11082639 and TGFBR2 rs3773651, on MetS and its individual components, with adjustment for the covariates of age and sex. Table 4 summarizes the results of GMDR analysis for two-way gene–gene interaction models with covariate adjustment to assess the effects of the two SNPs on MetS. As shown in Table 4, the two-way model involving SMAD2 rs11082639 and TGFBR2 rs3773651 was significant (P < 0.001). The effects of these two-way models remained significant after Bonferroni correction (P < 0.05/6 = 0.008). This finding indicated that a potential interaction between SMAD2 and TGFBR2 influences MetS. However, no two-way gene–gene interaction model influencing the individual components of MetS was obtained in this study.
Furthermore, we utilized multivariable logistic regression analysis with adjustment for age and sex to assess the two-way SMAD2 rs11082639 and TGFBR2 rs3773651 interaction models selected by the GMDR method (Table 5). Our analysis revealed that subjects with the CC genotype of SMAD2 rs11082639 and the AA genotype of TGFBR2 rs3773651 had a 2.5-fold higher risk of MetS than those with the T allele of SMAD2 rs11082639 and the AA genotype of TGFBR2 rs3773651 (Table 5). Subjects with the CC genotype of SMAD2 rs11082639 and the G allele of TGFBR2 rs3773651 also had a 5.85-fold higher risk of MetS than those with the T allele of SMAD2 rs11082639 and the AA genotype of TGFBR2 rs3773651 (Table 5). Finally, subjects with the T allele of SMAD2 rs11082639 and the G allele of TGFBR2 rs3773651 had a 1.55-fold higher risk of MetS than those with the T allele of SMAD2 rs11082639 and the AA genotype of TGFBR2 rs3773651 (Table 5).
Finally, statistical power analysis revealed that the present study had 99.9% power to detect associations of SMAD2 rs11082639 and TGFBR2 rs3773651 with MetS among the subjects with MetS and those without MetS.
Discussion
To date, our association study is the first to examine whether 141 tag SNPs in 8 TGF-β signaling pathway-associated genes, namely SMAD2, SMAD3, SMAD4, TGFB1, TGFB2, TGFB3, TGFBR1, and TGFBR2 genes, are significantly associated with the risk of MetS and its individual components independently, through gene–gene interactions, or both among Taiwanese individuals. Here, we report for the first time that the SMAD2 and TGFBR2 genes may play a key role in the development of MetS in a Taiwanese population. Notably, the significant association of two key SNPs, namely SMAD2 rs11082639 and TGFBR2 rs3773651, with MetS remained significant after Bonferroni correction (P < 0.0001). In addition, our data revealed that interactions between the SMAD2 and TGFBR2 genes may contribute to the etiology of MetS. Finally, our data revealed that the SMAD2 rs11082639 SNP was associated with the individual component of MetS, namely high waist circumference.
In the present study, we found a positive association of MetS with five SNPs in the SMAD2 gene, particularly the rs11082639 SNP. We also detected an association of SMAD2 rs11082639 with the dichotomous categorical variable of high waist circumference and the continuous variables of waist circumference and fasting glucose. The SMAD2 gene encodes the SMAD2 protein, which is recruited and phosphorylated by TGF-β receptors in response to TGF-β signals1,2. SMAD2 forms a heteromeric SMAD complex with SMAD4, which is subsequently translocated to the cell nucleus. Yang et al. reported that in Smad2-silenced cells in mice, gene expression related to lipogenesis was suppressed and gene expression related to β-oxidation was increased when Smad2 was inactivated by using an animal model of nonalcoholic steatohepatitis, one of the hepatic manifestations of MetS12. Their results indicated that the Smad signaling pathway is crucial for modulating lipid metabolism and lipid accumulation in hepatocytes through the suppression of lipogenesis-related genes and the induction of β-oxidation-related genes that promote the development of nonalcoholic steatohepatitis12. Several GWAS studies have been performed to investigate the genetic basis of MetS. For example, Kraja et al. performed a GWAS study on data from seven Caucasian cohorts and detected a significant association of triglyceride and glucose traits with the rs11820589 SNP in the BUD13 homolog (BUD13) gene and the rs12286037 SNP in the ZPR1 zinc finger (ZPR1) gene23. Moreover, another GWAS study by Kristiansson et al. implicated that the ZPR1 rs964184 SNP may be associated with lipid traits such as triglyceride and HDL in Finnish cohorts24.
In the present study, we found an association of MetS with four SNPs in the TGFBR2 gene, particularly the rs3773651 SNP. The TGFBR2 gene encodes the TGF-β type II receptor, which is a member of the Ser/Thr protein kinase family and is a key mediator of TGF-β signaling transduction1,2. Several association studies have identified that the TGFBR2 gene is associated with CAD17,20, congenital heart defects18,19, and various cancers25, which are some of the hallmarks of MetS. By analyzing glucose and insulin tolerance test results in an animal model of nonalcoholic steatohepatitis, Yang et al. demonstrated that silencing Tgfbr2 may contribute to MetS manifestations, including weight gain and insulin resistance; their finding indicated that Tgfbr2 is a potent mediator in the development of hepatic steatosis, hepatocyte death, inflammation, and fibrosis12.
Furthermore, we inferred the epistatic effects between the SMAD2 and TGFBR2 genes on MetS by using the GMDR approach. To the best of our knowledge, no other study has evaluated the interactions between these genes. In addition to determining the statistical significance of the interaction, we examined the potential biological mechanism underlying the interaction. The functional relevance of the interactive effects of SMAD2 and TGFBR2 on MetS remains to be elucidated. The SMAD2 and TGFBR2 genes encode the SMAD2 protein and TGF-β type II receptor, respectively, which are two core components of the TGF-β signaling pathway. Understanding the nature and extent of cross-talk between these two core components of the TGF-β signaling pathway is a critical future research direction. Our current understanding of the TGF-β signaling pathway is as follows1,2: First, the TGF-β ligand binds to the TGF-β type II receptor at the plasma membrane, resulting in the formation of the complex of the TGF-β type I receptor with the TGF-β type II receptor. Subsequently, the TGF-β type II receptor phosphorylates the TGF-β type I receptor. In turn, the activated TGF-β type I receptor phosphorylates SMAD2 and SMAD3 proteins. Finally, the phosphorylated SMAD2 and SMAD3 proteins form a complex with the SMAD4 protein, which is subsequently translocated into the nucleus for regulating the expression of specific target genes. Thus, regarding potential explanations for the biological effects of synergy between SMAD2 and TGFBR2, we speculate that the SMAD2 and TGFBR2 gene products may participate in a common pathogenic pathway, that is the TGF-β signaling pathway, leading to MetS.
This study has some limitations. The main weakness is that our findings require much more research to verify if the observations are replicated in diversified ethnic populations26,27,28. Moreover, in this study, a subject with MetS could be mistakenly classified as a subject without MetS because no records on the medications prescribed for treatment of dyslipidaemia, hypertension or diabetes were available. In the Taiwan Biobank, individuals were asked in a questionnaire whether a doctor had ever told them they had certain diseases such as hyperlipidemia, hypertension or diabetes. Thus, we further removed control subjects with a self-reported diagnosis of hyperlipidemia, hypertension or diabetes and then investigated the association between MetS and two key SNPs, namely SMAD2 rs11082639 and TGFBR2 rs3773651. The association of these two implicated SNPs with MetS remained significant after applying Bonferroni correction (Supplementary Table S5). Based on the candidate gene approach, the current findings are considered as only preliminary owing to the absence of supporting evidence from larger hypothesis-free GWAS studies23,24. Additional prospective clinical trials including other ethnic groups and conducting GWAS studies are warranted to thoroughly evaluate the association and interactions of the investigated genes with MetS and its individual components29,30,31.
In conclusion, we conducted an extensive analysis of the association and interactions of the TGF-β signaling pathway-associated genes with MetS and its individual components in Taiwanese subjects. Our findings indicated that the SMAD2 and TGFBR2 genes may affect the prevalence of MetS independently and through complex gene–gene interactions. Furthermore, the TGF-β signaling pathway-associated genes may be associated with the components of MetS. These findings contribute to accumulating evidence supporting that the TGF-β signaling pathway influences MetS. Further investigation with larger sample sizes is essential to provide more insights into the role of the TGF-β signaling pathway-associated genes investigated in this study.
Materials and Methods
Study population
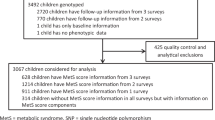
This study included Taiwanese subjects from the Taiwan Biobank. This biobank collected specimens and associated data from the general Taiwanese population with no history of cancer through recruitment centers across Taiwan during 2013–201532,33,34,35,36,37,38,39. Recruitment centers encompass regional and municipal hospitals, where advertisements were posted to recruit potential participants and incentives, such as free general health examinations and travel fee reimbursement, were offered to the participants32. This biobank is mainly funded by the Taiwanese government and aims to provide researchers with opportunities for collaboration to facilitate public health-related research concerning local common chronic diseases32. The study cohort consisted of 3,000 subjects. Individuals who could perform activities of daily living, were aged 30–70 years, and were self-reported as being of Taiwanese Han Chinese ancestry were included in this study33. Individuals with a history of cancer or nonresidents of Taiwan were excluded33. Ethical approval for this study was granted by the Institutional Review Board of the Taiwan Biobank before conducting the study. Each subject signed an approved informed consent form. All experiments were performed in accordance with relevant guidelines and regulations.
Metabolic syndrome
Measurements of metabolic traits including waist circumference, triglyceride, HDL cholesterol, systolic and diastolic blood pressure, and fasting glucose were obtained when subjects underwent general health examinations32,33,34. MetS was defined according to the International Diabetes Federation definition. Individual were considered to have MetS if they had central obesity (defined as waist circumference of ≥90 cm in male subjects and ≥80 cm in female subjects) and two or more of the following four components: (1) triglycerides ≥1.7 mmol/L; (2) HDL cholesterol <1.03 mmol/L in male subjects and <1.29 mmol/L in female subjects; (3) systolic blood pressure ≥130 mmHg or diastolic blood pressure ≥85 mmHg; and (4) fasting plasma glucose ≥5.6 mmol/L40. Two measurements of blood pressure were taken in both arms at least 10–15 minutes apart in the sitting position. These measurements were averaged to obtain the final blood pressure used in this study.
Genotyping
DNA was isolated from blood samples using a QIAamp DNA blood kit following the manufacturer’s instructions (Qiagen, Valencia, CA, USA). The quality of the isolated genomic DNA was evaluated using agarose gel electrophoresis, and the quantity was determined using spectrophotometry41. SNP genotyping was performed using custom Taiwan Biobank chips and was achieved using the Axiom Genome-Wide Array Plate System (Affymetrix, Santa Clara, CA, USA). To efficiently obtain maximal genetic information from the samples of subjects with Taiwanese Han Chinese ancestry, the custom Taiwan Biobank chips were designed using SNPs with minor allele frequencies (MAFs) ≥5% on the Axiom Genome-Wide CHB 1 Array (Affymetrix, Inc.), using SNPs in exons with MAFs >10% on the Human Exome BeadChip (Illumina, Inc., San Diego, CA, USA), and using SNPs previously reported in ancestry information panels, cancer studies, and pharmacogenetic studies33.
In this study, the SNP panel covered 261 SNPs from the following eight TGF-β signaling pathway-associated genes: SMAD2, SMAD3, SMAD4, TGFB1, TGFB2, TGFB3, TGFBR1, and TGFBR2 (Supplementary Table S1). Fifteen SNPs were excluded from further analysis due to failure to achieve the Hardy–Weinberg equilibrium (P < 0.05) or due to a genotyping call rate of <0.95. The genotyping results, including MAFs, P values for the Hardy–Weinberg equilibrium, and genotyping call rates, are shown in Supplementary Table S1. In addition, tag SNPs were identified using PLINK42, and an LD value (r²) of 0.8 was used as a threshold.
Statistical analysis
Categorical data were evaluated using the χ2 test. The Student’s t test was used to compare the difference in the means calculated from two continuous variables. To evaluate the association of the investigated SNPs with MetS, we conducted a logistic regression analysis to estimate the ORs and their 95% CIs, adjusting for covariates including age and sex43. Furthermore, we evaluated the association of the investigated SNPs with individual components of MetS by using logistic regression analysis, adjusting for age and sex44. We assessed whether the genotype frequencies were in the Hardy–Weinberg equilibrium by using the χ2 goodness-of-fit test with 1 degree of freedom (i.e., the number of genotypes subtracted from the number of alleles). Multiple testing was adjusted using Bonferroni correction. The criterion for significance was set at P < 0.05 for all tests. Data are presented as mean ± standard deviation.
We employed the GMDR method to investigate gene–gene interactions45. We tested two-way interactions using 10-fold cross-validation. GMDR software provides some output parameters, including the testing accuracy and empirical P values, to assess each selected interaction. Moreover, we used age and sex as covariates for gene–gene interaction models in our interaction analyses. Permutation testing provides empirical P values of prediction accuracy as a benchmark based on 1,000 shuffles. To correct for multiple testing, we applied a conservative Bonferroni correction factor for the number of tests employed in the GMDR analysis.
Based on the effect sizes in this study, the power to detect significant associations was evaluated using QUANTO software (http://biostats.usc.edu/Quanto.html).
Data Availability
The data that support the findings of this study are available from the Taiwan Biobank but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. To apply for access to these third party data please contact the Taiwan Biobank.
References
Kamato, D. et al. Transforming growth factor-beta signalling: role and consequences of Smad linker region phosphorylation. Cell Signal 25, 2017–2024, https://doi.org/10.1016/j.cellsig.2013.06.001 (2013).
Weiss, A. & Attisano, L. The TGFbeta superfamily signaling pathway. Wiley Interdiscip Rev Dev Biol 2, 47–63, https://doi.org/10.1002/wdev.86 (2013).
Toma, I. & McCaffrey, T. A. Transforming growth factor-beta and atherosclerosis: interwoven atherogenic and atheroprotective aspects. Cell Tissue Res 347, 155–175, https://doi.org/10.1007/s00441-011-1189-3 (2012).
Pickup, M., Novitskiy, S. & Moses, H. L. The roles of TGFbeta in the tumour microenvironment. Nat Rev Cancer 13, 788–799, https://doi.org/10.1038/nrc3603 (2013).
Loeffler, I. & Wolf, G. Transforming growth factor-beta and the progression of renal disease. Nephrol Dial Transplant 29(Suppl 1), i37–i45, https://doi.org/10.1093/ndt/gft267 (2014).
Zeng, L., Dang, T. A. & Schunkert, H. Genetics links between transforming growth factor beta pathway and coronary disease. Atherosclerosis 253, 237–246, https://doi.org/10.1016/j.atherosclerosis.2016.08.029 (2016).
Nikpay, M. et al. A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat Genet 47, 1121–1130, https://doi.org/10.1038/ng.3396 (2015).
Turner, A. W. et al. Functional Analysis of a Novel Genome-Wide Association Study Signal in SMAD3 That Confers Protection From Coronary Artery Disease. Arterioscler Thromb Vasc Biol 36, 972–983, https://doi.org/10.1161/ATVBAHA.116.307294 (2016).
Lin, H. M. et al. Transforming growth factor-beta/Smad3 signaling regulates insulin gene transcription and pancreatic islet beta-cell function. J Biol Chem 284, 12246–12257, https://doi.org/10.1074/jbc.M805379200 (2009).
Tan, C. K. et al. Smad3 deficiency in mice protects against insulin resistance and obesity induced by a high-fat diet. Diabetes 60, 464–476, https://doi.org/10.2337/db10-0801 (2011).
Yadav, H. et al. Protection from obesity and diabetes by blockade of TGF-beta/Smad3 signaling. Cell Metab 14, 67–79, https://doi.org/10.1016/j.cmet.2011.04.013 (2011).
Yang, L. et al. Transforming growth factor beta signaling in hepatocytes participates in steatohepatitis through regulation of cell death and lipid metabolism in mice. Hepatology 59, 483–495, https://doi.org/10.1002/hep.26698 (2014).
Martelossi Cebinelli, G. C., Paiva Trugilo, K. & Badaro Garcia, S. & Brajao de Oliveira, K. TGF-beta1 functional polymorphisms: a review. Eur Cytokine Netw 27, 81–89, https://doi.org/10.1684/ecn.2016.0382 (2016).
Ki, H. J. et al. Transforming growth factor-beta receptor 2 gene polymorphisms are associated with end-stage renal disease. Kidney Res Clin Pract 34, 93–97, https://doi.org/10.1016/j.krcp.2015.05.002 (2015).
Chen, Y. et al. Genetic variance of transforming growth factor beta2 gene in conotruncal heart defects. Biomarkers 22, 287–290, https://doi.org/10.1080/1354750X.2016.1217932 (2017).
Wilson, M. L., Desmond, D. H., Goodwin, T. M., Miller, D. A. & Ingles, S. A. Maternal and fetal variants in the TGF-beta3 gene and risk of pregnancy-induced hypertension in a predominantly Latino population. Am J Obstet Gynecol 201(295), e291–295, https://doi.org/10.1016/j.ajog.2009.05.038 (2009).
Tseng, Z. H. et al. Association of TGFBR2 polymorphism with risk of sudden cardiac arrest in patients with coronary artery disease. Heart Rhythm 6, 1745–1750, https://doi.org/10.1016/j.hrthm.2009.08.031 (2009).
Huang, F. et al. Association between TGFBR2 gene polymorphisms and congenital heart defects in Han Chinese population. Nutr Hosp 31, 710–715, https://doi.org/10.3305/nh.2015.31.2.8232 (2014).
Li, X. T. et al. Association of TGFBR2 rs6785358 Polymorphism with Increased Risk of Congenital Ventricular Septal Defect in a Chinese Population. Pediatr Cardiol 36, 1476–1482, https://doi.org/10.1007/s00246-015-1189-2 (2015).
Zou, M. et al. Autophagy inhibition of hsa-miR-19a-3p/19b-3p by targeting TGF-beta R II during TGF-beta1-induced fibrogenesis in human cardiac fibroblasts. Sci Rep 6, 24747, https://doi.org/10.1038/srep24747 (2016).
Liao, R. Y. et al. TGFBR1*6A/9A polymorphism and cancer risk: a meta-analysis of 13,662 cases and 14,147 controls. Mol Biol Rep 37, 3227–3232, https://doi.org/10.1007/s11033-009-9906-7 (2010).
Zhang, X. et al. The association of polymorphisms on TGFBR1 and colorectal cancer risk: a meta-analysis. Mol Biol Rep 39, 2567–2574, https://doi.org/10.1007/s11033-011-1009-6 (2012).
Kraja, A. T. et al. A bivariate genome-wide approach to metabolic syndrome: STAMPEED consortium. Diabetes 60, 1329–1339, https://doi.org/10.2337/db10-1011 (2011).
Kristiansson, K. et al. Genome-wide screen for metabolic syndrome susceptibility Loci reveals strong lipid gene contribution but no evidence for common genetic basis for clustering of metabolic syndrome traits. Circ Cardiovasc Genet 5, 242–249, https://doi.org/10.1161/CIRCGENETICS.111.961482 (2012).
Huang, Y. S., Zhong, Y., Yu, L. & Wang, L. Association between the TGFBR2 G-875A polymorphism and cancer risk: evidence from a meta-analysis. Asian Pac J Cancer Prev 15, 8705–8708 (2014).
Hsiao, T. J., Hwang, Y., Chang, H. M. & Lin, E. Association of the rs6235 variant in the proprotein convertase subtilisin/kexin type 1 (PCSK1) gene with obesity and related traits in a Taiwanese population. Gene 533, 32–37, https://doi.org/10.1016/j.gene.2013.10.016 (2014).
Hsiao, T. J. & Lin, E. A common rs7903146 variant of the transcription factor 7-like 2 gene is associated with type 2 diabetes mellitus and fasting glucose in a Taiwanese population. Diabetes Metab 43, 83–85, https://doi.org/10.1016/j.diabet.2016.05.003 (2017).
Lane, H. Y., Tsai, G. E. & Lin, E. Assessing gene-gene interactions in pharmacogenomics. Mol Diagn Ther 16, 15–27, https://doi.org/10.2165/11597270-000000000-00000 (2012).
Hsiao, T. J. & Lin, E. Evaluation of the glutamine 27 glutamic acid polymorphism in the adrenoceptor beta2 surface gene on obesity and metabolic phenotypes in Taiwan. J Investig Med 62, 310–315, https://doi.org/10.2310/JIM.0000000000000030 (2014).
Hsiao, T. J. & Lin, E. Association of a common rs9939609 variant in the fat mass and obesity-associated (FTO) gene with obesity and metabolic phenotypes in a Taiwanese population: a replication study. J Genet 95, 595–601 (2016).
Hsiao, T. J. & Lin, E. A Validation Study of Adiponectin rs266729 Gene Variant with Type 2 Diabetes, Obesity, and Metabolic Phenotypes in a Taiwanese Population. Biochem Genet 54, 830–841, https://doi.org/10.1007/s10528-016-9760-y (2016).
Fan, C. T., Lin, J. C. & Lee, C. H. Taiwan Biobank: a project aiming to aid Taiwan’s transition into a biomedical island. Pharmacogenomics 9, 235–246, https://doi.org/10.2217/14622416.9.2.235 (2008).
Chen, C. H. et al. Population structure of Han Chinese in the modern Taiwanese population based on 10,000 participants in the Taiwan Biobank project. Hum Mol Genet 25, 5321–5331, https://doi.org/10.1093/hmg/ddw346 (2016).
Lin, E. et al. Association and interaction of APOA5, BUD13, CETP, LIPA and health-related behavior with metabolic syndrome in a Taiwanese population. Sci Rep 6, 36830, https://doi.org/10.1038/srep36830 (2016).
Lin, E. et al. Effects of circadian clock genes and environmental factors on cognitive aging in old adults in a Taiwanese population. Oncotarget 8, 24088–24098, https://doi.org/10.18632/oncotarget.15493 (2017).
Lin, E. et al. Effects of circadian clock genes and health-related behavior on metabolic syndrome in a Taiwanese population: Evidence from association and interaction analysis. PLoS One 12, e0173861, https://doi.org/10.1371/journal.pone.0173861 (2017).
Lin, E. et al. Association and interaction effects of Alzheimer’s disease-associated genes and lifestyle on cognitive aging in older adults in a Taiwanese population. Oncotarget 8, 24077–24087, https://doi.org/10.18632/oncotarget.15269 (2017).
Lin, E. et al. The rs1277306 Variant of the REST Gene Confers Susceptibility to Cognitive Aging in an Elderly Taiwanese Population. Dement Geriatr Cogn Disord 43, 119–127, https://doi.org/10.1159/000455833 (2017).
Lin, E. et al. The ADAMTS9 gene is associated with cognitive aging in the elderly in a Taiwanese population. PLoS One 12, e0172440, https://doi.org/10.1371/journal.pone.0172440 (2017).
Liou, Y. J. et al. Gene-gene interactions of the INSIG1 and INSIG2 in metabolic syndrome in schizophrenic patients treated with atypical antipsychotics. Pharmacogenomics J 12, 54–61, https://doi.org/10.1038/tpj.2010.74 (2012).
Hsiao, T. J. & Lin, E. The ENPP1 K121Q polymorphism is associated with type 2 diabetes and related metabolic phenotypes in a Taiwanese population. Mol Cell Endocrinol 433, 20–25, https://doi.org/10.1016/j.mce.2016.05.020 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81, 559–575, https://doi.org/10.1086/519795 (2007).
Hsiao, T. J. & Lin, E. The Pro12Ala polymorphism in the peroxisome proliferator-activated receptor gamma (PPARG) gene in relation to obesity and metabolic phenotypes in a Taiwanese population. Endocrine 48, 786–793, https://doi.org/10.1007/s12020-014-0407-7 (2015).
Hsiao, T. J., Hwang, Y., Liu, C. H., Chang, H. M. & Lin, E. Association of the C825T polymorphism in the GNB3 gene with obesity and metabolic phenotypes in a Taiwanese population. Genes Nutr 8, 137–144, https://doi.org/10.1007/s12263-012-0304-8 (2013).
Lou, X. Y. et al. A generalized combinatorial approach for detecting gene-by-gene and gene-by-environment interactions with application to nicotine dependence. Am J Hum Genet 80, 1125–1137, https://doi.org/10.1086/518312 (2007).
Acknowledgements
The authors extend their sincere thanks to Vita Genomics, Inc. for this research. This work was supported by the Ministry of Economic Affairs in Taiwan (SBIR Grant S099000280249-154; E.L.), by Taipei Veterans General Hospital, Taiwan (Grants VGHUST103-G1-4-1, V105C-008, and V105E17-002-MY2-1; S.-J.T.), by the National Health Research Institutes, Taiwan (Grant NP-105-SP-04; Y.-L.L.), and by the Ministry of Science and Technology, Taiwan (Grant MST 102-2314-B-002-117-MY3; P.-H.K.).
Author information
Authors and Affiliations
Contributions
Study conception and design: E.L. and S.-J.T. Acquisition of data: P.-H.K., Y.-L.L., and A.C.Y. Analysis and interpretation of data: E.L. and S.-J.T. Draft manuscript: E.L. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, E., Kuo, PH., Liu, YL. et al. Transforming growth factor-β signaling pathway-associated genes SMAD2 and TGFBR2 are implicated in metabolic syndrome in a Taiwanese population. Sci Rep 7, 13589 (2017). https://doi.org/10.1038/s41598-017-14025-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14025-4
This article is cited by
-
Analysis of immunoinfiltration and EndoMT based on TGF-β signaling pathway-related genes in acute myocardial infarction
Scientific Reports (2024)
-
Exosome-Mediated Communication in Thyroid Cancer: Implications for Prognosis and Therapeutic Targets
Biochemical Genetics (2024)
-
ACSL4 accelerates osteosarcoma progression via modulating TGF-β/Smad2 signaling pathway
Molecular and Cellular Biochemistry (2024)
-
Runx1 is a key regulator of articular cartilage homeostasis by orchestrating YAP, TGFβ, and Wnt signaling in articular cartilage formation and osteoarthritis
Bone Research (2022)
-
Correlation between body mass index and prostate volume in benign prostatic hyperplasia patients undergoing holmium enucleation of the prostate surgery
BMC Urology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.