Abstract
Clinical treatment of the infections caused by various staphylococcal species differ depending on the actual cause of infection. Therefore, it is necessary to develop a fast and reliable method for identification of staphylococci. Raman spectroscopy is an optical method used in multiple scientific fields. Recent studies showed that the method has a potential for use in microbiological research, too. Our work here shows a possibility to identify staphylococci by Raman spectroscopy. We present a method that enables almost 100% successful identification of 16 of the clinically most important staphylococcal species directly from bacterial colonies grown on a Mueller-Hinton agar plate. We obtained characteristic Raman spectra of 277 staphylococcal strains belonging to 16 species from a 24-hour culture of each strain grown on the Mueller-Hinton agar plate using the Raman instrument. The results show that it is possible to distinguish among the tested species using Raman spectroscopy and therefore it has a great potential for use in routine clinical diagnostics.
Similar content being viewed by others
Introduction
Staphylococci are gram-positive bacteria inhabiting the skin and mucosal membranes of humans1,2. However, under certain circumstances (disruption of skin, diminution of immunity), they can cause infections of variable severity3. The most commonly found staphylococcal species in clinical material is Staphylococcus aureus. This pathogen can produce a broad range of virulence factors—haemolysins, proteases, leukocidins, toxic shock syndrome toxin, exfoliative toxins, enterotoxins plus immune-modulatory factors4—and can cause mild to severe infections including infections of skin and soft tissues, toxin-mediated food poisoning and toxic shock syndrome. After entering the bloodstream, S. aureus may become the cause of sepsis, endocarditis, osteomyelitis, meningitis or other life-threatening invasive diseases5. These infections are often hospital-acquired6 and caused by multiresistant strains6,7. Another coagulase-positive staphylococci that can rarely cause infections in humans (meningitidis, skin absceses) belong to the Staphylococcus intermedius group8,9.
Coagulase-negative staphylococci are, in contrast to S. aureus, saprophytic organisms living on human skin and are often found in clinical material as contaminants. However, in the recent years, they significantly contributed to the ever-increasing morbidity and mortality of nosocomial infections10. The majority of infections caused by coagulase-negative staphylococci are associated with biofilm formation that can occur on artificial materials and/or medical devices in the human body as well as on disrupted tissues of the patient2,11,12,13,14,15,16. These infections are difficult to treat and can be fatal. The most frequently isolated species from clinical specimens is Staphylococcus epidermidis 2. Other clinically important species that can be found in human clinical material include Staphylococcus haemolyticus (bloodstream infections, endocarditis, peritonitis and foreign-body infections)17,18,19,20, Staphylococcus lugdunensis and Staphylococcus saprophyticus (arthritis, urinary tract infections)17,21,22, Staphylococcus hominis, Staphylococcus caprae, Staphylococcus warneri, Staphylococcus capitis, Staphylococcus schleiferi, Staphylococcus xylosus and Staphylococcus auricularis (various infections)17,23, Staphylococcus sciuri (wounds, endocarditis)17,24,25, Staphylococcus simulans (cutaneous infections, osteoarticular infections, endocarditis)26 and Staphylococcus petrasii (ear infections)27.
As the character of infections caused by Staphylococcus aureus and coagulase-negative staphylococci differ, it is important to distinguish between these two groups and subsequently adjust the treatment. Therefore we tested the possibility to use a fast, relatively cheap, contactless, label-free spectroscopic method—Raman spectroscopy—for identification of the clinically most relevant staphylococcal species. In our comprehensive study on bacteria we have included 16 clinically relevant strains, with 277 sub-strains in order to cover most of the family of staphylococci. This could lead to the generation of a large reference database/library for Raman spectra of bacteria to unambiguously determine the identity of an unknown bacterial sample. Please note that our Raman spectral library presented here has been directed towards the clinical needs of the Department of Microbiology at St. Anne’s Faculty Hospital in Brno and can also be used as a starting point by other groups involved in clinical applications.
Raman spectroscopy is based on the shift between the frequency of incident and scattered light that is called the Raman effect. This shift is induced by molecular vibrations in the sample28,29 and makes Raman spectroscopy an useful tool for identification and characterization of biological systems30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50.
Materials and Methods
Microorganisms and sample preparation
The experiments included 277 staphylococcal strains: 62 Staphylococcus aureus strains, 8 Staphylococcus intermedius/pseudointermedius strains and 207 strains of coagulase-negative staphylococci (63 strains of Staphylococcus epidermidis, 21 strains of Staphylococcus haemolyticus, 21 strains of Staphylococcus hominis, 16 strains of Staphylococcus petrasii, 15 strains of Staphylococcus saprophyticus, 11 strains of Staphylococcus lugdunensis, 11 strains of Staphylococcus warneri, 9 strains of Staphylococcus sciuri, 8 strains of Staphylococcus schleiferi, 7 strains of Staphylococcus capitis, 7 strains of Staphylococcus xylosus, 6 strains of Staphylococcus auricularis, 6 strains of Staphylococcus caprae and 6 strains of Staphylococcus simulans). The majority of strains, excluding S. aureus and S. epidermidis strains, were kindly provided by Petr Petráš from the National Reference Laboratory for Staphylococci in Prague, Czech Republic. Furthermore, the set of analysed strains included eighteen reference strains (S. aureus CCM 7111, CCM 3953, CCM 4750 and CCM 4223, S. epidermidis CCM 7221 and CCM 2124, S. hominis CCM 2732, S. haemolyticus CCM 2729, S. capitis subsp. capitis CCM 2735, S. lugdunensis CCM 4068, S. caprae CCM 4546, S. schleiferi subsp. schleiferi CCM 4070, S. sciuri subsp. sciuri CCM 7040, S. saprophyticus subsp. saprophyticus CCM 2727, S. simulans CCM 2724, S. xylosus CCM 2725, S. warneri CCM 2731, S. intermedius CCM 4710) from the Czech Collection of Microorganisms (CCM) and one reference strain from the Czech National Collection of Type Cultures (CNCTC) – S. aureus CNCTC 7452. The other strains were clinical isolates stored in the Culture Collection of the Department of Microbiology, St. Anne’s Faculty Hospital in Brno, Czech Republic. All of the strains were identified using biochemical methods plus MALDI-TOF mass spectrometry and stored at −70 °C.
For the purpose of this experiment, the strains were thawed, inoculated onto Mueller-Hinton agar plates (MH, Oxoid, Basingstoke, United Kingdom) and cultivated for 24 hours at 37 °C. These conditions were selected in accordance with our previous work51.
Experimental setup
After cultivation, staphylococcal colonies were measured by a commercial Renishaw Raman spectrometer (Renishaw inVia Raman Spectrometer, Renishaw plc., Wotton-under-Edge, UK). Measurement settings were the same as described in our previous work51. Briefly, laser: single-mode diode, wavelength: 785 nm, microscope objective: Leica, Wetzlar, Germany, with numerical aperture 0.5, magnification: 500x, laser spot dimensions: approximately 2 μm × 10 μm, working distance: 0.5 mm, minimal number of measurements/strain: 10 (from at least 3 different bacterial colonies), spectral acquisition: 15 seconds. Each spectrum consists of 1015 points measured in the range 614–1724 cm−1.
The geometry described above ensures that the Raman signal is collected over an axial range of about 8 µm and, therefore, a contribution from cultivation media to the Raman spectra can be neglected51. Before each spectral acquisition, the laser was refocused onto a colony surface ensuring that the collected signal originates within the focal depth of the laser excitation and imaging optics.
On one day we acquired maximum of 200 Raman spectra. Measurements were performed in the same way on all measurement days. This applies also for the sample preparation. Reproducibility of microbial Raman spectra acquired this way was validated in our previous work51.
Data analysis
Raman spectroscopy typically suffers from a strong fluorescence background. Such background can be typically removed by various mathematical techniques52,53, such as polynomial baseline fitting, consequent numerical differentiation and integration, etc. It is even possible to suppress the fluorescence directly in the experiment using the frequency modulation of exciting laser54. Each method slightly disturbs the original spectrum and, therefore, may influence the quality of bacterial strain identification. We used two methods, namely Rolling-Circle Filter (RCF)55 (10 passes, 350 points circle radius) and iterative polynomial fitting (IPF)52 (maximum of 10 passes, 12th polynomial order). Subsequently, high-frequency noise was removed using Savistky-Golay filtering (2nd order, width 7points) and the spectra are normalized to the area of phenylalanine peak in the wavenumber range of 996–1009 cm−1 39.
Prior the spectral based identification we employed Principal Component Analysis (PCA) in order to extract the main features of the spectra and then use these features for bacterial strain identification. The whole ensemble of spectra is described by so-called loadings and principal component (PC) scores. The loadings represent an orthonormal coordinate base having the same dimension as is the number of measured Raman shifts and scores correspond to “coordinates” in this space. The PCA selects the loadings in a way that maximal variance of the original data is described by the first several scores. Therefore, it is typically sufficient to take into account only first 10–20 PC scores that would almost completely characterise the whole spectrum instead of the 1015 values corresponding to the intensities of each Raman shift.
In order to identify bacterial strains based on their Raman spectrum we used and compared 3 three methods typically used in computer science, namely Linear Discriminant Analysis (LDA)56, one nearest neighbor (1NN)57 and Support Vector Machine (SVM)58. These methods are often used in the field of computer vision or machine learning and were already used for Raman spectra identification53,59,60,61,62,63, too. They belong to the large group of supervised learning models, where the methods are initially “trained” using already known results.
In order to evaluate the performance of identification of staphylococci we used 5-fold cross-validation, i.e. the measured data set was randomly separated into 5 equally sized groups, the classification methods were trained using 80% of data and then the remaining 20% of data was classified using the trained model. For the identification we used MATLAB (Mathworks Inc., Natick, MA, USA) functions fitcdiscr, fitcknn, and fitcecoc (part of the Statistics and Machine Learning Toolbox of MATLAB). Furthermore, we optimised the number of PC scores used for bacterial identification in the range of 2–50.
Data Availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Results and Discussion
We collected Raman fingerprints from 277 staphylococcal strains (70 of coagulase-positive and 207 of coagulase-negative).
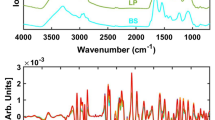
The averaged Raman spectra of each bacterial species (thick curves) are shown in Fig. 1. The fluorescent background was removed by the IPF method. The variance of spectra is marked by the gray area that borders 0,1st and 99,9th percentile of spectral intensity variations for a given Raman shift. Please note, that we used the percentiles instead of error bands based on stadard deviation of data since the measred spectral intensities are not normally distribured. The selected percentiles would correspond to commonly used 3σ interval for normally distributed data.
Averaged Raman spectra of all measured staphylococcal species (thick curves). The grayed area depicts the variations of measured spectral intensities corresponding to a given wavenumber. Border curves of this interval correpond to 0.1st (dashed) and 99.9th percentiles, respectively. Fluorescence spectral background was removed by the IPF method. The top-right inset compares both background removal methods (IPF and RC) on a single randomly selected spectrum of S. aureus.
The comparison of background removal methods can be seen in the inset of Fig. 1. A single, randomly selected spectrum of a S. aureus species is shown after filtering based on both IPF as well as RC background removal methods. Certain variations of less prominent spectral band might be seen but the positions and intensities of the most characteristic peaks are conserved.
The narrow phenylalalnine peak (at 1005 cm−1) is clearly present in all spectra and thus it is reasonable to use it as an internal standard for normalization. Furthermore, we observe quite strong variation of spectra even within one staphycoccal species which is depicted the separation of gray curves. One of the distinctive features is the presence or absence of peaks connected to carotenoids vibration in the wavenumber ranges 1110, 1160 and 1525 cm−1 corresponding to C-C-(CH3), =C-C=, and -C=C vibrations64. We see that certain species in our work do not exhibit those at all (such as Staphylococcus caprae, Staphylococcus (pseudo)intermedius, Staphylococcus schleiferi and Staphylococcus simulans) while the other strains exhibit medium or strong carotenoid signals. This is especially the case for Staphylococcus aureus, even though the carotenoid of this species varies strongly. Another feature noticable by naked eye is that Staphylococcus sciuri is the only species exhibiting peaks at 977 cm−1.
For identification of staphylococcal strains we used three methods: Linear Discriminant Analysis (LDA), one nearest neighbor (1NN) and Support Vector Machine (SVM). The accuracy of identification, i.e. relative count of correctly identified samples, depends both on the fluorescence background subtraction method as well as on the number of PC scores N considered for identification. Figure 2 shows the accuracy as a function of N used for identification for both IPF and RC background removal methods. One can see that 1NN and SVM exceed 98% accuracy for 10 PC scores and stay above this value up to N~30. Slightly better results are obtained for the IPF background removal method. Table 1 summarizes the best achieved results for each identification method as well as for both background removal methods.
Accuracy of staphycoccal strain identification for all three used methods (1 nearest neighbor, support vector machines and linear discriminant analysis) as a function of the number of used PCA scores N. Fluorescence background was removed using iterative polynomial fitting (a) or rolling circle (b) methods. Insets show magnified regions, with accuracy above 96%.
We can see that the highest level of identification is obtained using the 1NN algorithm that is applied on 14 PC scores and with the fluorescence background removal using IPF. Both 1NN and SVM give total successful identification around 99% with slightly better results for IPF background removal. LDA identification gives an accuracy of only around 87%.
Figure 3 shows the results of individual staphycoccal species identification in the form of the Confusion matrix65. We used the best approach shown in Table 1, i.e. the 1NN method, IPF background removal, and N = 14. Rows and columns of the Confusion matrix correspond to species identification by MALDI-TOF MS accompanied by biochemical testing (denoted as True Class) and species identification using Raman spectra (Predicted Class), respectively. The total number of correctly identified spectra is shown on the diagonal while the number of incorrectly identified strains is placed off-diagonal in a grayed background. These off-diagonal elements show how the spectra of a certain strains are mis-assigned to a different species. Furthermore, the rightmost columns show the sensitivity (True Positive Rate) and False Negative Rate), i.e. the relative count of Raman spectra measured from the given strains that was correctly or incorrectly identified using the 1NN method.
Confusion matrix showing the result of 5-fold cross-validated bacterial strain identification. Each row of the main part of table corresponds to bacterial species identified by MALDI-TOF MS plus biochemical methods (True Class) and each column corresponds to the bacterial identification predicted by the 1 nearest neighbor algorithm algorithm that employed iterative polynomial fitting background fluorescence removal and 14 PC scores. Numbers in the cells stand for correctly classified (diagonal) or missclassified (off-diagonal, red) spectra, respectively. The two rightmost columns show the sensitivity (True Positive Rate) and False Negative Rate), while two bottom columns show the Positive Predictive Value and the False Discovery Rate (values are rounded to integer values).
Similarly, the two bottom rows show the Positive Predictive Value and the False Discovery Rate, i.e. the relative count of properly identified spectra and spectra corresponding to different species that were identified as the one in the given column. Ideally, both True Positive Rate as well as Positive Predictive Value should be 100%. This was achieved for S. aureus, S. auricularis, S. (pseudo)intermedius, S.petrasii, S. sciuri, S. simulans and S. xylosus.
S. saprophyticus has a 100% Positive Predictive Value and a nearly non-zero False Negative Rate, i.e. no other spectra were identified as S. saprophyticus but one spectrum of this species (out of 150) was identified as S. epidermidis.
100% True Positive Rate and non-zero False Discovery Rate mean that all spectra of given species were identified correctly, but that some spectra of other species were incorrectly identified as the given species. This happened for S. caprae and S. heamolyticus. Probably the worst identification was obtained for S. capitis. However, both the False Negative Rate and the False Discovery Rate were still only 10 and 5 percent, respectively.
Even better results are achieved if we do not consider individual spectra but we look at the individual strains (10 spectra/strain). With this approach we get the overall success rate of 100%.
Moreover, we tested the performance of spectral identification on a reduced sample set without cross-validaton. Firstly, we randomly excluded approximately 20% of staphylococcal strains from the total set of measured spectra. We excluded at least one strain per species for identification of previously unknown strains. Consenquently, PCA was performed on the reduced spectral set and both the 1NN and SVN classificators were trained using 14 PC scores. In the next step, the excluded spectra were transformed into PC scores (using previously obtained loadings) that were identified using both methods. Depending on the seclection of excluded strains we achieved sucessfull identification of individual spectra in the range of 70–80%. However, if we consider only the strains that were completly mis-assigned to a single incorrect species by both methods we obtain that only up to 4% of excluded strains are completly mis-assigned. Thus, in this case we achieved also a very high accuracy of about 96%.
Our results show that it is possible to efficiently identify staphylococci using Raman spectroscopy. This is in good agreement with previous studies finding similar results for the identification of different species30,33,44,46,47. Recent studies also suggest that we can use Raman spectroscopy for the detection of antimicrobial resistance48,66 and other virulence factors like the ability to form biofilms37,41,43. These studies support the prospective use of Raman spectroscopy as a tool for microbial diagnostics. Moreover, the Raman fingerprints are highly reproducible. This was already proven for yeast spectra acquired in a time window of more than one year and the findings were supported by our recent work on S. aureus and S. epidermidis 38,51.
The high reproducibility coupled with the diagnostic potential presented here suggest the high potential of Raman spectroscopy for clinical diagnostics and that further research in this field is very promising. It is supported by the advantages of Raman spectroscopy including speed, low cost analyses and non-destructive nature. The non-destructive nature of the method allows for use of the sample in subsequent analyses. Raman spectroscopy does not require any time-consuming sample preparation nor the use of additional chemicals or materials, which is common for other routinely used biochemical methods or the commonly used MALDI-TOF mass spectroscopy.
Current disadvantages of Raman spectroscopy as a diagnostic tool include the absence of commercial databases for identification of bacterial spectra. This suggests the need for further investigations in this field that will help to build an automated diagnostic tool in the future. Also, in order to make a comparison of the spectra measured on our system with those measured on different Raman instruments we should consider that the quantum efficiency of the given detector and optical elements are wavelength dependent. This suggests that the data should be corrected according to the instrument response profile. In an ideal case the spectral sensitivity curve should be used so that the raw data from different systems can be transferred/evaluated. In many cases it is difficult to obtain the instrument response from the transmission and/or reflectivity of all optical elements in different Raman systems. Thus, we have corrected our data only for the quantum efficiency of the detector which can be readily obtained from the manufacturer. In the next step we used this corrected data for the analysis describe above to see any influence on the final data – we obtained the same results with only minor redistribution of off-diagonal elements in the confusion matrix. Also, the repeatability of our data collection has been proved by spectral identification on a reduced sample set with/without cross-validaton detailed in this section. However, we would like to note that for our “library” (developed for the Department of Microbiology at St. Anne’s Faculty Hospital in Brno) to be compared with spectra measured on different instruments the spectral sensitivity curves of given instruments should be provided. This can be obtained from a manufacturer on request.
Since Raman spectroscopy can also be used for the identification of certain virulence factors and antimicrobial resistance, it could make this method a very useful diagnostic tool providing a wide spectrum of characteristic information, in addition to strain identification, in one single measurement. That might significantly accelerate the diagnostic process.
In conclusion, analyses of Raman spectra acquired from 277 staphylococcal strains belonging to 16 species suggest that Raman spectroscopy can be used as a reliable tool for identification of staphylococci. We were able to achieve the total success rate of more than 99% for individual spectra and even 100% for a few individual strains.
Summary
The goal of the article was to assess Raman spectroscopy as a potential method for identification of 16 clinically important staphylococcal species since this method could accelerate the diagnostic process and lower the costs both for diagnostics and consequently treatment of patients.
References
Vandenbergh, M. F. & Verbrugh, H. A. Carriage of Staphylococcus aureus: epidemiology and clinical relevance. J. Lab. Clin. Med. 133, 525–534 (1999).
Piette, A. & Verschraegen, G. Role of coagulase-negative staphylococci in human disease. Vet. Microbiol. 134, 45–54 (2009).
Kocianova, S. et al. Key role of poly-g-DL-glutamic acid in immune evasion and virulence of Staphylococcus epidermidis. J. Clin. Invest. 115, 688–694 (2005).
Oogai, Y., Hashimoto, M., Kato, F., Sugai, M. & Komatsuzawa, H. Expression of Virulence Factors by Staphylococcus aureus Grown in Serum. Appl. Environ. Microbiol. 77, 8097–8105 (2011).
Lindberg, E. et al. Effect of lifestyle factors on Staphylococcus aureus gut colonization in Swedish and Italian infants. Clin. Microbiol. Infect. 17, 1209–1215 (2011).
van den Berg, S. et al. A human monoclonal antibody targeting the conserved staphylococcal antigen IsaA protects mice against Staphylococcus aureus bacteremia. Int. J. Med. Microbiol. 305, 55–64 (2015).
Cosgrove, S. E. The relationship between antimicrobial resistance and patient outcomes: mortality, length of hospital stay, and health care costs. Clin. Infect. Dis. 42(Suppl. 2), S82–S89 (2006).
Kelesidis, T. & Tsiodras, S. Staphylococcus intermedius is not only a zoonotic pathogen, but may also cause skin abscesses in humans after exposure to saliva. Int. J. Infect. Dis. 14, e838–e841 (2010).
Durdik, P. et al. Staphylococcus intermedius—rare pathogen of acute meningitis. Int. J. Infect. Dis. 14, e236–e238 (2010).
McCann, M. T., Gilmore, B. F. & Gorman, S. P. Staphylococcus epidermidis device-related infections: pathogenesis and clinical management. J. Pharm. Pharmacol. 60, 1551–1571 (2008).
Jansen, B., Hartmann, C., Schaumacher-Pedreau, F. & Peters, G. Late onset endopthalmitis associated with intraocular lens: a case of molecularly proved S. epidermidis aetiology. Br. J. Ophthalmol. 75, 440–441 (1991).
Verhoef, J. & Fleer, A. Staphylococcus epidermidis endocarditis and Staphylococcus epidermidis infection in an intensive care unit. Scand. J. Infect. Dis. Suppl. 41, 56–64 (1983).
Warren, J. W. Catheter-associated urinary tract infection. Int. J. Antimicrob. Agents 17, 299–303 (2001).
Rupp, M. E. & Archer, G. L. Coagulase-negative staphylococci: pathogens associated with medical progress. Clin. Infect. Dis. 19, 231–245 (1994).
Rupp, M. E. & Hamer, K. E. Effect of subinhibitory concentrations of vancomycin, cefazolin, ofloxacin, L-ofloxacin and D-ofloxacin on adherence to intravascular catheters and biofilm formation by Staphylococcus epidermidis. J. Antimicrob. Chemother. 41, 155–161 (1998).
Høiby, N. et al. ESCMID guideline for the diagnosis and treatment of biofilm infections 2014. Clin. Microbiol. Infect. 21, S1–S25 (2015).
Nanoukon, C. et al. Pathogenic features of clinically significant coagulase-negative staphylococci in hospital and community infections in Benin. Int. J. Med. Microbiol. 307, 75–82 (2017).
Perdoso, S. H. S. P. et al. Biofilm and toxin profile: A phenotypic and genotypic characterization of coagulase-negative staphylococci isolated from human bloodstream infections. Microb. Pathog. 100, 312–318 (2016).
Falcone, M. et al. Methicillin-Resistant Staphylococcal Bacteremia in Patients with Hematologic Malignancies: Clinical and Microbiological Retrospective Comparative Analysis of S. haemolyticus, S. epidermidis and S. aureus. J. Chemother. 16, 540–548 (2004).
Ertem, G. T. et al. Peritonitis due to teicoplanin-resistant Staphylococcus haemolyticus. Perit. Dial. Int. 30, 117–118 (2010).
Peel, T. N., Cole, N. C., Dylla, B. L. & Patel, R. Matrix-assisted laser desorption ionization time of flight mass spectrometry and diagnostic testing for prosthetic joint infection in the clinical microbiology laboratory. Diagn. Microbiol. Infect. Dis. 81, 163–168 (2015).
Lo, D. S., Shieh, H. H., Barreira, E. R., Ragazzi, S. L. & Gilio, A. E. High frequency of Staphylococcus saprophyticus urinary tract infections among female adolescents. Pediatr. Infect. Dis. J. 34, 1023–1025 (2015).
Delmas, J. et al. Evaluation of the Vitek 2 system with a variety of Staphylococcus species. J. Clin. Microbiol. 46, 311–313 (2008).
Marsou, R. et al. Distribution of Staphylococcus sciuri subspecies among human clinical specimens, and profile of antibiotic resistance. Res. Microbiol. 150, 531–541 (1999).
Giordano, N. et al. Erythema nodosum associated with Staphylococcus xylosus septicemia. J. Microbiol. Immun. Infect. 49, 134–137 (2016).
Shields, B. E., Tschetter, A. J. & Wanat, K. A. Staphylococcus simulans: An emerging cutaneous pathogen. JAAD Case Rep. 2, 428–429 (2016).
Pantůček, R. et al. Staphylococcus petrasii sp. nov. including S. petrasii subsp. petrasii subsp. nov. and S. petrasii subsp. croceilyticus subsp. nov., isolated from human clinical specimens and human ear infections. Syst. Appl. Microbiol. 36, 90–95 (2013).
Schie, I. W. & Huser, T. Methods and applications of Raman microspectroscopy to single-cell analysis. Appl. Spectrosc. 67, 813–828 (2013).
Read, D. S. & Whiteley, A. S. Chemical fixation methods for Raman spectroscopy-based analysis of bacteria. J. Microbiol. Meth. 109, 79–83 (2015).
Maquelin, K. et al. Prospective study of the performance of vibrational spectroscopies for rapid identification of bacterial and fungal pathogens recovered from blood cultures. J. Clin. Microbiol. 41, 324–329 (2013).
Afseth, N. K., Bloomfield, M., Wold, J. P. & Matousek, P. A. Novel approach for subsurface through-skin analysis of salmon using spatially offset Raman spectroscopy (SORS). Appl. Spectrosc. 68, 255–262 (2014).
Notingher, I. Raman spectroscopy cell-based biosensors. Sensors 7, 1343–1358 (2007).
Almarashi, J. F. M., Kapel, N., Wilkinson, T. S. & Telle, H. H. Raman spectroscopy of bacterial species and strains cultivated under reproducible conditions. Spectrosc. Int. J. 27, 361–365 (2012).
De Gelder, J., de Gussem, K., Vandenabeele, P. & Moens, L. Reference database of Raman spectra of biological molecules. J. Raman Spectrosc. 38, 1133–1147 (2007).
Martinelli, A. Effects of a protic ionic liquid on the reaction pathway during non-aqueous sol–gel synthesis of silica: A Raman spectroscopic investigation. Int. J. Mol. Sci. 15, 6488–6503 (2014).
Brauchle, E. & Schenke-Leyland, K. Raman spectroscopy in biomedicine—Non-invasive in vitro analysis of cells and extracellular matrix components in tissues. Biotechnol. J. 8, 288–297 (2013).
Samek, O., Al-Marashi, J. F. M. & Telle, H. H. The potential of Raman spectroscopy for the identification of biofilm formation by. Staphylococcus epidermidis. Laser Phys. Lett. 7, 378–383 (2010).
Rebrošová, K. et al. (2017) Differentiation between Staphylococcus aureus and Staphylococcus epidermidis strains using Raman spectroscopy. Future Microbiol. 12, 881–890 (2017).
Bernatová, S. et al. Following the mechanisms of bacteriostatic versus bactericidal action using Raman spectroscopy. Molecules 18, 13188–13199 (2013).
Samek, O. et al. Raman microspectroscopy of individual algal cells: Sensing unsaturation of storage lipids in vivo. Sensors 10, 8635–8651 (2010).
Sandt, C., Smith-Palmer, T., Pink, J., Brennan, L. & Pink, D. Confocal Raman microspectroscopy as a tool for studying the chemical heterogeneities of biofilms in situ. J. Appl. Microbiol. 103, 1808–1820 (2007).
Choo-Smith, L. P. et al. Investigating microbial (Micro)colony heterogeneity by vibrational spectroscopy. Appl. Environ. Microbiol. 67, 1461–1469 (2001).
Samek, O. et al. Candida parapsilosis Biofilm Identification by Raman Spectroscopy. Int. J. Mol. Sci. 15, 23924–23935 (2014).
Tien, N. I. et al. Diagnosis of bacterial pathogens in the dialysate of peritoneal dialysis patients with peritonitis using surface-enhanced Raman spectroscopy. Clin. Chim. Acta 461, 69–75 (2016).
Neugebauer, U., Rösch, P. & Popp, J. Raman spectroscopy towards clinical application: drug monitoring and pathogen identification. Int. J. Antimicrob. Agents 46, S35–S39 (2015).
Kotanen, C. N., Martinez, L., Alvarez, J. & Simecek, J. W. Surface enhanced Raman scattering spectroscopy for detection and identification of microbial pathogens isolated from human serum. Sens. Biosensing Res. 8, 20–26 (2016).
Pahlow, S. et al. Isolation and identification of bacteria by means of Raman spectroscopy. Adv. Drug Deliv. Rev. 89, 105–120 (2015).
Wulf, M. W. H. et al. The use of Raman spectroscopy in the epidemiology of methicillin-resistant Staphylococcus aureus of human- and animal-related clonal lineages. Clin. Microbiol. Infect. 18, 147–152 (2012).
Mathey, R. et al. Viability of 3 h grown bacterial micro-colonies after direct Raman identification. J. Microbiol. Methods 109, 67–73 (2013).
Schuster, K. C., Urlaub, E. & Gapes, J. R. Single-cell analysis of bacteria by Raman microscopy: spectral information on the chemical composition of cells and on the heterogeneity in a culture. J. Microbiol. Methods 42, 29–38 (2000).
Mlynáriková, K. et al. Influence of Culture Media on Microbial Fingerprints Using Raman Spectroscopy. Sensors 15, 29635–29647 (2015).
Liland, K. H., Almøy, T. & Mevik, B. H. Optimal choice of baseline correction for multivariate calibration of spectra. Appl. Spectrosc. 64, 1007–1016 (2010).
Cadusch, P. J., Hlaing, M. M., Wade, S. A., McArthur, S. L. & Stoddart, P. R. Improved methods for fluorescence background subtraction from raman spectra. J. Raman. Spectrosc. 44, 1587–1595 (2013).
De Luca, A. C., Mazilu, M., Riches, A., Herrington, C. S. & Dholakia, K. Online fluorescence suppression in modulated raman spectroscopy. Anal. Chem. 82, 738–745 (2010).
Brandt, N. N., Brovko, O. O., Chikishev, A. Y. & Paraschuk, O. D. Optimization of the rolling-circle filter for raman background subtraction. Appl. Spectrosc. 60, 288–293 (2006).
Fisher, R. A. The use of multiple measurements in taxonomic problems. Ann. Hum. Genet. 7, 179–188 (1936).
Sattlecker, M., Bessant, C., Smith, J. & Stone, N. Investigation of support vector machines and raman spectroscopy for lymph node diagnostics. Analyst 135, 895–901 (2010).
Altman, N. S. An Introduction to Kernel and Nearest-Neighbor Nonparametric Regression. Am. Stat. 46, 175–185 (1992).
Cortes, C. & Vapnik, V. Support-Vector Networks. Mach. Learn. 20, 273–297 (1995).
Harz, M. Micro-raman spectroscopic identification of bacterial cells of the genus staphylococcus and dependence on their cultivation conditions. Analyst 130, 1543–1550 (2005).
Seo, Y., Park, B., Hinton, A., Yoon, S. C. & Lawrence, K. C. Identification of staphylococcus species with hyperspectral microscope imaging and classification algorithms. Food. Measure. 10, 253–263 (2016).
Allen, V., Kalivas, J. H. & Rodriguez, R. G. Post-consumer plastic identification using raman spectroscopy. Appl Spectrosc 53, 672–681 (1999).
Dingari, N. C. et al. Development and comparative assessment of raman spectroscopic classification algorithms for lesion discrimination in stereotactic breast biopsies with microcalcifications. J. Biophotonics 6, 371–381 (2013).
Verma, S. P. et al. Resonance Raman spectra of beta-carotene in native and modified low-density lipoprotein. Biochem. Biophys. Res. Commun. 122, 867–875 (1984).
Fawcett, T. An introduction to ROC analysis. Pattern Recogn Lett 27, 861–874 (2006).
Spencer, A. R. et al. Staphylococcus aureus identification and antibiotic resistance determination using raman spectroscopy. J. Am. Coll. Surgeons 213, S49 (2011).
Acknowledgements
This work was supported by the Ministry of Health of the Czech Republic [grant numbers 16-29916 A, 16-31593 A]; the Czech Science Foundation [grant number GA15-20645S]; the Ministry of Education, Youth and Sports of the Czech Republic [LO1212, MUNI/A/0955/2016] and European Commission [ALISI ED0017/01/01].
Author information
Authors and Affiliations
Contributions
K.R., M.Š., O.S., P.Z., and F.R. are responsible for concept development, methodology, investigation and writing – original draft preparation. K.R., P.P. and F.R. were responsible for the species selection. S.B. and J.J. participated in methodology and performed the formal analysis. K.R., J.S. and P.P. participated in strain preparation. M.Š. and V.H. contributed to the data curation. They participated in writing – review & editing and visualisation. M.Š. is responsible for statistical analysis.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rebrošová, K., Šiler, M., Samek, O. et al. Rapid identification of staphylococci by Raman spectroscopy. Sci Rep 7, 14846 (2017). https://doi.org/10.1038/s41598-017-13940-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13940-w
This article is cited by
-
Nanotechnology in food and water security: on-site detection of agricultural pollutants through surface-enhanced Raman spectroscopy
Emergent Materials (2022)
-
Detection of multi-resistant clinical strains of E. coli with Raman spectroscopy
Analytical and Bioanalytical Chemistry (2022)
-
Rapid Identification of Different Pathogenic Spore-Forming Bacteria in Spice Powders Using Surface-Enhanced Raman Spectroscopy and Chemometrics
Food Analytical Methods (2022)
-
Microfluidic SERS devices: brightening the future of bioanalysis
Discover Materials (2022)
-
Raman microspectroscopy for microbiology
Nature Reviews Methods Primers (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.