Abstract
The goal of this work was to develop a mathematical model to predict Kaplan–Meier survival curves for chemotherapy combined with radiation in Non-Small Cell Lung Cancer patients for use in clinical trial design. The Gompertz model was used to describe tumor growth, radiation effect was simulated by the linear-quadratic model with an α/β-ratio of 10, and chemotherapy effect was based on the log-cell kill model. To account for repopulation during treatment, we considered two independent methods: 1) kickoff-repopulation using exponential growth with a decreased volume doubling time, or 2) Gompertz-repopulation using the gradually accelerating growth rate with tumor shrinkage. The input parameters were independently estimated by fitting to the SEER database for untreated tumors, RTOG-8808 for radiation only, and RTOG-9410 for sequential chemo-radiation. Applying the model, the benefit from concurrent chemo-radiation comparing to sequential for stage III patients was predicted to be a 6.6% and 6.2% improvement in overall survival for 3 and 5-years respectively, comparing well to the 5.3% and 4.5% observed in RTOG-9410. In summary, a mathematical model was developed to model tumor growth over extended periods of time, and can be used for the optimization of combined chemo-radiation scheduling and sequencing.
Similar content being viewed by others
Introduction
Non-Small Cell Lung Cancer (NSCLC) is a heterogeneous disease, which not only relates to its anatomical presentation, but also to the diverse biology1. As 40% of patients with NSCLC have locally advanced unresectable disease, combined chemotherapy and radiotherapy (CRT) is considered to be the first choice therapy for most of them2,3.
Though some gains in Overall Survival (OS) have been achieved over the past decades, long term survival in unresectable stage III NSCLC patients remains low and many efforts have been made to improve treatment. Mathematical modeling plays an important role for developing hypotheses to be tested in future clinical trials and for optimizing their design. Especially in the area of accelerated fractionation explored by the CHART and CHARTWEL trials, radiobiological models played a large role in trial design and estimating the therapeutic benefit4.
The trend to stratify patients further into smaller sub-groups and in-treatment adaptation approaches increase the importance of patient-specific modeling in lung cancer. The characterization of the effect of different treatment regimen is usually limited to the effect of radiation dose5,6, and sometimes include tumor volume7. However, it is well known that the combination and sequencing of chemotherapy with radiotherapy is playing an important role in the treatment of NSCLC8. There are few efforts to mathematically model these combination therapies, which would facilitate patient stratification and optimization of sequencing in combined chemoradiotherapy9.
In this paper, we develop a mathematical model to predict the Kaplan-Meier survival curve (KMsc), which is basically a cumulative survival frequency distribution curve, for a NSCLC patient population using a “Top-Bottom” methodology, i.e. using observable patient data as starting point. The aims are to
-
I.
Develop a NSCLC tumor growth model and patient death model using data of untreated patients, and compare it to clinical observations of tumor size and growth rates.
-
II.
Derive population distributions of radiation effect parameters using data of radiation-only trials and results from (I).
-
III.
Derive population distributions of chemotherapy effect parameters using data of sequential chemotherapy-radiation trials and results from (I) and (II).
The performance and viability of the model is evaluated and discussed with the published datasets.
Methods
Tumor growth modeling
Multiple methods of modeling the kinetics of tumor growth have been proposed10,11. One of the most popular models is exponential growth, which assumes the tumor can grow exponentially without any capacity constraints until the death of the patient. Exponential growth kinetics can describe tumor growth well when the tumor is relatively small, however, this is not the case for larger tumors considering the insufficient nutrition or vascularity of the tumor11. In order to account for the decreased growth rate with increased tumor volume, other models, e.g. the logistic and Gompertz model, were proposed. In this paper, we employed Gompertz growth, where the evolution of tumor cell number N (i.e. volume times cell density) with growth rate ρ is described by the following differential equation:
The growth rate \(\rho \) and carrying capacity \(K\) are the specific parameters determining the growth curve of a tumor. With this methodology, we are able to simulate one Gompertz growth curve for tumors with different initial volumes (e.g. different stages), enabling us to describe the clinical stages I through IV in one coherent framework.
In the Gompertz model, the volume doubling time is variable and not well defined as in the exponential growth model, where it is constant. In order to compare to data from the literature, we therefore used the same expression for the VDT as the exponential growth model, which is,
where t1 is the time of the first examination and t2 is the time of the second examination, \({v}_{t1}\) and \({v}_{t2}\) is the volume at t1 and t2 respectively. We assumed t2 occurs 1 year after t1 in this paper to compare to the literature values, which is similar to the observation periods used in most lung screening papers12,13.
Radiation effect modeling
The most commonly used tool for quantitative predictions of radiation effects is the linear quadratic (LQ) model14,15. The LQ model describes that the number of surviving cells after being irradiated by a certain dose of radiation takes the form of an exponential function with a linear and a quadratic term. In the typical dose range (under 10 Gy per fraction) and fractionation of clinical interest in stage III NSCLC, the LQ model shows good performance in terms of describing the radiation effect as a function of prescription dose. The formula can be expressed in differential equation as,
In our approach we assumed \(\alpha /\beta \) = 10, which is consistent with assessments from clinical trials for NSCLC patients16. The cell kill from radiation was determined by the patient-specific radiosensitivity parameter \({\rm{\alpha }}\) and \({\rm{\alpha }}/{\rm{\beta }}\). Radiosensitivity refers to the relative susceptibility of cells to the harmful effect of ionizing radiation.
Considering that cells with faster growth rate have shown to be more radiation-sensitive17,18,19, we also introduced a possible correlation between radio-sensitivity \(\alpha \) and the growth rate parameter \(\rho \).
Chemotherapy effect modeling
The log-cell kill model was used to simulate chemotherapy effects. The log-cell kill model subtracts a certain fraction of cancer cells based on the drug concentration regardless of the tumor size at the time of administration9,20,21. It can be expressed as,
where, \({\beta }_{c}\) represents the chemotherapy effect per dose, and \(C(t)\) is the drug concentration at a certain time point. In our model, the concentration was assumed as an exponential decay process,
Kaplan-Meier survival curve estimation
Death and tumor control conditions
In the literature the time of death is estimated to be about 41 doubling times, which corresponds to a tumor 13 cm in diameter22. In the present model, we therefore used the 13 cm diameter as the death condition. Besides the death from disease we implemented a 1.48% survival reduction according to life expectancy tables from the SEER data to account for unexpected natural death events.
To determine whether the tumor was controlled (i.e. no single clonogenic cell survives), we used the Poisson probability
which determines the probability of tumor control given N remaining cells. A random value was then generated between 0 and 1 to determine whether the specific tumor in the patient is controlled.
Initial volume
For the volume distribution in each stage, a log-normal distribution was assumed, as often seen in clinical patient series23. The range for each stage was defined based on the AJCC standard criteria24. The maximum of the volume distribution for stage I is 5 cm in diameter. For other stages, the volume can be any size, while it should be lower than the death condition (i.e. 13 cm). Considering the detection ability of the diagnostic technology, we use a diameter of 0.3 cm as the minimal detectable volume for all stages25. The tumor cell density was assumed to be 5.8 × 108 per cm3, as determined experimentally for lung cancer26. The choice of the density parameter should not be crucial in our model, as it simply scales everything in an equal manner and a different choice would not have an impact on the ability of the model to fit clinical data.
Monte Carlo Patient Population
To obtain a Kaplan-Meier survival curve, we used a Monte Carlo method to sample an initial patient population, where each patient has specific tumor and treatment properties, and survival time after treatment should be obtained for each patient. During the sampling process, normal distribution was used for growth and treatment effect parameters. The delay time, which is the time between diagnosis and the start of the treatment, was uniformly sampled from 2–3 weeks. In the fitting procedure, 10000 patients were sampled in each iteration in order to reduce the uncertainty. In the prediction studies, the same number as patients on the trial were sampled each time, which was repeated 100 times to estimate the uncertainty stemming from the limited population-volume of the specific trial.
Parameter estimation
We expressed the combined growth, chemotherapy and radiotherapy treatment for one patients as,
With this formula, we can derive the KMsc for a sampled patient population, that is for each patient we have the specific initial volume, the growth parameter, and the treatment response parameters. To derive the parameters based on the clinical observed KMsc, we formulized the cost function as,
where, \({{SF}}_{{model}}({Mo}{n}_{i})\) represent the survival fraction of patients who can survive longer than \({Mo}{n}_{i}\). The parameter of the model was derived by fitting to the observations from clinical trials, first only the growth term, then the radiation and chemotherapy terms separately. Consequently, the parameter estimation was a three stage process, as shown in Fig. 1, that is from top to bottom, the previously determined parameters were fixed and used as input for the next stage.
To derive the parameters that determine the tumor volume distribution and the growth pattern, the KMsc were fit to that of an untreated population collected in Detterbeck et al. in 2008 for stage I–IV disease22. The dataset includes the survival of 23954 patients with NSCLC who did not receive surgical resection, chemotherapy or radiotherapy.
With the resulting parameters being fixed we further derived the radiation effect parameter using a radiation-only clinical trial (RTOG 8808), with a prescribed dose of 60 Gy in 2 Gy fractions27.
For the parameters of the chemo-effect model, we used the 60 Gy arm of the RTOG 9410 trial which was designed to compare concurrent and sequential chemo-radiation therapy8. The enrolled patient numbers is over 200 per arm, prescription includes cisplatin at 100 mg/m3 on days 1 and 29 and vinblastine at 5 mg/m3 per week for 5 weeks. 60 Gy TRT started on day 50 and day 1 for the sequential and concurrent arm respectively. For the two drugs, we used the same \({\beta }_{c}\) for simplicity, with a unit of (mg/m3)−1. The half-life of the drugs was assumed to be 24 h, as measured in pharmacokinetic studies28,29.
Table 1 lists the complete list of the parameters that are used in the model. The subscripts of \({\rm{\mu }}\) and \({\rm{\sigma }}\) in the table represent the mean and sigma of the distribution for each variable. The optimization was constrained to avoid clearly implausible combinations of parameter values (as listed in Table 1). For example, the μ and σ for each parameter (including volume, \(\rho \), \(\alpha \) and \({\beta }_{c}\)) can not be negative, and the minimum detectable tumor diameter is ~3 mm. The model was implemented in python 3.5, for the fit process the lmfit package was applied using Powell’s method.
Results and Discussion
Tumor growth
Based on the survival curve of untreated population, the volume distribution and growth pattern for stage I–IV patients was derived. Considering that this procedure contains 13-dimensional (i.e. listed under the growth model section in Table 1) parameter space, which cannot be exhaustively explored in a heuristic fashion, we performed a two-stage optimization procedure to find a sensible and clinically relevant solution (i.e. within the constraints listed in Table 1). In the first stage, we used a pre-defined initial volume distribution taken from the literature (i.e. Stage I: 2.5 ± 2.5 cm, Stage II: 3.5 ± 3 cm, Stage III:6.6 ± 3 cm) to obtain the optimal parameters (i.e. carrying capacity K and growth parameter \(\rho \)) for the Gompertz growth model. Figure 2 illustrates the K and \(\rho \) surface with respect to the residuals for \({\rho }_{\mu }\) = 7 \(\times \) 10−5, revealing a valley with a minimum at K = 30 and \({\rho }_{\sigma }\) = 7.23 \(\times \) 10−3. In the second step we kept these optimal parameters (i.e. K and \(\rho \)) for the Gompertz growth model constant and re-optimized (i.e. finding the ones resulting in the minimal residual of the cost function) the initial volume distribution that was kept constant before.
Figure 3 shows the predicted Kaplan-Meier survival curves (KMsc) for the untreated populations for different stages together with the observed data. With the fitted initial volume distribution and the growth parameters (K and \(\rho \)), we obtained the distribution of volume doubling time (VDT) (based on equation 2) for patient populations of each stage during the first year after diagnosis. The one year interval was chosen because it is commonly used in most lung screening papers12,13. Table 2 lists the tumor volume distributions as predicted by the model fit for the median tumor diameter for stage I/II/III/IV, i.e. 1.2/3.5/6.9/9.7 cm, respectively. As an example, Figure 4 shows the volume and volume doubling time distributions for stage I patients. The values for stages I–III are close to observations from studies based on CT or radiographic imaging23,30. The estimated capacity is 30 cm in diameter, which has to be interpreted as a parameter to describe the slowing growth with increased tumor size, and not as a size to be reached or a realistic volume limit. Similarly, for the stage IV patients, the tumor volume here should not be interpreted as the solid volume of the primary tumor, but potentially multiple tumor nodules and metastases, and is more a description of “tumor load” in the entire patient. The median value of VDTs for the tumor were predicted to be 59/89/198/311 days for stage I/II/III/IV, which compares well to literature values12 , 31 even though deviations can be expected due to the dependence on the time interval32. For example, the mean VDT is between 93–452 days according to several publications12,31,33,34,35,36.
The exponential growth model was also implemented for comparison but resulted in about 40% lower median tumor volumes in stage III patients, not comparing well with the published data (e.g. i.e. Stage I: 2.5 ± 2.5 cm, Stage II: 3.5 ± 3 cm, Stage III:6.6 ± 3 cm)22,31 and not in line with published GTV (gross tumor volume) sizes at this stage37,38,39. Furthermore, the VDT is a constant of 80 days for all stages, which does not agree well with the literature especially for later stages (>100 days)12 and is not in agreement with the clinical findings that smaller tumors have a shorter VDT40.
Radiation Therapy
The radiation therapy response was modeled with the LQ model. Based on experimental observations, a correlation between the growth rate \(\rho \) and the radiosensitivity \(\alpha \) was considered. The underlying biological interpretation is that faster growing tumor cells spend a higher proportion of their time in mitosis, which impairs double strand break (DSB) repair and makes them more radiosensitive compared to slower growing ones17,18,19. In the Small-Cell variant of lung cancer this has also been shown in-vivo, as the growth rate measured by Ki-67 in patient samples correlated with the extent of volumetric response to CRT19. Figure 5 shows the distributions for \(\alpha \) and \(\rho \) on the left, and the survival fraction (i.e. fraction of cells retaining their reproductive integrity) distribution on the right. The survival fraction in Fig. 5c was obtained based on the LQ model from the derived \(\alpha \) distribution. As shown in Fig. 5b, the best fit is gained by assuming a slight correlation (coefficient = 0.87) between a patient’s growth rate and radiosensitvity. Figure 5b demonstrates the superior fit (i.e. lower residual value of the cost function) of the survival curve with correlation comparing to without correlation. The estimated median values for \(\alpha \) for the radiation effect was 0.16 Gy−1, which is similar to previous in vivo estimates of 0.16–0.1841. These estimated values for \(\alpha \) are below the common range of in vitro estimates (0.2–0.5 Gy−1)16,42,43. Figure 5c shows the distribution of SF2Gy (the cell survival fraction after 2 Gy), derived from the distribution of the radiosensitivity parameter \(\alpha \).
From clinical observations we know that the growth rate accelerates with the shrinkage of the tumor volume during radiation therapy, a phenomenon termed repopulation44,45. To account for this effect, most of the current models use the kickoff-repopulation approach, which introduces an exponential growth with a VDT of 3 days after a kick-off period of 28 days, counting from the start of therapy16. However, this phenomenon is in principle intrinsically included in the Gompertz model, as tumors shrinking from radiation cell kill grow faster according to Gompertz kinetics. We compared the two approaches in this study.
Figure 6 shows the results for the two methods. We found that both repopulation methods can describe the survival curve after radiation therapy well (see Fig. 6a). This confirms that the parameter-less Gompertz-repopulation can naturally account for repopulation (i.e. gradually decreased VDT as the tumor shrinks, see Fig. 6c) during radiation therapy. The estimated median values for the radiation effect \(\alpha \) were 0.16 for Gompertz repopulation and 0.17 for kickoff-repopulation respectively. These values agree well with clinical findings46,47, and underscore that Gompertz growth could naturally account for repopulation without additional parameters.
(a) Survival curves predicted with Gompertz and exponential repopulation, (b) illustration of the growth pattern and radiation effect of the model, and (c) illustration that the Gompertz model can naturally account for the repopulation during radiation therapy. Note that the VDTs here were calculated as the time in which the tumor reaches twice its current size.
Currently the exponential growth model is still the standard in radiotherapy effect modeling, evidenced for instance by the inclusion of an exponential growth based time factor in the BED model15,16. Sachs et al. described a similar model, and discussed that simple approaches such as the Gompertz growth and linear-quadratic models can still capture the complicated interactions present during radiation therapy48. The model described by Sachs et al. is similar with the present model. In a recent study, Jeong (2017) developed a mechanistic mathematical model for radiation therapy, which quantitatively predicted the differences in response seen in early stage lung cancer across the range of clinical dose and fractionation schemes49. They used a compartmental ‘tumourlet’-response model to account for the oxygen and tumor repopulation effect, which is more mechanistic and requires more parameters compared to the present model. Models such as this could be implemented with our chemotherapy and growth models to enable prediction of fractionation effects.
Chemotherapy
With consistent parameters for tumor growth and radiation response, we then proceeded to derive the chemo-effect parameter \({{\rm{\alpha }}}_{\mathrm{chemo}}\) using the sequential chemo-radiation (sCRT) arm of RTOG 9410. The sCRT included cisplatin at 100 mg/m3 on days 1 and 29 and vinblastine at 5 mg/m3 per week for 5 weeks with 60 Gy radiation therapy beginning on day 50, as shown in Figure. 7. To explain the observed improvement in survival of sCRT compared to radiation only, the resulting median chemotherapy effect parameter \({{\rm{\beta }}}_{{\rm{c}}}\) was determined to be 0.028/(mg/ml).
(a) Predicted Kaplan–Meier survival curve with the sequential Chemo-radiation and radiation only therapy for stage III patient comparing to the clinical trial RTOG 9410, and (b) an example patient with the growth and treatment response curve for sequential Chemo-radiation and radiation therapy, (c) the predicted overall survival with our model comparing to (d) the data from six trials summarized in Aupérin et al.54.
As our methodology has not been used before to derive quantitative cell kill parameters for chemotherapy agents, we verified the predictive power using published data. From published data of large cohorts, we know that with doublet chemotherapy in stage IV lung cancer patients, the median survival is about 6–10 months and 1-year OS is about 25–35%50,51,52. To test if our model would predict similar values, we generated cohorts of stage IV patients which received 4 cycles, which is the median number of cycles given51,52, of our chemotherapy simulation and predicted the improvement in survival. Our model predicted a median survival of 10.2 months, which is reasonably close to the observed survival.
The Gompertz model also explains the moderate repopulation (i.e. accelerated tumor growth) observed after the chemotherapy segment of sequential chemo-radiation, which was observed in clinical trials40,53. Our model predicted a median VDT of 12 days and a mean VDT of 41 days for a time interval of 100 days between two examinations after induction chemotherapy. Sharouni et al. measured the median and mean VDT of 29.4 and 45.8 days after induction chemotherapy, which they described as significantly lower than the normal VDT40. Also they found that small tumors have the shortest VDTs. This is a confirmation of the results given by our model.
Combined modality treatment
We simulated concurrent chemo-radiation (cCRT) by combining the parameters obtained above. We took the same chemo- and radiotherapy cell kill used in Fig. 7a for sCRT and shifted the RT section to start together with the first cycle of chemotherapy, mimicking the concurrent trial arm of RTOG-9410 (see Fig. 7b). The main goal was to determine if an additional radiosensitization parameter would be necessary to fit the observed survival curves, or if the increased cell kill in a shorter treatment time is sufficient to explain the superior clinical results of cCRT.
Figure 7c demonstrates that the combination of chemotherapy and radiotherapy in a concurrent fashion predicts OS improvements of 6.6% and 6.2% for 3 and 5-years for stage III patients, which compares well to the 5.3% and 4.5% observed in the cCRT arm (see Fig. 7d) of RTOG-94108.
For the overall survival prediction of concurrent and sequential CRT, our model performs well and predicts results observed in clinical trials (see Fig. 7). There is no need to add a radiosensitization parameter for the interaction between chemo- and radio-therapy, indicating that the reduction in treatment time and the resulting suppression of repopulation is the deciding factor for the superiority of cCRT compared to sCRT. Figure 8 demonstrates tumor cell load of a patient who will benefit from the concurrent compared to the sequential chemo-radiation regimen. With sCRT the tumor grows back after chemo- and radiotherapy, as the cell kill was not dense enough and allowed the tumor to re-grow. Since the tumor cell number was reduced to below 1 (high probability of tumor control) with Concurrent chemo-radiation therapy, the tumor is controlled according to our model.
The growth and treatment response curve for a specific patient with sequential and concurrent Chemo-radiation therapy, Parameters used for tumor simulation: Growth parameter \(\rho \): 0.008, \({V}_{{limit}}\): 30 cm in diameter, \(\alpha \) = 0.3 Gy−1, \(\alpha /\beta \): 10 Gy, delay time = 14 days, β c: 0.03 (mg/m3)−1.
Growth rate based Stratification of Concurrent and sequential CRT: an example application
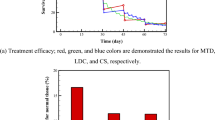
To demonstrate the application of the developed model in clinical practice, we investigated which population benefits most from concurrent chemo-radiation. For most patients who have unresectable disease, combined chemo and radiation therapy is considered as the first treatment of choice. However, it is unknown how to define criteria for selecting a subgroup of patients who would benefit from concurrent CRT compared to treating sequentially, which is predicted to cause less toxicity. We separated the patient population into subgroups according to the growth rate distribution (considering the median of the distribution and the growth parameter \({\rm{\rho }}\)). All parameters for treatment response are given in Table 3. The schedule of combined chemotherapy and radiation treatment was set as identical to RTOG 9410. The model predicted that for patients with a faster tumor growth (above the median), the benefit can be as high as 11.1% for stage III patients, but only 4.7% for those patients with a slower growth rate (below the median) for 3 year overall survival. The benefit can be as high as 14.4% for patients with a growth rate in the top 25th percentile. This could be important in the design of clinical trials evaluating treatment options for older and frail patients that cannot tolerate a concurrent regimen, verifying these predictions in patients might necessitate advanced imaging modalities (e.g. PET-MTV (PET derived metabolic tumor volume)) to determine the growth rate (e.g. utilizing the correlation between PET-MTV and tumor growth rate55).
Limitations and future directions
There are numerous models that describe tumor growth and treatment response. However, none of the standard models allows simulation and prediction of multi-modality regimen for locally-advanced NSCLC. We developed a mathematical model for the combined treatment of chemotherapy and radiation based on overall survival data from several clinical trials, with the purpose of improving and optimizing treatment strategies for future clinical trials. Notably, parameters in the model were derived from the clinical relevant outcome data. The tumor growth and patient death model can also be used as foundation for the simulation of other therapeutic interventions, such as targeted agents.
As every model has its inherent limitations, we would like to highlight the caveats of our approach.
-
(1)
The \({\rm{\alpha }}/{\rm{\beta }}\) -ratio used was 10 and the cell cycle effect and oxygen enhancement ratio for radiosensitivity was not considered. This might be necessary if one wants to investigate different fractionation regimen16, which was not considered in this study.
-
(2)
The Gompertz model in use was shown to fit the data and clinical observations better than the exponential model. However, there are other models with the same growth pattern, e.g. the logistic or the Bertalanffy model, which could be expected to have a similar modeling performance11.
-
(3)
Log-cell kill was used to model chemotherapy in the study, which is commonly used and a widely accepted approach. However, there is another main model in use for this purpose, the Norton-Simons model, which differs significantly56. We do not think this has an impact on the results of our study, as we investigate only one chemotherapy regimen and only study changes in timing between chemo- and radiotherapy. If the aim would be to investigate different chemotherapy regimen and their interaction with radiation, then the model in use could have an impact on the results.
-
(4)
There is no modeling of normal tissue toxicity, although natural deaths were considered. Toxicity from treatment can result in extra deaths, and RTOG 061757, which showed lower survival for the higher dose arm, has raised questions about the limits of dose escalation with concurrent CRT in the general NSCLC population. Whatever the exact reasons for the lower survival in this trial may be, toxicity might play a prominent role. Therefore models such as the one presented above should be used with care for regimen that are supposedly pushing the boundaries of normal tissue toxicity. However, a toxicity component could be added to the current model, which would necessitate the analysis of dose distributions for all patients, which were unavailable for these trials.
Conclusion
Combined CRT is a mainstay of treatment for locally-advanced NSCLC patients. We developed a mathematical model for the combined treatment of chemotherapy and radiation, with the purpose of improving and optimizing treatment strategies for future clinical trials. The established model can predict survival curves for populations of NSCLC patients with or without treatment. The parameter-less Gompertz repopulation can naturally account for repopulation during radiation therapy and after induction chemotherapy. The model provides a tool for the optimization of combined chemo-radiation scheduling and sequencing.
Change history
17 August 2018
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Chen, Z., Fillmore, C. M., Hammerman, P. S., Kim, C. F. & Wong, K.-K. Non-small-cell lung cancers: a heterogeneous set of diseases. Nature Reviews Cancer 15, 247–247 (2015).
Dincklage, von,J. J., Ball, D. & Silvestri, G. A. A review of clinical practice guidelines for lung cancer. Journal of Thoracic Disease 5, S607–S622 (2013).
Johnson, D. H. Locally Advanced, Unresectable Non-Small Cell Lung Cancer: New Treatment Strategies. CHEST 117, 123S–126S (2000).
Bentzen, S. M., Saunders, M. I. & Dische, S. From CHART to CHARTWEL in Non-small Cell Lung Cancer: Clinical Radiobiological Modelling of the Expected Change in Outcome. Clinical Oncology 14, 372–381 (2002).
Kong, F.-M., Haken, R. T., Eisbruch, A. & Lawrence, T. S. Non-Small Cell Lung Cancer Therapy-Related Pulmonary Toxicity: An Update on Radiation Pneumonitis and Fibrosis. Seminars in Oncology 32, 42–54 (2005).
Rosenzweig, K. E. et al. Results of a phase I dose‐escalation study using three‐dimensional conformal radiotherapy in the treatment of inoperable nonsmall cell lung carcinoma. Cancer 103, 2118–2127 (2005).
Willner, J., Baier, K., Caragiani, E., Tschammler, A. & Flentje, M. Dose, volume, and tumor control prediction in primary radiotherapy of non-small-cell lung cancer. International Journal of Radiation Oncology*Biology*Physics 52, 382–389 (2002).
Curran, W. J. et al. Sequential vs Concurrent Chemoradiation for Stage III Non-Small Cell Lung Cancer: Randomized Phase III Trial RTOG 9410. JNCI Journal of the National Cancer Institute 103, 1452–1460 (2011).
Grassberger, C. & Paganetti, H. Methodologies in the modeling of combined chemo-radiation treatments. Physics in Medicine & Biology 344–369, https://doi.org/10.1088/0031-9155/61/21/R344 (2016).
Benzekry, S. et al. Classical Mathematical Models for Description and Prediction of Experimental Tumor Growth. PLoS Comput Biol 10, e1003800–20 (2014).
Gerlee, P. The Model Muddle: In Search of Tumor Growth Laws. Cancer Research 73, 2407–2411 (2013).
Usuda, K., Saito, Y., Sagawa, M. & Sato, M. Tumor doubling time and prognostic assessment of patients with primary lung cancer. Cancer 74, 2239–2244 (1994).
Yankelevitz, D. F. et al. Overdiagnosis in chest radiographic screening for lung carcinoma. Cancer 97, 1271–1275 (2003).
Brenner, D. J. The Linear-Quadratic Model Is an Appropriate Methodology for Determining Isoeffective Doses at Large Doses Per Fraction. Seminars in Radiation Oncology 18, 234–239 (2008).
Fowler, J. F. 21 years of Biologically Effective Dose. BJR 83, 554–568 (2010).
Mehta, M. et al. A new approach to dose escalation in non–small-cell lung cancer. Radiation Oncology Biology 49, 23–33 (2001).
Lee, J. Y., Kim, M.-S., Kim, E. H., Chung, N. & Jeong, Y. K. Retrospective growth kinetics and radiosensitivity analysis of various human xenograft models. Lab Anim Res 32, 187–7 (2016).
Shibamoto, Y. et al. Proliferative activity and micronucleus frequency after radiation of lung cancer cells as assessed by the cytokinesis-block method and their relationship to clinical outcome. Clinical Cancer Research 4, 677–682 (1998).
Ishibashi, N. et al. Correlation between the Ki-67 proliferation index and response to radiation therapy in small cell lung cancer. Radiation Oncology 2017 12(12:1), 16 (2017).
Skipper, H. E. Perspectives in Cancer Chemotherapy: Therapeutic Design. Cancer Research 24, 1295–1302 (1964).
Powathil, G., Kohandel, M., Sivaloganathan, S., Oza, A. & Milosevic, M. Mathematical modeling of brain tumors: effects of radiotherapy and chemotherapy. Physics in Medicine & Biology 52, 3291–3306 (2007).
Detterbeck, F. C. & Gibson, C. J. Turning Gray: The Natural History of Lung Cancer Over Time. Journal of Thoracic Oncology 3, 781–792 (2008).
Wisnivesky, J. P., Yankelevitz, D. & Henschke, C. I. The Effect of Tumor Size on Curability of Stage I Non-small Cell Lung Cancers. CHEST 126, 761–765 (2004).
Edge, S. B. & Compton, C. C. The American Joint Committee on Cancer: the7th Edition of the AJCC Cancer Staging Manual and the Future of TNM. Ann Surg Oncol 17, 1471–1474 (2010).
Winer-Muram, H. T. et al. Volumetric Growth Rate of Stage I Lung Cancer prior to Treatment: Serial CT Scanning. Radiology 223, 798–805 (2002).
Switzer, P., Gerstl, B. & Greenspoon, J. Karyometry in the Estimation of Nuclear Population in Pulmonary Carcinomas. JNCI Journal of the National Cancer Institute 52, 1699–1704 (1974).
Sause, W. et al. Final Results of Phase III Trial in Regionally Advanced Unresectable Non-Small Cell Lung Cancer: Radiation Therapy Oncology Group, Eastern Cooperative Oncology Group, and Southwest Oncology Group. CHEST 117, 358–364 (2000).
Saeb-Parsy, K. Instant Pharmacology. (John Wiley & Sons, 1999).
Kumar, A. Vincristine and Vinblastine: A Review. IJMPS 6, 23–30 (2016).
Martini, N. et al. Survival after resection of stage II non-small cell lung cancer. The Annals of Thoracic Surgery 54, 460–466 (1992).
Geddes, D. M. The natural history of lung cancer: A review based on rates of tumour growth. British Journal of Diseases of the Chest 73, 1–17 (1979).
Mehrara, E., Forssell-Aronsson, E., Ahlman, H. & Bernhardt, P. Specific Growth Rate versus Doubling Time for Quantitative Characterization of Tumor Growth Rate. Cancer Research 67, 3970–3975 (2007).
Hasegawa, M. et al. Growth rate of small lung cancers detected on mass CTscreening. BJR 73, 1252–1259 (2014).
Steel, G. G. The growth Kinetics of Tumours. (Oxford University Press, 1977).
Fujimura, S., Suda, S., Yamauchi, A., Sato, H. & Sohara, Y. Tumor doubling time and PPD skin test reactivity in resectable lung cancer. J Jpn Lung Cancer Soc 19, 135–142 (1978).
Filderman, A. E., Shaw, C. & Matthay, R. A. Lung Cancer: Part I: Etiology, Pathology, Natural History, Manifestations, and Diagnostic Techniques. Investigative Radiology 21, 80 (1986).
Fox, J. et al. Quantification of Tumor Volume Changes During Radiotherapy for Non–Small-CellLung Cancer. International Journal of Radiation Oncology*Biology*Physics 74, 341–348 (2009).
Basaki, K. et al. Prognostic factors for survival in stage III non–small-cell lung cancer treated with definitive radiation therapy: Impact of tumor volume. International Journal of Radiation Oncology*Biology*Physics 64, 449–454 (2006).
Kozak, M. M. et al. Tumor Volume as a Potential Imaging-Based Risk-Stratification Factor in Trimodality Therapy for Locally Advanced Non-small Cell Lung Cancer. Journal of Thoracic Oncology 6, 920–926 (2011).
Sharouni, E, S. Y., Kal, H. B. & Battermann, J. J. Accelerated regrowth of non-small-cell lung tumours after induction chemotherapy. 89, 2184–2189 (2003).
Liu, F. et al. Tumor control probability modeling for stereotactic body radiation therapy of early-stage lung cancer using multiple bio-physical models. Radiotherapy and Oncology 122, 286–294 (2017).
Franken, N., Hovingh, S. & Rodermond, H. Radiosensitization with chemotherapeutic agents and hyperthermia: Effects on linear-quadratic parameters of radiation cell survival curves. J Cancer Sci Ther (2011).
Ohri, N., Dicker, A. P. & Lawrence, Y. R. Can Drugs Enhance Hypofractionated Radiotherapy? A Novel Method of Modeling Radiosensitization Using In Vitro Data. International Journal of Radiation Oncology*Biology*Physics 83, 385–393 (2012).
Shafiq, J., Hanna, T. P., Vinod, S. K., Delaney, G. P. & Barton, M. B. A Population-based Model of Local Control and Survival Benefit of Radiotherapy for Lung Cancer. Clinical Oncology 28, 627–638 (2016).
Kim, J. J. & Tannock, I. F. Repopulation of cancer cells during therapy: an important cause of treatment failure. Nature Reviews Cancer 5, 516–525 (2005).
Zhong, H. & Chetty, I. A note on modeling of tumor regression for estimation of radiobiological parameters. Med. Phys. 41, 081702 (2014).
Hedman, M., Björk-Eriksson, T., Brodin, O. & Toma-Dasu, I. Predictive value of modelled tumour control probability based on individual measurements of in vitro radiosensitivity and potential doubling time. BJR 86, 20130015 (2014).
Sachs, R. K., Hlatky, L. R. & Hahnfeldt, P. Simple ODE models of tumor growth and anti-angiogenic or radiation treatment. Mathematical and Computer Modelling 33, 1297–1305 (2001).
Jeong, J. et al. Modeling the Cellular Response of Lung Cancer to Radiation Therapy for a Broad Range of Fractionation Schedules. Clinical Cancer Research clincanres.3277, https://doi.org/10.1158/1078-0432.CCR-16-3277 (2017).
Booton, R. et al. A phase III trial of docetaxel/carboplatin versus mitomycin C/ifosfamide/cisplatin (MIC) or mitomycin C/vinblastine/cisplatin (MVP) in patients with advanced non-small-cell lung cancer: a randomised multicentre trial of the British Thoracic Oncology Group (BTOG1). Ann Oncol 17, 1111–1119 (2006).
Johnson, D. H. Evolution of Cisplatin-Based Chemotherapy in Non-Small Cell Lung Cancer: A Historical Perspective and The Eastern Cooperative Oncology Group Experience. CHEST 117, 133S–137S (2000).
Wozniak, A. J. et al. Randomized trial comparing cisplatin with cisplatin plus vinorelbine in the treatment of advanced non-small-cell lung cancer: a Southwest Oncology Group study. JCO 16, 2459–2465 (1998).
Chen, C. P., Weinberg, V. K., Jahan, T. M., Jablons, D. M. & Yom, S. S. Implications of Delayed Initiation of Radiotherapy: Accelerated Repopulation after Induction Chemotherapy for Stage III Non-small Cell Lung Cancer. Journal of Thoracic Oncology 6, 1857–1864 (2011).
Aupérin, A. et al. Meta-Analysis of Concomitant Versus Sequential Radiochemotherapy in Locally Advanced Non–Small-CellLung Cancer. JCO 28, 2181–2190 (2010).
Wang, J., Mahasittiwat, P., Wong, K. K., Quint, L. E. & Kong, F.-M. S. Natural growth and disease progression of non-small cell lung cancer evaluated with 18F-fluorodeoxyglucose PET/CT. Lung Cancer 78, 51–56 (2012).
Simon, R. & Norton, L. The Norton–Simon hypothesis: designing more effective and less toxic chemotherapeutic regimens. Nature Clinical Practice Oncology 3, 406–407 (2006).
Bradley, J. D. et al. Standard-dose versus high-dose conformal radiotherapy with concurrent and consolidation carboplatin plus paclitaxel with or without cetuximab for patients with stage IIIA or IIIB non-small-cell lung cancer (RTOG 0617): a randomised, two-by-two factorial phase 3 study. The Lancet Oncology 16, 187–199 (2015).
Author information
Authors and Affiliations
Contributions
C.G. (Changran Geng) and C.G. (Clemens Grassberger) designed and implemented the model, and analysed results. C.G. (Changran Geng) and C.G. (Clemens Grassberger) wrote the main manuscript text based on the discussion with H.P. C.G. (Clemens Grassberger) and H.P. co-supervised the project; All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Geng, C., Paganetti, H. & Grassberger, C. Prediction of Treatment Response for Combined Chemo- and Radiation Therapy for Non-Small Cell Lung Cancer Patients Using a Bio-Mathematical Model. Sci Rep 7, 13542 (2017). https://doi.org/10.1038/s41598-017-13646-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13646-z
This article is cited by
-
Predicting efficacy assessment of combined treatment of radiotherapy and nivolumab for NSCLC patients through virtual clinical trials using QSP modeling
Journal of Pharmacokinetics and Pharmacodynamics (2024)
-
A survey of deep causal models and their industrial applications
Artificial Intelligence Review (2024)
-
Interaction tumor-immune model with time-delay and immuno-chemotherapy protocol
Rendiconti del Circolo Matematico di Palermo Series 2 (2023)
-
Recurrent Continuous Adversarial Balancing for Estimating Individual Treatment Effect Over Time
International Journal of Computational Intelligence Systems (2023)
-
Iodine-125 brachytherapy suppresses tumor growth and alters bone metabolism in a H1299 xenograft mouse model
Medical Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.