Abstract
The order Chlamydiales are biphasic intracellular bacterial pathogens infecting humans and domesticated animals. Wildlife infections have also been reported, with the most studied example being Chlamydia pecorum infections in the koala, an iconic Australian marsupial. In koalas, molecular evidence suggests that spill-over from C. pecorum infected livestock imported into Australia may have had a historical or contemporary role. Despite preliminary evidence that other native Australian marsupials also carry C. pecorum, their potential as reservoirs of this pathogen and other Chlamydia-related bacteria (CRBs) has been understudied. Mucosal epithelial samples collected from over 200 native Australian marsupials of different species and geographic regions across Australia were PCR screened for Chlamydiales. Previously described and genetically distinct C. pecorum genotypes and a range of 16S rRNA genotypes sharing similarity to different CRBs in the broader Chlamydiales order were present. One 16S rRNA Chlamydiales genotype recently described in Australian ticks that parasitise native Australian marsupials was also identified. This study provides further evidence that chlamydial infections are widespread in native fauna and that detailed investigations are required to understand the influence these infections have on host species conservation, but also whether infection spill-over plays a role in their epidemiology.
Similar content being viewed by others
Introduction
Evidence that infections by obligate intracellular bacterial pathogens in the order Chlamydiales are generally prevalent in wildlife is increasing1, but their impact on the overall health of populations remains unclear1,2. The koala (Phascolarctos cinereus), an iconic native Australian marsupial, is the most characterised example of the detrimental effects that chlamydial infections can have on the health of individuals and populations2. Although asymptomatic infections are most common, C. pecorum infections in koalas can result in severe ocular, urinary tract and reproductive disease which, if left untreated, may progress to blindness, incontinence and infertility, respectively. The prevalence of koala C. pecorum infections varies from 10–90% in mainland populations, significantly contributing to the rapid decline of this threatened species in certain states of Australia2,3. Koalas also carry unique Chlamydia-related bacteria (CRBs), although the role these organisms play in disease is yet to be defined4.
The origin of C. pecorum infection in koalas is currently unclear. It is well known that domesticated livestock, such as sheep and cattle, are susceptible to C. pecorum infections. In Australia, genetically similar strains of livestock C. pecorum have been identified in koalas, in addition to koala-specific strains5,6,7,8,9. This body of molecular evidence has led to the hypothesis that at least some C. pecorum infections in koalas may be the result of historical or contemporary pathogen spill-over from livestock sources, consistent with the significant deleterious impact that these infections have on some koala populations2. Evidence to support this hypothesis is incomplete, however1.
Research into chlamydial prevalence across Australia’s native marsupial species and populations has been severely limited by geography, sample size and the logistical difficulties in sampling. Several marsupial species have been reported to carry Chlamydiaceae like C. pecorum, as well as both novel and previously described CRBs. C. pecorum has been reported in the greater glider, mountain brushtail possum and western barred bandicoot10,11; with C. pneumoniae also detected in western barred bandicoots12. Although some of these marsupials exhibited signs of chlamydiosis, such as ocular disease, there is little evidence that C. pecorum has the same pathogenic effect on these marsupials as it does on koalas. Reports such as these, suggest there may be complex and dynamic origins of Chlamydiales in marsupials, further complicating our understanding of this pathogen’s evolution. It is possible that C. pecorum is a naturally occurring organism within Australia’s marsupial species and the importation of livestock has selectively exacerbated its impact on koala populations1. More recently, preliminary investigations of tick species feeding on koalas and other Australian marsupials highlighted a diversity of novel CRBs, raising questions over the presence and impact of these novel bacteria on the marsupial host parasitised as well13.
To gain further insight into the prevalence and impact of chlamydial infections on Australian marsupial fauna, we conducted a broad-range Chlamydiales order-specific molecular survey. Chlamydial diversity was sampled from non-koala marsupials across a range of geographic regions in Australia alongside a small subset of wild koalas for comparison.
Results
Chlamydiales PCR positivity in Australian marsupials
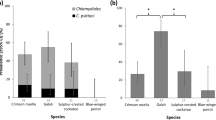
Pan-Chlamydiales 16S rRNA gene PCR screening of non-koala marsupial swabs from all regions sampled revealed 196/401 positive samples for Chlamydiales DNA, translating into a Chlamydiales positive rate of 48% for individual animals at one or more anatomical sites (111/231 animals; Table 1). A concurrent screening of 37 koalas tested (57 swab samples) from the East Coast revealed 100% PCR positivity for Chlamydiales DNA.
To determine the identity of the Chlamydiales detected in the marsupials screened in this study, a pan-Chlamydiales PCR amplifying an 800 bp fragment of the 16S rRNA gene was targeted. PCR products for 96 non-koala and 43 koala samples were obtained and directly sequenced. Results revealed that 18.7% (18/96) and 58.1% (25/43) of the PCR products for non-koala marsupials and koalas, respectively, returned single sequence results that allowed for direct sequence analysis. The remaining samples resulted in mixed sequences, despite repeated PCR amplification and sequencing efforts, suggesting the majority of samples contained DNA from multiple Chlamydiales organisms (data not shown). Nucleotide BLAST (nBLAST) analysis of the single sequence results revealed the presence of one previously described and six novel genotypes belonging to the order Chlamydiales 14,15 (Table 2, Fig. 1).
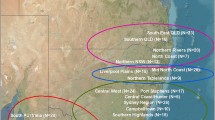
Distribution of Chlamydiales genotypes amongst Australian marsupials. Shaded areas represent sampling area for each region. Greyscale pie charts represent number of PCR positive and negative individuals and coloured pie charts represent the abundance and diversity of genotypes detected within each region (see key). Map was modified from (https://commons.wikimedia.org/wiki/File:Australia_states_blank.png) under the Creative Commons Attribution-Share Alike 3.0 Unported license (https://creativecommons.org/licenses/by-sa/3.0/), using Microsoft Windows Version 6.1 Paint and PowerPoint 2013.
Chlamydiales prevalence and genotype diversity varied by geography and marsupial host species
Common brushtail possums harboured the most diverse range of Chlamydiales, carrying four of the seven genotypes, with the remaining marsupial species carrying two or less genotypes. Chlamydial prevalence and diversity within non-koala marsupials was also noticeably larger at conjunctival sites (Table 1). PCR positivity of non-koala marsupials varied geographically, with Tasmania having the highest prevalence of 100%, followed by the East Coast sample collection (87%; 48/55) and the Northern Territory collection (26%; 41/154). In the minority of samples where we were able to resolve sequences, the distribution of 16S rRNA sequences across these different geographic locations varied. Only one genotype (Ca. Rhabdochlamydia porcellionis genotype 1) was detected in all three geographical locations (Fig. 1). The highest genotypic diversity was found in the East Coast collection where four of the seven different genotypes could be resolved. Three genotypes could each be resolved in the Tasmanian and Northern Territory cohorts (Tables 1 and 2).
Detection of C. pecorum 16S rRNA sequences in koalas and other marsupials
BLAST searching revealed the presence of two distinct C. pecorum 16S rRNA sequences in koalas and other marsupials in the East Coast sample set. The first of these partial sequences, detected in three common brushtail possums, one squirrel glider, one spotted tail quoll and 20 koalas was found to share 100% nucleotide identity to previously reported C. pecorum 16S rRNA sequences from cattle (P787–CP004035.1; E58–NR_102975.1) and koalas (MC/MarsBar - HQ457465.1; Table 2). For the purpose of this investigation, this 16S rRNA sequence will be referred to as Chlamydia pecorum 16S rRNA genotype P787. In addition to this previously described 16S rRNA genotype, a second novel 16S rRNA C. pecorum sequence was also detected in three common brushtail possums, two from Tasmania and one from the East Coast (Table 2). This sequence, designated Chlamydia pecorum genotype 1, had 99% identity to the top BLAST hit C. pecorum strain P787 (CP004035.1) described above, yet differed from this strain by three SNPs. The phylogenetic positioning of these two genetically distinct sequences was confirmed following Bayesian phylogenetic reconstruction (Fig. 2).
Phylogenetic relationships of Chlamydiaceae genotypes identified in Australian marsupials. Bayesian tree incorporating representative 16S rRNA sequences of each species of the genus Chlamydia from GenBank, as well as the three partial 16S rRNA genotypes identified in this study. Tree was built using 18 sequences of 588 bp under the HKY85 evolutionary model, posterior probability exceeding 0.75 is shown at internal nodes.
Confirmation of Chlamydia pecorum in non-koala marsupials and molecular typing using a C. pecorum-specific MLST scheme
In order to confirm the detection of C. pecorum in non-koala marsupials and to gain insight into the genetic relationships of these strains to those from other hosts, amplification of seven highly conserved C. pecorum MLST housekeeping genes was performed on the eight C. pecorum positive non-koala marsupial samples, as previously described8. Unfortunately, we were only successful in amplifying one of the seven C. pecorum MLST genes (gidA) from six of the non-koala marsupial samples. Analysis of these gidA gene sequence fragments and comparisons to previously described gidA sequences deposited in the C. pecorum PubMLST database16 revealed a 100% sequence match for the spotted tail quoll sample to C. pecorum gidA allele 31, an allele previously described in koalas. The squirrel glider sample was an exact match to C. pecorum gidA allele 22, which has previously been described in both sheep and cattle. However, the four common brushtail possum samples were all identified to have novel C. pecorum gidA allele 12 sequences, each encompassing one to two unique SNPs (Fig. 3), when compared to gidA alleles derived from sheep and cattle.
Phylogenetic relationships of C. pecorum gidA alleles identified in Australian marsupials. Bayesian tree incorporating six representative partial gidA alleles from PubMLST, as well as the six partial gidA alleles identified in this study. Marsupials identified with 16S rRNA as Chlamydia pecorum 16S rRNA genotype P787 are coloured green and those identified as Chlamydia pecorum genotype 1 are coloured blue. Tree was built using seven sequences of 474 bp under the HKY85 evolutionary model, posterior probability exceeding 0.75 is shown at internal nodes.
Detection of arthropod-associated Chlamydiales 16S rRNA sequences in Australian marsupials
The most geographically distributed CRB genotype (Ca. Rhabdochlamydia porcellionis genotype 1) detected in this study was 100% identical to a recently described novel Ca. Rhabdochlamydia porcellionis genotype (KX774317) detected in marsupial-feeding tick species removed from koalas13. This genotype is previously described as having 98% BLAST similarity to Ca. Rhabdochlamydia porcellionis (AY223862.1) and branches as a distinct lineage (Fig. 4). Ca. Rhabdochlamydia porcellionis genotype 1 was detected in two common brushtail possums, one short-eared possum and one northern brown bandicoot across the East Coast, Northern Territory and Tasmania (Fig. 1) and, additionally, in five koalas from the East Coast sample collection.
Beyond this previously described arthropod-associated Chlamydiales sequence, a novel Ca. Rhabdochlamydia crassificans strain was also detected in two northern quolls exclusive to the Northern Territory (Fig. 1). Ca. Rhabdochlamydia crassificans genotype 1 exhibited 99% BLAST identity to the closest match Ca. Rhabdochlamydia crassificans (AY928092.1), differing by seven SNPs (Fig. 4, Table 2). A sequence potentially representing a novel genus diverging from the previously described Ca. Rhabdochlamydiaceae (Fig. 4) was also detected in two common brushtail possums, exclusively to the Tasmanian region. This partial 16S rRNA genotype, Ca. Rhabdochlamydia porcellionis genotype 2, demonstrated 93% BLAST identity to Ca. Rhabdochlamydia porcellionis (HF933203.1) (Fig. 1, Table 2) as well as previously described novel Ca. Rhabdochlamydia genotypes identified in ticks13.
Phylogenetic relationships of Ca. Rhabdochlamydiaceae genotypes identified in Australian marsupials. Bayesian tree incorporating representative 16S rRNA sequences of each genus within the Ca. Rhabdochlamydiaceae from GenBank, as well as the three partial 16S rDNA genotypes identified in this study. The tree was built using seven sequences of 368 bp under the HKY85 evolutionary model, posterior probability exceeding 0.75 is shown at internal nodes.
Detection of other CRB 16S rRNA sequences in Australian marsupials
A partial 16S rRNA gene sequence potentially representing a novel genus within the family Chlamydiaceae was also detected in a northern quoll from the Northern Territory (Fig. 1). Fine-detailed phylogenetic analysis of this 16S rRNA sequence against other sequences belonging to species in the family Chlamydiaceae suggests that this novel sequence sits between the Chlamydia and Ca. Amphibiichlamydia genera (Fig. 2, Table 2). This genotype, Chlamydia-like genotype 1, exhibited 92% identity to Chlamydia pneumoniae (LN847058.1) 16S rRNA from a western barred bandicoot, as well as strains detected in koalas (CP001713.1), African clawed frogs (AF139200.1) and other western barred bandicoots (DQ444323.1).
To conclude the descriptions, we also detected an unusual partial 16S rRNA sequence in a sample from a spotted tail quoll from the East Coast, sharing 99% identity and two single nucleotide differences (Fig. 1) to a previously deposited sequence from the novel chlamydial pathogen of fish, Ca. Similichlamydia latridicola (KC686679.1).
Discussion
Koalas, an iconic Australian marsupial, and globally distributed domesticated animals such as sheep and cattle, are well-known hosts of C. pecorum 2,17. In the current study, a pan-Chlamydiales order specific 16S rRNA gene PCR screening strategy provided molecular evidence that a range of other Australian marsupials also carry C. pecorum. A distinct range of novel bacteria from the order Chlamydiales were also found, some of which have been previously reported in ticks18,19, including those found parasitising Australian marsupials13.
In this study, C. pecorum 16S rRNA genotype P787 and C. pecorum genotype 1 accounted for 44.4% of the partial 16S rRNA Chlamydiales genotypes obtained, suggesting that C. pecorum is more abundant among non-koala marsupials than previously thought. In this study the greatest diversity and abundance of C. pecorum was found in common brushtail possums, suggesting that some marsupial host species apart from koalas, may be more susceptible to C. pecorum infection than others. Possums have previously been identified as a carrier of C. pecorum 10. While we need to treat the geographic distribution data with caution since we were only able to resolve sequences for the minority of PCR-positive samples we detected, it is notable that the majority of C. pecorum positive samples were detected in the East Coast. This region strongly overlaps with the host range of koalas that are endemically infected with C. pecorum and, indeed, the majority of animal hospitals that were used for sampling in this study commonly receive high numbers of koalas into care with chlamydiosis20,21. Notably, animals sampled in the Northern Territory where koalas are not found, were free of C. pecorum. Inconsistent with this trend was the more surprising detection of C. pecorum positive samples from Tasmanian brushtail possums. Wild koalas do not inhabit Tasmania and, to the author’s knowledge, this report represents the first reported detection of chlamydial pathogens in Tasmanian fauna. Tasmania was last connected to mainland Australia during the Pleistocene (~25–8, 000 years ago)22 suggesting C. pecorum (i) has either been introduced to Tasmania by domestic species such as sheep and cattle, (ii) naturally occurs in possums from this region, and/or (iii) infections in Australian marsupials predated the separation of Tasmania from the mainland.
While only preliminary in nature, a combination of partial 16S rRNA Chlamydiales and C. pecorum gidA gene sequencing suggests that the C. pecorum strains detected may be genetically diverse and may differ from those previously detected in koalas and Australian livestock8,9. This was most apparent in the gidA sequences, where several previously described and novel gidA haplotypes were detected, suggesting the possibility of marsupial host specific strains and no favourable association with koalas or livestock based on the C. pecorum gidA data available to date. Unfortunately, we were not able to gain any further insight into the genetic diversity of these strains since we failed to amplify any of the remaining house-keeping genes utilised in previously described chlamydial MLST schemes (Jelocnik et al., 2013). While it is not entirely clear why this was the case, we expect it was due to the presence of only very low levels of C. pecorum DNA in these samples, combined with varying sensitivities of each of the individual PCR assays included in this scheme. Further confirmation of strain diversity and insight into the relationships of these strains is not possible without detailed genomics studies to compare these non-koala Australian marsupial C. pecorum strains to those from koalas, sheep and cattle.
In the absence of such data, a partial answer regarding the origin and impact of these infections in Australian marsupials may lie in the disease presentation of Chlamydia positive animals in this study, as well as the ecological niche that these animals occupy. All 231 individual non-koala marsupials screened in this study were void of any classical signs of chlamydiosis despite several being C. pecorum positive at either ocular or urogenital sites. However, the comparison koalas infected with the same strain (based on partial 16S rRNA gene sequence), ranged from being asymptomatic to presenting with severe chlamydiosis (data not shown). Previously, ocular and urogenital chlamydiosis was observed in approximately 20% of C. pecorum negative non-koala marsupials and C. pecorum positive test results could only be obtained from three quarters of animals showing classical signs of disease10,11. It is also interesting to note that in this and the previous study, only arboreal marsupials such as possums, gliders (and koalas) were thus far, found to be more often C. pecorum positive than ground-based marsupials, such as bandicoots which are more often C. pneumoniae positive10,11,12. For example, in our study, possums sampled from Tasmania had 100% Chlamydiales PCR positivity with C. pecorum strains identified twice. The quolls and bandicoots, predominantly sampled from the Northern Territory, had 27% Chlamydiales PCR positivity and no C. pecorum was identified; the Chlamydia-like genotype 1 identified from the quoll was, in fact, more similar to C. pneumoniae. While clearly too early yet to draw formal conclusions, we speculate that (i) C. pecorum infections are primarily asymptomatic in Australian marsupials and that (ii) ecological niche may play a role in predisposing individuals to infection. As to why koalas experience severe chlamydiosis, further comparative genomics analysis of koala and non-koala marsupial C. pecorum may reveal undetected C. pecorum strain diversity or other pathogenic agents, including other Chlamydiales that may predispose koalas to disease in a way that other marsupials are not.
Beyond C. pecorum, novel Chlamydiales were highly abundant amongst non-koala marsupials and made up approximately 50% of the genotypes retrieved in this screen (Table 2, Fig. 1). “Uncultured Chlamydiales” were previously described as the most abundant taxa in non-koala marsupials, even in smaller screens10,11, with the impact of these infections on animal health still unknown. Compared to the previous studies, the majority of the novel CRBs identified from marsupials in this study, however, are thought to primarily be limited to infections of arthropods13,18,23,24. Indeed, the predominant CRB sequence detected was a partial Ca. Rhabdochlamydia porcellionis 16S rRNA gene sequence (Genotype 1) in a range of marsupials that was 100% identical to a genotype we recently described in engorged marsupial feeding tick species Ixodes tasmani and Ixodes holocyclus removed from koalas13. In addition to this genotype being present in four non-koala marsupials distributed across all three geographic regions sampled, five of the 37 koalas we screened were also positive for this partial 16S rRNA gene sequence. Curiously, this genotype was found in marsupials in all three regions included in this study, yet we only found this genotype in ticks removed from koalas in Queensland, a sampling region included in the East Coast. Although the sample sizes were smaller, ticks removed from other marsupial species did not carry this genotype13. If this observation is true, this is surprising given that I. tasmani, the common marsupial tick is the most widely distributed Ixodes species and is found throughout mainland Australian as well as Tasmania. The fact that marsupials and their parasitising arthropods share the same CRBs could implicate ticks as a route of transmission for these novel Chlamydiae. This suggestion was also made by a recent study investigating the occurrence of CRBs in ticks and skin biopsies from suspected tick bite individuals in Finland. Multiple sequences obtained from skin biopsies were found to be most closely related to CRBs detected from ticks species found in Europe, and in some cases, Finnish ticks25. Further investigation into the pathogenesis and biology of novel CRBs is obviously warranted, especially given their presence in potential arthropod vectors.
Conclusions
This body of work further reinforces that chlamydial infections, including the previously described koala pathogen, C. pecorum, and other CRBs are widespread in Australian wildlife with some hosts more commonly infected than others. It appears that these infections do not cause traditional chlamydial diseases, certainly not like the debilitating chlamydial diseases that are observed in koalas, although longitudinal health studies would be required to ascertain this further. In the case of C. pecorum, what answers this holds about the origin and evolution of this pathogen in Australian marsupials is difficult to assess without further insight into the genetic differences that may exist between marsupial C. pecorum genotypes identified in this study with koala, sheep and cattle strains. Interestingly, the identification of a arthropod-associated Ca. Rhabdochlamydia porcellionis genotype in various marsupials, including koalas and koala-feeding tick species also suggests that arthropods are not the only host to this organism and further investigation into the pathogenesis and transmission of this and related CRBs is required.
Materials and Methods
Sampling
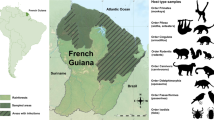
This study utilised a combination of (i) retrospective screening of wildlife samples collected as a part of other investigations and (ii) prospective sampling studies to investigate the prevalence of chlamydial infections in Australian marsupials from different geographic regions across Australia. Samples were obtained by swabbing the mucosal epithelia of conjunctiva, urogenital tract, rectal and penile tissues using Copan rayon tipped, aluminium-shafted applicator swabs (Interpath Services, Heidelberg West, Australia) for prospective sampling and both Copan rayon tipped, aluminium-shafted applicator swabs and Copan FLOQSwabs (Interpath Sevices) for retrospective sampling. Cumulatively, 401 swabs were opportunistically collected from 231 individual non-koala marsupials, comprised of 11 species across four Australian regions. The marsupial species, number of each marsupial and swab site are presented in Table 1. As a sample comparison set, 57 swabs from 37 koalas were opportunistically screened, as well.
All experimental protocols for the sampling of marsupials in this study were approved by the University of the Sunshine Coast Animal Ethics Committee (ANS1539). All methods were carried out in accordance with the 2013 Australian National Health and Medical Research Council ‘Australian code for the care and use of animals for scientific purposes’.
Retrospective sampling
Northern Territory
A total of 217 DNA extracts from 154 individual non-koala marsupials spanning four species were included from the Northern Territory (Table 1). DNA extracts of 91 mixed mucosal epithelial sites (eye, nose, cloaca) from 91 individuals, representing four marsupial species were collected as part of a large-scale study of small mammal health in the Northern Territory Australia26. Another 126 DNA extracts of ocular, urogenital and rectal mucosal epithelial sites from 63 individuals, representing two marsupial species were collected as part of a large-scale study of a reintroduction program of the northern quoll.
Prospective Sampling
East Coast
One hundred and eighteen swabs were opportunistically taken from various mucosal epithelial sites (eye/s, urogenital tract, rectum and penis) of 55 non-koala marsupials representing eight marsupial species (Table 1) presenting to collaborating Australian veterinarian and wildlife care centres in New South Wales and Queensland. A total of 57 swabs from 37 individual koalas were taken from eye and urogenital mucosal epithelial sites of 9 and 28 koalas presenting to Wildlife Rehabilitation Centre at Eumundi, Queensland and Port Macquarie Koala Hospital, New South Wales, respectively.
Tasmania
Sixty six swabs were collected from the left and right eye and the urogenital tract mucosal epithelium of 22 deceased, freeze-thawed common brushtail possums at the University of Tasmania (Table 1). The possums originated from commercial hunter-killed stock, collected from mixed agricultural-woodlands on private properties in Northern Tasmania.
DNA extraction
DNA extraction from swabs obtained from prospective sampling was performed using the QIAmp DNA mini kit (QIAGEN, Victoria, Australia). Swabs were vigorously shaken in TE (Tris-EDTA pH 8) buffer, followed by DNA extraction using the ‘DNA purification from tissues’ protocol as per the manufacturer’s instructions. DNA was stored at −20 °C until further use. DNA extractions performed on retrospective samples used the SIGMA Genelute bacterial DNA extraction kit in accordance with the manufacturer’s instructions.
Order specific 16S Chlamydiales PCR
To screen marsupials for Chlamydiales, an 800 bp fragment of the 16S rRNA gene partially covering the Chlamydiales signature sequence was amplified. PCR reactions consisted of 25 µl volumes containing 1.5 µl of each primer and 4 µl of template DNA, products were purified and sequenced following previously described methods13.
Chlamydia pecorum MLST PCR scheme
Samples with 16S rRNA sequences with a closest BLAST match to Chlamydia pecorum were subjected to C. pecorum MLST. The seven conserved C. pecorum specific housekeeping genes from the C. pecorum MLSA scheme was amplified following previously described methods8.
Sequence analysis and phylogenetic parameters
16S rRNA and gidA PCR products was purified using the Roche High Pure PCR Product Purification Kit (Roche, New South Wales, Australia) following the manufacturer’s instructions. Purified PCR products were sequenced by Macrogen Inc. (Seoul, Korea). Chromatograms of forward and reverse sequences were aligned in the Geneious R9.1.3 software package27. A consensus sequence was derived from each alignment and trimmed to the maximum length possible, gidA consensus sequences were trimmed to 474bp8. 16S rRNA gene consensus sequences, GenBank representatives of the closest BLAST match for each of the Chlamydiaceae, Ca. Rhabdochlamydiaceae and Ca. Parilichlamydiaceae families and an appropriate outgroup were then aligned in Geneious R9.1.3 using the MUSCLE plugin28 with default parameters. The alignments comprised of 18, 7 and 7 16S rDNA sequences trimmed to a length of 588, 368 and 701 bp, respectfully. For gidA, 12 sequences including consensus sequences, representatives of the closest PubMLST match16 from all possible host species and an appropriate outgroup were aligned in Geneious R9.1.3 using the MUSCLE plugin with default parameters. Bayesian phylogenies were constructed in Geneious R9.1.3 using the MrBayes plugin29 under the HKY85 substitution model. Run parameters included four Markov chain Monte Carlo (MCMC) chains with a million generations, sampled every 3,000 generations, and with the first 100,000 trees discarded as ‘burn-in’.
References
Burnard, D. & Polkinghorne, A. Chlamydial infections in wildlife–conservation threats and/or reservoirs of ‘spill-over’ infections? Vet. Microbiol. 196, 78–84 (2016).
Polkinghorne, A., Hanger, J. & Timms, P. Recent advances in understanding the biology, epidemiology and control of chlamydial infections in koalas. Vet. Microbiol. 165, 214–223 (2013).
Patterson, J. L. S. et al. The prevalence and clinical significance of chlamydial infection in island and mainland populations of Victorian koalas (Phascolarctos cinereus). J. Wildl. Dis. 51, 309–317 (2015).
Devereaux, L. N., Polkinghorne, A., Meijer, A. & Timms, P. Molecular evidence for novel chlamydial infections in the koala (Phascolarctos cinereus). Syst. Appl. Microbiol. 26, 245–253 (2003).
Jackson, M., Giffard, P. & Timms, P. Outer membrane protein a gene sequencing demonstrates the polyphyletic nature of koala Chlamydia pecorum isolates. Syst. Appl. Microbiol. 20, 187–200 (1997).
Marsh, J., Kollipara, A., Timms, P. & Polkinghorne, A. Novel molecular markers of Chlamydia pecorum genetic diversity in the koala (Phascolarctos cinereus). BMC microbiol. 11, 77 (2011).
Bachmann, N. L. et al. Comparative genomics of koala, cattle and sheep strains of Chlamydia pecorum. BMC Genomics 15, 667 (2014).
Jelocnik, M., Frentiu, F. D., Timms, P. & Polkinghorne, A. Multilocus sequence analysis provides insights into molecular epidemiology of Chlamydia pecorum infections in australian sheep, cattle, and koalas. J. Clin. Microbiol. 51, 2625–2632 (2013).
Jelocnik, M. et al. Evaluation of the relationship between Chlamydia pecorum sequence types and disease using a species-specific multi-locus sequence typing scheme (MLST). Vet. Microbiol. 174, 214–222 (2014).
Bodetti, T. J. et al. Wide range of Chlamydiales types detected in native Australian mammals. Vet. Microbiol. 96, 177–187 (2003).
Warren, K. et al. Ocular Chlamydiales infections of western barred bandicoots (Perameles bougainville) in Western Australia. J. Zoo. Wildl. Med. 36, 100–102 (2005).
Kumar, S. et al. Isolation and antimicrobial susceptibilities of chlamydial isolates from western barred bandicoots. J. Clin. Microbiol. 45, 392–394 (2007).
Burnard, D. et al. Novel Chlamydiales genotypes identified in ticks from Australian wildlife. Parasit. Vectors 10, 46 (2017).
Pillonel, T., Bertelli, C., Salamin, N. & Greub, G. Taxogenomics of the order Chlamydiales. Int. J. Syst. Evol. Microbiol. 65, 1381–1393 (2015).
Everett, K. D., Bush, R. M. & Andersen, A. Emended description of the order Chlamydiales, proposal of Parachlamydiaceae fam. Nov. And Simkaniaceae fam. Nov., each containing one monotypic genus, revised taxonomy of the family Chlamydiaceae, including a new genus and five new species, and standards for the identification of organisms. Int. J. Syst. Bacteriol. 2, 415–440 (1999).
Jolley, K.A., & Maiden, M. C. J. BIGSdb: Scalable analysis of bacterial genome variation at the population level. BMC Bioinformatics 11, 595 http://pubmlst.org/Chlamydiales/ (2010).
Walker, E., Lee, E. J., Timms, P. & Polkinghorne, A. Chlamydia pecorum infections in sheep and cattle: A common and under-recognised infectious disease with significant impact on animal health. Vet. J. 206, 252–260 (2015).
Pilloux, L. et al. High prevalence and diversity of Chlamydiales DNA within Ixodes ricinus ticks suggest a role of ticks as reservoir and vectors of Chlamydia-related bacteria. Appl. Environ. Microbiol. 81, 8177–8182 (2015).
Croxatto, A. et al. Presence of Chlamydiales DNA in ticks and fleas suggests that ticks are carriers of Chlamydiae. Ticks Tick-Borne Dis. 5, 359–365 (2014).
Griffith, J. E., Dhand, N. K., Krockenberger, M. B. & Higgins, D. P. A retrospective study of admission trends of koalas to a rehabilitation facility over 30 years. J. Wildl. Dis. 49, 18–28 (2013).
Gonzalez-Astudillo, V., Allavena, R., McKinnon, A., Larkin, R. & Henning, J. Decline causes of koalas in South East Queensland, Australia: A 17-year retrospective study of mortality and morbidity. Sci. Rep. 7, 42587 (2017).
Blom, W. M. & Alsop, D. B. Carbonate mud sedimentation on a temperate shelf: Bass basin, Southeastern Australia. Sediment. Geol. 60, 269–280 (1988).
Kostanjsek, R., Strus, J., Drobne, D. & Avgustin, G. ‘Candidatus Rhabdochlamydia porcellionis’, an intracellular bacterium from the hepatopancreas of the terrestrial isopod Porcellio scaber (Crustacea: Isopoda). Int. J Syst. Evol. Microbiol. 54, 543–549 (2004).
Corsaro, D. et al. Candidatus Rhabdochlamydia crassificans’, an intracellular bacterial pathogen of the cockroach Blatta orientalis (Insecta: Blattodea). Syst. Appl. Microbiol. 30, 221–228 (2007).
Hokynar, K. et al. Chlamydia-like organisms (CLOs) in Finnish Ixodes ricinus ticks and human skin. Microorganisms 4, 28 (2016).
Reiss, A. et al. Investigation of potential disease associated with Northern Territory Mammal declines. Murdoch University http://www.murdoch.edu.au/School-of-Veterinary-and-Life-Sciences/Our-courses/Postgraduate/Conservation-Medicine/Research/Investigating-the-role-of–disease-in-the-decline-of-small-mammal-species-in-the–Top-End-of-the-Northern-Territory/ (2015).
Kearse, M. et al. Geneious basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Edgar, R. C. Muscle: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Huelsenbeck, J. P. & Ronquist, F. Mrbayes: Bayesian inference of phylogeny. Bioinformatics 17, 754–755 (2001).
Acknowledgements
The authors would like to acknowledge Joanne Devlin and Jane Owens from the University of Melbourne for their roles in sample collection and DNA extractions; as well as Bruce Jackson of the Tasmanian DPI, Parks, Waters and the Environment and Sybil Sum for the collection of brushtail possum samples. We would like to thank the traditional owners of lands on which animals were sampled. The authors also wish to acknowledge the assistance of staff and participants from Wildlife Rehabilitation Centre at Eumundi, Australia Zoo Wildlife Hospital, Murdoch University and Northern Territory Flora and Fauna Division, North Queensland Wildlife Care Inc., Port Macquarie Koala Hospital, the Marthakal Rangers and the Territory Wildlife Park for assistance with fieldwork and collection of swabs. This project was supported by an Australian Research Council Discovery Project Grant awarded to PT and AP (DP130102066), an Australian Research Council Linkage grant awarded to JKW (LP LP150100722) and the Holsworth Wildlife Research Endowment - Equity Trustees Charitable Foundation awarded to DB. The sequenced retrieved from this study can be found under GenBank identifiers MF620049-MF620061.
Author information
Authors and Affiliations
Contributions
D.B. performed the majority of D.N.A. extractions, all molecular work, data analysis and was primarily responsible for preparation of the manuscript. W.H. and J.W. assisted with sample collection and D.N.A. extractions. M.J. assisted with molecular typing. A.G., A.R., S.F., S.C., J.C. and C.F. assisted with sample collection. A.P. and P.T. conceived the study and assisted with drafting of the manuscript. All authors have read and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Burnard, D., Huston, W.M., Webb, J.K. et al. Molecular evidence of Chlamydia pecorum and arthropod-associated Chlamydiae in an expanded range of marsupials. Sci Rep 7, 12844 (2017). https://doi.org/10.1038/s41598-017-13164-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13164-y
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.