Abstract
The generation of T cells from pluripotent stem cells (PSCs) is attractive for investigating T cell development and validating genome editing strategies in vitro. X-linked severe combined immunodeficiency (X-SCID) is an immune disorder caused by mutations in the IL2RG gene and characterised by the absence of T and NK cells in patients. IL2RG encodes the common gamma chain, which is part of several interleukin receptors, including IL-2 and IL-7 receptors. To model X-SCID in vitro, we generated a mouse embryonic stem cell (ESC) line in which a disease-causing human IL2RG gene variant replaces the endogenous Il2rg locus. We developed a stage-specific T cell differentiation protocol to validate genetic correction of the common G691A mutation with transcription activator-like effector nucleases. While all ESC clones could be differentiated to hematopoietic precursor cells, stage-specific analysis of T cell maturation confirmed early arrest of T cell differentiation at the T cell progenitor stage in X-SCID cells. In contrast, genetically corrected ESCs differentiated to CD4 + or CD8 + single-positive T cells, confirming correction of the cellular X-SCID phenotype. This study emphasises the value of PSCs for disease modelling and underlines the significance of in vitro models as tools to validate genome editing strategies before clinical application.
Similar content being viewed by others
Introduction
Pluripotent stem cells (PSCs), such as embryonic stem cells (ESCs) and induced PSCs (iPSCs), are attractive cells for the development of novel, patient-specific approaches in regenerative medicine, drug discovery and disease modelling. While ESCs are derived from the inner cell mass of mammalian blastocysts1, iPSCs are generated in vitro by the expression of defined transcription factors needed to convert a differentiated somatic cell into pluripotency2. Both cell types share common characteristics, such as their ability to grow indefinitely while maintaining pluripotency, and the ability to differentiate into somatic cell types, including blood and immune cells.
T cells are a key component of the adaptive immunity, which provides host protection against pathogens and cancer. Unlike other haematopoietic lineages, T cell development occurs outside the bone marrow in the thymus, a lymphoid organ that provides the optimal microenvironment to support T cell maturation3. Patients with hereditary defects in the T cell compartment can be severely immune deficient, and the underlying disorders are collectively called severe combined immunodeficiency (SCID)4. One of the most common forms is X-linked SCID (X-SCID), which is caused by mutations in the IL2RG gene5,6. IL2RG codes for the common gamma chain (GC), which is present in several interleukin receptors, such as the IL-2, IL-4, IL-7, IL-9, IL-15 and IL-21 receptors, and therefore essential for the development and function of lymphocytes7. The immune phenotype of X-SCID patients is characterized by the absence of T and NK cells in combination with poorly active B cells in their peripheral blood8. Because the early block in lymphopoiesis limits readily accessible patient material, X-SCID is difficult to study in patients. Moreover, the available mouse models fail to accurately recapitulate the human phenotype9. Thus, a stage-specific in vitro generation of T cells from PSCs is a valuable tool to better characterise the cellular phenotype of X-SCID.
X-SCID disease is of particular importance for the assessment of novel genome editing applications as gene therapy approaches for this disorder have been successfully validated in the clinic10,11. Retroviral IL2RG gene transfer in haematopoietic stem cells (HSCs) has been assessed in autologous settings in several clinical trials. The outcome of these studies has shown near complete immune reconstitution, with similar or even better outcome to that of mismatched allogeneic HSC transplantation12. While insertional mutagenesis led to the development of leukaemia in two early gene therapy trials involving first-generation gamma-retroviral vectors13,14, more recent trials with self-inactivating (SIN) vectors were successful without severe adverse events so far10. Additionally, a pre-clinical proof-of-concept study for zinc-finger nuclease (ZFN)-mediated correction of the IL2RG gene in HSCs demonstrated the feasibility of targeted gene editing in such multipotent cells15.
Designer nucleases are custom-made genome modifiers that have developed into indispensable tools for modelling human disease in vitro and for clinical applications16. The major classes of designer nucleases comprise ZFNs17, transcription activator-like effector nucleases (TALENs)18,19, and the clustered regularly interspaced short palindromic repeats (CRISPR)-Cas system20. These nucleases induce a site-specific DNA double strand break that activates one of the two major DNA repair pathways, non-homologous end joining (NHEJ) or homology-directed repair (HDR), which in turn can be harnessed either for gene disruption or gene targeting in the presence of a suitable donor DNA template21.
Although HSCs are the most relevant cell type for gene editing geared towards clinical translation, several restraints limit their use for detailed biological analyses, including the lack of robust protocols to culture and expand HSCs in vitro. PSCs, on the other hand, provide an unlimited source of stem cells that can be cloned and terminally differentiated into somatic cells of interest22. In several studies PSCs were combined with designer nuclease technology to provide proof-of-concept that genome engineering can be used to tackle disorders, including many haematopoietic disorders, such as chronic granulomatous disease23,24, β-thalassemia25 and haemophilia26. With regard to the in vitro generation of immune cells, PSCs have been successfully differentiated to myeloid cells23,24,27,28, but the in vitro production of lymphocytes has proven to be difficult. The differentiation of defined murine or human HSCs to T cells has been successfully achieved by cultivating the stem cells on a monolayer of murine OP9 bone marrow stroma cells expressing the notch delta-like 1 ligand (OP9-DL1)29. The same general setup was adapted to differentiate PSCs to T cell precursors, with the generation of few mature T cells alongside immature CD4 + /CD8 + double-positive (DP) T cells30,31,32,33. Recently, patient-specific IL2RG, JAK3 and WAS-deficient iPSC lines were corrected by designer nucleases and combined successfully with NK and T cell differentiation protocols34,35,36. However, these studies did not focus on stage-specific T cell development, which is particularly important when modelling X-SCID in order to demonstrate the role of GC in IL-7 and IL-2 dependent T cell development and T cell expansion7,37.
Here, we report a differentiation protocol, which enabled us to generate single-positive T cells from murine ESCs in a stage-specific manner. We verified the potential of this protocol for disease modelling by utilizing ESCs in which the murine Il2rg locus was replaced with a human IL2RG version harbouring the common G691A mutation in exon 5. After correction of the underlying mutation in IL2RG with the TALEN technology, the resulting ESCs were differentiated in the presence of IL-7 and IL-2 to single-positive CD4+ and CD8+ T cells, confirming correction of the cellular X-SCID phenotype in vitro. This study re-emphasises the significance of designer nucleases as a tool for targeted genetic engineering and demonstrates that pluripotent stem cells can be used to analyse in-depth the stage-specific differentiation of hematopoietic precursor cells into single-positive T cells.
Results
Generation of a humanised X-SCID mouse ESC model
To generate the humanised X-SCID mouse ESC line, the full-length mouse Il2rg locus with its promoter and enhancer elements was replaced by a full-length human IL2RG locus containing the common G691A point mutation in exon 5. This humanised cellular mouse model of X-SCID not only provides the mutant IL2RG target for site-specific endonucleases, but due to its location in the mouse genome, it also takes into account the chromatin structure and epigenetic factors that may affect gene targeting.
Genetic correction of IL2RG locus
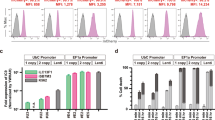
In previous studies, we compared the activities and toxicities of ZFNs and TALENs targeting exon 5 of the human IL2RG locus38,39. One specific TALEN pair combined high activity with little cytotoxic and was therefore chosen for this study. To correct the mutation in exon 5, we generated a therapeutic donor construct (GC donor) that contained a partial cDNA coding for exons 5 to 8, followed by the natural 3′-untranslated region (UTR) and a polyadenylation (pA) site (Fig. 1A). The inclusion of the natural 3′-UTR in combination with a viral pA site should stabilise the mRNA and improve transcriptional termination in order to enhance expression of the IL2RG gene product. To prevent cleavage of the donor by the TALEN pair, two silent mutations (sequence tag) were introduced into the left TALEN binding half-site. Furthermore, the donor harboured a phosphoglycerate kinase (PGK) promoter driven puromycin resistance cassette (PuroR) to enable enrichment of gene targeted cells. X-SCID ESCs were transfected with TALEN expression plasmids and the GC donor. Following puromycin selection, 100 resistant ESC clones were expanded and screened for the targeted integration event. Initial screening involved PCR-based confirmation of the 3′-junction and the presence of the donor (Figs 1B,S1). The expected PCR amplicon was detected in four out of 100 clones, indicative of successful integration of the GC donor into the endogenous IL2RG locus (Figure S1). Further genotyping involved a PCR-based confirmation of the correct 5′-junction in these four clones (Fig. 1B). The expected PCR amplicon of 1.1 kb was detected in all four clones, whereas the 1.9 kb product, indicative of the ‘SCID allele’, was detected only in non-transfected samples (mock) and cells transfected with the GC donor alone (Fig. 1B). To identify non-targeted random integration (RI) events, a PCR with a primer binding in the backbone of the GC donor was applied. Two of the four clones were negative, showing that these corrected clones (C16, C69) do not harbour additional integration events (Fig. 1B). For further proof of genetic correction, exon 5 was sequenced. Sequence analysis of both corrected clones confirmed correction of the point mutation and the presence of the sequence tag (Fig. 1C). In conclusion, we were able to genetically correct a point mutation in the human IL2RG gene in pluripotent stem cells using TALEN-mediated targeted integration of a partial cDNA into exon 5 of this locus.
Genetic correction of X-SCID. (A) Schematic of targeting strategy to correct the G691A mutation in IL2RG exon 5. The point mutation is highlighted in red and the TALEN target site in grey. The GC donor contains the therapeutic super-exon encompassing exons 5 to 8, a puromycin selection (PuroR) cassette, and homology arms to the endogenous IL2RG locus (indicated by interrupted lines). Arrows mark the binding sites of the primers used for PCR-based genotyping. (B) Genotyping. PCR-based genotyping was performed to detect the 5′-junction (primers 5′F/5′R), 3′-junction (primers 3′F/3′R; see also Figure S1) and randomly integrated (RI) donors (primers RIF/RIR; see also Table S1). Uncropped gel images shown in Figure S4B. (C) Sequence analysis of exon 5. The position of the mutated ‘A’ and the reversion to ‘G’ as well as the sequence tag (seq tag) are highlighted. C11, C16, C42, C69, corrected ESC clones; SCID, uncorrected X-SCID ESC clone; Donor, ESC clone transfected with donor alone.
Differentiation of ESCs to CD4 + /CD8 + double-positive T cells
In order to induce T cell development from haematopoietic stem cells (HSCs) in vitro, a combination of notch and IL-7 mediated signalling is essential29,40,41. Initial experiments with bone marrow derived lineage-negative (lin-) cells showed that co-cultivation with OP9-DL1 stroma cells enabled the differentiation of these multipotent stem cells through all CD4-/CD8- double-negative (DN) stages, as determined by flow cytometric analysis of CD25 and CD44 surface expression (Figure S2). Further cultivation of these early thymocytes on OP9-DL1 cells led to the generation of CD4+/CD8+ double-positive (DP) T cells, which expressed the beta-chain of the T cell receptor (TCRβ), indicating that these cells have successfully passed the β-selection checkpoint in vitro.
Pluripotent stem cells have the potential to differentiate to defined somatic cell types only when the right differentiation conditions are available at the right time. Differentiation of pluripotent stem cells to T cells includes an initial mesoderm induction, followed by the specification to haematopoietic precursor cells (HPCs) before proceeding to T cell differentiation (Fig. 2A). To this end, we have established an embryoid body (EB)-based differentiation protocol to generate HPCs from corrected and non-corrected X-SCID ESC clones, as well as from the wildtype (WT) ESC clone CCE1 as a control. In order to monitor the optimal time point for HPC induction, EBs were harvested and dissociated either directly after hematopoietic specification at day 7, or three days later at day 10. Enrichment of HPCs was monitored by assessing the expression of the surface markers CD41 and c-Kit (CD117)42 (Fig. 2B). For all ESC lines, comparable levels of CD41+or cKit+/CD41+ cells were detected at day 7 (Fig. 2C,D). However, the fraction of CD41+ or cKit+/CD41+ cells dropped sharply when harvested at day 10, and we were not able to differentiate these ‘late’ HPCs into T cells (not shown). This indicates the importance of proceeding to T cell differentiation directly after specification to hematopoietic differentiation at day 7, where a prominent cKit+/CD41+ cell population is evident.
Differentiation of ESCs to hematopoietic precursor cells. (A) Schematic overview of the differentiation protocol. Humanised mouse ESCs were kept in pluripotent state in mESC medium (precultivation). For EB formation and generation of hematopoietic precursor cells (HPCs), ESCs were cultured for either 7 or 10 days in EB medium. EBs were primed with mesoderm-inducing cytokines at day 3 (induction) and with haematopoiesis-inducing cytokines at day 6 (specification). (B) Qualitative assessment of in vitro generated HPCs. EBs were dissociated and stained for CD41 and c-Kit (CD117) at days 7 and 10. Flow cytometric analysis is shown for 7-AAD negative cells. Numbers indicate percentage of cells in each quadrant. (C,D) Quantitative assessment of in vitro generated HPCs. Shown are the average percentages with standard error of mean (S.E.M.) of generated CD41+ (C) or cKit+/CD41+ (D) cells harvested at days 7 or 10 (n = 3). SCID, uncorrected X-SCID ESC clone; C16 and C69, corrected ESC clones; WT, wildtype mouse ESC clone.
Because the IL2RG gene product is an integral part of the IL-7 receptor, it is essential for early T cell development. We therefore hypothesised that HPCs derived either from corrected ESC clones or WT ESCs will be able to differentiate to DP T cells, while cells derived from the non-corrected X-SCID ESC clone would be blocked at the DN1 stage (Fig. 3A). To verify this notion, day 7 HPCs were co-cultivated on a monolayer of OP9-DL1 cells, and T cell maturation profiled on a weekly basis by flow cytometry. We monitored the different developmental stages by determining surface expression of CD44 and CD25 within the CD4-/CD8- DN population: DN1 (CD44+/CD25-), DN2 (CD44+/CD25+), DN3 (CD44-/CD25+) and DN4 (CD44-/CD25-). As hypothesised, maturation of thymocytes derived from non-corrected HPCs was blocked at the DN1 stage (97%) and only detectable at week 2 of the analysis but not week 4 (Fig. 3B–D), suggesting that the derived T cell precursors go into apoptosis if differentiation is blocked at an early stage. Conversely, thymocytes derived from both corrected ESC clones (C16, C69) and the WT clone could progress into DN2, DN3, and DN4 stages (Fig. 3B,C ). Moreover, after 4 weeks of cultivation, the development of CD4+/CD8+ DP T cells became evident (Fig. 3B,D ), similarly to the T cells derived from lin- cells (Figure S2). Statistical analyses confirmed a difference in the fraction of generated DN1 T cells derived from the SCID clone as compared to all other clones at week 2 (Fig. 3C ) but not any significant differences when comparing any of the generated T cell subsets derived from the genetically corrected ESC lines with the WT clone (Fig. 3D ). In summary, these results demonstrate that our in vitro differentiation protocol allowed us to recapitulate the cellular X-SCID phenotype in vitro and enabled us to generate CD4+/CD8+ DP T cells in a stage-specific manner.
In vitro differentiation of ESC-derived hematopoietic precursors to CD4+/CD8+ double-positive T cells. (A) Schematic of early stage T cell development. ESCs derived hematopoietic precursor cells (HPCs; Fig. 2) were cultured on OP9-DL1 cells to provide DL-1 mediated notch signalling that is needed to commit differentiation of HPCs to the T cell lineage, including CD4-/CD8- double-negative (DN) early T cell progenitors (DN1), pro-T cells (DN2), which undergo V(D)J recombination to become pre-T cells (DN3), that in turn pass β-selection to develop into DN4 and finally CD4+/CD8+ double-positive (DP) T cells, which express the pan-lymphocyte marker CD3. Absence of the IL2RG gene product leads to an early block in T cell development at DN1 stage (red line). (B) Qualitative assessment of T cell maturation. Differentiation was analysed by flow cytometry after two and four weeks of co-cultivation on OP9-DL1 stroma cells by monitoring surface expression of CD44/CD25 and CD4/CD8 (see also Figure S2). (C,D) Quantitative assessment of T cell maturation. Shown is the average fraction of the various T cell subpopulations with standard error of mean (S.E.M.) of three independent experiments of cells analysed at week 2 (C) or week 4 (D). DN T cell subsets add up to 1.0 at week 2 (C), while at week 4 (D) also CD4 + /CD8 + DP (white columns) and some single-positive cells (not indicated, see Fig. 4) were present. Statistical differences in the fraction of DN1 cells at week 2 are indicated by **(p < 0.01) and ***(p < 0.001). No statistical differences between any of the generated T cell subsets derived from the gene-edited ESC lines and the WT clone was observed at week 4 (p > 0.05). Gating was applied in the following order: FSC/SSC and CD45 + /DAPI- (not shown) to assess CD4/CD8 expression; CD4-/CD8- to gate for DN1-DN4 stages assessed by CD25/CD44 expression. SCID, uncorrected X-SCID ESC clone; C16 and C69, corrected ESC clones; WT, wildtype ESC clone.
Generation of CD4+ and CD8+ single-positive T cells
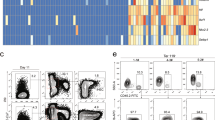
We further analysed the week 4 lymphocyte population (Fig. 3) for late T cell maturation stages by assessing the expression of the CD3 complex and the T cell receptor beta chain (TCRβ) (Fig. 4A). In combination with the CD4/CD8 expression patterns, this allowed us to determine the fraction of ‘DN’ T cells, immature single-positive (ISP) cells, ‘early DP’ and ‘late DP’ T cells, as well as CD4+ and CD8+ single positive (SP) T cells (Fig. 4B). When comparing the T cell maturation profiles of the two corrected ESC clones (C16, C69) with the wildtype (WT) cells, we neither observed significant qualitative (Fig. 4C) nor quantitative (Fig. 4D) differences between the generated T cell subsets derived from gene-edited (C16, C69) versus WT ESCs (p > 0.05). The lower numbers of overall produced T cells or the slightly higher percentage of single positive cells produced by the WT clone could be due to genetic differences between the WT cells and the two corrected X-SCID clones, which e.g. contain additional genetic elements in the IL2RG locus (Fig. 1A). Nonetheless, these results indicate that the corrected X-SCID cells were able to pass the positive selection checkpoint, which leads to upregulation of the CD3 complex before the cells differentiate into CD4+ and CD8+ single-positive (SP) thymocytes.
Assessment of in vitro generated T cells. (A) Schematic of late stage T cell maturation. T lymphocytes generated at week 4 on OP9-DL1 cells were characterised for their maturation stage based on the expression of CD4/CD8 as well as CD3 and T cell receptor beta-chain (TCRβ). CD4-/CD8- double-negative (‘DN’) T cells develop into immature single-positive (ISP) CD8+ cells that then differentiate into ‘early DP’ (CD4+ /CD8+ ) T cells. After passing positive selection, CD3 and TCRβ expression is upregulated. These ‘late DP’ then develop in CD4 + or CD8 + single positive (SP) T cells. (B) Qualitative assessment of late T cell maturation. Differentiation was analysed by flow cytometry by monitoring surface expression of CD4/CD8, CD3/TCRß, and CD3/CD8. A representative flow cytometry analysis for ESC clone C16 is shown. Gating was applied in the following order: FSC/SSC and CD45+/DAPI- (not shown) to assess CD4/CD8 expression; CD3 and TCRβ expression assessed on CD8+/CD4+T cells and CD3 expression assessed on CD8+ cells. (C) Relative assessment of T cell differentiation. Shown is the average fraction of the various T cell subsets with standard error of mean (S.E.M.) for both gene-edited clones (C16, C69) and the wildtype (WT) ESC clone. No statistical differences between any of the generated T cell subsets derived from the gene-edited ESC lines and the WT clone was observed (p > 0.05). (D) Quantitative assessment of the efficiency of T cell differentiation. Shown is the average of three independent experiments with S.E.M. for both corrected clones (C16, C69) and the WT ESC clone. No statistically significant differences between the number of T cell subsets generated from clones C16/C69 and WT was observed (p > 0.05).
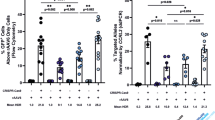
To assess whether the genetically corrected T cells are able to respond to IL-2, we activated the week 4 lymphocyte population with anti-CD3/CD28 beads and challenged them with antigen-presenting cells (irradiated splenocytes). After one week of co-culture in the presence or absence of IL-2, we assessed the proliferation capacity of the in vitro generated T cell population by labelling the cells with carboxyfluorescein succinimidyl ester (CFSE). Flow cytometric analysis revealed more cell divisions when cells were co-stimulated with IL-2 (Figure S3A). We also evaluated the fraction of CD8+ SP T cells by flow cytometry. While some CD8+ T cells were detectable after stimulation with anti-CD3/CD28 beads alone, IL-2 enhanced the proliferation of SP CD8+ cells considerably (Figure S3B) and induced a strong up-regulation of the IL-2 receptor alpha-chain (CD25) (Figure S3C). In conclusion, these supplementary experiments corroborated that genetic correction of the IL2RG locus restored IL-2 dependent proliferation of the T cell population.
Discussion
The therapeutic potential of haematopoietic stem cells has been extensively validated by allogeneic HSC transplantation for various indications. Moreover, retroviral gene therapy in autologous HSCs has been successfully adapted to treat several primary immunodeficiencies43, including X-SCID10, adenosine deaminase deficiency (ADA-SCID)44, and Wiskott-Aldrich Syndrome (WAS)45,46. While clinically relevant, HSCs are only of limited use for in vitro disease modelling. In this regard, PSCs have many advantages over HSCs, including the unlimited proliferation potential and the possibility to expand and screen individual clones47. Moreover, these cells hold great promise for the development of autologous cell therapies, personalised drug discovery and disease modelling. Here, we developed a cellular X-SCID disease model based on a murine knock-in ESC line that harbours a mutated copy of the human IL2RG locus. To model the cellular disease phenotype, we have established a T cell differentiation protocol that enabled us to differentiate early T cell precursors in a stage-specific manner to single positive T cells in vitro after TALEN-mediated correction of the IL2RG locus.
For clinical translation, it is imperative to identify designer nuclease with high activity, high specificity and low cytotoxicity and genotoxicity. We previously developed a TALEN architecture that combines high activity with high specificity and low cytotoxicity38, and this platform was successfully used to generate a TALEN pair to target exon 5 of IL2RG 39. However, targeting the human IL2RG locus with TALENs seems challenging. We have previously reported that the success rate of generating functional TALEN pairs to target IL2RG was considerably lower than producing TALENs that cleave the CCR5 locus or AAVS1 39. In this study, we used a murine ESC line that contains a mutated human IL2RG locus in lieu of the endogenous mouse Il2rg. The targeting efficiency after selection was only ~4% (4 out of 100 selected clones). Similarly, a recently reported TALEN-mediated gene targeting frequency in X-SCID patient-derived iPSCs was rather modest48. In contrast, ZFN-mediated gene targeting at the IL2RG locus in unselected multipotent HSCs reached ~6%15. Although the overall low gene targeting frequency in PSCs could be attributed to the potential differences in the chromatin structure at the IL2RG locus in PSCs as compared to HSCs, several studies in ESCs and iPSCs confirmed that even non-expressed genes in a closed chromatin structure could be easily targeted by ZFNs and TALENs49,50. On the other hand, it was shown that DNA methylation could have a negative impact on target site binding by TALENs51,52, which was not observed for ZFNs so far. Further studies will be needed to clarify this point.
Although T cell development is similar between humans and mice53, the available mouse disease models for X-SCID have failed to fully recapitulate the human phenotype54. This supports the notion that there are other, yet undiscovered molecular mechanisms, which differ between human and mouse. Interestingly, unlike the X-SCID mouse model, we did not observe leakage in T cell maturation in our in vitro differentiation system based on the IL2RG mutant mouse ESCs. Based on this observation, it will be interesting to generate novel mouse strains from our mutant and corrected ESC lines to address further the differences between in vivo and in vitro T cell development.
A detailed analysis of the different stages of T cell development in vitro is particularly valuable to study T cell maturation in a disease setting or to examine subtle changes in the epigenome or transcriptome during T cell differentiation. To our knowledge, this is the first report that recapitulates T cell differentiation in a disease context through all stages from early thymic progenitors (DN1) to CD4 + and CD8 + single positive T cells. Several studies described the generation of CD4+/CD8 + double positive T cells55,56 or even CD4+ and/or CD8+ single positive T cells30,31,32 from either normal murine or mouse ESCs or iPSCs. For X-SCID, Verma and colleagues validated the potential to differentiate gene edited patient derived iPSCs to NK cells and CD4+/CD8+ double positive T cells48, but failed to demonstrate stage-specific differentiation and maturation to single-positive T cells. Similarly, two other published studies demonstrated the feasibility to differentiate gene edited JAK3-deficient or WAS iPSCs to phenotypically mature T cells35,36 but did not assess the different stages of T cell development. Here, we show that PSCs can be differentiated in a stage-specific manner from early thymic progenitors (DN1) to single-positive CD4+ or CD8+ T cells. Using ESCs that contain the G691A point mutation in exon 5 of the human IL2RG locus, we demonstrate that hematopoietic differentiation of X-SCID ESCs was blocked at the DN1 stage, hence recapitulating the cellular in vivo phenotype in vitro. Moreover, HPCs derived from gene edited and WT ESCs went through all stages of T cell development, including β-selection, and could be differentiated to single-positive CD4+ or CD8+ T cells. Together, this study indicates that disease modelling based on PSC technology is not only a useful tool to study rare T cell disorders but, in combination with genome engineering tools, will allow researchers to analyse T cell differentiation and maturation step-by-step in an easily trackable in vitro system. A restriction in our hands, however, was that T cell development from PSCs seems to be a stochastic event, advocating for the need to identify the correct sub-population within the generated HPCs with high capacity for T cell differentiation.
In conclusion, our study provides evidence that naïve pluripotent stem cells can be differentiated in vitro in a stage-specific manner to CD3-positive CD8+ single-positive T cells. In our hands, the established system has been validated successfully for cellular disease models of X-SCID and RS-SCID33. Based on the developed protocol, it would be interesting to validate further this protocol to cover a broad range of T cell deficiencies, including mutations in the genes encoding ZAP70, CD3Z and CD45, in order to address different blocks in T cell development. Moreover, in combination with designer nucleases-based gene editing, the technology will also expedite the study of genotype-phenotype correlations and allow for screening of personalised patient-specific drugs by creating disease models in isogenic backgrounds.
Methods
Cells and cell culture
All cells were free of mycoplasma and kept at 37 °C with 5% CO2. OP9-DL1 cells (obtained from Juan Carlos Zúñiga-Pflücker) were cultured in OP9-DL1 medium [alpha-MEM (Life Technologies, supplement with 20% FBS (PAA), 100 U/ml of penicillin and 100 µg/ml of streptomycin (Sigma-Aldrich), 2 mM of L-glutamine (Merck Millipore) and 1 mM Na-pyruvate (PAA)]. All ESC lines were cultivated feeder-free on plates coated with 0.1% gelatine in ESGRO complete medium (Merck Millipore), supplement with 100 U/ml of penicillin and 100 µg/ml of streptomycin in addition to 4 µM of SB431542, 1 µM of PD0325901 and 3 µM of CHIR99021 (all Axon Medchem). Lineage-negative HSCs were isolated by flushing tibiae and femurs of C57BL/6 N mice, and purified by magnetic cell sorting (MACS) according to the manufacturer’s instructions (Lineage Cell Depletion Kit, Miltenyi Biotech). All procedures involving the usage of animals have been conducted according to institutional guidelines, national and state regulations, and approved by the Department for Rural Affairs and Consumer Protection of the Federal State of Baden-Wuerttemberg (MLR).
Plasmids
The TALEN expression plasmids were described before38,39. The GC donor plasmid generated by cloning the cDNA PCR product from exon 5–8, followed by the natural 3′-UTR of the IL2RG locus (kindly provided by Axel Schambach, Hannover Medical School). To improve transcriptional termination, the HSV polyA was cloned downstream of the 3′-UTR. The homology arms (HAR, HAL) were generated by PCR amplification of the IL2RG locus from genomic DNA of normal fibroblasts. Complete sequences and maps of plasmids can be obtained upon request.
Generation and genotyping of gene targeted mouse ESC clones
The humanised X-SCID mouse ESC line was generated by InGenious as follows: A 5.5 kb region of the human IL2RG gene was subcloned from a human BAC clone into shuttle vector pSP72 (Promega). MluI sites were engineered at the 5′ and 3′ ends by Red/ET recombineering. The G691A mutation was incorporated into exon 5 of the human IL2RG locus by overlap-extension PCR. A 14.4 kb region containing the mouse Il2rg locus was subcloned from a C57BL/6 BAC clone (RP23: 263O9) into the targeting vector. This region encompassed a 7.1 kb long 5′ homology arm, the 5.1 kb long coding region of the Il2rg gene, and the 3′ homology arm of 2.3 kb. The targeting vector also included a neomycin resistance gene for selection. After delivery of this vector to murine ESCs by electroporation, successfully targeted cells were enriched for by neomycin treatment. The aminoglycoside 3′-phosphotransferase cassette was flanked with LoxP sites, allowing the neomycin resistance gene to be excised from the genome via Cre-Lox recombination. Successfully gene targeted ESC clones were sequenced to confirm the presence of the mutant human IL2RG allele and deletion of the neomycin resistance cassette.
One day prior to the experiment, X-SCID ESCs were pre-treated with 10 nmol/ml of ROCK inhibitor (Wako, Japan). 2 × 10e6 cells were nucleofected with 1.25 µg of each TALEN expression plasmids and 2.5 µg of the GC donor using Mouse ES kit and program A-023 (Nucleofector I, Lonza). Selection with 0.4 µg/ml of puromycin (Roth, Germany) was applied for 5 days before individual resistance clones were isolated and expanded in 96-well plates. Once confluent, cells were lysed in 100 µl of Direct Lysis Buffer (PeqLab) supplement with protease K (20 µg/ml) (Thermo Scientific), incubated overnight at 55 °C, followed by heat inactivation at 86 °C for 45 min. 2 µl of the lysis mix was used as a template for the various PCR-based genotyping reactions. All PCR reactions were performed using Phire Hot Start II polymerase (Thermo Scientific, Germany) according to the manufacturer’s guidelines. The used primers are indicated in Supplementary Table 1.
Generation of CD4+ /CD8+ double-positive T cells
The protocol was modified from our previously published protocol33 as follows: For embryoid body (EB) formation, 1 × 10e5 mouse ESCs were kept in EB-medium [IMDM basic medium supplemented with 15% ES cult FBS (StemCell Technologies), 5% PFHMII medium (Life Technologies), 100 U/ml of penicillin and 100 µg/ml of streptomycin (Sigma-Aldrich), 2 mM L-glutamine (Merck Millipore), 50 µg/ml of vitamin C (Sigma-Aldrich), 150 mM monothioglycerol (Sigma-Aldrich), and 200 µg/ml of human transferrin (Sigma-Aldrich)] at 37 °C in 5% CO2 on a platform shaker at 80 rpm for a total of 7 to 10 days. Mesoderm induction was initiated at day 3 with rhBMP4, activin A, rhFGF-2 and rhVEGF (all at a final concentration of 5 ng/ml; all R&D Systems), followed by hematopoietic specification at day 6 with mIL-3 (10 ng/ml) and mSCF (40 ng/ml) (both Peprotech). To harvest the EBs, cells were incubated in 2 ml TrypLE (10X, Life Technologies) for 20 min at 37 °C incubator before being passed through 70 µm mesh. To induce T cell differentiation, the single cell solution containing HPCs was cultivated on a fresh monolayer of OP9-DL1 cells and primed with 2 ml of T cell medium [OP9DL1 complete medium supplemented with 1 ng/ml of mIL-7 and 5 ng/ml of rhFlt3-L (both Peprotech)]. Cells were harvested weekly by passing through a 70 µm mesh before being transferred to fresh monolayer of OP9-DL1 cells. The extent of T cell differentiation was assayed by flow cytometry as described below.
Generation of CD8+ single-positive T cells and cell proliferation assay
Mice splenocytes derived from C57BL/6 N mice were isolated and irradiated at 30 Gy prior to use. CD4+/CD8+ DP T cells were stained with 0.25 µM of carboxyfluorescein succinimidyl ester (CFSE) according to manufacturer’s instructions (Life Technologies). 5 × 10e5 stained cells were co-cultivated with the irradiated splenocytes at a ratio of 1:1 in 2 ml of T cell medium supplemented with anti-CD3/CD28 beads (Miltenyi Biotech) at a cell-to-bead ratio of 1:2 in the presence or absence of mIL-2 (100 ng/ml final volume). After 7 days, cells were harvested and analysed by flow cytometry as described below.
Flow cytometry
For flow cytometric analysis, cells were resuspended in FACS buffer (PBS supplemented with 2% FCS, 1 mM EDTA and 0.1% sodium azide (both Sigma-Aldrich)). EB-derived HPCs were treated with mouse Fc-blocking antibodies (BD Biosciences, Cat# 553142) for 5 minutes at 4 °C prior to staining with anti-CD41-PE (Cat# 12-0411-81) and anti-cKit-APC (Cat# 17-1171-81) antibodies, or corresponding isotype controls respectively (all eBioscience), and stained with 7-AAD for 2 min before analysis on BD Accuri C6 flow cytometer (BD Biosciences). ESC-derived T cells were stained with anti-CD45-APC-Cy7 (Cat# 557659), anti-CD4-PerCPR-Cy5.5 (Cat# 550954), anti-CD8-PE-Cy7 (Cat# 552877) (all BD Biosciences), anti-CD44-PE (Cat# 12-0441-81), anti-CD25-APC (Cat# 17-0251-81), anti-TCRß-FITC (Cat# 11-5961-82), anti-CD3e-PE (12-00331-82) (all eBioscience) antibodies and DAPI, before analysis on a FACSCanto II (BD Biosciences). All samples were evaluated using FlowJo software (Tree Star). To eliminate OP9-DL1 stroma cells from analysis, all samples were gated for CD45+ cells.
Statistical Analysis
All experiments were performed at least three times. Error bars represent standard error of mean (S.E.M.). Statistical significance was determined with a two-tailed Student’s t-test with equal variance.
Data Availability Statement
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676, doi:S0092-8674(06)00976-7 [pii] 10.1016/j.cell.2006.07.024 (2006).
Miller, J. F. The discovery of thymus function and of thymus-derived lymphocytes. Immunol Rev 185, 7–14 (2002).
Buckley, R. H. Molecular defects in human severe combined immunodeficiency and approaches to immune reconstitution. Annual review of immunology 22, 625–655, https://doi.org/10.1146/annurev.immunol.22.012703.104614 (2004).
Puck, J. M. et al. The interleukin-2 receptor gamma chain maps to Xq13.1 and is mutated in X-linked severe combined immunodeficiency, SCIDX1. Hum Mol Genet 2, 1099–1104 (1993).
Noguchi, M. et al. Interleukin-2 receptor gamma chain mutation results in X-linked severe combined immunodeficiency in humans. Cell 73, 147–157 (1993).
Rochman, Y., Spolski, R. & Leonard, W. J. New insights into the regulation of T cells by gamma(c) family cytokines. Nat Rev Immunol 9, 480–490, https://doi.org/10.1038/nri2580 (2009).
Conley, M. E. et al. X-linked severe combined immunodeficiency. Diagnosis in males with sporadic severe combined immunodeficiency and clarification of clinical findings. J Clin Invest 85, 1548–1554, https://doi.org/10.1172/JCI114603 (1990).
Mestas, J. & Hughes, C. C. Of mice and not men: differences between mouse and human immunology. J Immunol 172, 2731–2738 (2004).
Hacein-Bey-Abina, S. et al. A modified gamma-retrovirus vector for X-linked severe combined immunodeficiency. N Engl J Med 371, 1407–1417, https://doi.org/10.1056/NEJMoa1404588 (2014).
Gaspar, H. B. et al. Hematopoietic stem cell gene therapy for adenosine deaminase-deficient severe combined immunodeficiency leads to long-term immunological recovery and metabolic correction. Sci Transl Med 3, 97ra80, https://doi.org/10.1126/scitranslmed.3002716 (2011).
Touzot, F. et al. Faster T-cell development following gene therapy compared with haploidentical HSCT in the treatment of SCID-X1. Blood 125, 3563–3569, https://doi.org/10.1182/blood-2014-12-616003 (2015).
Hacein-Bey-Abina, S. et al. Insertional oncogenesis in 4 patients after retrovirus-mediated gene therapy of SCID-X1. J Clin Invest 118, 3132–3142, https://doi.org/10.1172/jci35700 (2008).
Howe, S. J. et al. Insertional mutagenesis combined with acquired somatic mutations causes leukemogenesis following gene therapy of SCID-X1 patients. J Clin Invest 118, 3143–3150, https://doi.org/10.1172/JCI35798 (2008).
Genovese, P. et al. Targeted genome editing in human repopulating haematopoietic stem cells. Nature 510, 235–240, https://doi.org/10.1038/nature13420 (2014).
Cornu, T. I., Mussolino, C. & Cathomen, T. Refining strategies totranslate genome editing to the clinic. Nat Med 23, 415–423, https://doi.org/10.1038/nm.4313 (2017). doi:.
Urnov, F. D., Rebar, E. J., Holmes, M. C., Zhang, H. S. & Gregory, P. D. Genome editing with engineered zinc finger nucleases. Nat Rev Genet 11, 636–646, https://doi.org/10.1038/nrg2842 (2010).
Mussolino, C. & Cathomen, T. TALE nucleases: tailored genome engineering made easy. Curr Opin Biotechnol 23, 644–650, https://doi.org/10.1016/j.copbio.2012.01.013 (2012).
Joung, J. K. & Sander, J. D. TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol 14, 49–55, https://doi.org/10.1038/nrm3486 (2013).
Doudna, J. A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096, https://doi.org/10.1126/science.1258096 (2014).
Carroll, D. Genome engineering with targetable nucleases. Annu Rev Biochem 83, 409–439, https://doi.org/10.1146/annurev-biochem-060713-035418 (2014).
Sterneckert, J. L., Reinhardt, P. & Scholer, H. R. Investigating human disease using stem cell models. Nat Rev Genet 15, 625–639, https://doi.org/10.1038/nrg3764 (2014).
Zou, J. et al. Oxidase-deficient neutrophils from X-linked chronic granulomatous disease iPS cells: functional correction by zinc finger nuclease-mediated safe harbor targeting. Blood 117, 5561–5572, https://doi.org/10.1182/blood-2010-12-328161 (2011).
Dreyer, A. K. et al. TALEN-mediated functional correction of X-linked chronic granulomatous disease in patient-derived induced pluripotent stem cells. Biomaterials 69, 191–200, https://doi.org/10.1016/j.biomaterials.2015.07.057 (2015).
Ma, N. et al. Transcription activator-like effector nuclease (TALEN)-mediated gene correction in integration-free beta-thalassemia induced pluripotent stem cells. J Biol Chem 288, 34671–34679, https://doi.org/10.1074/jbc.M113.496174 (2013).
Park, C. Y. et al. Targeted inversion and reversion of the blood coagulation factor 8 gene in human iPS cells using TALENs. Proc Natl Acad Sci USA 111, 9253–9258, https://doi.org/10.1073/pnas.1323941111 (2014).
Morishima, T. et al. Genetic correction of HAX1 in induced pluripotent stem cells from a patient with severe congenital neutropenia improves defective granulopoiesis. Haematologica 99, 19–27, https://doi.org/10.3324/haematol.2013.083873 (2014).
Lachmann, N. et al. Gene correction of human induced pluripotent stem cells repairs the cellular phenotype in pulmonary alveolar proteinosis. American journal of respiratory and critical care medicine 189, 167–182, https://doi.org/10.1164/rccm.201306-1012OC (2014).
Schmitt, T. M. & Zuniga-Pflucker, J. C. Induction of T cell development from hematopoietic progenitor cells by delta-like-1 in vitro. Immunity 17, 749–756 (2002).
Schmitt, T. M. et al. Induction of T cell development and establishment of T cell competence from embryonic stem cells differentiated in vitro. Nature immunology 5, 410–417, https://doi.org/10.1038/ni1055 (2004).
Timmermans, F. et al. Generation of T cells from human embryonic stem cell-derived hematopoietic zones. J Immunol 182, 6879–6888, https://doi.org/10.4049/jimmunol.0803670 (2009).
Kennedy, M. et al. T lymphocyte potential marks the emergence of definitive hematopoietic progenitors in human pluripotent stem cell differentiation cultures. Cell Rep 2, 1722–1735, https://doi.org/10.1016/j.celrep.2012.11.003 (2012).
Rahman, S. H. et al. Rescue of DNA-PK signaling and T-cell differentiation by targeted genome editing in a prkdc deficient iPSC disease model. PLoS Genet 11, e1005239, https://doi.org/10.1371/journal.pgen.1005239 (2015).
Menon, T. et al. Lymphoid regeneration from gene-corrected SCID-X1 subject-derived iPSCs. Cell Stem Cell 16, 367–372, https://doi.org/10.1016/j.stem.2015.02.005 (2015).
Chang, C. W. et al. Modeling Human Severe Combined Immunodeficiency and Correction by CRISPR/Cas9-Enhanced Gene Targeting. Cell Rep 12, 1668–1677, https://doi.org/10.1016/j.celrep.2015.08.013 (2015).
Laskowski, T. J. et al. Gene Correction of iPSCs from a Wiskott-Aldrich Syndrome Patient Normalizes the Lymphoid Developmental and Functional Defects. Stem Cell Reports 7, 139–148, https://doi.org/10.1016/j.stemcr.2016.06.003 (2016).
Boyman, O. & Sprent, J. The role of interleukin-2 during homeostasis and activation of the immune system. Nat Rev Immunol 12, 180–190, https://doi.org/10.1038/nri3156 (2012).
Mussolino, C. et al. A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Res 39, 9283–9293, https://doi.org/10.1093/nar/gkr597 (2011).
Mussolino, C. et al. TALENs facilitate targeted genome editing in human cells with high specificity and low cytotoxicity. Nucleic Acids Res 42, 6762–6773, https://doi.org/10.1093/nar/gku305 (2014).
Pui, J. C. et al. Notch1 expression in early lymphopoiesis influences B versus T lineage determination. Immunity 11, 299–308 (1999).
Radtke, F. et al. Deficient T cell fate specification in mice with an induced inactivation of Notch1. Immunity 10, 547–558 (1999).
McKinney-Freeman, S. L. et al. Modulation of murine embryonic stem cell-derived CD41 + c-kit + hematopoietic progenitors by ectopic expression of Cdx genes. Blood 111, 4944–4953, https://doi.org/10.1182/blood-2007-11-124644 (2008).
Williams, D. A. & Thrasher, A. J. Concise review: lessons learned from clinical trials of gene therapy in monogenic immunodeficiency diseases. Stem Cells Transl Med 3, 636–642, https://doi.org/10.5966/sctm.2013-0206 (2014).
Aiuti, A. et al. Gene therapy for immunodeficiency due to adenosine deaminase deficiency. N Engl J Med 360, 447–458, https://doi.org/10.1056/NEJMoa0805817 (2009).
Aiuti, A. et al. Lentiviral hematopoietic stem cell gene therapy in patients with Wiskott-Aldrich syndrome. Science 341, 1233151, https://doi.org/10.1126/science.1233151 (2013).
Hacein-Bey Abina, S. et al. Outcomes following gene therapy in patients with severe Wiskott-Aldrich syndrome. JAMA 313, 1550–1563, https://doi.org/10.1001/jama.2015.3253 (2015).
Ott de Bruin, L. M., Volpi, S. & Musunuru, K. Novel Genome-Editing Tools to Model and Correct Primary Immunodeficiencies. Front Immunol 6, 250, https://doi.org/10.3389/fimmu.2015.00250 (2015).
Menon, T. et al. Lymphoid Regeneration from Gene-Corrected SCID-X1 Subject-Derived iPSCs. Cell stem cell. https://doi.org/10.1016/j.stem.2015.02.005 (2015).
Hockemeyer, D. et al. Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat Biotechnol 27, 851–857, https://doi.org/10.1038/nbt.1562 (2009).
Hockemeyer, D. et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat Biotechnol 29, 731–734, https://doi.org/10.1038/nbt.1927 (2011).
Chen, S. et al. A large-scale in vivo analysis reveals that TALENs are significantly more mutagenic than ZFNs generated using context-dependent assembly. Nucleic Acids Res 41, 2769–2778, https://doi.org/10.1093/nar/gks1356 (2013).
Valton, J. et al. Overcoming transcription activator-like effector (TALE) DNA binding domain sensitivity to cytosine methylation. J Biol Chem 287, 38427–38432, https://doi.org/10.1074/jbc.C112.408864 (2012).
Dik, W. A. et al. New insights on human T cell development by quantitative T cell receptor gene rearrangement studies and gene expression profiling. The Journal of experimental medicine 201, 1715–1723, https://doi.org/10.1084/jem.20042524 (2005).
Kokron, C. M. et al. Searching for genes involved in the pathogenesis of primary immunodeficiency diseases: lessons from mouse knockouts. Journal of clinical immunology 17, 109–126 (1997).
Galic, Z. et al. T lineage differentiation from human embryonic stem cells. Proc Natl Acad Sci USA 103, 11742–11747, https://doi.org/10.1073/pnas.0604244103 (2006).
Galic, Z. et al. Generation of T lineage cells from human embryonic stem cells in a feeder free system. Stem Cells 27, 100–107, https://doi.org/10.1634/stemcells.2008-0813 (2009).
Acknowledgements
We thank Nadine Dannemann (deceased) and Shamim H. Rahman for experimental support, Karin Pike-Overzet, Sebastian Fuchs, and Luigi Naldini for scientific advice, Juan Carlos Zúñiga-Pflücker and Axel Schambach for materials, and Nils Craig-Müller for proof-reading of the manuscript. This work was supported by grants from the German Federal Ministry of Education and Research [ReGene-01GN1003D and BMBF-01EO0803 to T.C.], the European Commission’s 7th Framework Programme [PERSIST-222878 to A.J.T. and T.C., and Supersist-601958 to A.J.T.], The Wellcome Trust [104807/Z/14/Z to A.J.T.], and the Medical Research Council [studentship to C.P.].
Author information
Authors and Affiliations
Contributions
J.A. performed all genome editing and differentiation experiments. C.P. characterised the X-SCID ESC line. C.M. designed and generated the TALENs. T.C., S.J.H. and A.J.T. coordinated the project. J.A. and T.C. conceived the project, interpreted the data and wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
C.P. and S.J.H. are full-time employees of GlaxoSmithKline; T.C. is a consultant for TRACR Haematology. Research in T.C.’s lab is supported by Casebia Therapeutics, Cellectis and Miltenyi Biotec.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alzubi, J., Pallant, C., Mussolino, C. et al. Targeted genome editing restores T cell differentiation in a humanized X-SCID pluripotent stem cell disease model. Sci Rep 7, 12475 (2017). https://doi.org/10.1038/s41598-017-12750-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-12750-4
This article is cited by
-
CRISPR/Cas9-modified hematopoietic stem cells—present and future perspectives for stem cell transplantation
Bone Marrow Transplantation (2019)
-
Re-educating immunity in respiratory allergies: the potential for hematopoietic stem cell-mediated gene therapy
Journal of Molecular Medicine (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.