Abstract
Renal cell carcinoma (RCC) is one of the most lethal urologic cancers. About one-third of RCC patients already have distal metastasis at the time of diagnosis. There is growing evidence that Hox antisense intergenic RNA (HOTAIR) plays essential roles in metastasis in several types of cancers. However, the precise mechanism by which HOTAIR enhances malignancy remains unclear, especially in RCC. Here, we demonstrated that HOTAIR enhances RCC-cell migration by regulating the insulin growth factor-binding protein 2 (IGFBP2) expression. HOTAIR expression in tumors was significantly correlated with nuclear grade, lymph-node metastasis, and lung metastasis. High HOTAIR expression was associated with a poor prognosis in both our dataset and The Cancer Genome Atlas dataset. Migratory capacity was enhanced in RCC cell lines in a HOTAIR-dependent manner. HOTAIR overexpression accelerated tumorigenicity and lung metastasis in immunodeficient mice. Microarray analysis revealed that IGFBP2 expression was upregulated in HOTAIR-overexpressing cells compared with control cells. The enhanced migration activity of HOTAIR-overexpressing cells was attenuated by IGFBP2 knockdown. IGFBP2 and HOTAIR were co-expressed in clinical RCC samples. Our findings suggest that the HOTAIR-IGFBP2 axis plays critical roles in RCC metastasis and may serve as a novel therapeutic target for advanced RCC.
Similar content being viewed by others
Introduction
Renal cell carcinoma (RCC), which accounts for about 3% of all cancers in adults, is the most lethal of all urological malignancies1. One-third of RCC patients already have metastases at the time of diagnosis, and 20–30% of patients treated by radical nephrectomy will suffer metastasis or recurrence2. The prognosis of metastatic RCC is poor: the median survival is about 13 months3. Although recent developments in targeted therapy have improved survival rates for metastatic RCC, most patients still succumb to the disease. Therefore, new therapeutic approaches and prognostic factors are needed to treat advanced RCC.
Although numerous lncRNAs (non-coding RNAs longer than 200 nucleotides)4 have been identified as factors in cancer progression and the development and spread of metastases5, lncRNAs also regulate a wide variety of cell functions in normal tissue. Since many lncRNAs are differentially expressed in specific organs, tissues, or cancer types, lncRNAs are potential prognostic markers4.
Hox antisense intergenic RNA (HOTAIR), a lncRNA that acts as an oncogenic molecule in various types of cancer, is localized to the HOXC gene cluster. HOTAIR interacts with PRC2 (polycomb repressive complex 2) to enhance H3K27 trimethylation, and thereby decreases the expression of a large number of genes6. Several groups, including our laboratory, have reported that high HOTAIR expression is correlated with a poor prognosis in several types of cancer, including breast7, colorectal8, cervical9, non-small lung cell10, and gastric cancer11. However, the underlying mechanism by which HOTAIR is involved in malignancy remains uncertain. Many downstream molecules of HOTAIR have been identified: in breast cancer, HOTAIR increases cancer invasiveness and metastasis in a manner dependent on PRC27. In esophageal squamous cell carcinoma, HOTAIR decreases WIF-1 expression and activates the Wnt/β-catenin signaling pathway, thus promoting cell migration12. In cervical cancer, HOTAIR promotes tumor growth and invasion by targeting the Notch pathway13. However, there are few reports addressing HOTAIR’s molecular mechanism in RCC.
Insulin growth factor-binding protein 2 (IGFBP2) belongs to a family of six IGF-binding proteins, IGFBP1–6. These proteins bind to IGF1 and IGF2. The IGFBP2 expression is elevated in many cancer types, in both tumor cells and plasma14,15,16. Although conventionally known as the IGF regulatory protein, IGFBP was recently demonstrated to have pro-tumorigenic activity that is independent of IGF signaling in glioma cells: IGFBP2 contributes to cancer progression by enhancing MMP2 (matrix metalloprotease 2) gene transcription and, in turn, tumor-cell invasion17. IGFBP2 also binds integrin alpha 5 and activates pathways downstream of integrin, increasing cell motility18. Exogenous IGFBP2 promotes glioma-cell proliferation and invasion capability via the ERK pathway, which is activated by integrin β1 signaling19. However, it is not known how IGFBP2 is regulated in cancer cells, or whether IGFBP2 has oncogenic activity in RCC.
In this study, we analyzed correlations between HOTAIR expression and clinical characteristics in 64 RCCs. We clarified HOTAIR’s role in RCC and identified IGFBP2 as a molecule downstream of HOTAIR that is involved in RCC migratory capacity and prognosis.
Results
HOTAIR expression and clinicopathological characteristics in RCC
To evaluate correlations between HOTAIR expression and clinical characteristics, we examined the HOTAIR expression in 64 RCCs and their corresponding normal renal tissues using quantitative real-time PCR. We analyzed clinicopathological features such as age, gender, stage, T stage, N stage, M stage, nuclear grade, and vascular invasion, and measured the tumor HOTAIR expression relative to that in corresponding normal tissues. The cut-off point was determined according to the survival receiver operating characteristic (ROC) curve; tumors with HOTAIR levels at least 1.2-fold higher than that in the corresponding normal tissue were defined as high-expression, and those with HOTAIR levels below this threshold were defined as low-expression (Fig. 1A). We found that HOTAIR expression was associated with vascular invasion, nuclear grade, lymph-node metastasis, and distant metastasis (Table 1). Next, we analyzed the relationship between HOTAIR expression and patient prognosis using the Kaplan-Meier method. HOTAIR expression was significantly associated with a shorter survival in RCC patients (Fig. 1B). These results suggested that high HOTAIR expression indicates a poorer prognosis in RCC. To confirm this correlation, we searched TCGA and obtained the gene expression data and patient prognosis for 521 samples. Again, we found that high HOTAIR expression was significantly correlated with a poor prognosis (Fig. 1C). These data indicated that HOTAIR plays a critical role in RCC.
HOTAIR expression in RCCs (A) HOTAIR expression in 64 RCCs and their corresponding normal renal tissues was measured by quantitative real-time PCR. (B) Kaplan-Meier curves for cancer-specific survival according to HOTAIR expression in 64 RCC patients. (C) Kaplan-Meier curves for overall survival ratio according to HOTAIR expression in the RCC patient dataset obtained from TCGA (n = 521). Differences between groups were compared by log-rank test.
Enhanced RCC-cell migration by HOTAIR in vitro
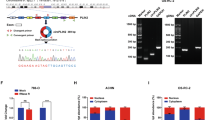
Since high HOTAIR expression was significantly associated with metastasis and prognosis in patients with RCC, we used two human RCC cell lines to generate cells that stably overexpressed HOTAIR (Fig. 2A,B). These HOTAIR-transduced cells overexpressed HOTAIR at levels 6-fold higher (ACHN-HOTAIR) and 30-fold higher (A498-HOTAIR) than did empty vector (EV)–transduced cells. In Caki-1 cells, which had higher endogenous HOTAIR expression than the ACHN or A498 RCC cell lines (Supplementary Fig. S1), knocking down HOTAIR by transfection with small interfering RNA (siRNA) decreased the HOTAIR expression by 55–66% (Fig. 2C).
HOTAIR enhanced the migration of RCC cells in vitro (A,B) HOTAIR expression in ACHN and A498 cells transduced with a HOTAIR-overexpressing (HOTAIR) or empty vector (EV), determined by real-time PCR. (C) HOTAIR expression levels in HOTAIR-targeted siRNA-transfected Caki-1 cells, determined by real-time PCR. (D,E, and F) Cell-proliferation CCK-8 assays in the indicated cell lines. (G,H, and I) Two-chamber assays using HOTAIR-overexpressing (G,H) or knockdown cells (I) in arbitrary units (a.u.). (J to L) Wound-healing assays. *P < 0.05, Mann-Whitney U test.
We assayed the effect of HOTAIR on cell proliferation using a cell-counting kit and found that the HOTAIR overexpression in ACHN and A498 cells did not alter their proliferation compared to control cells (Fig. 2D,E). HOTAIR siRNA knockdown did not affect cell proliferation in Caki-1 cells (Fig. 2F).
We evaluated HOTAIR’s role in migratory capability with a two-chamber assay and found that the migrated cells included significantly more HOTAIR-transduced than EV-transduced ACHN and A498 cells (Fig. 2G,H, and Supplementary Figure S2), and significantly fewer HOTAIR-knockdown than control Caki-1 cells (Fig. 2I). Similarly, a wound-healing assay showed that migration capacity was significantly enhanced by HOTAIR overexpression, but diminished by its knockdown (Fig. 2J–L). These data indicated that HOTAIR plays a critical role in the RCC cells’ migratory capacity.
Effect of HOTAIR overexpression on ACHN cells in vivo
To evaluate the malignant characteristics of HOTAIR-expressing cancer cells in vivo, we assayed their tumorigenicity using NOD/Shi-scid-IL2Rγnull (NOG) mice. When injected subcutaneously, HOTAIR-overexpressing cells grew faster than control cells (Fig. 3A). HOTAIR overexpression in cells introduced by tail-vein injection (Fig. 3B,C) or orthotopic xenograft (Fig. 3D,E) significantly increased the number of lung metastases. These data suggested that HOTAIR overexpression enhanced the colonization and metastasis of RCC cells in vivo.
In vivo effects of HOTAIR overexpression in ACHN cells (A) Tumorigenicity was assayed by injecting HOTAIR-overexpressing or control cells subcutaneously into NOG mice; graph shows average tumor volume and standard error. (B) HOTAIR-overexpressing or control cells injected into the tail vein of NOG mice; 24 weeks later, metastatic pulmonary nodules were counted microscopically. (C) Representative images of lung metastatic nodules resulting from tail-vein injections; the dotted circles indicate metastatic foci. (D) Mice received orthotopic xenografts of HOTAIR-overexpressing or control cells, and pulmonary nodules were counted microscopically 14 weeks later. (E) Representative images of lung metastatic nodules in the orthotopic xenograft assay; the dotted circles indicate metastatic foci*.P < 0.05, Mann–Whitney U test.
Induction of IGFBP2 expression by HOTAIR
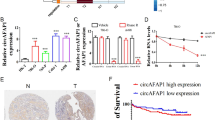
Since our results showed that HOTAIR regulates cancer invasiveness and metastasis both in vitro and in vivo, we identified invasiveness-related genes that were upregulated in our microarray data for HOTAIR-overexpressing cells and were reported to be potentially oncogenic. After a search of the literature for each of the 11 genes that were upregulated most strongly in HOTAIR-overexpressing cells, as identified by the rank product algorithm, we chose IGFBP2 for further analysis because it has been identified as a key factor in malignancy, especially in cancer invasiveness (Fig. 4A)17,18,19. Western blots and real-time PCR results confirmed that IGFBP2 was upregulated in the ACHN-HOTAIR cells (Fig. 4B,C). Together, these results suggested that IGFBP2 acts downstream of HOTAIR in RCC.
s migration in HOTAIR-expressing cells was attenuated by IGFBP2 knockdown (A) Heatmap showing the expression profiles of 11 genes in HOTAIR-overexpressing (HOTAIR1–3) and control (EV1–4) ACHN cells as determined by microarray analysis using the rank product algorithm. Red and green indicate higher and lower expression, respectively. (B, C) IGFBP2 expression was determined by real-time PCR (B) and western blotting (C) in HOTAIR-overexpressing ACHN cells (HOTAIR) and control cells (EV). (D) IGFBP2 expression was determined by real-time PCR in ACHN-EV cells transfected with control siRNA, ACHN-HOTAIR cells transfected with control siRNA, and ACHN-HOTAIR cells transfected with siRNA against HOTAIR. (E) IGFBP2 level in ACHN cells was measured by western blotting. (F) ACHN cells were transfected with the indicated siRNA, and cell proliferation was assayed 72 h later by CCK-8. (G, H) Migration measured by two-chamber assay (G) and wound-healing assay (H) in ACHN cells transfected with siRNA against IGFBP2 as indicated in (D); a. u., arbitrary units. *P < 0.05 by Mann-Whitney U test. Unprocessed original scans of blots are shown in Supplementary Figure S6.
Attenuation of the enhanced migration of HOTAIR-overexpressing cells by IGFBP2 knockdown
To test whether IGFBP2 regulates the migratory ability downstream of HOTAIR, we transfected ACHN-HOTAIR cells with siRNAs to knockdown the IGFBP2 gene, and confirmed that the IGFBP2 expression was significantly reduced in the transfected cells compared to control cells (Fig. 4D,E). Cell-proliferation assays revealed that 72 h after transfection, the proliferation was slightly reduced in the IGFBP2-knockdown cells compared to cells transfected with control siRNA (Fig. 4F). Cell migration was clearly reduced in IGFBP2-knockdown cells compared to cells transfected with control siRNA, in both the two-chamber (Fig. 4G) and scratch assay (Fig. 4H). We also tested HOTAIR-overexpressing A498 cells. As with ACHN cells, IGFBP2 was upregulated by HOTAIR overexpression in A498 cells, and the ability of these cells to migrate was dependent on IGFBP2 (Supplementary Figure S3, A to C). These results suggested that the migration of HOTAIR-overexpressing cells was mainly dependent on IGFBP2.
Correlation of HOTAIR expression with IGFBP2 in RCC tissues
To determine whether HOTAIR expression was correlated with IGFBP2 expression in human RCC tissues, we extracted the total RNA from resected RCC tissues and measured the IGFBP2 and HOTAIR expression by real-time PCR. We found that the IGFBP2 expression was positively correlated with the HOTAIR expression in RCC tissues (Table 2, Fisher’s exact test).
We used in situ hybridization (ISH) and immunohistochemistry (IHC) to confirm the HOTAIR and IGFBP2 co-expression in RCC tissues. Figure 5 shows representative serial sections that underwent hematoxylin and eosin (HE) staining, ISH for HOTAIR, and IHC for IGFBP2. The section contained a typical clear-cell RCC with a low nuclear grade, and another with sarcomatoid differentiation and a high nuclear grade (Fig. 5A). In the low-grade tumor area, ISH and IHC detected only slight levels of HOTAIR and IGFBP2 (Fig. 5D,E,I,J). Higher levels of HOTAIR and IGFBP2 were detected in the high-grade tumor area (Fig. 5G–J).
HOTAIR and IGFBP2 colocalization in high-grade tumor areas (A) Whole-section, low-magnification view of an HE-stained renal-cell carcinoma containing both low-grade and high-grade tumors. (B) Immunohistochemistry (IHC) for IGFBP2, at low magnification. (C to E) Serial sections of the low-grade tumor area at high magnification, showing HE staining (C), ISH for HOTAIR (D), and IHC for IGFBP2 (E). (F to H) Serial sections of the high-grade tumor area at high magnification, showing HE staining (F), ISH for HOTAIR (G), and IHC for IGFBP2 (H). The black arrowheads indicate dot-like positive staining of HOTAIR. (I and J) Quantification of areas positive for HOTAIR (ISH, red) and IGFBP2 (IHC, brown) using ImageJ software. Total eight areas were randomly selected from low- and high-grade tumor areas. *P < 0.001.
IGFBP2 is a prognostic factor in RCC
Finally, we investigated whether IGFBP2 was correlated with RCC prognosis. In our dataset, high IGFBP2 expression determined by real-time PCR was significantly associated with a poor prognosis (Fig. 6A). To confirm this correlation, we searched TCGA and obtained the gene expression data and patient prognosis for 521 samples. Again, we found that high IGFBP2 expression was significantly correlated with a poor prognosis (Fig. 6B). Interestingly, the group with high levels of both HOTAIR and IGFBP2 had the poorest prognosis (Supplementary Fig. S4). The prognosis for the HOTAIRlow-IGFBP2high group was almost the same as that for the HOTAIRlow-IGFBP2low group. (Fig. 6C). These data indicated that the regulation of IGFBP2 by HOTAIR is critical in RCC development.
IGFBP2 is prognostic for RCC (A) Kaplan-Meier analysis for cancer-specific survival according to the IGFBP2 expression of in 64 RCC patients. The cut-off point was the median value in the current sample. (B) Kaplan-Meier curves for overall survival ratio according to IGFBP2 expression in the RCC patient dataset obtained from TCGA (n = 521). (C) Kaplan-Meier analysis for the cancer-specific survival rate of the four groups; differences between the groups were assessed by log-rank tests.
Discussion
In this study, we demonstrated that HOTAIR is involved in RCC. HOTAIR-overexpressing cells grew faster in an in vivo tumorigenicity assay but grew at a similar rate to control cells in vitro, suggesting that HOTAIR is involved in colonization capacity. Tail-vein injection and orthotopic xenograft assays indicated that HOTAIR induced colonization capacity in the lung. HOTAIR was associated with distant and lymph-node metastases in RCC patients. Collectively, these data suggested that HOTAIR plays critical roles in RCC’s ability to invade and colonize. A previous study showed that HOTAIR knockdown in RCC cell lines decreased their migration in vitro 20, which is compatible with our present study.
We also demonstrated that the presence of both HOTAIR and IGFBP2 in RCC is a prognostic marker in RCC, suggesting that the HOTAIR-IGFBP2 axis may represent a specific therapeutic target for RCC. Because the HOTAIRlow-IGFBP2high group had a better prognosis than the HOTAIRhigh-IGFBP2high group, we speculate that HOTAIR-dependent IGFBP2 expression is important for the progression of RCC. IGFBP2 overexpression is well documented in tumors, including glioma21, breast cancer22,23, ovarian cancer24, prostate cancer25, colorectal cancer26, gastric cancer27, lung cancer28, leukemia29, and astrocytoma30. Recent studies implicate elevated IGFBP2 in a shorter overall survival time31,32,33. In glioma cells, IGFBP2’s interaction with integrin alpha 5 is essential for cell mobility18, and the IGFBP2/integrin/ILK/NF-κB signaling pathway drives cancer progression in vivo 34. Although the precise molecular mechanisms of cell invasion in RCC are unknown, IGFBP2 may be critical for this progression.
Several reports suggest that HOTAIR affects the expression of numerous genes involved in various cellular functions, especially those related to epithelial-mesenchymal transition (EMT)7. HOTAIR acts in esophageal squamous cell carcinoma by activating the Wnt/β-catenin signaling pathway12. In gastric cancer, inhibiting HOTAIR reverses the EMT process and reduces invasiveness mediated by MMP1 and MMP335. In breast cancer, TGFβ1 treatment induces EMT via HOTAIR expression36. In the present study, we found that HOTAIR expression did not affect EMT-related genes (Supplementary Fig. S5). On the other hand, several reports indicate HOTAIR’s involvement in pathways unrelated to EMT. Suppressing HOTAIR upregulates the HOXA5 level and downregulates MMP2 and MMP9, which are critical players in tumor invasion and metastasis37. A recent report demonstrated that HOTAIR’s targets, including EMT-related genes, strongly depend on the type of cancer cell in urothelial cancer38. Collectively, HOTAIR primarily affects cell migration in RCC not via EMT signaling, but rather through an IGFBP2-related pathway, although it is not clear how HOTAIR regulates IGFBP2 expression.
Here, we newly identified IGFBP2 as a downstream molecule of HOTAIR, which is involved in the migratory capability of RCC cells. Our findings suggest that the HOTAIR-IGFBP2 axis plays critical roles in the progression of RCC, and presents a novel therapeutic target for advanced RCC treatment.
Materials and Methods
Ethics statement
This study was conducted according to the principles expressed in the Declaration of Helsinki and was approved by the Ethics Committees at the Miyagi Cancer Center Research Institute (Natori, Japan). Experimental protocols involving animals were approved by the Miyagi Cancer Center Animal Care and Use Committee.
Clinical samples
Sixty-four RCC samples and their corresponding normal renal tissues were obtained from patients who underwent nephrectomy or partial nephrectomy at the Miyagi Cancer Center between 2007 and 2015. All specimens ware immediately frozen and stored at −80 °C until RNA extraction. The Institutional Review Board of the Miyagi Cancer Center approved this study protocol, and each patient provided informed, written consent. No patient received any neoadjuvant therapy, such as molecularly targeted therapy, before surgery. All tumor tissues were diagnosed pathologically as clear cell RCC. The significance of differences between two groups was estimated by Fisher’s exact test.
Validation with TCGA data
Clinical and pathological data were manually extracted from reports downloaded from the TCGA Data Portal (https://gdc-portal.nci.nih.gov/); gene expression profiles and clinical data from 521 patients with clear cell renal carcinoma were available from the website. Tumors with HOTAIR expression of at least 0.3 FPKM (fragments per kilobase of exon per million fragments mapped) were classified as high HOTAIR, and those with less than 0.3 FPKM as low HOTAIR. Tumors with IGFBP2 expression of at least 6 FPKM were classified as high IGFBP2, and those with less than 6 FPKM as low IGFBP2. The Kaplan-Meier method for gene expression-specific survival analysis was performed using package “survival” in statistical computing language R, and the statistical significance of survival curves between different expression levels (high- and low-) was evaluated by log-rank test.
RNA preparation, cDNA synthesis, and quantitative real-time RT-PCR
Total RNA was extracted from frozen samples and RCC cell lines by ISOGENE (Nippon Gene, Tokyo, Japan) according to the manufacturer’s protocol. For all samples, cDNAs were synthesized from 1.0 μg of total RNA with the PrimeScript 1st Strand cDNA Synthesis Kit (TaKaRa Bio, Shiga, Japan) following the manufacturer’s protocol.
HOTAIR expression was quantified as previously described39. The primer sequences used: for GAPDH, F 5′-TGAAGGTCGGAGTCAACGG-3′ and R 5′-AGAGTTAAAAGCAGCCCTGGTG-3′; for HOTAIR, F 5′-GGTAGAAAAAGCAACCACGAAGC-3′ and R 5′-ACATAAACCTCTGTCTGTGAGTGCC-3′; and for IGFBP2, F 5′-AGCCCAAGAAGCTGCGACCAC-3′ and R 5′-CTGCCCGTTCAGAGACATCTTGC-3′. HOTAIR expression was normalized to GAPDH expression in each sample. The HOTAIR expression in each RCC was determined relative to that in the corresponding normal renal tissue.
Cell lines and cell culture
Three clear-cell RCC cell lines (ACHN, A498, and Caki-1) were obtained from the Japanese Cancer Research Resources Bank, Tokyo, Japan (JCRB). The cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco/Life technologies, Carlsbad, USA) with 10% FBS and 1% penicillin-streptomycin. The cells were incubated in a humidified incubator at 37 °C and 5% CO2.
Retroviral transfection
Human HOTAIR cDNA (obtained from Addgene, Cambridge, MA, USA) was amplified by PCR and inserted into the pBabe-hygro vector (pBabe-HOTAIR). The recombinant retrovirus was produced with the Platinum-A packaging-cell line (Plat-A, kindly provided by Prof. Kitamura, Tokyo University) as described previously40. Briefly, Plat-A cells were transfected with pBabe-HOTAIR or a pBabe-hygro empty vector (EV). Fugene 6 (Roche Applied Science) and Opti-MEM I (Gibco/Life Technologies Co.) were added following the manufacturer’s protocol. The retrovirus-containing supernatant was collected 48 h after transfection and passed through a 0.45-μm filter. ACHN cells were infected with the recombinant retroviruses and then selected with hygromycin.
siRNA transfection
The HOTAIR siRNAs #1 (FLJ41747HSSS160170), #2 (FLJ41747HSSS160171), the IGFBP2 siRNAs #1 (HSS142627) and #2 (HSS142629), and non-silencing control siRNA (12935–300) were purchased from Invitrogen (Carlsbad, CA, USA). The siRNA transfections were performed using Lipofectamine RNAiMAX Reagent (Life Technologies, Carlsbad, CA, USA) in antibiotic-free medium for 48 h. The efficiency of siRNA knockdown was confirmed by real-time PCR (HOTAIR and IGFBP2) and western blotting (IGFBP2).
Cell-proliferation assay
Cell proliferation was detected with the Cell Counting Kit-8 (CCK-8) from Dojindo Laboratories (Kumamoato, Japan) according to the manufacturer’s protocol. Cells were seeded in 96-well plates (1 × 103 cells per well) in normal cell-growth medium. The relative numbers of viable cells at 0, 24, 48, 72, and 96 h of incubation were assessed according to the absorbance of optical density at 450 nm detected by microplate reader (VersaMax, Molecular Devices, Sunnyvale, CA, USA).
Two-chamber assay
Cell migration was assessed using a two-chamber assay with a cell-culture insert (8-μm pore size, BD Biosciences, San Jose, CA, USA) in 24-well plates. Cells (5 × 104) were plated in each insert in serum-free medium. The bottom well contained medium with 10% FBS. After 48 h, the bottom of the insert was stained with Diff-Quick (Dade Behring Inc, Newark, DE, USA), and the cells that had invaded through the membrane to the lower surface were counted.
Wound-healing assay
Cells were seeded in 24-well plates in normal cell-growth medium, incubated until confluent, and treated with 5 μg/ml mitomycin C (Wako, Osaka, Japan) for 3 h. A yellow pipette tip was used to make a straight scratch, simulating a wound. The medium was changed to serum-free medium. After a 24 h incubation, the area occupied by cells that had migrated into the scratch area was measured using ImageJ software.
Western blotting
Western blotting was performed as previously described with minor modifications39. In brief, whole-cell lysates were prepared in RIPA buffer supplemented with a proteinase-inhibitor cocktail. The lysates were clarified by centrifugation, the supernatant was boiled, and separated by SDS-PAGE. The following antibodies were used for western blots: anti-α-tubulin (Santa Cruz Biotechnology, Santa Cruz, CA, USA, B-5–1–2) and anti-IGFBP2 (Santa Cruz Biotechnology, Santa Cruz, CA, USA, C-18).
Animals
Six-week-old female NOG mice were purchased from the Central Institute for Experimental Animals (CIEA, Kawasaki, Japan). The mice were housed in a cleanroom at the animal care facility at Miyagi Cancer Research Center with standard temperature, humidity, and timed lighting, and given mouse chow/water ad libitum.
In vivo proliferation assay by subcutaneous injection
NOG mice were given subcutaneous injections of 1.0 × 104 ACHN cells transfected with a HOTAIR-overexpression (n = 8) or empty vector (n = 8) with 0.2 ml PBS. Tumors were measured weekly with a digital caliper, and tumor volume was calculated using the modified ellipsoid formula 1/2 (length × width2). The animals were sacrificed 8 weeks after the injection.
In vivo metastasis assays by orthotopic xenograft and tail-vein injection
For in vivo experimental metastasis assays, 1 × 104 ACHN-HOTAIR (n = 4) or ACHN-EV (n = 4) cells were injected into the tail vein of NOG mice. The mice were sacrificed 24 weeks later, and pulmonary nodules were assessed macroscopically and microscopically. The harvested organs were processed into paraffin-embedded sections, stained with HE, and examined pathologically to confirm the presence of metastatic lung tumors. In orthotopic xenograft implants, 1 × 105 ACHN-HOTAIR (n = 4) or ACHN-EV (n = 3) cells in 20 μl matrigel were injected into the subcapsular area of the left kidney of the NOG mice. The mice were sacrificed 14 weeks later and the lungs were collected and analyzed. Pulmonary nodules were counted under a microscope.
Microarray analysis
A microarray analysis was performed to search for genes regulated by HOTAIR. We used the SurePrint G3 Human GE 8 × 60 K Microarray (Agilent Technologies, Santa Clara, CA, USA) to profile the gene expression in control and HOTAIR-overexpressing ACHN cells. The obtained gene expression data were expressed in logarithmic scale, and heat maps were generated using the rank product algorithm41. The analyses were performed using R software. The dataset was uploaded to the Gene Expression Omnibus (GEO) database (GSE100107).
In situ hybridization by RNAscope
In situ hybridization for HOTAIR was performed using the RNAscope 2.5 HD assay kit according to the manufacturer’s protocol (Advanced Cell Diagnostics, Hayward, CA, USA)42. In brief, formalin-fixed, paraffin-embedded 7-μm tissue sections were deparaffinized and pretreated with protease for hybridization with HOTAIR target mixture probes. After hybridization, a series of signal-amplification steps were hybridized to the target probes followed by color development with Fast Red. Positive staining was identified as red punctate dots in the cell. Negative control probes (DapB) and positive control probes (PPIB) were also tested for each slide to check the quality of hybridization.
Immunohistochemistry
For IGFBP2 immunostaining, paraffin-embedded, formalin-fixed 3-μm tissue sections from human RCC patients were deparaffinized in xylene and rehydrated by washing with a series of ethanol dilutions and distilled water. Heat-induced epitope retrieval was performed by microwaving the sections in a pH-9.0 target-retrieval solution (Dako, Produktionsvej, Denmark). Endogenous peroxidases were blocked with 0.3% H2O2. The sections were incubated with anti-human IGFBP2 for 12 h (1:1,000) at 37 °C, then with mouse LINKER (Dako) for 15 min, followed by incubation with the secondary antibody and development with 3,3′-diaminobenzidine (DAB) chromogen (EnVision Detection Systems Peroxidase/DAB, Rabbit/Mouse, Dako).
Statistical analysis
All data from both in vitro and in vivo analyses were analyzed for statistical significance by Mann-Whitney U test, and P < 0.05 was considered significant. All statistical analyses were performed using JMP 11 (SAS Institute, Cary, NC, USA) and GraphPad Prism software (Version 6.05; GraphPad Software Inc., La Jolla, CA, USA).
References
Rini, B. I. & Bernard Escudier, S. C. C. Renal cell carcinoma. lancet 373, 1119–1132 (2009).
Motzer, R. J., Bander, N. H. & Nanus, D. M. Renal-cell carcinoma. N Engl J Med 335, 865–875 (1996).
Cohen, H. T. & McGovern, F. J. Renal-cell carcinoma. N Engl J Med 353, 2477–2490 (2005).
Yarmishyn, A. A. & Kurochkin, I. V. Long noncoding RNAs: a potential novel class of cancer biomarkers. Front Genet 6, 145 (2015).
Gibb, E. A., Brown, C. J. & Lam, W. L. The functional role of long non-coding RNA in human carcinomas. Mol Cancer 10, 38 (2011).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Gupta, R. A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Kogo, R. et al. Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res 71, 6320–6326 (2011).
Kim, H. J. et al. Long non-coding RNA HOTAIR is associated with human cervical cancer progression. International journal of oncology 46, 521–530 (2015).
Nakagawa, T. et al. Large noncoding RNA HOTAIR enhances aggressive biological behavior and is associated with short disease-free survival in human non-small cell lung cancer. Biochemical and biophysical research communications 436, 319–324 (2013).
Endo, H. et al. Enhanced expression of long non-coding RNA HOTAIR is associated with the development of gastric cancer. PloS one 8, e77070 (2013).
Ge, X. S. et al. HOTAIR, a prognostic factor in esophageal squamous cell carcinoma, inhibits WIF-1 expression and activates Wnt pathway. Cancer science 104, 1675–1682 (2013).
Lee, M. et al. The long non-coding RNA HOTAIR increases tumour growth and invasion in cervical cancer by targeting the Notch pathway. Oncotarget 7, 44558–44571 (2016).
Myers, A. L. et al. IGFBP2 modulates the chemoresistant phenotype in esophageal adenocarcinoma. Oncotarget 6, 25897–25916 (2015).
Hawsawi, Y. et al. Deregulation of IGF-binding proteins -2 and -5 contributes to the development of endocrine resistant breast cancer in vitro. Oncotarget, 32129-32143 (2016).
Gao, S. et al. IGFBP2 Activates the NF-kappaB Pathway to Drive Epithelial-Mesenchymal Transition and Invasive Character in Pancreatic Ductal Adenocarcinoma. Cancer Res, 6543-6554 (2016).
Wang, H. et al. Insulin-like growth factor binding protein 2 enhances glioblastoma invasion by activating invasion-enhancing genes. Cancer Res 63, 4315–4321 (2003).
Wang, G. K., Hu, L., Fuller, G. N. & Zhang, W. An interaction between insulin-like growth factor-binding protein 2 (IGFBP2) and integrin alpha5 is essential for IGFBP2-induced cell mobility. The Journal of biological chemistry 281, 14085-14091 (2006).
Han, S., Li, Z., Master, L. M., Master, Z. W. & Wu, A. Exogenous IGFBP-2 promotes proliferation, invasion, and chemoresistance to temozolomide in glioma cells via the integrin beta1-ERK pathway. Br J Cancer 111, 1400–1409 (2014).
Chiyomaru, T. et al. Long non-coding RNA HOTAIR is targeted and regulated by miR-141 in human cancer cells. The Journal of biological chemistry 289, 12550–12565 (2014).
Zheng, S. et al. DNA hypermethylation profiles associated with glioma subtypes and EZH2 and IGFBP2 mRNA expression. Neuro-oncology 13, 280–289 (2011).
Busund, L. T. et al. Significant expression of IGFBP2 in breast cancer compared with benign lesions. Journal of clinical pathology 58, 361–366 (2005).
Probst-Hensch, N. M. et al. IGFBP2 and IGFBP3 protein expressions in human breast cancer: association with hormonal factors and obesity. Clinical cancer research: an official journal of the American Association for Cancer Research 16, 1025–1032 (2010).
Lancaster, J. M. et al. High expression of insulin-like growth factor binding protein-2 messenger RNA in epithelial ovarian cancers produces elevated preoperative serum levels. International journal of gynecological cancer: official journal of the International Gynecological Cancer Society 16, 1529–1535 (2006).
Ambrosini-Spaltro, A., Farnedi, A., Montironi, R. & Foschini, M. P. IGFBP2 as an immunohistochemical marker for prostatic adenocarcinoma. Applied immunohistochemistry & molecular morphology: AIMM 19, 318–328 (2011).
Mishra, L., Bass, B., Ooi, B. S., Sidawy, A. & Korman, L. Role of insulin-like growth factor-I (IGF-I) receptor, IGF-I, and IGF binding protein-2 in human colorectal cancers. Growth hormone & IGF research: official journal of the Growth Hormone Research Society and the International IGF Research Society 8, 473–479 (1998).
Zhang, L. et al. Expression of IGFBP2 in gastric carcinoma and relationship with clinicopathologic parameters and cell proliferation. Digestive diseases and sciences 52, 248–253 (2007).
He, Y. et al. Aberrant expression of proteins involved in signal transduction and DNA repair pathways in lung cancer and their association with clinical parameters. PloS one 7, e31087 (2012).
Kuhnl, A. et al. High expression of IGFBP2 is associated with chemoresistance in adult acute myeloid leukemia. Leukemia research 35, 1585–1590 (2011).
Becher, O. J., Peterson, K. M., Khatua, S., Santi, M. R. & MacDonald, T. J. IGFBP2 is overexpressed by pediatric malignant astrocytomas and induces the repair enzyme DNA-PK. Journal of child neurology 23, 1205–1213 (2008).
Liou, J. M. et al. Plasma insulin-like growth factor-binding protein-2 levels as diagnostic and prognostic biomarker of colorectal cancer. The Journal of clinical endocrinology and metabolism 95, 1717–1725 (2010).
Inman, B. A. et al. Insulin-like growth factor binding protein 2: an androgen-dependent predictor of prostate cancer survival. European urology 47, 695–702 (2005).
Guo, C. et al. Insulin-like growth factor binding protein-2 level is increased in blood of lung cancer patients and associated with poor survival. PloS one 8, e74973 (2013).
Holmes, K. M. et al. Insulin-like growth factor-binding protein 2-driven glioma progression is prevented by blocking a clinically significant integrin, integrin-linked kinase, and NF-kappaB network. Proceedings of the National Academy of Sciences of the United States of America 109, 3475–3480 (2012).
Xu, Z. Y. et al. Knockdown of long non-coding RNA HOTAIR suppresses tumor invasion and reverses epithelial-mesenchymal transition in gastric cancer. International journal of biological sciences 9, 587–597 (2013).
Padua Alves, C. et al. Brief report: The lincRNA Hotair is required for epithelial-to-mesenchymal transition and stemness maintenance of cancer cell lines. Stem cells (Dayton, Ohio) 31, 2827–2832 (2013).
Liu, X. H. et al. The long non-coding RNA HOTAIR indicates a poor prognosis and promotes metastasis in non-small cell lung cancer. BMC cancer 13, 464 (2013).
Heubach, J. et al. The long noncoding RNA HOTAIR has tissue and cell type-dependent effects on HOX gene expression and phenotype of urothelial cancer cells. Mol Cancer 14, 108 (2015).
Mochizuki, M. et al. CD271 regulates the proliferation and motility of hypopharyngeal cancer cells. Scientific reports 6, 30707 (2016).
Kitamura, T. et al. Retrovirus-mediated gene transfer and expression cloning: powerful tools in functional genomics. Experimental hematology 31, 1007–1014 (2003).
Breitling, R., Armengaud, P., Amtmann, A. & Herzyk, P. Rank products: a simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS letters 573, 83–92 (2004).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. The Journal of molecular diagnostics: JMD 14, 22–29 (2012).
Acknowledgements
This work was supported by JSPS KAKENHI Grant Numbers 15K09055 and 16K07132, Japan Agency for Medical Research and Development, and Novartis Pharma Research Grants. This work was partly supported by the Biomedical Research Core of Tohoku University School of Medicine.
Author information
Authors and Affiliations
Contributions
H.K., K.T., and K.S. designed the study, interpreted the results, analyzed the data and wrote and edited the paper. S.K., T.T., I.S. and Y.A. provided the surgical specimens and provided the histopathologic findings. H.K. and R.S. performed in vivo animal experiments. H.K., M.N, M.M., and K.Y. performed in vitro experiments. H.K., K.S, T.O., and W.F. analyzed and calculated TCGA dataset. All authors are aware of the content in this paper and have read and edited the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Katayama, H., Tamai, K., Shibuya, R. et al. Long non-coding RNA HOTAIR promotes cell migration by upregulating insulin growth factor–binding protein 2 in renal cell carcinoma. Sci Rep 7, 12016 (2017). https://doi.org/10.1038/s41598-017-12191-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-12191-z
This article is cited by
-
Hybridisation chain reaction-based visualisation and screening for lncRNA profiles in clear-cell renal-cell carcinoma
British Journal of Cancer (2022)
-
Noncoding RNA actions through IGFs and IGF binding proteins in cancer
Oncogene (2022)
-
Long noncoding RNAs as tumorigenic factors and therapeutic targets for renal cell carcinoma
Cancer Cell International (2021)
-
Long non-coding RNAs in genitourinary malignancies: a whole new world
Nature Reviews Urology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.