Abstract
Gibberellins (GAs) participate in controlling various aspects of basic plant growth responses. With the exception of bryophytes, GA signalling in land plants, such as lycophytes, ferns and angiosperms, is mediated via GIBBERELLIN-INSENSITIVE DWARF1 (GID1) and DELLA proteins. To explore whether this GID1-DELLA mechanism is present in pines, we cloned an orthologue (PtGID1) of Arabidopsis AtGID1a and two putative DELLA proteins (PtDPL; PtRGA) from Pinus tabuliformis, a widespread indigenous conifer species in China, and studied their recombinant proteins. PtGID1 shares with AtGID1a the conserved HSL motifs for GA binding and an N-terminal feature that are essential for interaction with DELLA proteins. Indeed, A. thaliana 35S:PtGID1 overexpressors showed a strong GA-hypersensitive phenotype compared to the wild type. Interactions between PtGID1 and PtDELLAs, but also interactions between the conifer-angiosperm counterparts (i.e. between AtGID1 and PtDELLAs and between PtGID1 and AtDELLA), were detected in vivo. This demonstrates that pine has functional GID1-DELLA components. The Δ17-domains within PtDPL and PtRGA were identified as potential interaction sites within PtDELLAs. Our results show that PtGID1 has the ability to interact with DELLA and functions as a GA receptor. Thus, a GA-GID1-DELLA signalling module also operates in evolutionarily ancient conifers.
Similar content being viewed by others
Introduction
Gibberellins (GAs) are a class of phytohormones that function in a wide range of basic plant growth responses. These GA-mediated responses include seed germination, stem elongation, shade avoidance (in competitive interactions), leaf expansion, pollen maturation, and induction of flowering1,2,3,4, but GAs can also induce temporary growth arrest under adverse environmental conditions5. Regarding our understanding of the GA signalling pathways in planta, progress has been made in the past few years primarily in rice6 and Arabidopsis thaliana 7.
Angiosperms use the GA-GID1 (GIBBERELLIN-INSENSITIVE DWARF1)-DELLA pathway, which involves the nuclear GA receptor GID18, the repressor DELLA protein9,10, and the F-box protein GID2/SLY1, which degrades the repressor DELLA protein to trigger GA-mediated downstream responses5,11,12. Although it has not yet been thoroughly studied, GA signalling in conifers should also follow the GID1-DELLA pathway; this molecule is present in early vascular plants such as the lycophyte Selaginella moellendorffii 13, but not in non-vascular bryophytes14. Hence, GID1-mediated GA signalling likely appeared after the divergence of vascular plants from mosses, which took place ~430 million years ago15.
Arabidopsis thaliana possesses at least 10 similar DNA sequences for GID1, of which only three (AtGID1a, AtGID1b, and AtGID1c) encode proteins that function as GA receptors16; conversely, there is only one GID1 gene in rice (OsGID1). AtGID1b is unique in that it can also interact with DELLAs in the absence of GA17, indicating that there are GA-independent pathways for this interaction. However, GID1 overexpression leads to a GA overdose phenotype, where plants are tall with long, light green leaves, fewer tillers, and dramatically reduced fertilities8. Leaf expansion, and stem and root elongation are reduced in gid1-knockout mutants, which is consistent with the GA-deficient phenotype18.
At the molecular level, GID1 displays an alpha/beta-hydrolase fold characteristic of hormone-sensitive lipases (HSLs), and its GA-binding pocket also corresponds to the substrate-binding site of HSLs; however, the N-terminal lid is specific to GID1 proteins19. The primary function of this movable N-terminal lid is to stabilise GA within the GID1 binding site20. The N-terminal lid also participates in GA-dependent interactions with DELLA proteins20, leading to the formation of GA-GID1-DELLA protein complexes5.
Rice contains only a single DELLA protein, SLR1, whose causative recessive mutation is responsible for the ‘slender rice’ constitutive GA response phenotype21. In contrast, A. thaliana has five DELLA family members with partly overlapping functions: GA-INSENSITIVE (GAI), REPRESSOR OF ga1-3 (RGA), and three RGA-like proteins (RGL1, RGL2, and RGL3)22. A 17-amino-acid deletion(Δ17), DELLAVLGYKVRSSEMA, within the DELLA region turns proteins into constitutive repressors of GA signalling, wherein DELLA fails to interact with GID1 in the presence of GA, conferring a GA-insensitive dwarf phenotype9,23.
The typical molecular features of the GA-signalling regulatory DELLA proteins include the conserved N-terminal DELLA and TVHYNP motifs, which are required for GA binding and for inactivation of DELLA proteins6, and the conserved C-terminal GRAS domain, which interacts with GID119,22.
Recent findings suggest that DELLA controls the GA signalling pathway through antagonistic regulation of the GA-positive regulator SCARECROW-LIKE 3 (SCL3) promoter sequence via co-regulatory intermediary proteins24. DELLAs also have a more general function in adapting plant growth to environmental conditions. For example, salt-activated signalling pathways (via abscisic acid and ethylene) enhance the growth-repressing effects of DELLAs25.
The objective of the present study was to investigate whether the GA-GID1-DELLA module is also present in gymnosperms. Our study species was Pinus tabuliformis, an economically and ecologically important indigenous hard pine of Northern China26. Characteristic of conifers, P. tabuliformis has a long growing cycle, a huge genome (Mean DNA content = 25.7 ± 0.13 Gb)27, and a large evolutionary separation from angiosperms28. Our primary long-term research focus is the characterisation of the regulatory program underlying pine cone development. We have previously studied the patterns of expression for GA metabolism genes and found that GA plays different roles in the early and late stages of cone development26. We have also isolated and identified the MADS-box genes and their potential regulators related to reproductive development on a genome-wide basis28. Furthermore, we have systematically isolated the pine homologues of the functional genes from the miRNA pathway involved in cone development29. In these previous studies, we discovered that the GA biosynthesis pathway has diverged remarkably between conifers and angiosperms26. To determine which components of the GA signalling pathway are conserved between conifers and angiosperms, we assessed the potential involvement of a conifer GA-GID1-DELLA module for cone development in Pinus tabuliformis.
Results
The P. tabuliformis orthologue of AtGID1a and DELLA proteins
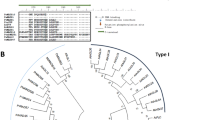
Based on P. tabuliformis transcriptome data (SRA 056887) and a large collection of high-quality ESTs that was obtained, assembled de novo and characterised30, we screened and cloned five GID1 homologue sequences: PtGID1, PtGIDI-L1, PtGIDI-L2, PtGIDI-L3, and PtBSU1. Phylogenetic reconstruction revealed that only PtGID1 was the true orthologue of AtGID1a and OsGID1 (Figure S1). The PtGID1 gene encodes a 357-amino-acid-long polypeptide, and has the same gene structure as the A. thaliana and rice genes, containing one intron and two exons. PtGID1 shares homology with the consensus sequence of the HSL family, including the conserved HSL motifs HGG and GXSXG. Multiple sequence alignment revealed that 16 of the 17 essential sites in PtGID1 are identical to those found within OsGID1, AtGID1a, and AtGID1c, while valine (V) at position C326 (following OsGID1’s amino acid sequence as the reference sequence in the alignment) is replaced by isoleucine (I) (Fig. 1, Δ). Nevertheless, both amino acids have similar hydrophobic properties. The results of comparative sequence analysis with functional GID1 from angiosperms suggested that the isolated PtGID1 gene might indeed encode the functional GA receptor for GA-dependent signalling pathways in pine. Homology modelling for PtGID1 further showed that the PtGID1 core domain is similar to that of OsGID1 based on crystal structure-derived information for the rice orthologue (Figure S2). The GA binding site of PtGID1 has one additional α-helix compared to OsGID1. However, this N-terminal extension (N-Ex) is the same as in AtGID1a20, demonstrating that the structure of the GA-binding sites of PtGID1, OsGID1, and the AtGID1s are highly conserved during evolution. These findings indicate that the PtGID1 gene might encode fully functional GA signal transduction capability.
Multiple sequence alignment of protein sequences encoded by PtGID1-like genes and A. thaliana and O. sativa GID1 genes. Black and gray boxes indicate identical or similar residues. The 17 arrows at the top indicate the residues essential for the gibberellic acid binding activity of GID1 in angiosperms. The two asterisks at the bottom indicate sites that are not identical for PtGID1-like genes within the conserved sequence region found in angiosperms. The triangle indicates similar residues in AtGID1b, AtGID1c, and PtGID1, and in OsGID1 and AtGID1a. As in rice and Arabidopsis, three conserved amino acids (S, D, and H) shape the catalytic triad in the HSL family (Nakajima et al.16; Ueguchi-Tanaka et al.8). Two of them (S and D) are conserved in PtGID1, while the third (H) is replaced by V or I. There are 13 functional domains (TWVLIS, LDR, FFHGGSF, HS, IYD, YRR, DGW, GDSSGGNI, GNI, MF, LDGKYF, WYW, and GFY) in GID1 (Hirano et al.14). Within TWVLIS, W is replaced by F, while all other domains are conserved. We confirmed the presence of all amino acid residues essential for GA binding encoded within the cloned PtGID1 gene.
We identified two candidates from high-quality P. tabuliformis ESTs30 and designated the encoded proteins as PtDPL and PtRGA based on their specific DELLA and GRAS domains, which were indicative of DELLA proteins. Phylogenetic reconstruction revealed a greater evolutionary distance between PtRGA and angiosperms than between PtRGA and the spike moss Selaginella (Fig. 2A). However, it has already been shown that the spike moss protein (SmDELLA1) is capable of interacting with its GID2 protein as a component in the GID-dependent GA signalling pathway. Functionally characterised angiosperm DELLA proteins have a conserved domain that is 27 amino acids in length, starting with the aspartic acid, glutamic acid, leucine, leucine, and alanine (“DELLA”) sequence that lends the protein family its specific name (Fig. 2B). This sequence can then interact with the GID1 protein N-Ex sequence. In the absence of the Δ17-domain (Fig. 2B), GID1 is unable to recognise DELLA, leading to the interruption of GA signal transmission31. In P. tabuliformis, seven amino acid sites within the DELLA Δ17-domains of both PtDPL and PtRGA are identical to those within the respective angiosperm proteins (Fig. 2B). Thus, these residues may play a crucial role in GID1 recognition and its interactions, as they have been evolutionarily conserved.
Comparative sequence analysis for 11 DELLA proteins or homologues in angiosperms with or without vascular tissue (rice, Arabidopsis, Selaginella, and moss) and conifers (pine). (A) Phylogenetic analysis of DELLA proteins or homologues. The maximum likelihood tree is based on the complete amino acid sequences of DELLA proteins from P. tabuliformis, A. thaliana, P. patens, S. moellendorffii, and O. sativa. The species, and gene names and IDs are displayed on the right side of each branch. Bootstrap values were obtained by running 1,000 bootstrap replicates. The horizontal branch lengths are proportional to the estimated number of amino acid substitutions per residue. The arrows indicate P. tabuliformis genes isolated in the present study. (B) DELLA domain sequence alignment for DELLA homologues in A. thaliana, O. sativa, P. patens, S. moellendorffii, and P. tabuliformis. Black and gray boxes indicate identical or similar residues, respectively. Asterisks at the bottom represent identical residues. The DELLA Δ17 domain range is depicted by a black line at the bottom. The sizes of the letters above the sequence alignment represent the residue frequency at each site for the 11 studied gene sequences. The arrows indicate the two P. tabuliformis genes isolated for this study.
PtGID1 expression is regulated by GA
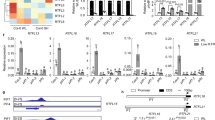
The GA metabolism genes in P. tabuliformis were isolated and identified in a previous study, in which PtGA20ox and PtGA3ox were shown to catalyse the penultimate and final steps, respectively, and ent-kaurenoic acid oxidase (PtKAO) was shown to be expressed upstream26. We measured the expression levels of PtGA3ox, PtGA20ox, PtKAO, and PtGID1 in P. tabuliformis after GA3, GA4+7, or treatment with the GA biosynthesis inhibitor paclobutrazol (PAC) (Fig. 3). PAC treatment resulted high expression levels of PtGA3ox. PtKAO2 mRNA levels were significantly lower than in non-treated pine needles when exogenous GA was present. This indicated that GA and PAC treatment affected the GA feedback mechanism. PtGID1 had dramatically lower expression levels following the application of either GA3 or GA4+7 than non-treated pine needles, and higher expression levels after PAC treatment. This showed that PtGID1 had functional GA signal transduction capability.
PtGID1 expression is decreased in P. tabuliformis after GA treatment. Relative expression levels of the GA biosynthesis genes PtGA3ox1, PtGA3ox2, PtGA20ox1, PtKAO1, and PtKAO2 and the GA receptor PtGID1, determined by real-time RT-PCR in 2-month-old pine after treatment with 50 μM GA3, 50 μM GA4+7, or 50 μM PAC. The expression levels were normalised to 18S rRNA. The data are shown as the means ± SD of biological triplicates.
PtGID1 confers a GA-sensitive phenotype
To establish that PtGID1 genes encode a GA receptor function in vivo, we generated A. thaliana transformants overexpressing the PtGID1 gene under the control of the constitutive 35S promoter. Seeds from wild-type A. thaliana, and from three lines of PtGID1-overexpressing A. thaliana (PtGID1-1, PtGID1-10, and PtGID1-12), were sown with medium containing low PAC concentrations (20 μM). Wild-type growth was suppressed under these conditions, but most seeds from the three PtGID1-overexpressor lines germinated (Fig. 4A). Because the three PtGID1-overexpressor lines had a higher rate of germination than the wild type, PtGID1 overexpressors presumably have a greater capability to perceive endogenous GA than the wild type. After increasing PAC concentration to 40 μM, growth was repressed in all cases (control case, Fig. 4B). Wild-type growth was repressed with the application of exogenous GA3, but PtGID1 overexpressors germinated normally (Fig. 4C). This indicated that PtGID1 overexpressors are better able to detect GA and are more efficient in GA signal transduction than the wild type.
PtGID1 overexpression rescues Arabidopsis plants grown on medium containing the GA biosynthesis inhibitor PAC. Wild-type (WT) plants and three lines of PtGID1 plants (PtGID1-1, PtGID1-10, and PtGID1-12) were sown on MS-agar with (A) 20 μM PAC, (B) 40 μM PAC, or (C) 40 μM PAC and 1 μM GA3, and incubated at 22 °C. Scale bar, 1 cm.
The opposing functions of DELLA proteins and GA in regulating root growth have been reported32. However, it remains unclear whether PtGID1 (which degrades DELLA proteins) exhibits sensitive GA-binding activity and influences root growth. To investigate this question, we created AtGID1a overexpressors. Figure 5A shows root growth of the wild type, AtGID1a overexpressor, and three independent lines of the PtGID1-overexpressing plants (PtGID1-1, PtGID1-10, and PtGID1-12). All plants were grown on GA3-free medium or on medium supplemented with 0.2, 0.5, or 1 μM GA3. We observed greater root elongation in all lines with increasing exogenous GA3 concentrations, but root elongation in the wild type was significantly and consistently lower than in the AtGID1 and PtGID1 overexpressors (Fig. 5B). This indicated a GA-hypersensitive phenotype in the transformants and confirmed that PtGID1 is a GA receptor in P. tabuliformis.
PtGID1 overexpression in Arabidopsis plants promotes root elongation under GA application. (A) Representative 6-day-old seedling primary roots of WT, PtGID1-1, PtGID1-10, and PtGID1-12 seedlings grown on MS-agar with GA3-free, 0.2, 0.5, or 1 μM GA3. Scale bar, 1 cm (n = 36). (B) Mean lengths in cm (mean ± SD, n = 36) of WT, PtGID1-1, PtGID1-10, and PtGID1-12 seedlings grown on MS-agar with GA3-free, 0.2, 0.5, or 1 μM GA3 are presented.
In A. thaliana, it has been suggested that DELLA proteins suppress plant growth to enhance survival in saline environments5,9,33. We therefore investigated the role of the GA receptor in growth and salt tolerance by promoting the degradation of DELLAs (Fig. 6A). We found that all three PtGID1 overexpressor lines were less viable than the wild type under conditions of high salt stress (Fig. 6B). Thus, we surmised that PtGID1 was involved in the GA-GID1-DELLA module, and that the PtGID1-GA protein complex stimulates growth by promoting the degradation of DELLAs.
Non-survival rates of A. thaliana PtGID1 overexpressors at toxic salt concentrations. (A) Representation of survival among WT, PtGID1-1, PtGID1-10, and PtGID1-12 transformants grown on high-salt (150 mM) medium. Photographs were taken 20 days after plants had been transferred to high-salt medium. Live plants are green; dead plants are white. (B) Numbers of WT, PtGID-1, PtGID-10, and PtGID-12 plants that failed to grow on high-salt medium (expressed as total number of dead plants and rate of non-survival in %).
In the GA metabolism pathway of Arabidopsis, GA 20-oxidase (AtGA20ox) and GA 3-oxidase (AtGA3ox) catalyse successive steps in the synthesis of bioactive GAs34. However, AtGA2ox also deactivates bioactive GAs35. We assessed the levels of mRNA expression for AtGA20ox, AtGA3ox, and AtGA2ox in wild-type plants, as well as in GID1-transgenic plants (Fig. 7). We found that both AtGID1 overexpressors and PtGID1 overexpressors contained lower levels of AtGA3ox1, AtGA3ox2, AtGA20ox1, and AtGA20ox2 transcripts than wild-type plants. In contrast, GID1 overexpressors exhibited highly increased levels of AtGA2ox2 and AtGA2ox4 expression. Thus, the overexpression of PtGID1, like AtGID1, increased the sensitivity of Arabidopsis to GA.
PtGID1 overexpressors upregulate GA 2-oxidase transcript levels. Relative expression levels of GA biosynthesis AtGA3ox1, AtGA3ox2, AtGA20ox1, and AtGA20ox2 transcripts and GA deactivation of AtGA2ox2, AtGA2ox4 gene transcripts levels (determined by real-time RT-PCR) in 8-d-old seedlings of WT, AtGID1, PtGID1-1, PtGID1-10, and PtGID1-12 plants. The expression levels were normalised to 18S rRNA. The data are shown as the means ± SD of biological triplicates.
Angiosperms and evolutionarily distant conifers share similar GID-DELLA modules
While previous studies have thoroughly elucidated the interaction between AtGID1 and DELLA proteins, to date no such studies have focused on conifers. We performed yeast two-hybrid assays, and observed that PtDPL and PtRGA had self-activating functions, which is consistent with previous reports36. We also performed bimolecular fluorescence complementation (BiFC) experiments to test whether PtGID1 and DELLA proteins interact in vivo. Each was fused to the reporter yellow fluorescence protein (YFP) which lacked either its C-terminal or its N-terminal end; thus, the YFP signal could be detected only upon interaction between PtGID1 and DELLA (Fig. 8). Fluorescence signals indicating interactions between PtGID1/AtGID1 and PtDPL, PtRGA, and AtGAI were observed in the nucleus, indicating that PtGID1 could interact with PtDPL and PtRGA (Fig. 8C,D), and that AtGID1 interacted with AtGAI (Fig. 8I). Moreover, AtGID1 could also interact with PtDPL and PtRGA (Fig. 8J,K), and PtGID1 could interact with AtGAI (Fig. 8B), suggesting that conifers and angiosperms have similar GID1-DELLA interaction patterns. To gain further insight into the sites of interaction for DELLA proteins in P. tabuliformis, we removed the identified Δ17-domains from PtDPL, PtRGA, and AtGAI to generate the mutated Ptdpl, Ptrga, Atgai sequences, respectively. We cloned these mutated sequences into expression vectors using overlap-extension PCR technology for site-directed mutagenesis. We observed no fluorescence signals corresponding to interactions between PtGID1/AtGID1 and Ptdpl, Ptrga, and Atgai (Fig. 8E,F,G,L,M,N). This supported our hypothesis that the Δ17-domain contains the site necessary for interaction. In particular, the seven amino acid residues within this Δ17 domain that are conserved between P. tabuliformis and angiosperms may be the core sites for GID1-DELLA interaction in conifers.
BiFC analysis of the GID1-DELLA interaction in nuclei of transfected A. thaliana. AtGID1 and PtGID1 were expressed and interactions tested with wild-type and mutant A. thaliana and P. tabuliformis DELLA proteins (AtGAI, Atgai; PtDPL, Ptdpl; PtRGA; Ptrga), respectively. Bright-field image, YFP fluorescence image, and the merged image are each displayed for expression of PtGID1 alone (A); co-expression with PtDPL (B), AtGAI (C), PtRGA (D), Δ17-domain mutant Ptdpl (E), Δ17-domain mutant Atgai (F), and Δ17-domain mutant Ptrga (G); Bright-field image, YFP fluorescence image, and the merged image are each displayed for expression of AtGID1 alone (H); co-expression with PtDPL (I), AtGAI (J), PtRGA (K), Δ17-domain mutant Ptdpl (L), Δ17-domain mutant Atgai (M), and Δ17-domain mutant Ptrga (N).
Discussion
We cloned five sequences from P. tabuliformis that had high sequence homology to GID1 proteins, and confirmed that only PtGID1 is orthologous to the A. thaliana and rice GID1 genes. PtGID1 shares the conserved motifs HGG and GXSXG with the HSL protein family, as well as the essential sites for binding GA and interacting with DELLA proteins. The 3D protein structure of PtGID1 is similar to that of OsGID1, while PtGID1 has an additional α-helix compared to OsGID1 and an N-ex sequence similar to AtGID1a. Thus, during the course of evolution, ancient GID1-like receptors from lycophytes and mosses have developed a pocket into which the GA molecule fits37. An additional innovation in conifers and angiosperms is the amino acid ‘lid’ in GID1 that holds the GA molecule in place.
Our data show that the conifer GID1 gene is derived from genes in the HSL family, as their essential sites for protein structure and function are highly conserved in the conifer GID1 protein. This led us to hypothesise that PtGID1 has the ability to interact with DELLA, and that it may also function as a GA receptor similar to GIDs in angiosperms. Moreover, when P. tabuliformis pine needles were treated with GA, the level of PtGID1 expression decreased dramatically compared to the non-treated control. In other words, a common GA-GID1-DELLA signalling module may also operate in conifers.
As part of this potential GA-GID1-DELLA signalling module, we also identified two DELLA proteins from P. tabuliformis, which we termed PtDPL and PtRGA. These two DELLA proteins have a variant of the DELLA domain when compared to DELLA proteins from other higher plants/angiosperms, which is capable of interacting with the respective GID1 N-Ex. DELLA proteins are of great importance in the evolution of the GA signalling pathway: they have played a role in interacting with GID1 and regulating downstream signal transduction since the divergence from lycophytes. Seven amino acid residues within the Δ17 domain may be the core sites for GID1-DELLA interaction in P. tabuliformis. As these amino acid residues are also identical in angiosperms, they may play a crucial role in the interaction with GID1.
We also identified a GA-hypersensitive phenotype in PtGID1 overexpressors. This indicated an increase in the ratio of inactive GID1-DELLA complex to active DELLA repressor38, and suggested that these mutants had an increased ability to stimulate the GA signalling pathway. In support of this theory, the PtGID1-overexpressor A. thaliana mutant exhibited greater root elongation compared to the wild type. This effect is consistent with a GA overdose phenotype, and is comparable to the tall OsGID1 and AtGID1 overexpressor phenotypes8,16 and to the enhanced stem elongation phenotype induced by overexpressing PttGID in aspen39. These PtGID1 overexpressors had a reduced ability to endure salt toxicity compared to the wild type, indicating reduced salt tolerance. This supported the notion that DELLA proteins were degraded by overdosing GA-GID1 in PtGID1 overexpressors. This finding supports the notion that DELLA proteins help to enhance survival in saline environments by repressing plant growth25.
The expression levels of the GA biosynthesis genes AtGA3ox and AtGA20ox were downregulated and GA deactivation of AtGA2ox genes was significantly increased in PtGID1 overexpressors compared with the wild type. These effects were the same as those observed in AtGID1 overexpressors. Thus, the changes in the expression of GA metabolism genes in PtGID1-transgenetic Arabidopsis, and the PtGID1 gene in P. tabuliformis, showed that PtGID1 is a GA receptor with biological function.
To demonstrate the existence of functional GID1-DELLA interaction in P. tabuliformis, we investigated several GA signalling models (Fig. 8). Using BiFC assays, we detected interactions between AtGID1 and the AtDELLA proteins, as well as between PtGID1and the PtDELLA proteins. There were no interactions in the species with Δ17-domain-mutated DELLA proteins; however, there were interactions between GID1 and DELLAs from angiosperm and conifer species. This led us to conclude that (1) conifer P. tabuliformis has functional GID1-DELLA components; (2) conifers and angiosperms have the same patterns of GID1-DELLA interaction; and (3) the Δ17-domain from PtDPL/PtRGA contains sites necessary for interaction. Although only half of the amino acid residues within the Δ17-domain of PtDPL and PtRGA are identical to those within the respective A. thaliana and rice genes, the GID-DELLA interaction does exist in the P. tabuliformis GA signalling pathway. Therefore, the GID1-mediated GA signalling cascade appeared after the divergence of vascular plants from the moss lineage14. The GA-GID1-DELLA signalling pathway has been gradually modified over the course of lycophyte, fern, conifer, and angiosperm evolution by changes within DELLA and GID proteins.
In conclusion, PtGID1 acts as the GA receptor in P. tabuliformis. It is capable of interacting with DELLA proteins, and has the ability to recognise GA to form the GA-GID1-DELLA signalling module. This suggests that the GA signalling pathway operates in conifers, and is present in other vascular plants of substantially different evolutionary ages, such as in lycophytes, ferns, and angiosperms.
Materials and Methods
Plant material
Pinus tabuliformis tree cones were collected from genetically distinct trees selected at random in a primary clonal seed orchard located in Xingcheng City, Liaoning Province, China30. Details about this seed orchard can be found in a previous study40. Seeds from wild-type A. thaliana Col, and from transformants in which the PtGID1 gene was overexpressed, were used in this study. All genotypes were in the Columbia background. Prior to germination, seeds were washed in 84% hydrogen peroxide and alcohol, then washed again five times with sterile water. All seeds were germinated on Murashige and Skoog (MS)-agar supplemented with 20 μM PAC, 40 μM PAC, or 40 μM PAC and 1 μM GA3, and incubated for 15 or 25 d at 22 °C. For root growth experiments, all seeds were grown on MS-agar supplemented with 0, 0.2, 0.5, or 1 μM GA3 and stacked vertically in a growth chamber (22 °C; 16-h photoperiod). Root length was measured from root tip to the base of the hypocotyl. For salt tolerance experiments, seeds were thoroughly washed as described above and grown on MS-agar medium for 6 d. Subsequently, seedlings were transferred to MS-agar supplemented with 150 mM NaCl and incubated at 22 °C for 20 d. For gene expression analysis, pine seedlings were irrigated with 50 μM GA3, 50 μM GA4+7, or 50 μM PAC for 2 d.
Identification, cloning, and in silico protein structural analysis of GID1 and DELLA genes from P. tabuliformis
The protein-encoding gene sequences of GID1 in A. thaliana, AtGID1a (AT3G05120), AtGID1b (AT3G63010), and AtGID1c (AT5G27320), and in rice, OsGID1 (Os05g0407500), were used to screen the P. tabuliformis reference transcriptome30 for homologous EST sequences. Full-length coding sequences from P. tabuliformis were cloned based on high homology. Next, we performed a reverse database search at The Arabidopsis Information Resource (TAIR) by contrasting these GID1-homologous sequences from P. tabuliformis with those from the A. thaliana genome, using TBLASTN to obtain homologous sequences in A. thaliana. Subsequently, we carried out multiple alignment of full-length protein sequences using MUSCLE41,42. We used these protein sequences to build a maximum-likelihood (ML) phylogenetic tree based on the JTT model to identify the true orthologues of GID1. SWISS-MODEL (http://swissmodel.expasy.org) was used to generate 3D protein models43 on the basis of the known X-ray crystal structure profile for rice GID1.
The protein-coding gene sequences of DELLAs in A. thaliana, AtRGA (AT2G01570), AtGAI (AT1G14920), AtRGL2 (AT3G03450), AtRGL3 (AT5G17490), and AtRGL1 (AT1G66350), were used to screen the P. tabuliformis reference transcriptome30 for homologous EST sequences. The P. tabuliformis full-length mRNA sequences of hits were obtained, and were further translated in silico. We constructed an ML phylogenetic tree, including the P. tabuliformis sequences as well as full-length protein sequences from rice, Physcomitrella patens, and S. moellendorffii (OsSLR1 [Os03g49990], PpGAL1 [XP_001754090], PpGAL2 [XP_001774314], and SmDELLA1 [XP_00296024]).
Overexpression of the PtGID1 gene in A. thaliana
Full-length cDNAs for PtGID1 containing suitable restriction enzyme sites at both ends were prepared by PCR, then inserted into a pBI121 vector containing the constitutive 35S promoter. The SpeI site was used for cloning PtGID1 and obtaining the p35S-PtGID1 fragment. The p35S-PtGID1 fragment was introduced into wild-type A. thaliana plants by Agrobacterium-mediated transformation44. Expression of the transgene in A. thaliana plants was confirmed by PCR. The primers used are listed in Table S1. Transgenic plants were grown in a greenhouse under a constant day length of 16 h.
BiFC and infiltration in A. thaliana leaf tissue
For the bimolecular fluorescence complementation (BiFC) assay, AtGID1 and PtGID1 genes containing appropriate restriction sites at both ends were cloned into the pSPYNE vector, using the StuI-XhoI sites for AtGID1 and PtGID1, to produce pSPYNE-GID1 plasmids. Similarly, the entire coding regions of PtDPL, PtRGA, AtGAI, Ptdpl, Ptrga, and Atgai sequences were cloned into the pSPYCE vector, using the StuI-XhoI site to produce pSPYCE-DELLA and pSPYCE-della plasmids. Table S1 lists the primers that were used. All expression vectors were introduced into A. tumefaciens LBA4404. Agrobacteria were incubated, harvested, and resuspended in agroinfiltration buffer (0.2 mM acetosyringone, 10 mM MgCl2, and 10 mM MES). Agroinfiltration buffer was mixed with an equal volume of the protein mixture and injected into A. thaliana leaves using a syringe. Seventy two hours after infiltration, images were taken using a Leica TCS SP5 confocal microscope.
Gene expression analysis
Total RNA was extracted using the TRIzol reagent (Invitrogen, California, USA) from 8-d-old Arabidopsis seedlings or 2-month-old P. tabuliformis pine needles. RNA yield was determined using a NanoDrop 2000 spectrophotometer (Thermo Scientific, USA), and its integrity was evaluated using agarose gel electrophoresis with ethidium bromide staining. Total RNA (0.5 μg) was reverse transcribed into cDNA in a GeneAmp PCR System 9700 (Applied Biosystems, USA). A 1-μl aliquot of cDNA was used in 10-μl reactions in the GeneAmp PCR System 9700 (Applied Biosystems, USA) using the LightCycler 480 II Real-time PCR Instrument (Roche, Swiss). Each sample was run in triplicate. At the end of the PCR cycles, melting curve analysis was performed to validate the generation of the expected PCR product. The gene-specific primers used are listed in Table S2. mRNA expression levels were normalised to 18S rRNA and were calculated using the 2−ΔΔCt method45.
References
Davies, P. J. Plant Hormones:Physiology, Biochemistry, and Molecular Biology [833] Kluwer Academic, Dordrecht (1995).
Ogawa, M. et al. Gibberellin Biosynthesis and Response During Arabidopsis Seed Germination. Plant Cell. 15, 1591–1604 (2003).
Djakovic-Petrovic, T., de Wit, M., Voesenek, L. A. & Pierik, R. DELLA Protein Function in Growth Responses to Canopy Signals. Plant J. 51, 117–126 (2007).
Sun, T. P. Gibberellin-GID1-DELLA: A Pivotal Regulatory Module for Plant Growth and Development. Plant Physiol. 154, 567–570 (2010).
Harberd, N. P., Belfield, E. & Yasumura, Y. The Angiosperm gibberellin-GID1-DELLA Growth Regulatory Mechanism: How an “Inhibitor of an Inhibitor” Enables Flexible Response to Fluctuating Environments. Plant Cell. 21, 1328–1339 (2009).
Ueguchi-Tanaka, M. et al. Molecular Interactions of a Soluble Gibberellin Receptor, GID1, with a Rice DELLA Protein, SLR1, and Gibberellin. Plant Cell. 19, 2140–2155 (2007).
Sun, T. P. Gibberellin Metabolism, Perception and Signaling Pathways in Arabidopsis. Arabidopsis Book. 6, e103 (2008).
Ueguchi-Tanaka, M. et al. Gibberellin Insensitive DWARF1 Encodes a Soluble Receptor for Gibberellin. Nature. 437, 693–698 (2005).
Peng, J. et al. The Arabidopsis GAI Gene Defines a Signaling Pathway that Negatively Regulates Gibberellin Responses. Genes Dev. 11, 3194–3205 (1997).
Silverstone, A. L., Ciampaglio, C. N. & Sun, T. The Arabidopsis RGA Gene Encodes a Transcriptional Regulator Repressing the Gibberellin Signal Transduction Pathway. Plant Cell. 10, 155–169 (1998).
Ueguchi-Tanaka, M., Nakajima, M., Motoyuki, A. & Matsuoka, M. Gibberellin Receptor and its Role in Gibberellin Signaling in Plants. Annu Rev Plant Biol. 58, 183–198 (2007).
Itoh, H., Ueguchi-Tanaka, M. & Matsuoka, M. Molecular Biology of Gibberellins Signaling in Higher Plants. Int Rev Cell Mol Biol. 268, 191–221 (2008).
Hayashi, K. et al. Endogenous Diterpenes Derived From Ent-Kaurene, a Common Gibberellin Precursor, Regulate Protonema Differentiation of the Moss Physcomitrella Patens. Plant Physiol. 153, 1085–1097 (2010).
Hirano, K. et al. The GID1-mediated Gibberellin Perception Mechanism is Conserved in the Lycophyte Selaginella Moellendorffii but Not in the Bryophyte Physcomitrella Patens. Plant Cell. 19, 3058–3079 (2007).
Kenrick, P. & Crane, P. R. The Origin and Early Evolution of Plants On Land. Nature. (1997).
Nakajima, M. et al. Identification and Characterization of Arabidopsis Gibberellin Receptors. Plant J. 46, 880–889 (2006).
Yamamoto, Y. et al. A Rice Gid1 Suppressor Mutant Reveals that Gibberellin is Not Always Required for Interaction Between its Receptor, GID1, and DELLA Proteins. Plant Cell. 22, 3589–3602 (2010).
Griffiths, J. et al. Genetic Characterization and Functional Analysis of the GID1 Gibberellin Receptors in Arabidopsis. Plant Cell. 18, 3399–3414 (2006).
Shimada, A. et al. Structural Basis for Gibberellin Recognition by its Receptor GID1. Nature. 456, 520–523 (2008).
Murase, K., Hirano, Y., Sun, T. P. & Hakoshima, T. Gibberellin-Induced DELLA Recognition by the Gibberellin Receptor GID1. Nature. 456, 459–463 (2008).
Ikeda, A. et al. Slender Rice, a Constitutive Gibberellin Response Mutant, is Caused by a Null Mutation of the SLR1 Gene, an Ortholog of the Height-Regulating Gene GAI/RGA/RHT/D8. Plant Cell. 13, 999–1010 (2001).
Hirsch, S. & Oldroyd, G. E. GRAS-domain Transcription Factors that Regulate Plant Development. Plant Signal Behav. 4, 698–700 (2009).
Dill, A., Jung, H. S. & Sun, T. P. The DELLA Motif is Essential for Gibberellin-Induced Degradation of RGA. Proc Natl Acad Sci USA 98, 14162–14167 (2001).
Yoshida, H. et al. DELLA Protein Functions as a Transcriptional Activator through the DNA Binding of the Indeterminate Domain Family Proteins. Proc Natl Acad Sci USA 111, 7861–7866 (2014).
Achard, P. et al. Integration of Plant Responses to Environmentally Activated Phytohormonal Signals. Science. 311, 91–94 (2006).
Niu, S., Yuan, L., Zhang, Y., Chen, X. & Li, W. Isolation and Expression Profiles of Gibberellin Metabolism Genes in Developing Male and Female Cones of Pinus Tabuliformis. Funct Integr Genomics. 14, 697–705 (2014).
Joyner, K. L., Wang, X. R., Johnston, J. S., Price, H. J. & Williams, C. G. DNA Content for Asian Pines Parallels New World Relatives. Canadian Journal of Botany. 79, 192–196 (2001).
Niu, S. et al. A Transcriptomics Investigation Into Pine Reproductive Organ Development. New Phytol. 209, 1278–1289 (2016).
Niu, S. H. et al. Identification and Expression Profiles of sRNAs and their Biogenesis and Action-Related Genes in Male and Female Cones of Pinus Tabuliformis. BMC Genomics. 16, 693 (2015).
Niu, S. H. et al. Transcriptome Characterisation of Pinus Tabuliformis and Evolution of Genes in the Pinus Phylogeny. BMC Genomics. 14, 263 (2013).
Willige, B. C. et al. The DELLA Domain of GA INSENSITIVE Mediates the Interaction with the GA INSENSITIVE DWARF1A Gibberellin Receptor of Arabidopsis. Plant Cell. 19, 1209–1220 (2007).
Fu, X. & Harberd, N. P. Auxin Promotes Arabidopsis Root Growth by Modulating Gibberellin Response. nature. 421, 740–743 (2003).
Fleet, C. M. & Sun, T. P. A DELLAcate Balance: The Role of Gibberellin in Plant Morphogenesis. Curr Opin Plant Biol. 8, 77–85 (2005).
Phillips, A. L. et al. Isolation and Expression of Three Gibberellin 20-Oxidase cDNA Clones From Arabidopsis. Plant Physiol. 108, 1049 (1995).
Thomas, S. G., Phillips, A. L. & Hedden, P. Molecular Cloning and Functional Expression of Gibberellin 2- Oxidases, Multifunctional Enzymes Involved in Gibberellin Deactivation. P Natl Acad Sci USA 96, 4698–4703 (1999).
Dill, A., Thomas, S. G., Hu, J., Steber, C. M. & Sun, T. The Arabidopsis F-box Protein SLEEPY1 Targets Gibberellin Signaling Repressors for Gibberellin-Induced Degradation. Plant Cell. 16, 1392 (2004).
Ross, J. J. & Reid, J. B. Evolution of Growth-Promoting Plant Hormones. FUNCT Plant Biol. 37, 795–805 (2010).
Ariizumi, T., Murase, K., Sun, T. P. & Steber, C. M. Proteolysis-Independent Downregulation of DELLA Repression in Arabidopsis by the Gibberellin Receptor GIBBERELLIN INSENSITIVE DWARF1. Plant Cell. 20, 2447–2459 (2008).
Mauriat, M. & Moritz, T. Analyses of GA20ox- and GID1-over-expressing Aspen Suggest that Gibberellins Play Two Distinct Roles in Wood Formation. PLANT J. 58, 989–1003 (2009).
Li, W., Wang, X. & Li, Y. Stability in and Correlation Between Factors Influencing Genetic Quality of Seed Lots in Seed Orchard of Pinus Tabuliformis Carr. Over a 12-Year Span. Plos One. 6, e23544 (2011).
Edgar, R. C. MUSCLE: A Multiple Sequence Alignment Method with Reduced Time and Space Complexity. BMC Bioinformatics. 5, 113 (2004).
Edgar, R. C. MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL Workspace: A Web-Based Environment for Protein Structure Homology Modelling. Bioinformatics. 22, 195–201 (2006).
Hiei, Y., Ohta, S., Komari, T. & Kumashiro, T. Efficient Transformation of Rice (Oryza Sativa L.) Mediated by Agrobacterium and Sequence Analysis of the Boundaries of the T-DNA. Plant J. 6, 271–282 (1994).
Livak, K. J. & Schmittgen, T. D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 25, 402–408 (2001).
Acknowledgements
This work was supported by “The Fundamental Research Funds for the Central Universities (BLX2014-23); The National Natural Science Foundation of China (31370657, 31770713) and China Postdoctoral Science Foundation (2015M570042).
Author information
Authors and Affiliations
Contributions
R.D. performed experiments, analyzed data and drafted the manuscript. S.N., Y.L. and X.S. participated in the samples preparation and provided helpful advices. I.P. and Y.A.E. helped to approved the final manuscript. W.L. planned and designed the research.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Du, R., Niu, S., Liu, Y. et al. The gibberellin GID1-DELLA signalling module exists in evolutionarily ancient conifers. Sci Rep 7, 16637 (2017). https://doi.org/10.1038/s41598-017-11859-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-11859-w
This article is cited by
-
Phylogeny and expression patterns of ERF genes that are potential reproductive inducers in hybrid larch
BMC Genomics (2024)
-
OsCSN2 orchestrates Oryza sativa L. growth and development through modulation of the GA and BR pathways
Functional & Integrative Genomics (2024)
-
Molecular Landscape of Bolting in Spinach Explored Through Gene Expression Profiling
Journal of Plant Growth Regulation (2024)
-
Transcriptome analysis of the genes regulating phytohormone and cellular patterning in Lagerstroemia plant architecture
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.