Abstract
Methylated non-CpGs (mCpH; H means A, C, and T) have emerged as key epigenetic marks in mammalian embryonic stem cells (ESCs) and neurons, regulating cell type-specific functions. In these two cell types, mCpHs show distinct motifs and correlations to transcription that could be a key in understanding the cell type-specific regulations. Thus, we attempted to uncover the underlying mechanism of the differences in ESCs and neurons by conducting a comprehensive analysis of public whole genome bisulfite sequencing data. Remarkably, there were cell type-specific mCpH patterns around methylated CpGs (mCpGs), resulted from preferential methylation at different contexts by DNA methyltransferase (DNMT) 3a and 3b. These DNMTs are differentially expressed in ESCs and brain tissues, resulting in distinct mCpH motifs in these two cell types. Furthermore, in ESCs, DNMT3b interacts with histone H3 tri-methylated at lysine 36 (H3K36me3), resulting in hyper-methylation at CpHs upon actively transcribed genes, including those involved in embryo development. Based on the results, we propose a model to explain the differential establishment of mCpHs in ESCs and neurons, providing insights into the mechanism underlying cell type-specific formation and function of mCpHs.
Similar content being viewed by others
Introduction
DNA methylation, the addition of a methyl group on the fifth carbon at cytosine, is one of the most important epigenetic modifications. It preferentially occurs at CpG (cytosine followed by guanine) sites in mammalian cells, regulating cell development and maintenance1, 2. In contrast, the methylation at non-CpG (CpH; H includes A, C, and T) sites had been barely detected in mammalian cells. However, recent improvements in sequencing technology enabled researchers to determine that significant amounts of methylated CpHs (mCpHs) exist in mammalian pluripotent stem cells and non-dividing cells (i.e., neurons). The mCpHs are involved in cell type-specific regulation such as embryonic stem cell differentiation or synaptogenesis3,4,5,6,7,8. Thus, mCpH, as mCpG, has emerged as a key epigenetic factor, especially in pluripotent stem cells and neuron.
In these cells, both mCpGs and mCpHs are induced by DNA methyltransferase3a and 3b (DNMT3a and DNMT3b, respectively), with allosteric cooperation of DNMT3l9. Since these methyltransferases show much higher affinities at CpGs than CpHs, the mCpHs are spatially dependent on mCpGs3, 6, 10,11,12,13. Despite the dependency, mCpHs regulate several cell type-specific functions independently to the methylation at CpGs. In the brain, mCpHs gradually increase with age, similar with the progression of synaptogenesis5; whereas mCpGs do not. In addition, the methyl-CpG binding protein 2 (MeCP2), the mutation of which causes Rett syndrome, binds to not only mCpGs, but also mCpHs14. Considering that postnatal onset of Rett syndrome coincides with the emergence of mCpH in neurons, MeCP2-related neuro-diseases could be governed by mCpHs rather than mCpGs15. Additionally, in ESCs, the reduction of mCpHs leads to the decrease of differentiation capacity16. Thus, even though mCpHs are spatially correlated to mCpGs, they play important roles on cell type-specific processes independently of mCpGs.
One of the underlying mechanisms by which mCpHs govern cell type-specific processes is their differential distribution between the cell types. For example, the CpH methylation primarily occurs at CAG motif in ESCs8, 17, while at CAC motifs in neurons5, 18, implying that at least two distinct mechanisms for formation (and/or function) of the mCpHs exist10. In addition, CpHs tend to be hyper-methylated in actively transcribed genes in ESCs, while hypo-methylated in neurons5, 8. Since DNA methylation tends to repress gene expression in most cell types19, the positive correlation between CpH methylation and gene expression in ESC has been an enigma among researchers8, 10. Thus, mCpH shows different distribution and potential role in gene expression between ESCs and neurons. However, what causes the differences remains unknown.
In this study, to uncover the mechanism of the differential CpH methylation, we analysed the whole genome bisulfite sequencing (WGBS) data of human ESCs, neurons, and brain tissues. In addition, we included WGBS data of DNMT knockout human and mouse ESCs to investigate the role of DNMTs on cell type-specific methylation5, 12, 20,21,22,23,24,25. Through a comprehensive analysis, we found that CpH methylation pattern near CpGs (within ± 100 base pair (bp) from CpGs) is highly distinguishable between ESCs and neurons. Further analyses uncovered that DNMT3a and DNMT3b, differentially expressed in those two cell types, preferentially methylate cytosine in different contexts, resulting in distinct motifs and patterns of mCpHs in ESCs and neurons. Additionally, in ESCs, the DNMT3b preferentially interacts with H3K36me3, resulting in hypermethylation at CAGs on actively transcribed gene-body regions. The methylated CAGs were enriched in genes related to embryo development, indicating possible role of mCpHs on embryogenesis. Based on these results, we suggest a differential CpH methylation model that could explain distinct features and functions of mCpHs in ESCs and neurons. Altogether, our results provide new insights on cell type-specific formation and function of mCpHs in mammals.
Results
The integrative approach for WGBS read-aligning successfully reproduced known characteristics of mCpH in ESCs and neurons
To analyse methylation in both CpG and CpH contexts, we aligned WGBS reads generated from various experiments (S. Table 1). Three bisulfite read aligners, Bismark26, BSMAP27, and BS-seeker228, were used for read aligning, and the outputs were integrated as previously described29. This integration method has proven to improve methylation detection accuracy and reduce artifacts from experimental settings29.
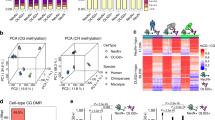
Statistic features of the regenerated data coincided with those obtained in previous studies (Fig. 1a)3, 5,6,7,8, 10. In human samples, although most of the CpGs were hyper-methylated (75–85%), the average methylation levels at CpHs (mCpH level) were distinct across cell types. It was mostly abundant in neurons (>5%), abundant in ESCs derived from male (H1) and matured brain tissues (>1%), detectable in ESCs derived from female (H9), early-passage ESCs (HUES64), and immature brain tissues (0 to 5 year-old; about 1%), and mostly undetectable in other tissues (The heart, spleen, and lung; <0.5%). In addition, we observed an increase of mCpH levels along with brain aging5, and lower mCpH level in H9 than in H1, as previously reported8, 17, 30. Remarkably, the WGBS samples were clearly clustered based on the tissue of origin (or cell types) by both CpG and CpH methylation patterns, indicating our integrated dataset well represents differential methylation pattern among cell types (Fig. 1b).
Human methylome from integrated WGBS data. (a) Genome-wide average methylation level at CpGs (red line) and CpHs (blue bar) extracted from WGBS data. NEUR and Br represent neurons and brain tissues, respectively. The numbers followed by Br- means age of samples (d and y mean day and year, respectively). H1, H9, and HUES64 are human embryonic stem cell lines. Numbers at the end of the labels represent biological replicates. (b) WGBS samples hierarchically clustered by genome-wide CpG (left) and CpH (right) methylation levels. The whole genome was divided into 1 k-bp-long blocks and the average methylation levels of blocks were used to calculate the Pearson’s correlation coefficient (PCC). Colours in the x-axis represents the same cell (or tissue) types (blue: brain and neurons, red: ESCs, and yellow: other tissues).
Lastly, we confirmed the known difference of mCpHs between ESC and neuron. The motif abundant at hyper-methylated CpHs (\({{Me}}_{i}\)>0.5, See Method section) was “CAG” in all ESC samples, and “CAC” in all neuron and brain samples (S. Figure 1a), as previously reported5, 6, 8, 17, 18. In addition, the mCpH level in gene-body regions was positively correlated to FPKMs in ESCs (H1 and HUES64), whereas it was negatively correlated in adult brain (S. Figure 1b, Spearman’s rank correlation coefficient (ρ) equals to 0.37, 0.3, and −0.29, respectively).
Altogether, our integrated methylome successfully reproduced the known characteristics of CpH methylation in mammalian cells. Since the mCpH levels in early brains (Br-35d and −2y) and somatic cells are extremely low, we excluded these samples from further analysis and focused on methylation in ESCs, neurons, and adult brains. In addition, we considered that the characteristic of mCpHs in brain samples represents that in neurons, since mCpHs are mostly absent in other cells of brain5.
Cell type-specific CpH methylation patterns are observed in mCpG-proximal region (±100 bp)
Using the WGBS dataset, we analysed correlation between CpG and CpH methylation. We divided human whole genome (hg19) into 1 k-bp blocks and compared average methylation level at CpGs and CpHs in each block. The genome-wide CpG and CpH methylation levels were positively correlated as previously reported3, 6, 11, 18 (S. Figure 2a; Pearson’s correlation coefficients (PCC) were from 0.27 to 0.44). In further analysis with blocks in which CpGs are hemi-methylated (difference of mCpG level between DNA strands > 0.5), mCpH levels were significantly higher at the same strand with highly methylated CpGs, compared to those at the opposite strand, indicating that CpG and CpH methylation correlated in a strand-specific manner (S. Figure 2b,c). Additionally, the correlation between mCpG and mCpH was significantly higher at exons, promoter regions, and putative enhancer regions, implying that the co-regulation of CpG and CpH methylation mainly occurs in transcription-regulatory regions (S. Figure 2d).
Next, we attempted to measure distance required so that mCpG and mCpH could be co-regulated. First, we measured the probability that mCpHs appear at each distance from methylated or un-methylated CpGs (Fig. 2a). The probability showed a clear peak near mCpGs in both ESCs and neurons, indicating the concentration of mCpHs near mCpGs. The tendency was confirmed with a subset of CpGs, around which (±500 bp) no other CpGs exist (S. Figure 3a). Second, we measured the correlation between mCpGs and surrounding mCpHs. The correlation coefficients were also significantly higher near CpGs (S. Figure 3b) supporting that CpG and CpH methylation could be co-regulated when these are closely positioned. Thus, we concluded that methylation at CpHs is highly dependent to that at proximal CpGs, when the distance between CpGs and CpHs is less than ± 100 bp, which sufficiently covers the probability and correlation coefficient peaks.
Cell type-specific CpH methylation around CpGs. (a) Probability that mCpHs exist around methylated CpGs (metylation level >0.8, red), and un-methylated CpGs (methylation level <0.2, black). The identification of (un)methylated CpHs is described in the Method section. To calculate the probabilities, the surrounding (±500 bp) of every CpG divided into 10-bp-long blocks, and blocks containing more than one mCpH were counted, given the condition of CpGs (methylated or un-methylated). H1 represent H1 sample replicate 1 (H1.1), and NEUR represents neuron sample replicate 1 (NEUR.1). (b) The accumulated CpH methylation levels at each distance from methylated CpGs (red bars), and un-methylated CpGs (black bars). Blue arrows highlight mCpH level peak at -4 bp from mCpG in ESC, and 8 bp-periodicity of the peaks in neuron. (c) Heatmap showing the Pearson’s correlation coefficients (PCC) across mCpG-proximal (±100 bp) mCpH levels of the samples. H9 represents H9 replicate 1 (H9.1), HUES64 represents HUES64 replicate 1 (HUES64.1).
We analysed methylation pattern at CpHs within ± 100 bp from mCpGs (Fig. 2b). In ESCs, a clear CpH methylation peak was observed at −4 bp from mCpG as previously reported31. On the other hand, in neurons and brains, there was an 8–10 bp periodicity among mCpH peaks starting from mCpGs. This has been suggested as a mark for CpG methylation by DNMT3a-DNMT3l enzyme complex32, indicating that mCpGs and mCpHs are co-regulated by DNMT3a in neurons and brain tissues. Remarkably, the methylation pattern at mCpG-proximal CpHs was conserved among same cell types but clearly distinguishable between ESCs and neurons (Fig. 2c), implying that different methylation mechanisms exist between the two cell types. Altogether, our data indicated that the CpHs are methylated along with CpGs, but show different methylation patterns in ESCs and neurons.
DNMT3a and DNMT3b preferentially methylate CAC and CAG contexts, respectively
To uncover what causes the distinct methylation patterns at mCpG-proximal CpHs, and the known differences of mCpHs in ESC and neuron (S. Figure 1a and b), we analysed DNMT knockout human and mouse ESCs.
As a first step, we analysed mouse ESCs (mESCs) in which DNMT1, and both DNMT3a and DNMT3b are knocked out (D1-KO and D3-KO mESC, respectively)23. The mCpH level was largely decreased in D3-KO mESC, indicating that DNMT3a and DNMT3b are mainly responsible for CpH methylation (S. Figure 4a). Interestingly, in D1-KO mESC, the mCpH level was higher near mCpGs, and showed clear peaks with 180 bp interval, close to the nucleosome positioning pattern (Fig. 3a). Since both mCpGs and mCpHs are mainly introduced by DNMT3s in D1-KO sample, this result indicates that CpHs are methylated along with CpGs, by DNA walking of DNMT3s. In addition, the correlation between mCpG and mCpH levels was higher in D1-KO mESC and lower in D3-KO mESC, compared to that in wild type mESC (S. Figure 4b; PCC = 0.4, 0.07, and 0.2, respectively). Altogether, our data confirmed that DNMT3a and DNMT3b methylate CpHs, in correlated way to the methylation at CpGs.
Methylation at cytosines in different contexts by DNMT3a and DNMT3b in mouse (a,c) and human (b) ESCs. (a) The window-slided (window size = 10 bp) average CpH methylation levels around mCpG (methylation level >0.8) in WT, DNMT1-KO, and DNMT3-KO mouse ESC. (b) Bar graphs showing the average methylation levels at CpHpH (blue) and CpHpG (red) contexts in wild type (WT), DNMT3a-knock out (D3a-KO), DNMT3b-knock out (D3b-KO), and DNMT3a/b-knock out (D3-DKO) HUES64 cell line. WT ESC is HUES64.2. (c) Average methylation level at CAC, CAT, CAA, and CAG contexts in mouse ESCs, in which DNMT1, DNMT3a, and DNMT3b were knocked out (TKO), and DNMT3a and DNMT3b were reintroduced (TKO+3a and TKO+3b), respectively24. Red and grey bars respresent average mCpH level in mCpG-proximal regions (mCpG level>0.5) and whole genome, respectively.
Next, we analysed the contribution of DNMT3a and DNMT3b to CpH methylation in human ESC (HUES64 cell line). We divided the CpHs into two groups, asymmetric (CpHpH; CpH followed by A, C, or T), and symmetric (CpHpG; CpH followed by G) CpHs6, 33. Remarkably, in the DNMT3a knockout sample, the methylation level at CpHpH contexts decreased more than that at CpHpG contexts, whereas the opposite was observed in the DNMT3b-knockout sample (Fig. 3b). Similarly, in DNMT3b-knocked out H9 cells20, the methylation level at CpHpG contexts decreased more than that at CpHpH contexts (S. Figure 4c). Further analyses indicated that the methylation level at CAC tri-nucleotides was largely decreased by DNMT3a-knockout, whereas that at CAGs was decreased by DNMT3b-knockout (S. Figure 4d). The methylation level at CAAs was also reduced by the DNMT3b-knockout, even though the methylation level in wild type was not as high as that at CACs or CAGs. The methylation level in other contexts did not show significant difference.
We confirmed similar tendency in mouse ESC samples, generated by Dr. Tuncay Baubec’s group24. They knocked out DNMT1, DNMT3a, and DNMT3b from mouse ESCs and then reintroduced DNMT3a and DNMT3b, respectively, to measure de novo methylation by each methyltransferase. With this sample set, we confirmed preferential de novo methylation at mCpG-proximal CpHs by the DNMTs (Fig. 3c). Remarkably, the cytosines at CACs are more methylated by DNMT3a than by DNMT3b, whereas those in CAGs are mostly methylated by DNMT3b. The methylation at CAAs increased as DNMT3b re-induced in lower level compared to that at CAGs, as shown in human ESC samples. Thus, we concluded that both DNMT3a and DNMT3b are responsible for the methylation at CpHs, especially near mCpGs, but they methylate cytosines in different contexts; DNMT3a primarily methylate CAC tri-nucleotides, while DNMT3b methylates CAG tri-nucleotides.
The differential activities of DNMT3a and DNMT3b cause distinct features of mCpHs in ESCs and neurons
To understand how the differential activities of DNMT3a and DNMT3b affect cell type-specific features of mCpH, we analysed WGBS, RNA-seq, and ChIP-seq data of human ESCs, neurons, and brain tissues.
First, by combining expression data from 143 human adult brains and 33 human ESCs (S. Table 2), we confirmed that DNMT3b is highly expressed in ESCs, whereas DNMT3a is in adult brains (Fig. 4a). This result is coincided with previous result34, reporting that DNMT3b is actively transcribed in early ESCs, but decreased as cell differentiated. Remarkably, the abundant motif at mCpHs in DNMT3b-knockout human ESC was not CAG, but CAC (S. Figure 1a), indicating that the hyperactivity of DNMT3b causes the preferential methylation at CAGs in ESC. In addition, the methylation pattern at mCpG-proximal CpHs in DNMT3b-knockout ESC was more similar to that in the brains and neurons, than that in wild type ESCs (Fig. 4b). Thus, our data revealed that the differentially enriched mCpH motifs in ESCs and neurons, as well as the distinct mCpG-proximal mCpH patterns in the two cell types, are caused by the differential activities of DNMT3a and DNMT3b.
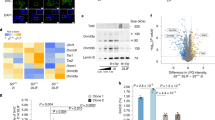
Cell-type specific features of mCpHs induced by differential activity of DNMT3s. (a) Boxplot representing the FPKM and RPKM of DNMT3a and DNMT3b in 36 ESCs (H1 and H9) and 143 adult brain tissues (from 13 to 40 year-old) of human. The boxes include values from 25% to 75% in order. The P-value was calculated from Wilcox-rank t-test. Specifics for the data are described in S. Table 2. (b) Hierarchical clustering results by mCpG-proximal CpH methylation pattern. Shown in bottom represents abundant DNA motif near mCpHs (−1 bp to 4 bp) in mCpG-proximal region. (c) Methylation levels around highly expressed (red, higher 20%), lowly expressed (blue, lower 20%), and total (yellow) genes. The gene bodies and surround (±20% of those) were normalized into 1k-bins and extracted the average methylation levels at each bin, and processed by sliding window (window size = 30 bp). The transcription star site (TSS) and transcription terminate site (TTS) are in 142’th and 857’th bins, respectively, described as black dotted lines. (d) H3K36me3 peaks and mCpH levels around gene-body region of ubiquitin specific peptidase 22 (USP22), an epigenetic modifier that regulates embryonic stem cell differentiation60. This is printed by IGV61.
Next, we attempt to understand whether the activity of DNMT3s affects the correlation between CpH methylation and gene expression. In contrast with neurons and brains, the highly transcribed genes in ESC showed higher mCpH level in their gene-body regions (Fig. 4c). This tendency remained in DNMT3a-knockout ESC, however, almost disappeared in DNMT3b-knockout ESC. In addition, the hyper-methylation in highly expressed gene-body regions was greatly focused on CpHpG contexts, at which methylation mainly occurs by DNMT3b. Thus, we concluded that DNMT3b causes the positive correlation between mCpH level and gene expression level in ESCs.
To explore further, we focused on previous reports that the DNMT3b interacts with tri-methylated histone 3 lysine 36 (H3K36me3), enriched in highly expressed genes24, 35, assuming that the preferential interaction between DNMT3b and H3K36me3 causes the hyper-methylation at CpHs in highly expressed gene-body regions. To confirm this, we analysed the methylation at CACs and CAGs upon 7 histone modifications, known as markers of transcription regulatory regions such as enhancer, promoter, or gene-body (S. Table 3)36. Among the histone marks, the H3K36me3 was highly positively correlated with methylation at CAGs, compared to the methylation at CACs (S. Figure 5a; PCC equals to 0.34 and 0.15, respectively). The positive correlation became weak as DNMT3b knocked out (PCC equals to 0.21). In addition, we confirmed that CpH methylation is more positively correlated with the H3K36me3 overlapping rate (Method section), than with gene expression level (S. Figure 5b; Spearman’s rank correlation coefficient (ρ) equals to 0.73 and 0.33, respectively). Altogether, we concluded that the interaction between DNMT3b and H3K36me3 leads to the positive correlation between CpH methylation and gene expression in ESC.
We confirmed this with mouse ESCs, by analysing the mCpH level and expression level of genes in samples that DNMTs and SET domain containing 2 (SETD2), a catalyser of H3K36me37, were knocked out24 (S. Figure 5c). The mCpH level tended to be high at actively expressed gene bodies in wild type ESCs (ρ = 0.12), whereas the tendency disappeared as DNMT1, 3a, and 3b are knocked out (Corr. = −0.04). Remarkably, the positive correlation was recovered by reintroducing DNMT3b (Corr. = 0.02), whereas it was not by reintroducing DNMT3a (Corr. = −0.06). The positive correlation in DNMT3b-reintroduced sample disappeared as removing H3K36me3 marks by SETD2-knockout (Corr. = −0.15). Altogether, the results indicated that the positive correlation between CpH methylation and gene expression in ESC is caused by the activity of DNMT3b, preferentially methylates CAGs upon H3K36me3 marks that enriched in actively transcribed gene-body regions.
Lastly, we attempted to explore whether the methylation by interaction between H3K36me3 and DNMT3b is involved in ES-specific functions. We screened 418 genes of which CAGs in gene-body regions are hyper-methylated (M ei > 0.05, top 3% of all genes), and more than half of the gene-body regions are covered by H3K36me3 marks (\({histone\; overlapping\; rate}\) > 0.5, S. Figure 6a). The enriched gene ontology terms (GO terms, S. Table 4) of the genes were clustered into several representative biological processes such as mRNA metabolism, chromosome organization, and cellular response to DNA damage stimulus (S. Figure 6b, S. Table 5). These have been identified as being regulated by H3K36me3 marks in previous studies38,39,40, indicating that our data represents the properties of H3K36me3-enriched genes. Remarkably, our gene set included 13 genes related to embryo development (GO:0009790, P-value < 7.54 × 10−4, S. Table 6). The gene-body regions of the genes were clearly marked by both H3K36me3 marks and mCpHs in ESC, whereas these were not marked by mCpHs in brain tissue (Fig. 4d, S. Figure 7). The mCpHs disappeared in DNMT3b-knocked out ESC, indicating that these are introduced by DNMT3b. Thus, our data implied that the mCAGs, outcome of the preferential interaction between H3K36me3 and DNMT3b, marks genes involved in embryo development.
Discussion
Through comprehensive analysis of WGBS data, we uncovered the differential mechanism of CpH methylation in the two representative mCpH-containing mammalian cell types, ESC and neuron10.
WGBS has been considered as the only method to extract reliable information about mCpH, since other experiments such as microarray data41 or reduced representation bisulfite sequencing (RRBS)42 are mainly targeting CpG dinucleotides43. However, this method is financially and timely consuming, so that researchers had to deduce results from insufficient number of samples. The integrative read aligning strategy solved this problem by facilitating the use of public data with improved accuracy and reduced experimental artifacts. Through this integrative approach, we employed 21 human WGBSs and 8 mouse WGBSs including neurons, brains, ESCs, DNMT-knockout ESCs, and other somatic cells. The quality of the resulting dataset was verified by reproducing the known characteristics of mCpH in each cell type, and by clustering the samples into their originated cell types by genome-wide methylation pattern. The enlarged sample set by integrating public data, with improved detection quality, contributed to the statistical robustness of the following results.
Through analysing genome-wide methylation pattern, we confirmed that mCpHs are spatially correlated with mCpGs. Especially, these were co-regulated by DNMT3 when the distance between CpG and CpH is under ± 100 bp. This tendency was observed in both ESCs and neurons. Remarkably, however, there were distinct methylation patterns at CpHs in CpG-proximal regions ( ±100 bp from CpGs) between ESCs and neurons. Along with previously reported difference of the mCpHs in ESCs and neurons10, 31, 32, this distinct methylation pattern could be a marker for distinguishing the two cell types, and be a measure for the process of ESC differentiation to neuron.
We further explored that the differential distribution of mCpHs is caused by differential activities of DNMT3a and DNMT3b. By comparing genome-wide mCpH level in wild type and DNMT knockout ESCs, we found that DNMT3a preferentially methylates CAC contexts, whereas DNMT3b methylates CAG contexts. The differential targeting of DNMT3a and 3b, combined with their differential expression in ESCs and the brains, is suggested as the main reason of the distinct mCpH distribution in the two cell types. In ESCs, the hyperactivity of DNMT3b results in preferential methylation at CAGs, whereas in neurons, the DNMT3a is highly expressed, resulting in hypermethylation at CACs. Decisively, in DNMT3b-knockout ESCs, mCpH was enriched in CAC context than CAG, indicating that the enriched mCpH motif, CAG, is caused by DNMT3b. Additionally, the mCpG-proximal CpH methylation pattern in DNMT3b-knockout ESC was more similar to that in brain tissues and neurons, than to that in wild type ESCs, implying that the mCpG-proximal mCpH pattern in ESC is induced by the hyperactivity of DNMT3b. On the other hand, in neuron, the hyperactivity of DNMT3a results in preferential methylation at CAC motifs. Considering that MeCP2, a DNA-binding protein that broadly related to neuro-diseases, preferentially binds methylated CACs44, the CpH methylation by DNMT3a could be a key regulator of neuron-specific cellular processes. Altogether, we concluded that the differential activity of DNMT3a and DNMT3b induces the distinct distribution of mCpHs in ESCs and neurons. The molecular-level mechanism of the differential targeting by the enzymes remains as further study subject.
Finally, the positive correlation between CpH methylation and gene expression in ESCs was explained by the activity of DNMT3b. In a recent study24, DNMT3b showed preferential interaction with H3K36me3 mark in highly expressed gene bodies in ESCs. In our analysis, the methylated CAGs (mCAGs) showed spatial correlation with H3K36me3, and significantly accumulated in highly expressed gene-body region in human ESC. Decisively, the positive correlation between mCpH level and expression level disappeared when either DNMT3b or SETD2, catalysing H3K36me3 marks, was knocked out, indicating that the preferential interaction between DNMT3b and H3K36me3 marks causes the positive correlation between CpH methylation level and gene expression. Additionally, we confirmed that the methylation at CAGs, mediated by the interaction between DNMT3b and H3K36me3, marks genes involved in embryo development. This implies that the DNMT3b- H3K36me3 interaction, and/or the outcome of the interaction could be involved in ES-specific cellular processes. For example, recent study found that mCpGs, mediated by the interaction between DNMT3b and H3K36me3, prevent spurious transcription initiation in mouse ESC35. The mCAGs could be involved in fine-tuning of the process by the clearly marking actively transcribed gene-body regions, even though further study is necessary for proving it. On the other hand, in neuron and brain, the methylation at CpHs in gene-body regions could be hindered by other transcription factors (TFs) in same way with that at CpGs45, since DNMT3a, actively transcribed in neuron, showed little preference at gene-body regions or H3K36me3 marks24.
Based on these results, we suggest a differential CpH methylation mechanism by DNMT3a and DNMT3b in ESCs and neurons, which instructs further research directions (Fig. 5). Altogether, this study uncovered the mechanism underlying the differential distribution and functions of mCpH in ESCs and neurons, and provides insightful information regarding the cell type-specific CpH methylation mechanism.
Proposed differential methylation mechanism between ESCs and neurons. The illustration describes two main discovery about differential CpH methylation in ESCs and neurons. First, the DNMT3b is actively transcribed in ESCs, primarily methylating CAG contexts, whereas the DNMT3a is more transcribed in neuron, methylating CAC contexts. The methylated CACs are recognized by proteins related to brain diseases, such as MeCP244. Second, the DNMT3b preferentially interacts with H3K36me3 marks, abundant in actively transcribed gene-body regions, whereas DNMT3a showed little preference. The filled and unfilled circles represents methylated and unmethylated statuses, respectively.
Methods
WGBS data analysis
WGBS data were downloaded from NCBI Gene Expression Omnibus (GEO). We performed quality control by fastqx-toolkit46 with two steps. First, we trimmed tails of the reads of which quality score is under 20. Then, we removed reads of which length became less than half after the trimming or the average read quality is still under 20. Then, we mapped the high-quality bisulfite sequenced reads into the reference genomes (hg19 for human samples and mm10 for mouse samples) by three bisulfite-read aligners, Bismark26, BSMAP27, and BS-seeker228. Then, we removed duplicated reads by using Picard (http://broadinstitute.github.io/picard) (for the output of Bismark and BS-seeker2) and Samtools47 (for the output of BSMAP), following the suggestion from the manual of each bisulfite-read aligner. Then, we extracted cytosine positions covered by more than 5 reads, aligned by more than two aligners.
Methylation detection
Methylation levels at the cytosines were calculated as read-depth weighted average of those from individual aligner29. Then, the non-conversion rate was subtracted from the methylation levels at each cytosine, based on the previous statistical model5.
Where \({{Me}}_{i}\) is methylation level at cytosine \(i\), \({t}_{{ij}}\) is aligned read number at cytosine \(i\) by bisulfite-read aligner \({\rm{j}}\), and \({M}_{{ij}}\) is unconverted read number at cytosine \(i\) by bisulfite-read aligner \({\rm{j}}\). In case the \({{Me}}_{i}\) is under 0, we set M ei = 0.
Identification of methylated cytosines
Since the distribution of the methylation levels at CpGs and CpHs are different, we used different strategy for defining methylated or unmethylated status of the two contexts. The mCpG was defined as \(M{e}_{i}\,\geqq \,0.8\). The mCpH, used for calculating the probability that mCpH exists around CpGs (Result 2), was identified by a previously introduced method6. We used binomial distribution to detect the methylated cytosine loci. At every detected cytosine loci (\({\rm{i}}\)), we calculated the probability that methylated reads (\({k}_{i}\)) occur out of total read number (\({n}_{i}\)) based on binomial distribution with the success probability (\({\rm{p}}\)) as bisulfite non-conversion rate. If the p-value was under a certain threshold, we determined the cytosine loci as truly methylated.
In addition, we calculated false discovery rate (FDR) to eliminate false positive. We created an artificial methylome for every WGBS sample, in which read depth at each cytosine was equal to real data. Here, methylated read depth was generated following binomial distribution, using the bisulfite non-conversion rate as success probability. We calculated FDR by identifying methylated cytosines in artificial data, and finally set the p-value as 10−5, by which the FDR was under 0.01 within all WGBS samples.
Correlation analysis
We divided whole-genome into 1 k-bp blocks to compare the methylation pattern at CpG and CpH. Then, the correlation between CpG and CpH methylation levels was calculated across blocks that contain more than 10 CpGs and CpHs. In addition, to compare the mCpHs in Cis/Trans-strand to mCpG, we extracted blocks where the difference of mCpG level is over 0.5. Additionally, we measured the correlation between mCpG and mCpH levels in genic regions, using blocks of which more than 500 bp was covered by the genic regions. Genic regions were defined as follows: promoter defined as transcription start site ± 5000 bp, intragenic as all the regions of transcription start site (TSS) to transcription termination site (TTS), and intergenic as complementary of intragenic. The position information of TSS, TTS, and exon regions were obtained from RefSeq annotation. Lastly, to analyse the correlation based on the distance between CpGs and CpHs, we measured average mCpH level of every 10 bp from CpGs, to ±500 bp.
Gene expression
RNA-seq data were downloaded from NCBI GEO, and pre-processed along the same line with the WGBS data. The pre-processed reads were aligned by Tophat248, and the fragments per kilo-base of transcript per million mapped reads (FPKM) was extracted by Cufflink49. In addition, we collected reads per kilobase of transcript per million mapped reads (RPKMs), and FPKMs of DNMT3a and DNMT3b in brains50 and ESCs51,52,53, respectively, for statistical analysis (S. Table 2).
Histone marks
Histone ChIP-seq data were downloaded and mapped into the reference genome by Bowtie54. Then, MACS255 was used for peak calling with default parameters (P < 0.00001, mfold > 10). In addition, H3K27ac and H3K4me1 peaks were combined for defining putative enhancer regions. The rate of gene-body region overlapped by each histone mark was calculated as follow.
Gene ontology analysis
We analysed enriched gene ontology terms (GO terms) to explore approximate function of genes marked by both H3K36me3 and mCAGs. First, we selected 418 genes whose gene-body regions are highly methylated (mCAG level > 0.05, top <3% out of 25919 unduplicated genes), and are enriched with H3K36me3 marks (\({histone\; overlapping\; rate}\) > 0.5). Then, GOrilla56 was used for identifying enriched GO terms with P-value <10−3. The genes and enriched GO terms in HUES64 are described in S. Tables 4 and 5, respectively. Then, we used REVIGO57 for clustering GO terms, with similarity >0.5. The specifics for the clustered GO terms are described in S. Table 6. In all steps, we considered genes that length >1000 bp. In addition, gene symbols that start with “LOC” and “Rik” were not included in analysis since those are not recognized by GOrilla.
Statistics
The Wilcoxon signed rank test were used for determining significance, with P < 0.05. In addition, the Pearson’s correlation coefficient (PCC) was used for measuring correlation. All the boxes in boxplot boxes include values from 25% to 75% in order. The number of data points for drawing Figures and Extended Data Figures are described in S. Tables 4 and 5, respectively.
Data availability
The data that support the findings of this study are available in Gene Expression Omnibus (GEO). The identifiers for WGBS data are GSE1625625, GSE1731225, GSE479665, GSE4671021, GSE4664422, GSE6327812, GSE3226820, GSE6145723, GSE3020658, and GSE5741324, and those for RNA-seq data are GSE1625625, GSE479665, GSE3028058, GSE3056759, GSE2439952, and GSE7574853, and those for ChIP-seq data is GSE1625625. Statistics of the data are described in S. Tables 1 and S3. In addition, the genome-wide CpG and CpH methylome processed by three bisulfite-read aligners are browsable at Openlooper (https://openlooper.hgc.jp/), with identifiers described in S. Table 1.
References
Holliday, R. & Pugh, J. E. DNA modification mechanisms and gene activity during development. Science 187, 226–232 (1975).
Day, K. et al. Differential DNA methylation with age displays both common and dynamic features across human tissues that are influenced by CpG landscape. Genome Biol 14, R102, doi:https://doi.org/10.1186/gb-2013-14-9-r102 (2013).
Ziller, M. J. et al. Genomic distribution and inter-sample variation of non-CpG methylation across human cell types. PLoS Genet 7, e1002389, doi:https://doi.org/10.1371/journal.pgen.1002389 (2011).
Haines, T. R., Rodenhiser, D. I. & Ainsworth, P. J. Allele-specific non-CpG methylation of the Nf1 gene during early mouse development. Dev Biol 240, 585–598, doi:https://doi.org/10.1006/dbio.2001.0504 (2001).
Lister, R. et al. Global epigenomic reconfiguration during mammalian brain development. Science 341, 1237905, doi:https://doi.org/10.1126/science.1237905 (2013).
Guo, J. U. et al. Distribution, recognition and regulation of non-CpG methylation in the adult mammalian brain. Nature neuroscience 17, 215–222, doi:https://doi.org/10.1038/nn.3607 (2014).
Lister, R. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73, doi:https://doi.org/10.1038/nature09798 (2011).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322, doi:https://doi.org/10.1038/nature08514 (2009).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nature reviews. Genetics 11, 204–220, doi:https://doi.org/10.1038/nrg2719 (2010).
He, Y. & Ecker, J. R. Non-CG Methylation in the Human Genome. Annu Rev Genomics Hum Genet 16, 55–77, doi:https://doi.org/10.1146/annurev-genom-090413-025437 (2015).
Arand, J. et al. In vivo control of CpG and non-CpG DNA methylation by DNA methyltransferases. PLoS Genet 8, e1002750, doi:https://doi.org/10.1371/journal.pgen.1002750 (2012).
Liao, J. et al. Targeted disruption of DNMT1, DNMT3A and DNMT3B in human embryonic stem cells. Nat Genet 47, 469–478, doi:https://doi.org/10.1038/ng.3258 (2015).
Holz-Schietinger, C. & Reich, N. O. The inherent processivity of the human de novo methyltransferase 3A (DNMT3A) is enhanced by DNMT3L. J Biol Chem 285, 29091–29100, doi:https://doi.org/10.1074/jbc.M110.142513 (2010).
Gabel, H. W. et al. Disruption of DNA-methylation-dependent long gene repression in Rett syndrome. Nature 522, 89–93, doi:https://doi.org/10.1038/nature14319 (2015).
Chen, L. et al. MeCP2 binds to non-CG methylated DNA as neurons mature, influencing transcription and the timing of onset for Rett syndrome. Proc Natl Acad Sci USA 112, 5509–5514, doi:https://doi.org/10.1073/pnas.1505909112 (2015).
Butcher, L. M. et al. Non-CG DNA methylation is a biomarker for assessing endodermal differentiation capacity in pluripotent stem cells. Nat Commun 7, 10458, doi:https://doi.org/10.1038/ncomms10458 (2016).
Laurent, L. et al. Dynamic changes in the human methylome during differentiation. Genome research 20, 320–331, doi:https://doi.org/10.1101/gr.101907.109 (2010).
Xie, W. et al. Base-resolution analyses of sequence and parent-of-origin dependent DNA methylation in the mouse genome. Cell 148, 816–831, doi:https://doi.org/10.1016/j.cell.2011.12.035 (2012).
Siegfried, Z. & Simon, I. DNA methylation and gene expression. Wiley Interdiscip Rev Syst Biol Med 2, 362–371, doi:https://doi.org/10.1002/wsbm.64 (2010).
Martins-Taylor, K., Schroeder, D. I., LaSalle, J. M., Lalande, M. & Xu, R. H. Role of DNMT3B in the regulation of early neural and neural crest specifiers. Epigenetics 7, 71–82, doi:https://doi.org/10.4161/epi.7.1.18750 (2012).
Wen, L. et al. Whole-genome analysis of 5-hydroxymethylcytosine and 5-methylcytosine at base resolution in the human brain. Genome Biol 15, R49, doi:https://doi.org/10.1186/gb-2014-15-3-r49 (2014).
Ziller, M. J. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481, doi:https://doi.org/10.1038/nature12433 (2013).
Li, Z. et al. Distinct roles of DNMT1-dependent and DNMT1-independent methylation patterns in the genome of mouse embryonic stem cells. Genome Biol 16, 115, doi:https://doi.org/10.1186/s13059-015-0685-2 (2015).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247, doi:https://doi.org/10.1038/nature14176 (2015).
Chadwick, L. H. The NIH Roadmap Epigenomics Program data resource. Epigenomics 4, 317–324, doi:https://doi.org/10.2217/epi.12.18 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572, doi:https://doi.org/10.1093/bioinformatics/btr167 (2011).
Xi, Y. & Li, W. BSMAP: whole genome bisulfite sequence MAPping program. BMC bioinformatics 10, 232, doi:https://doi.org/10.1186/1471-2105-10-232 (2009).
Guo, W. et al. BS-Seeker2: a versatile aligning pipeline for bisulfite sequencing data. BMC genomics 14, 774, doi:https://doi.org/10.1186/1471-2164-14-774 (2013).
Lee, J. H., Park, S. J. & Kenta, N. An integrative approach for efficient analysis of whole genome bisulfite sequencing data. BMC genomics 16(Suppl 12), S14, doi:https://doi.org/10.1186/1471-2164-16-S12-S14 (2015).
Zvetkova, I. et al. Global hypomethylation of the genome in XX embryonic stem cells. Nat Genet 37, 1274–1279, doi:https://doi.org/10.1038/ng1663 (2005).
Shirane, K. et al. Mouse oocyte methylomes at base resolution reveal genome-wide accumulation of non-CpG methylation and role of DNA methyltransferases. PLoS Genet 9, e1003439, doi:https://doi.org/10.1371/journal.pgen.1003439 (2013).
Ferguson-Smith, A. C. & Greally, J. M. Epigenetics: perceptive enzymes. Nature 449, 148–149, doi:https://doi.org/10.1038/449148a (2007).
Patil, V., Ward, R. L. & Hesson, L. B. The evidence for functional non-CpG methylation in mammalian cells. Epigenetics 9, 823–828, doi:https://doi.org/10.4161/epi.28741 (2014).
Watanabe, D., Suetake, I., Tada, T. & Tajima, S. Stage- and cell-specific expression of Dnmt3a and Dnmt3b during embryogenesis. Mech Dev 118, 187–190 (2002).
Neri, F. et al. Intragenic DNA methylation prevents spurious transcription initiation. Nature 543, 72–77, doi:https://doi.org/10.1038/nature21373 (2017).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how, and why? Mol Cell 49, 825–837, doi:https://doi.org/10.1016/j.molcel.2013.01.038 (2013).
Zhang, Y. et al. H3K36 histone methyltransferase Setd2 is required for murine embryonic stem cell differentiation toward endoderm. Cell Rep 8, 1989–2002, doi:https://doi.org/10.1016/j.celrep.2014.08.031 (2014).
Dominguez, D. et al. A high-resolution transcriptome map of cell cycle reveals novel connections between periodic genes and cancer. Cell Res 26, 946–962, doi:https://doi.org/10.1038/cr.2016.84 (2016).
Chantalat, S. et al. Histone H3 trimethylation at lysine 36 is associated with constitutive and facultative heterochromatin. Genome research 21, 1426–1437, doi:https://doi.org/10.1101/gr.118091.110 (2011).
Wagner, E. J. & Carpenter, P. B. Understanding the language of Lys36 methylation at histone H3. Nat Rev Mol Cell Biol 13, 115–126, doi:https://doi.org/10.1038/nrm3274 (2012).
Morris, T. J. & Beck, S. Analysis pipelines and packages for Infinium HumanMethylation450 BeadChip (450k) data. Methods 72, 3–8, doi:https://doi.org/10.1016/j.ymeth.2014.08.011 (2015).
Meissner, A. et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic acids research 33, 5868–5877, doi:https://doi.org/10.1093/nar/gki901 (2005).
Bock, C. Analysing and interpreting DNA methylation data. Nature reviews. Genetics 13, 705–719, doi:https://doi.org/10.1038/nrg3273 (2012).
Lagger, S. et al. MeCP2 recognizes cytosine methylated tri-nucleotide and di-nucleotide sequences to tune transcription in the mammalian brain. PLoS Genet 13, e1006793, doi:https://doi.org/10.1371/journal.pgen.1006793 (2017).
Domcke, S. et al. Competition between DNA methylation and transcription factors determines binding of NRF1. Nature 528, 575–579, doi:https://doi.org/10.1038/nature16462 (2015).
Blankenberg, D. et al. Manipulation of FASTQ data with Galaxy. Bioinformatics 26, 1783–1785, doi:https://doi.org/10.1093/bioinformatics/btq281 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079, doi:https://doi.org/10.1093/bioinformatics/btp352 (2009).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14, R36, doi:https://doi.org/10.1186/gb-2013-14-4-r36 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28, 511–515, doi:https://doi.org/10.1038/nbt.1621 (2010).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399, doi:https://doi.org/10.1038/nature11405 (2012).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74, doi:https://doi.org/10.1038/nature11247 (2012).
Yang, L., Duff, M. O., Graveley, B. R., Carmichael, G. G. & Chen, L. L. Genomewide characterization of non-polyadenylated RNAs. Genome Biol 12, R16, doi:https://doi.org/10.1186/gb-2011-12-2-r16 (2011).
Chu, L. F. et al. Single-cell RNA-seq reveals novel regulators of human embryonic stem cell differentiation to definitive endoderm. Genome Biol 17, 173, doi:https://doi.org/10.1186/s13059-016-1033-x (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359, doi:https://doi.org/10.1038/nmeth.1923 (2012).
Liu, T. Use model-based Analysis of ChIP-Seq (MACS) to analyze short reads generated by sequencing protein-DNA interactions in embryonic stem cells. Methods Mol Biol 1150, 81–95, doi:https://doi.org/10.1007/978-1-4939-0512-6_4 (2014).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC bioinformatics 10, 48, doi:https://doi.org/10.1186/1471-2105-10-48 (2009).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6, e21800, doi:https://doi.org/10.1371/journal.pone.0021800 (2011).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495, doi:https://doi.org/10.1038/nature10716 (2011).
Deng, X. et al. Evidence for compensatory upregulation of expressed X-linked genes in mammals, Caenorhabditis elegans and Drosophila melanogaster. Nat Genet 43, 1179–1185, doi:https://doi.org/10.1038/ng.948 (2011).
Sussman, R. T. et al. The epigenetic modifier ubiquitin-specific protease 22 (USP22) regulates embryonic stem cell differentiation via transcriptional repression of sex-determining region Y-box 2 (SOX2). J Biol Chem 288, 24234–24246, doi:https://doi.org/10.1074/jbc.M113.469783 (2013).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14, 178–192, doi:https://doi.org/10.1093/bib/bbs017 (2013).
Acknowledgements
This work was partially supported by Japan Society for the Promotion of Science (JSPS) KAKENHI Grant Number 16H01403. Computational resources were provided by the supercomputer system at the Human Genome Center, the Institute of Medical Science, the University of Tokyo. We thank to Dr. Alexander Meissner, in Harvard Stem Cell Institute, USA, for feedbacks and advices.
Author information
Authors and Affiliations
Contributions
J.-H.L. designed and conducted the data analyses. J.-H.L. and S.-J.P. wrote the manuscript. K.N. conceived and designed the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, JH., Park, SJ. & Nakai, K. Differential landscape of non-CpG methylation in embryonic stem cells and neurons caused by DNMT3s. Sci Rep 7, 11295 (2017). https://doi.org/10.1038/s41598-017-11800-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-11800-1
This article is cited by
-
Investigating neural differentiation of mouse P19 embryonic stem cells in a time-dependent manner by bioinformatic, microscopic and transcriptional analyses
Molecular Biology Reports (2023)
-
Differential expression of m5C RNA methyltransferase genes NSUN6 and NSUN7 in Alzheimer’s disease and traumatic brain injury
Molecular Neurobiology (2023)
-
Developmental and Injury-induced Changes in DNA Methylation in Regenerative versus Non-regenerative Regions of the Vertebrate Central Nervous System
BMC Genomics (2022)
-
Sperm cryopreservation and DNA methylation: possible implications for ART success and the health of offspring
Journal of Assisted Reproduction and Genetics (2022)
-
Proteins in DNA methylation and their role in neural stem cell proliferation and differentiation
Cell Regeneration (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.