Abstract
Interpretations of genetic data concerning the prehistory of Europe have long been a subject of great debate, but increasing amounts of ancient and modern DNA data are now providing new and more informative evidence. Y-chromosome resequencing studies in Europe have highlighted the prevalence of recent expansions of male lineages, and focused interest on the Bronze Age as a period of cultural and demographic change. These findings contrast with phylogeographic studies based on mitochondrial DNA (mtDNA), which have been interpreted as supporting expansions from glacial refugia. Here we have undertaken a population-based resequencing of complete mitochondrial genomes in Europe and the Middle East, in 340 samples from 17 populations for which Y-chromosome sequence data are also available. Demographic reconstructions show no signal of Bronze Age expansion, but evidence of Paleolithic expansions in all populations except the Saami, and with an absence of detectable geographical pattern. In agreement with previous inference from modern and ancient DNA data, the unbiased comparison between the mtDNA and Y-chromosome population datasets emphasizes the sex-biased nature of recent demographic transitions in Europe.
Similar content being viewed by others
Introduction
Our understanding of European prehistory has been revolutionized by the availability of new DNA sequencing technologies1, which have allowed the unbiased characterization of sequence variation in modern and ancient human genomes. Genome-wide ancient DNA (aDNA) data have shown a clear discontinuity between Paleolithic hunter-gatherers and Neolithic farmers2,3,4,5. Patterns of diversity suggested low Paleolithic population sizes, with regional differences among Western and Scandinavian groups6. This picture has been further refined by the study of the DNA of ancient Yamnaya herders from the region of the Pontic-Caspian steppe, the apparent source of Bronze Age migrations into Europe and Asia7,8,9, and a debated region of origin for Indo-European languages10. These studies highlight an introgression into Europe at around 4.5 thousand years ago (KYA) from the East, followed by development of genetic structure in Bronze Age Europe7,8.
Ideas on European prehistory have been strongly influenced by studies not only of autosomal DNA, but also of uniparentally-inherited markers, which can provide information about sex-biased processes11. Analyses of aDNA show that today’s most frequent Y-chromosome haplogroup (R1b-M269) is very rare in Europe until 4.5 KYA5 (see summary elsewhere12), while it is present in all the Yamnaya samples8,9. This had initially suggested a major introgression of males from the Pontic-Caspian steppe; however, the R1b sublineage (R1b-L11) now common in Europe has not so far been found among Yamnaya sequences13. In contrast to Y data, patterns of ancient mtDNA reveal a period of widespread turnover of lineages in the Late Glacial period ~14.5 KYA14,15 and support the picture of a discontinuity between Paleolithic hunter-gatherers and Neolithic farmers seen in autosomal data, but contain no apparent signal of the Bronze Age steppe expansion16. Ancient DNA data on uniparentally-inherited markers therefore suggest a strong sex-bias in recent demographic changes in Europe.
Resequencing of the male-specific region of the Y chromosome (MSY)17 in modern European populations also emphasizes the importance of Bronze-Age transition12,18,19,20. Demographic reconstructions support an expansion starting ~2.1–4.2 KYA, and times-to-most-recent-common-ancestor (TMRCAs) of three major haplogroups, including R1b, are estimated between 3.5 and 7.3 KYA12. Many mtDNA studies have been undertaken in modern European samples, but most concentrate on particular segments of the mitochondrial genome21, consider the European continent as a single unit18,19, or have taken a phylogeographic approach, focusing on specific lineages of interest22,23,24,25,26,27. General conclusions have been that current mtDNA variation represents re-expansions from glacial refugia. However, population-based whole mtDNA resequencing studies in Europe are lacking, so a systematic comparison of the demographic histories of males and females has yet not been possible.
Here, we carry out a population-based study, resequencing mtDNA in a set of 17 populations for which MSY resequencing data are already available. The pattern observed in mtDNA is strikingly different from that in MSY, compatible with expansion after the Last Glacial Maximum, and emphasizing the male-specific nature of the Bronze Age expansion in Europe.
Results
To allow a comparison between female and male histories, we resequenced the mitochondrial genomes of 340 European and Middle Eastern individuals belonging to 17 populations (Table S1) that were previously analyzed for MSY12.
We constructed a maximum parsimony (MP) tree (Fig. 1a; see also median-joining network in Figure S1) based on the mtDNA coding region only, which is best suited for reliable phylogenetic inference due to its relatively low content of recurrently mutating sites (Figure S2 presents a median-joining network based on the entire sequence). We also determined haplogroups from sequences (Table S1): their frequencies and geographical distributions (Fig. 1b) are broadly consistent with previous data21.
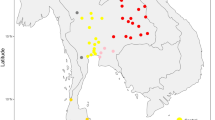
Phylogenies and geographical distributions of European mtDNA and MSY lineages. (a) Maximum-parsimony mtDNA tree, based on resequencing of 15,447 bp of the coding region. Branch lengths are proportional to molecular divergence among haplotypes, and colours indicate haplogroups. Point estimates of TMRCAs in KY are given in parentheses after haplogroup names; see also Table 1. (b) Map with pie-charts showing frequencies of mtDNA haplogroups (defined and coloured as in part (a)) in 17 populations from Europe and the Near East. Population abbreviations are as follows: bas: Spanish Basque country; bav: Bavaria (Germany); CEU: Utah residents with Northern and Western European ancestry from the CEPH collection (France); den: Denmark; eng: England;41,42 fri: Frisia (Netherlands); gre: Greece; hun: Hungary; ire: Ireland; nor: Norway; ork: Orkney;41,42 pal: Palestinians; saa: Saami (Finland); ser: Serbia; spa: central Spain; TSI: Toscani in Italia (Italy); tur: Turkey. Map from Mountain High Map Frontiers™ version 94.01 (Mountain High Maps® Copyright © 1993 Digital Wisdom®, Inc.; www.digiwis.com/dwi_frl.htm). (c) Maximum-parsimony MSY tree, based on resequencing of 3.7 Mb in each individual12. Branch lengths are proportional to molecular divergence among haplotypes, and colours indicate haplogroups. Point estimates of TMRCAs are given in parentheses after haplogroup names. (d) Map with pie-charts showing frequencies of MSY haplogroups (defined and coloured as in part (c)) in 17 populations from Europe and the Near East12. Population abbreviations are as in part (b). Map from Mountain High Map Frontiers™ version 94.01 (Mountain High Maps® Copyright © 1993 Digital Wisdom®, Inc.; www.digiwis.com/dwi_frl.htm).
In the overall dataset, the main haplogroups observed are H (34.1%), U (17.9%), T (13.5%), J (9.1%), K (7.3%), and V (5.3%). The remaining 12.6% of the dataset is comprised of many minor haplogroups; these include (in the Palestinian and Spanish samples) two examples of each of the lineages L2 and L3, typically found in sub-Saharan Africa28 (see also YRI data in Table S1).
Visual inspection of the distributions of haplogroups (Fig. 1b) shows the Saami to be a clear outlier with low diversity, dominated by haplogroups U5 and V, and including a single example of haplogroup D, found mostly in north and east Asia29. In the remaining populations there is no obvious geographical pattern, in agreement with previous observations based on analyses of specific segments of the mitochondrial genome21,30.
This phylogeography of mitochondrial genomes was compared with that of MSY, based on resequencing of 3.7 Mb of Y-DNA in the same set of samples12 (Fig. 1c,d). The geographical distributions of haplogroups are more localized for MSY (Fig. 1d) than for mtDNA (Fig. 1b): for example, R1b is at particularly high frequency in the northwest, R1a in north and central Europe, and J2 in the south. Although the two MP trees have very different scales, due to the much smaller number of nucleotides analyzed in mtDNA compared to MSY (with a ratio of 1:250), both show deep-rooting clades (mtDNA: haplogroups U, K and T2; MSY: E1b-M35, G2a-L31, I2-P215, J2-M172, L and T), as well as star-like clades, indicative of population expansions. For mtDNA, these clades are H, J1, T1, V, representing 51.7% of our sample, while for MSY they are I1-M253, N1c-M178, R1a-M198 and R1b-M269 (taken together, 64%). However, the major difference is in the relative lengths of the internal branches, which indicate that the expansions of mtDNA lineages are more ancient than those of MSY lineages. This is supported by TMRCA point estimates (based on the entire mitochondrial genome) for the star-like haplogroups (Fig. 1a,c; Table 1), which are all ≤6 KYA (post-Neolithic) for the MSY12, but >13 KYA (Paleolithic) for mtDNA.
To consider populations, rather than lineages, we reconstructed demographic histories by using Bayesian Skyline Plots (BSPs) based on mtDNA sequences (Fig. 2). As expected from their unusual haplogroup composition, the Saami also represent an outlier here, showing a steady decline in effective population size that becomes more marked at around 5 KYA. All other populations show a signature of Paleolithic expansion, between 13 and 20 KYA. The Turkish and Palestinian samples differ from the majority in showing considerably more ancient population expansion, at >40 KYA. These patterns contrast sharply with the BSPs for MSY12 in the same populations (Fig. 2), which in most cases (13/17 populations) show demographic histories featuring a minimum effective population size around 3 KYA (late Bronze Age for many of the populations studied), followed by rapid expansion to the present. In all comparisons except those in Basques and Danish, current point estimates of effective population size are higher for mtDNA than for MSY.
We also calculated diversity indices for each population (Table 2). In agreement with the observation of a limited number of haplogroups in Saami, and the corresponding BSP, this population shows the lowest value for all diversity measures. The highest values are seen in the Palestinian and Turkish samples, which again is concordant with the ancient population growth seen in the BSPs. We observe negative values of both D and FS for all populations except the Saami, which can indicate population growth; however, both values are significant for only twelve of the remaining populations. At a glance, there appears to be more diversity in southern than northern populations (Fig. 1b; Table 2). To formally test this, we carried out a correlation analysis between genetic diversity (number of polymorphic sites, and nucleotide diversity) and latitude, longitude and overland distance from the Franco-Cantabrian and Near-Eastern glacial refugia (Table S2). When all populations are included, both measures show a statistically significant correlation with latitude and distance from the Near-Eastern refugium, but not with longitude. These correlations are lost when we remove the outlier populations described above (Saami, Turkish and Palestinian), demonstrating the lack of any pattern of mtDNA diversity in most of Europe.
Discussion
The study of mtDNA in Europe has a long history and has been influential in developing hypotheses about the origins of modern Europeans15,31. Early population-based studies involved sequencing of the control region32, and later the typing of specific haplogroup-defining SNPs in the coding region21. Analyses of whole mitochondrial genome sequences have focused on specific haplogroups, often in the framework of the re-peopling of Europe after the last glacial maximum23,24,25. At the population scale, the only previous studies have considered Europe at the continental level18,19, and with low sample sizes (n = 86 and 81 respectively), and patchy population coverage.
Here, we have carried out whole mtDNA sequencing in a European and Middle Eastern population-based sample set of 340 individuals, in which MSY resequencing12 had previously been undertaken. The spectrum of haplogroups we observe (Table S1, Fig. 1a) is compatible with previously published data21. The population-based design of this study and the unbiased nature of variant ascertainment means that European mtDNA and MSY diversity can be compared fairly. The phylogenies and demographic reconstructions concur in showing a marked difference between female and male population histories, with Paleolithic expansions for mtDNA contrasting with Bronze Age expansions for MSY. While this is in agreement with continental-level differences observed previously18,19, here we also show that this difference holds for most of the individual populations, and reflects a lack of geographical pattern in Europe. The most ancient mtDNA expansions we detect, dating close to the early peopling of Eurasia (40–50 KYA), are in the Near and Middle East. This difference in timing of European female and male lineage expansions is mirrored in the Indian subcontinent, where a recent analysis33 shows that mtDNA expansions reflect processes in the pre-Holocene era, while MSY expansions are mostly in the last 10 KYA, with marked male-driven spread from Central Asia during the Bronze Age.
Since mtDNA analysis has forensic utility34, it is worth noting that the 340 individuals carry only 318 distinct haplotypes based on complete mitochondrial genomes (Figure S2; Table S3), emphasizing the relatively low discriminatory power of mtDNA sequencing, even at its maximum resolution. We observe 12 identical pairs of haplotypes, three trios, and one example of a haplotype found five times in the dataset. These cases represent within-population sharing (in Danish, Norwegian, Orcadian, Frisian, Basque and Saami), with one exception, a haplotype within hg V shared between two Saami and a Spanish individual (Figure S2b) – reflecting a connection previously noted35. As reflected in diversity measures (Table 2) the Saami have particularly high haplotype sharing (two pairs, two trios, one quintet). These findings emphasize the importance of large and appropriate reference databases in forensic analysis.
The outlier status of the Saami in our dataset of 17 populations is clear not only from the high frequencies of closely related mtDNA sequences (Figure S2), but also haplogroups for both mtDNA and MSY that are rare elsewhere in the dataset (Fig. 1b,d), as well as examples of MSY sequences36 and Y-STR haplotypes36 that are found in more than one individual. These features are in agreement with the lack of growth in effective population size seen in the BSP (Fig. 2). Population-based genome-wide SNP analysis37 and whole-genome sequencing of a single individual38 also show the Saami to be genetically differentiated compared to Europeans, and to carry East Asian ancestry components.
Our data are consistent with ancient DNA data14,15,16 in supporting sex-biased processes in recent European demographic changes: patterns of modern mtDNA diversity show no signal of the Bronze Age expansion, while much of the modern European MSY diversity has been shaped by this process12. However, the modern data differ in showing no clear signal of the Neolithic transition that has been highlighted in ancient mitochondrial and autosomal data5,16. This could be due to drift, which is important in shaping the observed patterns of diversity in uniparental markers, and also sampling effects.
Much progress has been made in understanding the prehistory of the European continent since the first classical genetic data were interpreted in favour of agriculturally-mediated demic diffusion39. A wealth of both modern and ancient DNA data is now available, and has highlighted previously unsuspected past migration and expansion events, with a sex-biased aspect supported by our population-based resequencing approach. However, there is still much future work to be done in increasing sample sizes and geographical coverage, and in fully integrating the ancient and modern data to test explicitly the complex scenarios they suggest.
Materials and Methods
DNA samples and sequencing
DNA samples from 340 individuals belonging to 17 populations (20 individuals each) across Europe and the Near East were used for analysis, as in our previous study of MSY diversity12. Populations were as follows: Greek, Serbian, Hungarian, German [Bavaria], Spanish Basque, central Spanish, French (Centre d’Etude du Polymorphisme Humain [CEPH] collection in Utah, USA, with ancestry from Northern and Western Europe40 [CEU]), Italian (Toscani in Italia40 [TSI]), Dutch [Frisia], Danish, Norwegian, Finnish [Saami], English41,42 [Herefordshire and Worcestershire, Gloucestershire, Oxfordshire, Forest of Dean], Orcadian41,42, Irish, Turkish and Palestinian. Twenty individuals from each of two additional HapMap40 population samples, CHB (Han Chinese in Beijing, China), and YRI (Yoruba in Ibadan, Nigeria) were included to provide variant validation data. All methods were carried out in accordance with relevant guidelines and regulations, and all experimental protocols were approved by the University of Leicester Research Ethics Committee. Informed consent was obtained from all subjects (University of Leicester Research Ethics Committee reference: maj4-cb66).
We generated three datasets using parallel sequencing strategies (based on Illumina HiSeq, Illumina MiSeq and Ion Torrent PGM technologies [Table S4]) and bioinformatic pipelines (Table S5), and validated variants by comparison with independent sequence and SNP-genotype data (Table S6). These three datasets were merged for all the subsequent evolutionary analyses. The 380 sequences are available in Supplementary Dataset S1.
Tree construction and haplogroup prediction
A maximum parsimony (MP) tree was constructed from coding-region sequences (positions 576-16,023) via MEGA643, using the Subtree-Pruning-Regrafting (SPR) algorithm44 with search level 0 in which the initial trees were obtained by the random addition of sequences (10 replicates). Branch lengths were calculated using the average pathway method44 and are proportional to the number of mutations. FigTree v1.4.045 was used for tree visualization. Haplogroups were predicted using HaploGrep246, and their phylogenetic coherence was verified using the tree, with manual examination of possible ‘phantom’ mutations, as inferred using Haplogrep246 (Table S7).
Median-joining networks47 based on either coding region or whole mtDNA sequences were constructed using Network 5.0.0.0, and represented using Network Publisher 2.1.1.2. Polymorphic sites were weighted according to their evolutionary rates using the parameters suggested in the literature48.
Haplogroups were defined according to Phylotree1649 by using HaploGrep50 and their relative frequencies represented as pie-charts plotted on a geographical map.
TMRCA estimation
TMRCAs of nodes of interest were estimated51 using BEAST v1.8.0. MCMC samples were based on 50,000,000 generations, logging every 1000 steps, with the first 5,000,000 generations discarded as burn-in. Three runs were combined for analysis using LogCombiner. We used an exponential growth coalescent tree prior, HKY substitution model, and an uncorrelated relaxed clock with a lognormal distribution for mutation rate (2.21 ± 0.17 × 10−8 mutations/nucleotide/year52). TMRCAs were estimated in a single run including all 17 populations and assigning samples to specific clades in agreement with the MP tree shown in Fig. 1. For this analysis, the entire mitochondrial genome was considered: given the timeframe of interest (<40 KYA), the rate and its standard error were adjusted by using the calculator52 in Soares et al. (2009) to infer the rate at each of the nodes of interest (see Table 1). The median of this distribution of values was used for estimating TMRCAs for the haplogroups of interest.
Bayesian skyline plots
BSPs51 were generated using BEAST v1.8.0. MCMC samples were based on 30,000,000 generations, logging every 1000 steps, with the first 3,000,000 generations discarded as burn-in. We used a piecewise linear skyline model with ten groups, a HKY substitution model, and an uncorrelated relaxed clock with a lognormal distribution for mutation rate (2.21 ± 0.17 × 10−8 mutations/nucleotide/year52) and a generation time of 30 years53,54. For this analysis, the entire mitochondrial genome was considered: given the timeframe of interest (<40 KYA) the rate and its standard error were adjusted by using the calculator52 in Soares et al. (2009) to infer the rate at each of the nodes of interest (see Table 1). The median of this distribution of values was used for estimating TMRCAs for the haplogroups of interest.
Intrapopulation diversity and geographical correlation
The number of polymorphic sites per population (S), nucleotide diversity, Tajima’s D 55, and Fu’s FS 56 were calculated57 using Arlequin 3.5. Correlations of S, and of nucleotide diversity, with latitude, longitude, and distances from glacial refugia were examined using the function cor.test of the package stats within R. The locations Anamur, Turkey (36.1°, 32.8°) and Fleurac, France (45.0°, 1.0°) were taken as proxies for the centres of the Near-Eastern and Franco-Cantabrian refugia respectively. Distances account for geographical barriers, and were estimated using the land transport distance tool at www.freemaptools.com.
Data availability
All data generated during this study are included in this article (and its Supplementary Information files).
References
Veeramah, K. R. & Hammer, M. F. The impact of whole-genome sequencing on the reconstruction of human population history. Nat Rev Genet 15, 149–162 (2014).
Skoglund, P. et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012).
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014).
Lazaridis, I. et al. Genomic insights into the origin of farming in the ancient Near East. Nature 536, 419–424 (2016).
Fu, Q. et al. The genetic history of Ice Age Europe. Nature 534, 200–205 (2016).
Sanchez-Quinto, F. et al. Genomic affinities of two 7,000-year-old Iberian hunter-gatherers. Curr Biol 22, 1494–1499 (2012).
Allentoft, M. E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Mathieson, I. et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528, 499–503 (2015).
Gimbutas M. Proto-Indo-European culture: the Kurgan culture during the fifth, fourth and third millennia B.C. In: Indo-European and Indo-Europeans (ed^(eds Cardona G, Hoenigswald HM, Senn A). University of Pennsylvania Press (1970).
Batini, C. & Jobling, M. A. Detecting past male-mediated expansions using the Y chromosome. Hum Genet 136, 547–557 (2017).
Batini, C. et al. Large-scale recent expansion of European patrilineages shown by population resequencing. Nat Commun 6, 7152 (2015).
Balanovsky, O. et al. Genetic differentiation between upland and lowland populations shapes the Y-chromosomal landscape of West Asia. Hum Genet 36, 437–450 (2017).
Posth, C. et al. Pleistocene mitochondrial genomes suggest a single major dispersal of Non-Africans and a late glacial population turnover in Europe. Curr Biol 26, 827–833 (2016).
Richards, M. B., Soares, P. & Torroni, A. Palaeogenomics: mitogenomes and migrations in Europe’s past. Curr Biol 26, R243–246 (2016).
Brandt, G. et al. Ancient DNA reveals key stages in the formation of central European mitochondrial genetic diversity. Science 342, 257–261 (2013).
Jobling, M. A. & Tyler-Smith, C. Human Y-chromosome variation in the genome-sequencing era. Nat Rev Genet 18, 485–497 (2017).
Karmin, M. et al. A recent bottleneck of Y chromosome diversity coincides with a global change in culture. Genome Res 25, 459–466 (2015).
Lippold, S. et al. Human paternal and maternal demographic histories: insights from high-resolution Y chromosome and mtDNA sequences. Investig Genet 5, 13 (2014).
Poznik, G. D. et al. Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences. Nat Genet 48, 593–599 (2016).
Richards, M. et al. Tracing European founder lineages in the near eastern mtDNA pool. Am J Hum Genet 67, 1251–1276 (2000).
Achilli, A. et al. The molecular dissection of mtDNA haplogroup H confirms that the Franco-Cantabrian glacial refuge was a major source for the European gene pool. Am J Hum Genet 75, 910–918 (2004).
Behar, D. M. et al. The Basque paradigm: genetic evidence of a maternal continuity in the Franco-Cantabrian region since pre-Neolithic times. Am J Hum Genet 90, 486–493 (2012).
Olivieri, A. et al. Mitogenomes from two uncommon haplogroups mark late glacial/postglacial expansions from the near east and neolithic dispersals within Europe. PLoS ONE 8, e70492 (2013).
Pala, M. et al. Mitochondrial DNA signals of late glacial recolonization of Europe from near eastern refugia. Am J Hum Genet 90, 915–924 (2012).
Pereira, L. et al. High-resolution mtDNA evidence for the late-glacial resettlement of Europe from an Iberian refugium. Genome Res 15, 19–24 (2005).
Pereira, J. B. et al. Reconciling evidence from ancient and contemporary genomes: a major source for the European Neolithic within Mediterranean Europe. Proc Biol Sci 284, 20161976 (2017).
Rosa, A. & Brehem, A. African human mtDNA phylogeography at-a-glance. J Anthropol Sci 89, 25–58 (2011).
Stoneking, M. & Delfin, F. The human genetic history of East Asia: weaving a complex tapestry. Curr Biol 20, R188–193 (2010).
Richards, M. et al. Paleolithic and neolithic lineages in the European mitochondrial gene pool. Am J Hum Genet 59, 185–203 (1996).
Pinhasi, R., Thomas, M. G., Hofreiter, M., Currat, M. & Burger, J. The genetic history of Europeans. Trends Genet 28, 496–505 (2012).
Richards, M. B., Macaulay, V. A., Bandelt, H. J. & Sykes, B. C. Phylogeography of mitochondrial DNA in western Europe. Ann Hum Genet 62, 241–260 (1998).
Silva, M. et al. A genetic chronology for the Indian Subcontinent points to heavily sex-biased dispersals. BMC Evol Biol 17, 88 (2017).
Melton, T., Holland, C. & Holland, M. Forensic Mitochondrial DNA Analysis: Current Practice and Future Potential. Forensic Sci Rev 24, 101–122 (2012).
Torroni, A. et al. A signal, from human mtDNA, of postglacial recolonization in Europe. Am J Hum Genet 69, 844–852 (2001).
Hallast, P. et al. The Y-chromosome tree bursts into leaf: 13,000 high-confidence SNPs covering the majority of known clades. Mol Biol Evol 32, 661–673 (2015).
Huyghe, J. R. et al. A genome-wide analysis of population structure in the Finnish Saami with implications for genetic association studies. Eur J Hum Genet 19, 347–352 (2011).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Menozzi, P., Piazza, A. & Cavalli-Sforza, L. L. Synthetic maps of human gene frequencies in Europeans. Science 201, 786–792 (1978).
International HapMap Consortium. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52-58 (2010).
Winney, B. et al. People of the British Isles: preliminary analysis of genotypes and surnames in a UK-control population. Eur J Hum Genet 20, 203–210 (2012).
Leslie, S. et al. The fine-scale genetic structure of the British population. Nature 519, 309–314 (2015).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30, 2725–2729 (2013).
Nei M, Kumar S. Molecular Evolution and Phylogenetics. Oxford University Press (2000).
Rambaut A. Fig.Tree. Tree Figure Drawing Tool, version 1.4.0. Available at: http://tree.bio.ed.ac.uk/software/figtree/. (ed^(eds) (2006-2012).
Weissensteiner, H. et al. HaploGrep 2: mitochondrial haplogroup classification in the era of high-throughput sequencing. Nucleic Acids Res 44, W58–W63 (2016).
Bandelt, H.-J., Forster, P. & Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16, 37–48 (1999).
Kong, Q. P. et al. Distilling artificial recombinants from large sets of complete mtDNA genomes. PLoS ONE 3, e3016 (2008).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum Mutat 30, E386–394 (2009).
Kloss-Brandstatter, A. et al. HaploGrep: a fast and reliable algorithm for automatic classification of mitochondrial DNA haplogroups. Hum Mutat 32, 25–32 (2011).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol 29, 1969–1973 (2012).
Soares, P. et al. Correcting for purifying selection: an improved human mitochondrial molecular clock. Am J Hum Genet 84, 740–759 (2009).
Fenner, J. N. Cross-cultural estimation of the human generation interval for use in genetics-based population divergence studies. Am J Phys Anthropol 128, 415–423 (2005).
Moorjani, P. et al. A genetic method for dating ancient genomes provides a direct estimate of human generation interval in the last 45,000 years. Proc Natl Acad Sci USA 113, 5652–5657 (2016).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Fu, Y. X. & Li, W. H. Statistical tests of neutrality of mutations. Genetics 133, 693–709 (1993).
Excoffier, L. & Lischer, H. E. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10, 564–567 (2010).
Acknowledgements
We thank Eduardo Arroyo-Pardo, Walter Bodmer, Gianpiero Cavalleri, Peter de Knijff, Berit Myhre Dupuy, Turi King, Adolfo López de Munain, Ana López-Parra, Aphrodite Loutradis, Jelena Milasin, Andrea Novelletto, Aslıhan Tolun, and Bruce Winney for assistance with DNA samples, and helpful comments; Lorna Gregory and the Oxford Genomics Centre for library preparation, target enrichment and sequencing; Rita Neumann and Pierpaolo Maisano Delser for help with IonTorrent sequencing; NUCLEUS Genomic Services at the University of Leicester for access to Illumina MiSeq sequencing; and two anonymous reviewers for helpful comments on the manuscript. CB, PH, DZ and MAJ were supported by a Wellcome Trust Senior Fellowship, grant number 087576. AS was supported by the Finnish Foundations’ Professor Pool (Paulo Foundation). This research used the ALICE High Performance Computing Facility at the University of Leicester.
Author information
Authors and Affiliations
Contributions
Conceived the project: M.A.J., C.B. Provided DNA samples: H.A.E., H.P., A.S. Designed the sequencing experiments: P.H., C.B., D.Z., M.A.J. Carried out the sequencing: C.B., Å.J.V. Carried out data analysis: C.B., P.H., D.Z., Å.J.V., J.H.W. Wrote the manuscript: C.B., M.A.J. All authors read the manuscript and provided critical input.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Batini, C., Hallast, P., Vågene, Å.J. et al. Population resequencing of European mitochondrial genomes highlights sex-bias in Bronze Age demographic expansions. Sci Rep 7, 12086 (2017). https://doi.org/10.1038/s41598-017-11307-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-11307-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.