Abstract
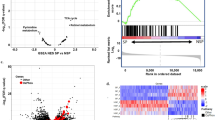
Prostate cancer (PCa) is one of the leading cancers in men in the USA. Lack of experimental tools that predict therapy response is one of the limitations of current therapeutic regimens. Mitochondrial dysfunctions including defective oxidative phosphorylation (OXPHOS) in cancer inhibit apoptosis by modulating ROS production and cellular signaling. Thus, correction of mitochondrial dysfunction and induction of apoptosis are promising strategies in cancer treatment. We have used Fluorescence Lifetime Imaging Microscopy (FLIM) to quantify mitochondrial metabolic response in PCa cells by tracking auto-fluorescent NAD(P)H, FAD and tryptophan (Trp) lifetimes and their enzyme-bound fractions as markers, before and after treatment with anti-cancer drug doxorubicin. A 3-channel FLIM assay and quantitative analysis of these markers for cellular metabolism show in response to doxorubicin, NAD(P)H mean fluorescence lifetime (τm) and enzyme-bound (a2%) fraction increased, FAD enzyme-bound (a1%) fraction was decreased, NAD(P)H-a2%/FAD-a1% FLIM-based redox ratio and ROS increased, followed by induction of apoptosis. For the first time, a FRET assay in PCa cells shows Trp-quenching due to Trp-NAD(P)H interactions, correlating energy transfer efficiencies (E%) vs NAD(P)H-a2%/FAD-a1% as sensitive parameters in predicting drug response. Applying this FLIM assay as early predictor of drug response would meet one of the important goals in cancer treatment.
Similar content being viewed by others
Introduction
Fluorescence Lifetime Imaging Microscopy (FLIM) is a powerful tool to assess the metabolic state of cells and tissues under different pathophysiological conditions using the auto-fluorescent properties of metabolic co-enzymes NAD(P)H and FAD1, 2. Both are predominantly located in the mitochondrial tricarboxylic acid cycle (TCA) and electron transfer chain (ETC) – FAD exclusively – producing ATP and reactive oxygen species (ROS) as part of the apoptosis pathway; NAD(P)H is also present in the cytosolic glycolysis pathway, important for highly proliferating cells like cancer1, 3, 4. The glycolysis pathway in the cytosol releases NADH and contributes to the free NADH pool. Mitochondrial oxidative phosphorylation (OXPHOS) activity consumes NADH (increased NADH-enzyme-bound fraction) and produces FAD (diminished FAD enzyme-bound fraction). Both the co-enzymes in their reduced (NAD(P)H and FADH2) and oxidized (NAD(P)+ and FAD) forms participate in the cellular oxidation-reduction reactions critical for cell physiology. Since the fluorescence spectra of NADH and NADPH cannot be readily distinguished5, we have chosen to use NAD(P)H through this manuscript.
The fluorescence lifetimes of NAD(P)H and FAD are sensitive to changes in pH, temperature, their conformational state and proximity to quenchers6. These co-enzymes exist in “free” or “enzyme-bound” states during cellular metabolic activity. FLIM is a very sensitive tool which allows discriminating the lifetimes and their relative fractions of free and enzyme-bound states of the co-enzymes from their fluorescence lifetime decay curve. Typically, fitting of lifetime decays of NAD(P)H and FAD are based on a two-component exponential decay model6. The shorter (0.4 ns) and longer (2.4 ns) lifetimes of NAD(P)H represent the “free” and “enzyme-bound” components, respectively. On the other hand, the shorter (0.12 ns) and longer (3.38 ns) lifetimes of FAD represent the “enzyme-bound” and “free” components, respectively. As mentioned above, NAD(P)H auto-fluorescence signals come from the cytosol and mitochondria whereas, FAD signals mostly originate from the mitochondria thus, both are regarded as reporters of metabolic activity1, 3, 7; their ratios are used as a marker of cellular redox states8. In this manuscript, enzyme-bound fractions of NAD(P)H (a2%) and FAD (a1%) and their lifetimes (τm) are used to quantify these metabolic changes, in particular with our novel NAD(P)H-a2%/FAD-a1% FLIM-based redox ratio. The commonly used intensity-based redox ratio [NAD(P)H/FAD], based predominantly on signals driven by mitochondrial OXPHOS is defined as a reduction of this ratio, due to the conversion of fluorescent NAD(P)H to non-fluorescent NAD+ and conversion of non-fluorescent FADH2 to fluorescent FAD. We have instead used the above FLIM-based NADH-a2%/FAD-a1% ratio to avoid potential intensity-related artefacts (due to photo-bleaching, fluctuations in illumination sources, etc.) as a sensitive indicator of mitochondrial redox state. Here, an increase in metabolic activity is defined as an increase of this ratio, due to the increase of the fluorescent NAD(P)H enzyme-bound lifetime fraction a2% and decrease of the fluorescent FAD enzyme-bound fraction a1%9.
We are also introducing a new assay for the auto-fluorescent biomarker, tryptophan (Trp), which has been linked to cancer investigations10,11,12,13,14. Trp, an essential amino acid, is a precursor of niacin, which in turn is a precursor of NAD(P)H15. Increased Trp catabolism and increased indoleamine 2,3-dioxygenase (IDO) activity in the kynurenine pathway are linked to cancer development and progression16. Therefore, probing Trp is clinically relevant. In general, Trp fluorescence intensity and lifetime mainly provide information on the protein composition, protein structure of which they are a part of and changes in overall cellular microenvironment. Trp is therefore, used as a marker for protein abundance. Like NAD(P)H and FAD, Trp also has shorter (0.5 ns–2.5 ns) and longer (3.1 ns) lifetimes which represent the “protein-bound” (as residues in proteins) and “free” (free amino acid) lifetime components, respectively. One of the major applications of FLIM is the measurement of Förster resonance energy transfer or FRET17,18,19,20,21. In FLIM-FRET measurements, FRET events are identified if there is reduction in the donor lifetime, because of quenching of its fluorescence in the presence of the acceptor. Trp-NAD(P)H is a known FRET pair6, 13, 22, 23. The NAD(P)H-interacting enzymes carrying Trp residues from different metabolic pathways are potentially responsible for the quenching of Trp resulting in FRET. The efficiency of energy transfer or E% is calculated from the following equation E = 1 − τm/τ0, where τm is the mean lifetime of Trp in cells and τ0 = 3.1 ns of unquenched Trp lifetime measured from Trp in solution. We have demonstrated that Trp-NAD(P)H interactions can be used as a reporter of metabolic activity13, 23.
In cancer there is re-programming of cellular metabolism. Mitochondrial OXPHOS is largely defective and cellular energy demands are met through a hyperactive glycolytic pathway, referred to as the Warburg Effect24. This characteristic shift in cancer cell metabolism has been linked with cell proliferation, progression and metastasis24,25,26. Mutations, deletions and reductions in mtDNA have been linked to defective OXPHOS activity, dysregulated production of ROS and hyperactive glycolytic pathway27. In PCa, reduced mtDNA and mitochondrial dysfunction have been demonstrated27,28,29. It is observed that African-American (AA) respond poorly to therapy as compared with Caucasian-American (CA) PCa patients. Although, the reason for this disparity could be multi-factorial, reduced mtDNA and mitochondrial dysfunction has been identified as the underlying cause28. Also, reduced mtDNA and defective OXPHOS have been linked with the resistance and/or attenuation to apoptosis in cancer cells30, 31. Therefore, induction of cell death by correction of impaired mitochondrial OXPHOS activity is a promising strategy in cancer treatment. Doxorubicin is one of the potent anti-cancer drugs known to induce apoptosis32, 33. In a recent study, it was shown that doxorubicin targets OXPHOS for apoptosis induction29. In the study, mitochondrial response to 10 µM doxorubicin treatment for 24 hr in colon and PCa cells showed increase in mitochondrial ROS, increase in cytochrome c release, increase in caspase-3 activation and increase in the percentage of apoptotic cells. All these markers show that there is induction of the mitochondrial apoptotic pathway eventually leading to cancer cell death.
The direct link between defective OXPHOS activity in cancer cell metabolism and apoptosis having been demonstrated in published work, we have focused in this manuscript on the metabolic responses to doxorubicin drug treatment of aggressive (African-American, E006AA) and more responsive (Caucasian-American, LNCaP) PCa cells using our FLIM-FRET assay, hypothesizing that FLIM-based markers should show the correction of impaired OXPHOS. The goals were to identify (i) the molecular events associated with the changes in mitochondrial energy metabolism/OXPHOS activity upon doxorubicin treatment, and (ii) the early predictors of drug response by following the lifetimes (τ1, τ2, τm) and their relative fractions (a1%, a2%) of NAD(P)H, FAD, Trp; NAD(P)H-a2%/FAD-a1% FLIM-based redox ratio, NAD(P)H-Trp FRET interactions (E%), E% vs NAD(P)H-a2%/FAD-a1% median correlation along-with biochemically quantifying the generation of ROS and caspase-3 activity. The FLIM-based redox ratio results of this study clearly demonstrate lower metabolic activity before doxorubicin treatment and early molecular changes after treatment associated with increased OXPHOS after 5 time-points each with a 15 min interval up to 60 min, preceding the effector caspase-3 activation and apoptosis. Based on our FLIM method for quantitative analysis of mitochondrial energy metabolism, we differentiate between the (1) Pre-Apoptotic, (2) Responsive and (3) Slow responder PCa cells. Previous studies have utilized NAD(P)H and FAD lifetime changes for the evaluation of metabolic variations in cancer34,35,36. For the first time, we demonstrate that along with NAD(P)H and FAD lifetime changes, Trp-quenching due to Trp-NAD(P)H FRET interactions, and E% vs NAD(P)H-a2%/FAD-a1% median correlation are sensitive parameters in predicting the drug response of PCa cells.
Results
Induction of mitochondrial OXPHOS activity upon doxorubicin treatment
Metabolic changes in the mitochondria were investigated in LNCaP and E006AA PCa cells before and after doxorubicin treatment using our 3-channel FLIM-FRET assay. FLIM images were acquired at time zero as control and upon treatment with doxorubicin at 15 min intervals for 60 min. Doxorubicin-treated LNCaP cells showed a more impaired cellular morphology like cell shrinkage and some blebbing (Fig. 1a–e), whereas E006AA cells (Fig. 1g–k) morphologically appeared to be less affected by the cytotoxic effects of doxorubicin during the treatment time course. The mean lifetimes of NAD(P)H at the mitochondrial loci increased over the time course (0–60 min) in both cell lines upon treatment with doxorubicin (Fig. S2). Mitochondrial FAD mean lifetimes showed an increase in E006AA cells over the same time course, but no appreciable change in LNCaP cells (Fig. S3). Before analyzing the FLIM-based redox state, the frequency distribution of the two individual enzyme-bound fractions of NAD(P)H-a2% (Fig. 1f and l) and FAD-a1% (Fig. 2f and l) were examined.
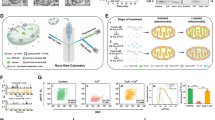
Increase in the enzyme-bound fraction of NAD(P)H-a2% upon doxorubicin treatment in PCa cells. Upper panel (a–f) LNCaP and lower panel (g–l) E006AA (AA) cells. PCa cells: LNCaPs (n = 26) with 4400–4890 ROIs, and E006AA cells (n = 28) with 5585–6372 ROIs, each ROI of 2 × 2 pixels, were analyzed for 5 time points (0, 15, 30, 45 & 60 min). The representative (a–e and g–k) color coded images and (f and l) the histograms showed increase in the enzyme-bound fraction of NAD(P)H-a2% with time (0–60 min) upon treatment with 1 µM doxorubicin when compared with untreated 0 min time point control (Ctrl). Greater magnitude of response was seen in LNCaP vs E006AA cells (60% vs 23%) from 0 to 60 min.
Decrease in the enzyme-bound fraction of FAD-a1% upon doxorubicin treatment in PCa cells. Upper panel (a–f) LNCaP and lower panel (g–l) E006AA (AA) cells. PCa cells: LNCaPs (n = 26) with 4400–4890 ROIs, and E006AA cells (n = 28) with 5585–6372 ROIs, each ROI of 2 × 2 pixels, were analyzed for 5 time points (0, 15, 30, 45 & 60 min). The representative (a–e and g–k) color coded images and (f and l) the histograms showed decrease in the enzyme-bound fraction of FAD-a1% with time (0–60 min) upon treatment with 1 µM doxorubicin when compared with untreated 0 min time point control (Ctrl).
The mitochondrial OXPHOS activity utilizes NAD(P)H and produces FAD. In other words, there is more enzyme-bound NAD(P)H-a2% and less enzyme-bound FAD-a1% fractions. In agreement with this, NAD(P)H-a2% increased over the time course while FAD-a1% decreased, exhibiting an increase in OXPHOS activity with doxorubicin treatment over time, suggesting normalization of an impaired mitochondrial function and opening the apoptotic pathway, mentioned in the introduction. The peak change of NAD(P)H-a2% from 0 min control to 60 min after doxorubicin treatment, was an 60% increase in LNCaP and 23% increase in E006AA cells (Fig. 1f and l); FAD-a1% decreased by 8.3% in LNCaP and 9.1% in E006AA cells (Fig. 2f and l).
In line with above observation, an increase of the FLIM redox ratio was observed with doxorubicin treatment over time in both PCa cell lines (Fig. 3), mainly driven by the increase of NAD(P)H-a2% and less so by the decrease of FAD-a1% (Figs 1f and 1, 2f and l). The peak change of FLIM-based redox ratio from 0 min control to 60 min after doxorubicin treatment was an 89% increase in LNCaP (Fig. 3a) and a 38% increase in E006AA (Fig. 3b). Increase in all these parameters indicates overall increase in OXPHOS activity after doxorubicin treatment. Hence, these FLIM parameters can be used as early predicators of drug-response.
Increase in mitochondrial NAD(P)H-a2/FAD-a1 Redox Ratio upon doxorubicin treatment in PCa cells. Mitochondrial OXPHOS activity uses NAD(P)H (more enzyme bound-a2%) and produces FAD (less enzyme bound-a1%). The histograms showed increase in mitochondrial NAD(P)H-a2/FAD-a1 Redox Ratio in (a) LNCaP and (b) E006AA (AA) cells for the same ROIs (mentioned above) with time (0–60 min) upon treatment with 1 µM doxorubicin compared with 0 min untreated control (Ctrl). The data suggests that there is increased mitochondrial OXPHOS activity with doxorubicin treatment. Greater magnitude of response was seen in LNCaP vs E006AA cells (89% vs 38%) from 0 to 60 min.
Increased E% in the Trp-NAD(P)H FRET interaction correlates with increased metabolic activity upon doxorubicin treatment
We have previously demonstrated that increased Trp-NAD(P)H FRET interaction and increased E% upon stimulation with glucose in cancer cells as an indicator of increased metabolic activity13, 37. This E% increase correlates with the increase in the enzyme-bound fraction of NAD(P)H (a2%). In the current study, we have extended the analysis to correlate with the FLIM-based redox ratio, showing that E% increased with rising redox states. Figure 4 tracks median E% by Field-of-view (FoV) covering >5000 Regions of Interest (ROIs) and correlates E% with the NAD(P)H-a2%/FAD-a1% redox ratio. E% increased concurrently with the increase in the FLIM-based redox ratio in both PCa cells suggesting that E% - NAD(P)H-a2%/FAD-a1% median correlation can be used to predict the drug-response in PCa cells.
Increase in E% with increase in mitochondrial NAD(P)H-a2/FAD-a1 Redox Ratio upon doxorubicin treatment in PCa cells. Correlation plots showed increase in median E% w.r.t. median NAD(P)H-a2/FAD-a1 redox ratio from control (Ctrl) to doxorubicin treatment groups with time (0–60 min) in both (a) LNCaP and (b) E006AA (AA) cells which suggests increased interaction of Trp (residues in proteins/enzymes) with NAD(P)H with increased mitochondrial OXPHOS activity.
Increase in the mitochondrial oxidative stress upon doxorubicin treatment
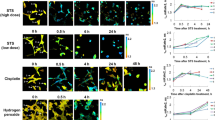
Increase in the production of ROS induces oxidative stress which results in oxidization of cellular macromolecules, inhibits protein function and promotes cell death38. Increased ROS is shown to be involved in doxorubicin induced apoptosis in cancer cells29. Usually, such studies have used 24–48 hr time points to assess induction of ROS and apoptosis. The current study’s objective inter alia was to identify early predictors of drug response and we therefore, correlated the observed changes in the NAD(P)H, FAD and Trp lifetimes and their relative fractions with the induction of ROS over 0–60 min period of doxorubicin treatment. MitoSOX Red is a selective live cell dye for detecting superoxide – the predominant ROS in the mitochondria. We observed increase in ROS as early as 15 min of doxorubicin treatment, which gradually increased over time (Fig. 5). Representative images of untreated 0 min control (Fig. 5a) and doxorubicin treated for 60 min (Fig. 5b) are shown. Clusters of adjoining cells (n = ~100) were grouped together to represent individual ROIs as numbered in Fig. 5c. Each ROI was quantified from 0–60 min and shows a 1.5- to 5.4-fold increase in ROS over time after doxorubicin treatment (Fig. 5d). Therefore, generation of ROS, which indicates beginning of oxidative cellular damage, correlates with the increase in the FLIM parameters in the early phase of doxorubicin treatment.
Increase in mitochondrial ROS upon doxorubicin treatment. LNCaP cells were labeled with MitoSox Red. After imaging of the control cells for 0 min time point, 1 µM doxorubicin was added on stage and proceeded with time lapse imaging at 15 min intervals up to 60 min. Representative images of cells (n = ~100) (a) 0 min control (Ctrl), (b) increased ROS at 60 min doxorubicin treatment, (c) auto-contrasted image with clusters of adjoining cells (n = ~100) grouped together to represent individual ROIs as numbered, (d) were quantified and showed a 1.5-to 5.4-fold increase in ROS over time after doxorubicin treatment.
Induction of apoptosis upon doxorubicin treatment
Doxorubicin has been shown to induce apoptosis in various types of cancer cells32, 33. However, the early changes in the mitochondrial NAD(P)H, FAD and Trp lifetimes prior to apoptosis upon doxorubicin treatment were not defined. Apoptosis is a long process with a cascade of upstream and downstream events, ultimately causing the activation of the effector caspase-3 which executes the final steps of apoptosis39. Apoptosis was therefore, analyzed by measuring caspase-3 activity by a DEVDase assay in both PCa cell lines. We have extended the time period to the point of detecting caspase activity to show that our shorter FLIM assay actually predicts the later apoptotic changes. No caspase-3 activity was detected at earlier time points (0–60 min) of doxorubicin treatment. At 12 hr and 24 hr time points post doxorubicin treatment, caspase-3 activity in both LNCaPs and E006AA cells occurred (Fig. 6). Based on the −/+ activation of effector caspase-3, we have categorized these phases as “Pre-Apoptotic” and “Apoptotic or Responsive” phases, respectively.
Increase in caspase-3 activity with time upon doxorubicin treatment in PCa cells. (a) LNCaP and (b) E006AA (AA) PCa cells were treated with 1 µM doxorubicin for different time points and assessed for the effector caspase-3 activity by DEVDase assay. There was no increase in caspase-3 activity in earlier time points 0 min–1 hr (Pre-Apoptotic Phase), which increased significantly ***p < 0.001 and ****p < 0.0001 at 12 hr and 24 hr (Apoptotic Phase) with doxorubicin treatment as compared to control group.
Indicators of doxorubicin drug-response
Metabolic changes in the “apoptotic phase” after 24 hr doxorubicin treatment in LNCaP vs E006AA cells showed increases in the mean lifetime of NAD(P)H and FAD (not shown), increases in NAD(P)H-a2% (33.3% vs 36.4%, Fig. S4a and d), decreases in FAD-a1% (10.3% vs 25.0%, Fig. S4b and e) and increases in NAD(P)H-a2%/FAD-a1% FLIM-based redox ratio (30% vs 60%, Fig. S4c and f). LNCaP cells exhibited morphological signs of apoptosis accompanied by cell shrinkage, condensed cytoplasmic organelles and extensive plasma membrane blebbing (Fig. 7b) and were termed as doxorubicin “Responsive” cells. In contrast, these morphological abnormalities were not so prominent in E006AA cells and therefore were termed as doxorubicin “Slow Responders” (Fig. 7e). Correlation of medians of E% and NAD(P)H-a2%/FAD-a1% showed decrease in E% in LNCaPs doxorubicin “Responsive” cells (Fig. 7c). Interestingly, we saw no change in E% in E006AA doxorubicin “Slow Responders” as compared to the control with the increase in redox ratio (Fig. 7f), which suggests that these cells had a slower response to treatment.
Median correlation of E% vs NAD(P)H-a2%/FAD-a1% an indicator of doxorubicin responsiveness. Upper panel (a–c) LNCaP and lower panel (d–f) E006AA (AA) PCa cells. Representative (a,b and d,e) NAD(P)H photon images show apoptotic cell morphology in (b) LNCaPs-doxorubicin “Responsive” cells as compared to the (a) LNCaP control cells; whereas the (d) E006AA control and (e) E006AA-doxorubicin “Slow Responders” look morphologically similar. The median correlation of E% vs NAD(P)H-a2%/FAD-a1% showed (c) decrease in E% in LNCaPs-doxorubicin “Responsive” cells and (f) no change in E% in E006AA-doxorubicin “Slow Responders” with increase in NAD(P)H-a2%/FAD-a1% ratio.
Applying the categorization of “Pre-Apoptotic” (0–60 min of treatment) and “Responsive”, “Slow Responders” (after 24 hr treatment) to NAD(P)H-τm, NAD(P)H-a2%, FAD-a1%, NAD(P)H-a2%/FAD-a1% FLIM-based redox ratio, similar trends were observed across these 3 categories (Table 1). Only the median correlation of E% with NAD(P)H-a2%/FAD-a1% parameter was able to differentiate these 3 categories and depict the heterogeneity in their doxorubicin drug responses showing that E% increases in the “Pre-Apoptotic” phase, decreases in the “Responsive” cells and no change in the “Slow Responders”.
FLIM E% calculation is commonly based on Trp-τm [(τ1 *a1%) + (τ2 *a2%)] which contains, both, the fractions of bound and free Trp and relating more closely to intensity-based E% calculations, which cannot differentiate between bound and free moieties. Extending the analysis to Trp-τm and Trp photons counts in these 3 categories tracks the Trp quenching by FRET events occurring in categories. The “Pre-Apoptotic” phase showed decrease in Trp-τm and Trp photons counts in both PCa cells which reflects increased Trp quenching (see Fig. S5a–d) with increase in metabolic activity; LNCaP-“Responsive” cells showed increase in Trp-τm and increasing trends in Trp photon counts (p = 0.054) which reflects less Trp quenching (see Fig. S5e–f) in the apoptotic cells, whereas the E006AA “Slow Responder” cells showed no appreciable change in Trp-τm or in Trp photon counts (p = 0.1) from control (see Fig. S5g–h). Our results, demonstrates that Trp-quenching and median E% correlation with NAD(P)H-a2%/FAD-a1% can be used as sensitive indicators to track a drug response in cancer in pre-apoptotic, apoptotic and slow responder cells, which is one of the most sought after goals in treatment to be able to identify and track a treatment response in heterogeneous cancer cell populations.
Discussion
In the current study, we have investigated the metabolic response of a potent anti-cancer drug doxorubicin on two PCa cell lines African American (E006AA or AA) and Caucasian American (LNCaP) by FLIM of NAD(P)H, FAD and Trp in the mitochondria. Differences in all applied parameters are observed between CA and AA PCa cell lines, the poorer response to treatment in more aggressive AA PCa’s being attributed to the lower mtDNA28. Mitochondrial dysfunction and defective OXPHOS activity are recognized as the cause of this disparity27. It is interesting to note, that inhibition of the OXPHOS complex I causes a reduction in the release of mitochondrial cytochrome c, which contributes to the inhibition of caspase activation and apoptosis29. Although, the exact role of OXPHOS in regulating apoptosis is not clearly understood, our previous findings suggest that restoration of mitochondrial function induces ROS production leading to permeabilization of mitochondrial membrane and apoptosis induction29, 40. Therefore, correction of the impaired mitochondrial OXPHOS activity and subsequent restoration of the apoptosis induction are promising strategies in cancer treatment. Since molecular changes precede morphological or phenotypic changes, this investigation identifies the molecular changes as the early predictors of drug response using our FLIM-FRET assay.
Our results in these two cell lines match those of patient-based data28, 41,42,43, except that our assay could be applied at an earlier stage of PCa for diagnosis and treatment. The assay identifies morphological changes, level of cytotoxicity and magnitude of metabolic response upon doxorubicin treatment in LNCaP vs E006AA cells in the “Pre-Apoptotic” phase with the peak change of 60% vs 23% increase in NAD(P)H-a2% (see Fig. 1f and l) and 8.3% vs 9.1% decrease in FAD-a1% fractions (Fig. 2f and l) and 88.88% vs 38.46% increase in the NAD(P)H-a2%/FAD-a1% (Fig. 3a and b) FLIM-based redox from 0 min to 60 min of treatment. These metabolic changes were accompanied with the increased Trp-NAD(P)H interaction resulting in increased E% which correlated with NAD(P)H-a2%/FAD-a1% redox ratio (Fig. 4). Also, there was increased ROS generation (Fig. 5) followed by induction of apoptosis (Fig. 6).
Both LNCaP and E006AA cells showed induction of apoptosis at 12 hr and 24 hr. We compared changes in the FLIM-FRET parameters in the “Pre-Apoptotic” (0–60 min) and apoptotic (~24 hr) phases to delineate the differences and interpret the doxorubicin drug-induced metabolic response in these phases. Interestingly, two groups emerged: LNCaP doxorubicin “Responsive” and E006AA doxorubicin “Slow Responder” PCa cells (Table 1). The LNCaP doxorubicin- “Responsive” cells showed morphological signs of apoptosis (Fig. 7b) whereas in the E006AA doxorubicin “Slow Responder” cells such morphological changes were not observed (Fig. 7e). However, at this stage, it is difficult to conclude that whether the E006AA cells were resistant to apoptosis or whether they had a slower response to treatment; which may be influenced by variation in experimental culture conditions, drug dose and duration of treatment; real-time mapping of apoptosis and FLIM-FRET metabolic responses to drug treatment within the same cell, at the same time would help to elucidate and correlate these molecular changes with apoptosis. In a recent study, FLIM changes and ratiometric caspase biosensor activity in an antibiotic induced apoptosis has been reported35. Similar to ours, studies have shown increase in OXPHOS with staurosporine antibiotic induced apoptosis in colon cancer and HeLa cells35, 44.
The current study provides spatial and temporal resolution of “Responsive” and “Slow Responder” cells and demonstrates Trp-quenching and median E% correlation with NAD(P)H-a2%/FAD-a1% as sensitive indicators to track the doxorubicin drug response in PCa cells. Using the E% and NAD(P)H-a2%/FAD-a1% median correlation we were able to categorize PCa cells into “Pre-Apoptotic”, “Responsive” and “ Slow Responders”, one of the most sought after goals in cancer treatment and follow up. Future direction of this study would be real-time mapping of the identified FLIM-FRET parameters with the generation of ROS and apoptosis in in vitro spheroid and organoid cultures and later in PCa mouse model to assess heterogeneity and track the treatment response.
Another potential application of the FLIM assay relates to elevated mitochondrial glutaminolysis in PCa compared to normal tissue45, 46. The enzyme Glutaminase (GLS) coverts glutamine to glutamate; glutamate is subsequently converted to α-ketoglutarate by glutamate dehydrogenase using NAD(P)+ as an co-enzyme and producing non-enzyme bound (free) NAD(P)H, contributing to the free NAD(P)H pool, while mitochondrial OXPHOS contributes to the enzyme-bound NAD(P)H fraction1, 15. PCa’s dysregulation in glycolysis, glutaminolysis and OXPHOS pathways compared to normal cause levels of free and enzyme-bound NAD(P)H fractions to vary. Using the FLIM assay to measure the altered levels of the NAD(P)H fractions, we could differentiate normal from PCa cells that is dependent on mitochondrial glutaminolytic pathway.
In conclusion, this study shows FLIM & FRET has the capability to detect earlier molecular changes associated with the correction in mitochondrial OXPHOS activity before the onset of apoptosis with doxorubicin treatment in PCa cells. Hence, our investigation provides molecular basis that induction of cell death by correction of impaired mitochondrial OXPHOS activity are promising strategies in cancer treatment. We observed these changes as early as 15 min of doxorubicin treatment accompanied with the increase in ROS. Also, this study reports for the first time, Trp-quenching due to Trp-NAD(P)H interactions (FRET), and E% vs NAD(P)H-a2%/FAD-a1% median correlation are sensitive parameters in tracking the drug response of PCa cells. Therefore, these parameters can be used as early predictors of drug response and in assessing the efficacy of anti-cancer drug treatment.
Materials and Methods
Cell Culture
Different PCa cell lines from African-American (E006AA) and Caucasian-American (LNCaP) origins have been used in this study. The E006AA (or AA) cells were maintained in high-glucose Dulbecco’s Modified Eagle Medium (Life Technologies) supplemented with 10% cosmic calf serum (Hyclone), 1% Penicillin-Streptomycin (Life Technologies), and 4 mM Sodium Pyruvate (Life Technologies). The LNCaP cells were maintained in RPMI 1640 (Life Technologies) supplemented with 10% cosmic calf serum (Hyclone) and 1% Penicillin-Streptomycin (Life Technologies). All cells were maintained in the cell culture incubator, at 37 °C with 5% CO2.
FLIM Instrumentation, Processing and Analysis
Our 3-channel FLIM imaging system consists of a Zeiss LSM 780 confocal/multiphoton (MP) laser scanning system coupled to the Zeiss inverted epi-fluorescence microscope, which is controlled with the ZEN software (Carl Zeiss, Inc). Multiphoton excitation of Trp, NAD(P)H and FAD was achieved by using an ultrafast (150 fs) tunable Ti:sapphire laser (680–1060 nm), operating at 80 MHz repetition rate (Chameleon Vision II, Coherent, Inc.). To excite the NAD(P)H and Trp we used 740 nm (NAD(P)H: 2-photon ecx, ET480/40 em; Trp: 3-photon exc, HQ360/40 em)13 and 890 nm for FAD (2-photon exc, 540/40 em). The fluorescence decay per pixel was measured using 3-channel SPC-150 TCSPC board (Becker & Hickl, GmbH) where the SPCM software was used to acquire the FLIM data (v. 8.91). Details on the methodology of our FLIM set up can be found elsewhere9, 47. A Zeiss 40 × 1.3NA oil, (EC Plan-Neuofluar, UV transmission is 60% at 340 nm) objective lens was used to focus the light on the sample and collect the emission for 60 s. The average power at the specimen plane (7 mW) and the acquisition time was chosen to reduce any photodamage to the cells.
After simultaneous acquisition of FLIM images for Trp, NAD(P)H and FAD, the florescence lifetime images were fitted for 2-components using SPCImage software (v. 5.5, Becker & Hickl). Number of parameters was generated including photon images, τ1, τ2, a1%, a2%, and χ2 for each pixel of each channel. Since, NAD(P)H signal matches with the mitochondrial morphology (Fig. S1), the mitochondrial Regions of Interest (ROI)s were thresholded by 2 × 2 pixels/ROI using the NAD(P)H photon image. The generated mitochondrial ROIs were used for further FLIM data analysis. The exported results from multiple samples and FOVs were further analyzed as described elsewhere9, 47.
Fluorescence Lifetime Imaging Microscopy and Doxorubicin Treatment
For imaging, LNCaP and E006AA PCa cells were plated onto 25 mm round #1.5 glass coverslips (Thermo Scientific), in their recommended growth medium and grown to 70–80% confluence. Fluorescence lifetime imaging was acquired in phenol-free or in FluoroBrite-DMEM (Thermo Fisher Scientific) growth medium with the microscope-heated stage maintained at 37 °C and under the flow of humidified blood-gas mixture (5% CO2). After imaging of the control FoVs, 1 µM doxorubicin was added in the growth medium on stage and FLIM data was again acquired at 15 min interval up to 60 min.
Quantification of the Optical Redox Ratio
For the optical redox ratio calculation, we have used a FLIM-based method which uses NAD(P)H-a2%/FAD-a1% redox pairs9. Our preferred FLIM-based NAD(P)H-a2%/FAD-a1% ratio avoids potential intensity-related artefacts, such as photo-bleaching or fluctuations of illumination levels.
Quantification of Mitochondrial ROS
LNCaP cells plated on glass coverslips were incubated with 500 nM MitoSOX Red reagent (Molecular Probes, Invitrogen) in HBSS Buffer with 5 mM Glucose for 45 min at 37 °C in the cell culture incubator. Live cell imaging was implemented in growth medium with heated stage maintained at 37 °C and under humidified gas flow on the Leica TCS SP5X using 40x water immersion, NA 1.1, excitation with white light laser 510 nm at 59% laser intensity and emission was collected for 550–650 nm range with 750 V PMT gain. After imaging of the control FoVs, 1 µM doxorubicin was added in the growth medium on stage and time lapse imaging was done at 15 min interval for 60 min. Images were exported as “tiff” and Region of Interests (ROIs) for groups of cells were drawn and quantified (n = ~100 cells) in Fiji.
Quantification of the Caspase Activity
PCa cells were treated with 1 µM doxorubicin for different time points as described in Figure legends. Treated Cells (LNCaP & E006AA) were washed with PBS, and the whole cell lysates were made in the sample lysis buffer (50 mM HEPES, 1% Triton X-100, 0.1% CHAPS, 1 mM dithiothreitol (DTT), and 0.1 mM EDTA). 40 μg of protein was added to a reaction mixture containing 30 μM fluorogenic peptide substrates, DEVD-AFC, 50 mM of HEPES, pH 7.4, 10% glycerol, 0.1% CHAPS, 100 mM NaCl, 1 mM EDTA, and 10 mM DTT, and incubated at 37 °C for 90 min. Fluorescence from 7-amino-4-trifluoromethylcoumarin (AFC) was monitored in a spectrofluorimeter (Biotek Synergy 2) using excitation wavelength 400/30 nm and emission wavelength 508/20 nm. The results are presented as fold activation over the control48.
Statistical Analysis
Results are expressed as mean ± SEM. Differences between groups were analyzed for statistical significance using a Student’s t-test and one-way ANOVA analysis wherever applicable. The threshold for the significance level or p-value of the test was 0.05.
Data Availability
All data generated or analyzed in this study are included in this manuscript (and in the Supplementary Information).
References
Heikal, A. A. Intracellular coenzymes as natural biomarkers for metabolic activities and mitochondrial anomalies. Biomark. Med. 4, 241–263 (2010).
Gukassyan, V. & Heikal, A. A. (ed.). Natural Biomarkers for Cellular Metabolism: Biology, Techniques, and Applications. (CRC Press, Taylor & Francis Group, Boca Raton, Florida, USA, 2014).
Chance, B., Legallais, V. & Schoener, B. Metabolically linked changes in fluorescence emission spectra of cortex of rat brain, kidney and adrenal gland. Nature 195, 1073–1075 (1962).
Mayevsky, A. & Rogatsky, G. G. Mitochondrial function in vivo evaluated by NADH fluorescence: from animal models to human studies. Am. J. Physiol. Cell Physiol. 292, C615–40 (2007).
Blacker, T. S. & Duchen, M. R. Investigating mitochondrial redox state using NADH and NADPH autofluorescence. Free Radic. Biol. Med. 100, 53–65 (2016).
Lakowicz, J. R. (ed.). Principles of fluorescence spectroscopy. (Springer, USA, 2006).
Richards-Kortum, R. & Sevick-Muraca, E. Quantitative optical spectroscopy for tissue diagnosis. Annu. Rev. Phys. Chem. 47, 555–606 (1996).
Walsh, A. J. et al. Quantitative optical imaging of primary tumor organoid metabolism predicts drug response in breast cancer. Cancer Res. 74, 5184–5195 (2014).
Wallrabe, H., Svindrych, Z., Alam, S. R., Siller, K. H. & Periasamy, A. Quantitative FLIM analysis of discrete redox states in cancer cells. Free Radic. Biol. Med. (under review) (2017).
Diagaradjane, P., Yaseen, Ma., Yu, J., Wong, M. S. & Anvari, B. Autofluorescence characterization for the early diagnosis of neoplastic changes in DMBA/TPA-induced mouse skin carcinogenesis. Lasers Surg. Med. 37, 382–395 (2005).
Mujat, C. et al. Endogenous optical biomarkers of normal and human papillomavirus immortalized epithelial cells. Int. J. Cancer 122, 363–371 (2008).
Li, D., Zheng, W. & Qu, J. Y. Two-photon autofluorescence microscopy of multicolor excitation. Opt. Lett. 34, 202–204 (2009).
Jyothikumar, V., Sun, Y. & Periasamy, A. Investigation of tryptophan-NADH interactions in live human cells using three-photon fluorescence lifetime imaging and Förster resonance energy transfer microscopy. J. Biomed. Opt. 18, 60501 (2013).
Wiggins, T., Kumar, S., Markar, S. R., Antonowicz, S. & Hanna, G. B. Tyrosine, phenylalanine, and tryptophan in gastroesophageal malignancy: A systematic review. Cancer Epidemiology Biomarkers and Prevention 24, 32–38 (2015).
Nelson, D. & Cox, M. Lehninger Principles of Biochemistry. (W. H. Freeman, New York, USA, 2004).
Platten, M., Wick, W. & Van Den Eynde, B. J. Tryptophan catabolism in cancer: Beyond IDO and tryptophan depletion. Cancer Research 72, 5435–5440 (2012).
Sekar, R. B. & Periasamy, A. Fluorescence resonance energy transfer (FRET) microscopy imaging of live cell protein localizations. J. Cell Biol. 160, 629–633 (2003).
Periasamy, A. & Clegg, R. M. (ed.). FLIM Microscopy in Biology and Medicine. (CRC Press, Taylor & Francis Group, Boca Raton, Florida, USA, 2010).
Sun, Y., Day, R. N. & Periasamy, A. Investigating protein-protein interactions in living cells using fluorescence lifetime imaging microscopy. Nat. Protoc. 6, 1324–1340 (2011).
Rehman, S., Gladman, J. T., Periasamy, A., Sun, Y. & Mahadevan, M. S. Development of an AP-FRET based analysis for characterizing RNA-protein interactions in myotonic dystrophy (DM1). PLoS One 9, e95957 (2014).
Sun, Y. & Periasamy, A. In Advanced Fluorescence Microscopy SE - 6 (ed. Verveer, P. J.) 1251, 83–107 (Springer, New York, USA, 2015).
Torikata, T., Forster, L. S., Johnson, R. E. & Rupley, Ja Lifetimes and NADH quenching of tryptophan fluorescence in pig heart cytoplasmic malate dehydrogenase. J. Biol. Chem. 254, 3516–3520 (1979).
Jyothikumar, V., Sun, Y. & Periasamy, A. In Natural Biomarkers for Cellular Metabolism (eds Gukassyan, V. & Heikal, A. A.) (CRC Press, Taylor & Francis Group, Boca Raton, Florida, USA, 2014).
Warburg, O. On Respiratory Impairment in Cancer Cells. Science (80-.). 124, 269–70 (1956).
Pecqueur, C., Oliver, L., Oizel, K., Lalier, L. & Vallette, F. M. Targeting metabolism to induce cell death in cancer cells and cancer stem cells. Int. J. Cell Biol. 805975 (2013).
Zhao, Y., Butler, E. B. & Tan, M. Targeting cellular metabolism to improve cancer therapeutics. Cell Death Dis. 4, e532 (2013).
Chaudhary, A. K. et al. Mitochondrial dysfunction-mediated apoptosis resistance associates with defective heat shock protein response in African–American men with prostate cancer. Br. J. Cancer 114, 1–11 (2016).
Koochekpour, S., Marlowe, T., Singh, K. K., Attwood, K. & Chandra, D. Reduced Mitochondrial DNA Content Associates with Poor Prognosis of Prostate Cancer in African American Men. PLoS One 8, e74688 (2013).
Yadav, N. et al. Oxidative phosphorylation-dependent regulation of cancer cell apoptosis in response to anticancer agents. Cell Death Dis. 6, e1969 (2015).
Dey, R. & Moraes, C. T. Lack of oxidative phosphorylation and low mitochondrial membrane potential decrease susceptibility to apoptosis and do not modulate the protective effect of Bcl-x(L) in osteosarcoma cells. J. Biol. Chem. 275, 7087–7094 (2000).
Chandra, D., Liu, J. W. & Tang, D. G. Early mitochondrial activation and cytochrome c up-regulation during apoptosis. J. Biol. Chem. 277, 50842–50854 (2002).
Tehranian, N. et al. Combination effect of PectaSol and Doxorubicin on viability, cell cycle arrest and apoptosis in DU-145 and LNCaP prostate cancer cell lines. Cell Biol. Int. 36, 601–610 (2012).
Tacar, O., Sriamornsak, P. & Dass, C. R. Doxorubicin: An update on anticancer molecular action, toxicity and novel drug delivery systems. J. Pharm. Pharmacol. 65, 157–170 (2013).
Rück, A., Hauser, C., Mosch, S. & Kalinina, S. Spectrally resolved fluorescence lifetime imaging to investigate cell metabolism in malignant and nonmalignant oral mucosa cells. J. Biomed. Opt. 19, 96005 (2014).
Sergeevaa, T. F. et al. Relationship between intracellular pH, metabolic co-factors and caspase-3 activation in cancer cells during apoptosis. Biochim Biophys Acta 1864, 604–611 (2017).
Shah, A. T., Heaster, T. M. & Skala, M. C. Metabolic Imaging of Head and Neck Cancer Organoids. PLoS One 12 (2017).
Rehman, S. et al. Investigation of prostate cancer cells using NADH and Tryptophan as biomarker: Multiphoton FLIM-FRET microscopy. In Progress in Biomedical Optics and Imaging - Proceedings of SPIE (ed. Periasamy Ammasi; So Peter T. C.; König Karsten) 9712 (2016).
Circu, M. L. & Aw, T. Y. Reactive oxygen species, cellular redox systems, and apoptosis. Free Radical Biology and Medicine 48, 749–762 (2010).
Fulda, S. & Debatin, K.-M. Extrinsic versus intrinsic apoptosis pathways in anticancer chemotherapy. Oncogene 25, 4798–4811 (2006).
Yadav, N. et al. Transformations of the macromolecular landscape at mitochondria during DNA-damage-induced apoptotic cell death. Cell Death Dis. 5, e1453–12 (2014).
Powell, I. J. et al. The predictive value of race as a clinical prognostic factor among patients with clinically localized prostate cancer: a multivariate analysis of positive surgical margins. Urology 49, 726–731 (1997).
Godley, P. A. et al. Racial differences in mortality among Medicare recipients after treatment for localized prostate cancer. J. Natl. Cancer Inst. 95, 1702–1710 (2003).
Chornokur, G., Dalton, K., Borysova, M. E. & Kumar, N. B. Disparities at Presentation, Diagnosis, Treatment, and Survival in African American Men, Affected by Prostate Cancer. Prostate 71, 985–997 (2011).
Yu, J.-S., Guo, H.-W., Wang, C.-H., Wei, Y.-H. & Wang, H.-W. Increase of reduced nicotinamide adenine dinucleotide fluorescence lifetime precedes mitochondrial dysfunction in staurosporine-induced apoptosis of HeLa cells. J. Biomed. Opt. 16, 36008 (2011).
Pan, T. et al. Elevated expression of glutaminase confers glucose utilization via glutaminolysis in prostate cancer. Biochem. Biophys. Res. Commun. 456, 452–458 (2015).
Chen, L. & Cui, H. Targeting Glutamine Induces Apoptosis: A Cancer Therapy Approach. Int. J. Mol. Sci. 9, 22830–22855 (2015).
O’Melia, M. J., Wallrabe, H. & Svindrych, Zdenek Rehman, Shagufta Periasamy, A. FLIM data analysis of NADH and Tryptophan autofluorescence in prostate cancer cells. In Proc. SPIE 9712, Multiphoton Microscopy in the Biomedical Sciences XVI, 97122E (eds Periasamy, A. & So, P.) (2016).
Srivastava, P. et al. Neem oil limonoids induces p53-independent apoptosis and autophagy. Carcinogenesis 33, 2199–2207 (2012).
Acknowledgements
We acknowledge funding from the National Institutes of Health (NIH) EB020843 (AP), CA160685 (DC) and the University of Virginia (AP).
Author information
Authors and Affiliations
Contributions
S.R.A., H.W. and A.P. conception, design of research and interpreted the results of the experiment; S.R.A. and H.W. performed experiments; S.R.A. and A.P. drafted and approved the final version of the manuscript; S.R.A., H.W., and Z.S. analyzed the data; A.K.C. and D.C. provided cells and the Fig. 6; S.R.A. and K.C. cell culture.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alam, S.R., Wallrabe, H., Svindrych, Z. et al. Investigation of Mitochondrial Metabolic Response to Doxorubicin in Prostate Cancer Cells: An NADH, FAD and Tryptophan FLIM Assay. Sci Rep 7, 10451 (2017). https://doi.org/10.1038/s41598-017-10856-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-10856-3
This article is cited by
-
Generative adversarial network enables rapid and robust fluorescence lifetime image analysis in live cells
Communications Biology (2022)
-
Characterization of mitochondrial dysfunction due to laser damage by 2-photon FLIM microscopy
Scientific Reports (2022)
-
Imaging Technologies for Microfluidic Biochips
BioChip Journal (2022)
-
FLIM as a Promising Tool for Cancer Diagnosis and Treatment Monitoring
Nano-Micro Letters (2021)
-
Lapatinib inhibits doxorubicin induced migration of HER2-positive breast cancer cells
Inflammopharmacology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.