Abstract
Euglena gracilis (E. gracilis) has been proposed as one of the most attractive microalgae species for biodiesel and biomass production, which exhibits a number of shapes, such as spherical, spindle-shaped, and elongated. Shape is an important biomarker for E. gracilis, serving as an indicator of biological clock status, photosynthetic and respiratory capacity, cell-cycle phase, and environmental condition. The ability to prepare E. gracilis of uniform shape at high purities has significant implications for various applications in biological research and industrial processes. Here, we adopt a label-free, high-throughput, and continuous technique utilizing inertial microfluidics to separate E. gracilis by a key shape parameter-cell aspect ratio (AR). The microfluidic device consists of a straight rectangular microchannel, a gradually expanding region, and five outlets with fluidic resistors, allowing for inertial focusing and ordering, enhancement of the differences in cell lateral positions, and accurate separation, respectively. By making use of the shape-activated differences in lateral inertial focusing dynamic equilibrium positions, E. gracilis with different ARs ranging from 1 to 7 are directed to different outlets.
Similar content being viewed by others
Introduction
Euglena gracilis (E. gracilis) is a species of single-celled eukaryotic microalgae that can grow both autotrophically and heterotrophically on non-agricultural land. It has been attracting great industrial and academic interests for its ability to synthesize valuable products with commercial applications, such as paramylon, wax esters, and various types of nutrients (i.e. vitamins, minerals, fatty acids, and amino acids). E. gracilis accumulates paramylon, an insoluble β-1,3-glucan, under aerobic conditions, which has been shown to be able to stimulate immunomodulating activities1, treat and prevent infectious, oncologic and allergic diseases (i.e. human immunodeficiency virus-HIV2, colon cancer3, and atopic dermatitis4), and fabricate nanomaterials (i.e. nanofibers)5. Wax esters, mainly composed of C14:0 saturated fatty acid, myristic acid, and myristyl alcohol6, synthesized under anaerobic conditions, holds promise for biodiesel and jet fuel production as an alternative to fossil fuels. Moreover, E. gracilis has been reported to be capable of producing dietary supplements α-tocopherol and β-carotene7, as well as assay vitamin B12 in body fluids8, waste water quality9, and ecotoxicity of engineered nanoparticles10. Thus, microalgae such as E. gracilis may be able to impact a large number of fields, including renewable bioenergy, nutritional and functional food, environment, and health-care.
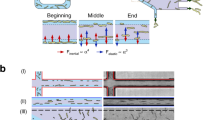
The variation in shape represents one of the most important noticeable characteristics for E. gracilis. The microalga can vary in shape dramatically, ranging from nearly spherical (aspect ratio, AR = 1) to elongated cylindrical (AR > 5) with most cells appearing spindle-shaped. The shape of E. gracilis has been demonstrated to be closely correlated to biological clock, photosynthesis, respiration, cell cycle, and environmental factors (Fig. 1). When grown under the synchronizing effect of a repetitive daily light-dark cycle, E. gracilis displays two shape changes per day11: the cell population increases in length up until the middle of the light cycle, exhibiting an elongated cylindrical shape, and decreases in length for the remainder of the 24-hour period, exhibiting a nearly spherical shape at the end of the dark cycle. These shape variations reflect a circadian rhythm, and have been demonstrated to be controlled by the biological clock12. Photosynthetic and respiratory reactions have also been reported to be involved in daily E. gracilis shape changes11, 13: photosynthetic electron transport pathways are necessary for the shape change from spherical to elongated, while respiratory pathways are involved in both the spherical to elongated and the elongated to spherical shape changes. Shape is also a cell cycle biomarker for E. gracilis, which reproduces by longitudinal binary fission, and a decrease in cell length could be a signal for the onset of mitosis. Moreover, the shape can vary in response to a variety of physical and chemical treatments, including light, temperature, pH, and cation concentration14,15,16,17.
In order to develop metabolically and genetically engineered E. gracilis with superior characteristics for low-cost biodiesel and biomass production, it would be important to prepare shape-synchronized E. gracilis cell populations adopting relatively uniform shape and size at high purities. The study of a homogeneous E. gracilis population allows for a better understanding of the biochemistry and molecular biology of valuable derivatives (i.e. paramylon and wax esters) production, along with nuclei and subcellular organelles (i.e. mitochondrion, chloroplasts, flagella, and pellicles). Shape-based separation of E. gracilis can provide an enriched cell population at a single stage, with the same status or phase. This would improve the investigation of correlated physiological and metabolic signatures by gene expression profiling. To date, sequencing the genome of E. gracilis has proved problematic due to its complexity18. More importantly, the shape-based separation, enrichment, and synchronization of E. gracilis open venues for a wide range of downstream applications, including biochemical assays, mutational analysis, and next generation sequencing.
However, shape has not been considered in traditional separation methods, such as pore-based filtration and centrifugation. Several microfluidic techniques have been utilized for shape-based separation of particles. Some techniques are based on internal channel or microstructure topology induced microflow, such as hydrodynamic filtration (HDF)19, and deterministic lateral displacement (DLD)20, 21. But these techniques require complex networks of microscale structures (i.e. branched channels and post arrays, respectively, for HDF and DLD), and require low flow rates (i.e. <10 µL/min). Other techniques integrate microfluidic flow with externally applied electrical22 or magnetic23 fields. However, these techniques require more device fabrication and external setup, along with the operation at relatively slow flow rates. More recently, viscoelastic fluids24, 25 have been demonstrated as another effective approach to sort spherical and peanut shaped microparticles due to the balance between elastic lift and inertial forces. Although this technique is high-throughput and simple-to-integrate, a precise control over the rheological properties of suspending medium is needed, limiting its practical applications.
Here, we utilize inertial focusing techniques26, 27 to separate E. gracilis by shape in a label-free, high-throughput, and cost-effective manner. Under the combined effects of two inertial lift forces (i.e. a shear-gradient lift force and a wall-effect lift force), particles migrate to dynamic equilibrium positions across fluid streamlines. Inertial microfluidics has been extensively studied to focus, order, and separate spherical particles, and mammalian cells with relatively constant size and shape, yet there have been few reports about inertial effects on shaped biological and synthetic particles. Hur et al.28 characterized the inertial effects on polymeric particles with various shapes, such as cylindrical, doublet-shaped, and disk-shaped. Masaeli et al.29 demonstrated the ability of inertial microfluidics to separate polystyrene beads and yeast cells based on shape. We recently showed that inertial focusing can be utilized together with secondary flows to focus E. gracilis cells varying in shape to a single focal stream30. However, it has not been shown whether inertial lift forces can be used to separate microalgae based on shape. In this work, we investigate the behaviors of E. gracilis cells with different ARs ranging from 1 to 7 under inertial effects in a straight microchannel region, an expansion region, and outlets. As the dynamic inertial focusing equilibrium positions are dependent on E. gracilis cell AR, E. gracilis with particular ARs migrate to distinct lateral equilibrium positions, and exit from different outlets. To the best of our knowledge, this is the first time that separation and enrichment of a single species of microalgal cells by shape has been reported using microfluidics.
Results
The microfluidic device used in the experiment consists of an inlet, a high aspect ratio (height/width >2) straight microchannel, a gradually expanding region, and five branched outlets each terminated with fluidic resistors, i.e. serpentine channels (see Fig. 2). The Reynolds number, Re = ρνD h /μ, is a dimensionless parameter describing the ratio between inertial and viscous forces26. Here, ρ, ν, and μ are the density, mean velocity, and dynamic viscosity of the fluid, respectively. D h = 2WH/W+H is the hydraulic diameter of the channel, where W and H are the channel width and height, respectively. At finite Re, the movement and position of target E. gracilis cells in the three regions indicated by sequence numbers i, ii, and iii, are distinct: (i) E. gracilis cells are randomly distributed close to the inlet; (ii) E. gracilis cells with various shapes generally migrate to two equilibrium positions along the long faces of the microchannel due to the balance between a shear-gradient lift force and a wall-effect lift force, and long rod-shaped cells migrate to positions closer to the channel centerline than spherical ones; (iii) the differences in shape-dependent lateral equilibrium positions are enhanced while cells continue along their focusing streamlines (Fig. 2a). Finally, E. gracilis cells with variable shapes are directed to different outlets: spherical cells exit from the outlets closest to the channel sidewalls, while longer rod-shaped cells are collected from the outlet closer to the channel centerline. Figure 2b presents schematically and experimentally the E. gracilis cells with various ARs at the inlet, 4-cm downstream of the inlet, and expansion region.
Shape-based separation of E. gracilis with different aspect ratios due to shape-dependent lateral inertial focusing equilibrium positions. (a) A schematic view of the inertial microfluidic channel and separation principles. (b) A top view (upper) showing the structure and dimensions of the microfluidic device which consists of an inlet, a straight rectangular microchannel, an expansion region, and five outlets with fluidic resistors. Schematics (lower left) and superimposed experimental images (lower right) illustrating the distribution of E. gracilis cells with different ARs at the inlet, downstream, and expansion region. Scale bar = 10 µm.
Lateral equilibrium positions depend on E. gracilis cell aspect ratio
E. gracilis cells with different ARs have different lateral (Y-) inertial focusing equilibrium positions. The effect of cell AR on lateral equilibrium positions is investigated by flowing E. gracilis cells with different ARs ranging from 1 to 7 at three flow rates, 300, 500, and 800 µL/min. The corresponding Re are 77, 128 and 205, respectively. We chose seven levels of AR with a spacing of 1 in order to give readers a higher definition view of the dynamics and lateral equilibrium position distributions for E. gracilis cells having different shapes. We chose a straight rectangular channel region, 4-cm downstream of the inlet (Fig. 3a), for the measurement in order to ensure effective inertial focusing and ordering effects. The violin plots visualize the distribution of lateral focusing distributions for E. gracilis with different ARs, and its probability density at different values. In all three flow conditions, there are two sets of lateral equilibrium positions symmetric about the channel centerline for E. gracilis cells with all ARs (Fig. 3c–e). However, spherical E. gracilis cells migrate to locations closer to the walls compared to cells that have higher AR. In contrast, long rod-shaped cells occupy equilibrium positions much closer to the channel centerline than cells that have lower AR. The averaged lateral positions demonstrate that the distance to the channel centerline decreases with increasing AR (Fig. 3f). Similar AR-dependent inertial focusing behaviors were observed in a 50-µm-wide, 90-µm-high microchannel (see Supplementary Fig. S1). These results agree with previous numerical analysis by Gossett et al.31 and experimental studies by Masaeli et al.29: there are usually two preferred equilibrium positions along the longer channel faces in a rectangular microchannel, and rod-like microparticles migrate to a stable position closer to the channel centerline than that of spherical ones with the same volume. With finite inertia, ellipsoid E. gracilis cells undergo rotational motion in the microchannel. Compared to lower AR E. gracilis cells, higher AR ones possess inertial focusing equilibrium positions further from the channel wall due to their larger rotational diameter, resulting in larger wall-effect lift force in the near-wall region, which is much stronger than the balancing shear-gradient force, and directs the cells further away from the wall29.
Lateral (Y-) focusing distributions depend on E. gracilis cell aspect ratio (AR). (a) An illustration of a straight rectangular channel region used for measuring the lateral positions of E. gracilis flowing through the microfluidic channel. (b) Superimposed experimental images showing the lateral inertial focusing equilibrium positions for E. gracilis with different ARs at Re = 77. Scale bar = 20 µm. (c–e) Violin plots of lateral positions for E. gracilis cells with ARs ranging from 1 to 7 for Re = 77, 128 and 205, respectively. (f) Averaged equilibrium positions are plotted as a function of cell AR for three Re. The error bars indicate the standard deviations for at least 100 measurements.
Expansion region enhances the differences in lateral equilibrium positions
A gradually expanding region is designed downstream of the straight rectangular microchannel to enhance the differences in AR-dependent lateral inertial focusing equilibrium positions, enabling high-purity shape-based separation of E. gracilis cells at the outlets. E. gracilis cells with different ARs are inertially focused to two lateral equilibrium positions after flowing through the straight microchannel, while the expansion region gradually enhances the differences in lateral positions (see Supplementary Movie S1). We choose an expansion region, 4.25-cm downstream of the inlet (Fig. 4a) for the measurement (channel width is around 335 µm). Similar to the straight rectangular microchannel region, the distance to the channel centerline decreases with increasing AR at the expansion region (Fig. 4c and d). The differences in lateral equilibrium positions for E. gracilis cells with different ARs at the expansion region is greater than the differences in the straight region (Fig. 3). Relative differences in averaged lateral positions between E. gracilis cells with AR = 1 and others (i.e. AR = 2–7) at straight region and expansion region are compared and presented in Supplementary Fig. S2. For example, at Re = 77 the averaged lateral position difference between E. gracilis cells with AR = 1 and AR = 7 is 45.1 µm at the expansion region, which is 8.5 times as large as that at the straight channel region (i.e. 5.3 µm). This expansion in distance is expected given the corresponding spreading of the focusing streamlines by 8.375-fold due to channel width expansion. This observation agrees with the previous experimental study by Hur et al.28, in which an expansion region was used to separate and enrich mammalian cells based on deformability.
The gradually expanding region enhances the differences in AR-dependent lateral positions while maintaining E. gracilis cells in the focusing streamlines. (a) An illustration of an expansion region used for measuring the lateral positions of E. gracilis flowing through the microfluidic channel. (b) A superimposed experimental image showing the lateral positions for E. gracilis with different ARs at Re = 77 (half of the expansion region is shown). Scale bar = 20 µm. (c,d) Spread plots and box plots of lateral positions for E. gracilis cells with different ARs ranging from 1 to 7 for Re = 77 and 205, respectively.
Separation and enrichment of E. gracilis by shape at the outlets
By taking advantages of differences in AR-dependent lateral equilibrium positions demonstrated above, we have performed shape-based separation and enrichment of E. gracilis cells with different ARs ranging from 1 to 7. Collection of cells with different ARs at different outlets with a flow rate of 300 µL/min (Re = 77) is shown (Fig. 5a–c). Each outlet has a fluidic resistor (i.e. a serpentine microchannel) with high resistance, which is designed to minimize the flow ratio distortion caused by any small variation (i.e. tubing length and debris clogging) in the outlet fluidic resistance28. E. gracilis cells with lower ARs (i.e. 1 and 2) are found to mostly exit from outlets 1 and 5 (see also Supplementary Movie S2), while the cells with higher AR (i.e. 6 and 7) are mainly captured from outlet 3 (see also Supplementary Movie S3). We use two parameters to quantify the separation efficiency: purity represents the proportion of cells with a specific AR at the inlet and each outlet; enrichment factor is defined as the proportion of cells with a specific AR in a given outlet to its proportion at the inlet. We have collected lower AR cells (AR = 1 and 2) at outlet 1 and outlet 5 with a purity of 92.6% and 96.8%, respectively; and higher AR cells (AR = 6 and 7) at outlet 3 with purity of 82.4% (Fig. 5d). Moreover, the enrichment factors for E. gracilis cells with AR = 1 at outlets 1 are 3.1 and 3.3, respectively, and the enrichment factors for cells with AR = 6 and 7 at outlet 3 are 5.9 and 7.8, respectively (Fig. 5e). E. gracilis cells with a middle range ARs (i.e. 3–5) are more likely to exit from outlets 2 and 4, with up to 87% purity and around 1.5 enrichment factor.
Shape-based separation of E. gracilis cells with different ARs ranging from 1 to 7 at the outlets at Re = 77. (a) An illustration of five outlets with high fluidic resistance (i.e. serpentine channels). (b) Snap-shot pictures comparing E. gracilis proportion at the inlet and outlets. (c) Superimposed experimental images showing that E. gracilis with different ARs are more likely to exit from different outlets: spherical shaped and long rod-shaped cells exit from outlets closer to the sidewall and centerline, respectively. Scale bar = 40 µm (d) A comparison of purity for E. gracilis cells with different ARs ranging from 1 to 7 at inlet and each outlet. (e) Bar graphs of enrichment factors for E. gracilis cells with different ARs for each outlet. At least 100 cells were measured for inlet and each outlet.
Discussion
Cell viability tests were carried out using FDA and PI, which show that E. gracilis cells remain highly viable after flowing through the microchannel under inertial effects (see Supplementary Fig. S3). Moreover, since the rapid processing of the device does not lead to significant effect on gene expression over the time course of a separation28, the proposed platform is desirable for various downstream applications where transcriptomic analyses are required. We utilized dilute cell suspension (i.e. 105 cells/mL) in the experiment in order to avoid particle-particle interactions, leading to the defocusing of cells at the expanding region, which adversely affect cell separation at the outlets32. It has been shown that particle concentration is an important factor affecting focusing behavior and accuracy. Inertially influenced particle-particle hydrodynamic interactions lead to multiple particle strains in the lateral direction as concentration is increased33. Moreover, the broadening of focused particle streams at expanding regions appeared at high particle suspensions32. At the expanding region, particles slow down and move closer due to expanding streams, which lead to an increase in particle–particle hydrodynamic interactions.
The throughput of the device can reach 800 µL/min (~1,300 cells/sec), which could be improved by optimizing cell concentration and Re, and using the design in a parallel network. Note that fluidic resistors located in the respective outlets could be tuned to capture enriched fractions of cells having particular particle shapes by adjusting the amount of fluid that splits into each outlet28. Further improvement and optimization of the channel parameters, i.e. number and lateral positions of outlets, flow rates (or Re) are also expected to increase enrichment factors for E. gracilis with specific AR(s). Moreover, the purity and enhancement factor could be increased by using the devices in a cascade configuration. Therefore, we are able to obtain high-purity E. gracilis cell samples with a specific AR (or shape) according to the specific application requirements.
In this work, we demonstrate passive, label-free, and high-throughput separation of E. gracilis microalgae by shape using inertial microfluidics. Under the combined effects of two inertial lift forces, a shear-gradient lift force and a wall-effect lift force, E. gracilis cells with various ARs migrate to different AR-dependent lateral dynamic equilibrium positions when flowing through a high aspect ratio rectangular microchannel. The distance to the channel centerline is shown to decrease with increasing cellular AR. The expansion region downstream of the straight microchannel enhances the differences in equilibrium positions, while maintaining cells in the focusing streamlines. Effective shape-based separation and enrichment of E. gracilis cells are achieved at the outlets: cells with lower ARs (i.e. 1 and 2), higher ARs (i.e. 6 and 7), and middle range ARs (i.e. 3–5) are found to mostly exit from the outlets closest to the channel walls, the outlet in the channel center, and the ones between the center and the sidewalls, respectively. Since the cell AR (or shape) of E. gracilis is an important biological, metabolic, and environmental marker, this capability enables a large range of industrial, research and clinical applications. Moreover, by integrating with metabolic and genetic engineering techniques, it is envisioned that this platform can be used as a powerful tool to develop enhanced E. gracilis cells with desirable properties34,35,36,37.
Methods
Microfluidic device design and fabrication
The microfluidic channel was fabricated with polydimethylsiloxane (PDMS) using standard soft-lithography techniques38. The inlet enables a relatively smooth pressure transition which reduces recirculations, where optional filters could also be placed to prevent the channel from be clogged by aggregates. The inertial focusing channel was designed to ensure that cells reach two lateral equilibrium positions along the longer faces of rectangular channel at the end. The basic parameter design rules, i.e. channel length required for focusing, flow rate, and channel cross-section can be found in our previous review articles26, 27. The whole microchannel has a uniform height of 90 µm. The width and length of the straight channel is 40 µm and 4.2 cm, respectively. It is preferable to have a smooth transition from the inertial focusing microchannel to the downstream expanding region that first gradually increases to avoid the formation of vortices that can trap and disturb cell trajectories, but then rapidly extends the cross-sectional area of the expanding region in a short distance as one moves along the direction of the flow, to sufficiently expand the distance between differentially-focused cells. In this regard, the expansion region was designed by increasing the angle between the channel wall and flow direction by 2° per 100 µm28. The serpentine microchannels serving as fluidic resistors at the outlets is composed of 20 U-shaped periods with a channel width of 40 μm and a total path length of around 2.5 cm.
Cell culture
Euglena gracilis Z (NIES-48) was obtained from Microbial Culture Collection at National Institute for Environmental Studies (NIES), Japan. E. gracilis cells were cultured autotrophically in 50 ml tubes using Cramer-Myers (CM) medium39 at pH = 5.5. The cultures were exposed to a continuous light with an intensity of 100 μmol photons m−2 s−1, aerated with filter-sterilized mixture of carbon dioxide and air in a volumetric ratio of 15% at a flow rate of 1 L/min, and incubated at a constant temperature of 29 °C. Due to biological, metabolic, and environmental differences, the cultured E. gracilis cells have different shapes and sizes. Viability of E. gracilis cells was examined using fluorescence-based live-dead assays with fluorescein diacetate (FDA, Sigma-Aldrich) and propidium iodide (PI, Sigma-Aldrich), which can stain viable and dead cells, respectively (see Supplementary Fig. S4). The concentrations of FDA and PI are 10 µg/mL and 20 µg/mL, respectively, in phosphate buffered saline (PBS, Thermo Fisher Scientific).
High-speed imaging and data analysis
E gracilis suspensions with a concentration of around 105 cells/mL were injected into the microchannel at flow rates ranging from 300 to 800 µL/min using a syringe pump (Harvard Apparatus, MA, USA). The motion and behaviors of E. gracilis cells were monitored and recorded using an inverted microscope (Nikon Ti, Japan) equipped with a high-speed camera (Vision Research, NJ, USA). The high-speed images were captured with an exposure time of 1 µs, and frame rates were varied according to the channel region for measurement. Recorded videos and image sequences were processed using Phantom Camera Control Software (PCC) (https://www.phantomhighspeed.com/) and a custom-built MATALB routine (MathWorks, MA, USA). The code determines the center, maximum/minimum length (or AR) of individual E. gracilis cells in each image frame. The cell lateral position is calculated by measuring the distance between the cell center and the channel centerline.
References
Novak, M. & Vetvicka, V. β-Glucans, History, and the Present: Immunomodulatory Aspects and Mechanisms of Action. Journal of Immunotoxicology 5, 47–57 (2008).
Koizumi, N. et al. Anti-HIV (human immunodeficiency virus) activity of sulfated paramylon. Antiviral Research 21, 1–14 (1993).
Watanabe, T. et al. Antitumor activity of the beta-glucan paramylon from Euglena against preneoplastic colonic aberrant crypt foci in mice. Food & function 4, 1685–1690 (2013).
Sugiyama, A. et al. Oral administration of paramylon, a beta-1,3-D-glucan isolated from Euglena gracilis Z inhibits development of atopic dermatitis-like skin lesions in NC/Nga mice. The Journal of veterinary medical science 72, 755–763 (2010).
Shibakami, M., Tsubouchi, G., Nakamura, M. & Hayashi, M. Preparation of carboxylic acid-bearing polysaccharide nanofiber made from euglenoid β-1,3-glucans. Carbohydrate Polymers 98, 95–101 (2013).
Furuhashi, T. et al. Wax ester and lipophilic compound profiling of Euglena gracilis by gas chromatography-mass spectrometry: toward understanding of wax ester fermentation under hypoxia. Metabolomics 11, 175–183 (2015).
Mokrosnop, V. M., Polishchuk, A. V. & Zolotareva, E. K. [Accumulation of alpha-tocopherol and beta-carotene in Euglena gracilis Cells under Autotrophic and Mixotrophic Culture Conditions]. Prikladnaia biokhimiia i mikrobiologiia 52, 230–236 (2016).
Ross, G. I. M. Vitamin B(12) Assay in Body Fluids using Euglena gracilis. Journal of Clinical Pathology 5, 250–256 (1952).
Park, K. Y., Lim, B. R. & Lee, K. Growth of microalgae in diluted process water of the animal wastewater treatment plant. Water science and technology: a journal of the International Association on Water Pollution Research 59, 2111–2116 (2009).
Brayner, R. et al. ZnO Nanoparticles: Synthesis, Characterization, and Ecotoxicological Studies. Langmuir 26, 6522–6528 (2010).
Lonergan, T. A. Regulation of Cell Shape in Euglena gracilis: I. Involvement of the Biological Clock, Respiration, Photosynthesis, and Cytoskeleton. Plant Physiology 71, 719–730 (1983).
Goto, K., Laval-Martin, D. & Edmunds, L. Biochemical modeling of an autonomously oscillatory circadian clock in Euglena. Science 228, 1284–1288 (1985).
Lonergan, T. A. The Photosynthesis and Cell Shape Rhythms can be Naturally Uncoupled from the Biological Clock in Euglena gracilis. Journal of Experimental Botany 37, 1334–1340 (1986).
Buetow, D. E. In eLS (John Wiley & Sons, Ltd, 2001).
Murray, J. M. Control of cell shape by calcium in the euglenophyceae. Journal of cell science 49, 99–117 (1981).
Lonergan, T. A. Regulation of cell shape in Euglena gracilis. II. The effects of altered extra- and intracellular Ca2 + concentrations and the effect of calmodulin antagonists. Journal of cell science 71, 37–50 (1984).
Pogo, A. O. & Arce, A. Synchronization of cell division in Euglena gracilis by heat shock. Experimental Cell Research 36, 390–397 (1964).
O’Neill, E. C. et al. The transcriptome of Euglena gracilis reveals unexpected metabolic capabilities for carbohydrate and natural product biochemistry. Molecular bioSystems 11, 2808–2820 (2015).
Sugaya, S., Yamada, M. & Seki, M. Observation of nonspherical particle behaviors for continuous shape-based separation using hydrodynamic filtration. Biomicrofluidics 5, 024103 (2011).
Beech, J. P., Holm, S. H., Adolfsson, K. & Tegenfeldt, J. O. Sorting cells by size, shape and deformability. Lab on a Chip 12, 1048–1051 (2012).
Holm, S. H., Beech, J. P., Barrett, M. P. & Tegenfeldt, J. O. Separation of parasites from human blood using deterministic lateral displacement. Lab on a Chip 11, 1326–1332 (2011).
Valero, A. et al. Tracking and synchronization of the yeast cell cycle using dielectrophoretic opacity. Lab on a Chip 11, 1754–1760 (2011).
Zhou, R., Bai, F. & Wang, C. Magnetic separation of microparticles by shape. Lab on a Chip 17, 401–406 (2017).
Lu, X. & Xuan, X. Elasto-Inertial Pinched Flow Fractionation for Continuous Shape-Based Particle Separation. Analytical Chemistry 87, 11523–11530 (2015).
Lu, X., Zhu, L., Hua, R.-m & Xuan, X. Continuous sheath-free separation of particles by shape in viscoelastic fluids. Applied Physics Letters 107, 264102 (2015).
Di Carlo, D. Inertial microfluidics. Lab on a Chip 9, 3038–3046 (2009).
Amini, H., Lee, W. & Di Carlo, D. Inertial microfluidic physics. Lab on a Chip 14, 2739–2761 (2014).
Hur, S. C., Henderson-MacLennan, N. K., McCabe, E. R. B. & Di Carlo, D. Deformability-based cell classification and enrichment using inertial microfluidics. Lab on a Chip 11, 912–920 (2011).
Masaeli, M. et al. Continuous Inertial Focusing and Separation of Particles by Shape. Physical Review X 2, 031017 (2012).
Li, M. et al. Inertial focusing of ellipsoidal Euglena gracilis cells in a stepped microchannel. Lab on a Chip 16, 4458–4465 (2016).
Gossett, D. R. et al. Inertial Manipulation and Transfer of Microparticles Across Laminar Fluid Streams. Small 8, 2757–2764 (2012).
Lee, W., Amini, H., Stone, H. A. & Di Carlo, D. Dynamic self-assembly and control of microfluidic particle crystals. Proceedings of the National Academy of Sciences 107, 22413–22418 (2010).
Humphry, K. J., Kulkarni, P. M., Weitz, D. A., Morris, J. F. & Stone, H. A. Axial and lateral particle ordering in finite Reynolds number channel flows. Physics of Fluids 22, 081703 (2010).
Wakisaka, Y. et al. Probing the metabolic heterogeneity of live Euglena gracilis with stimulated Raman scattering microscopy. Nature Microbiology 1, 16124 (2016).
Yamada, K. et al. Efficient selective breeding of live oil-rich Euglena gracilis with fluorescence-activated cell sorting. Scientific Reports 6, 26327 (2016).
Guo, B. et al. High-throughput, label-free, single-cell, microalgal lipid screening by machine-learning-equipped optofluidic time-stretch quantitative phase microscopy. Cytometry Part A, n/a–n/a (2017).
Lei, C. et al. High-throughput label-free image cytometry and image-based classification of live Euglena gracilis. Biomedical optics express 7, 2703–2708 (2016).
Qin, D., Xia, Y. & Whitesides, G. M. Soft lithography for micro- and nanoscale patterning. Nat. Protocols 5, 491–502 (2010).
Cramer, M. & Myers, J. Growth and photosynthetic characteristics of euglena gracilis. Archiv für Mikrobiologie 17, 384–402 (1952).
Acknowledgements
This work is primarily supported by ImPACT Program of the Council of Science, Technology and Innovation (Cabinet office, Government of Japan), and partly by Burroughs Wellcome Fund. We thank Dr. Soojung Claire Hur for help with device design, Dr. Osamu Iwata and Dr. Hideyuki Suzuki in Euglena Co., Ltd. for help with cell culture. We also thank ISNC microfabrication facilities, California NanoSystems Institute for mold fabrication.
Author information
Authors and Affiliations
Contributions
M.L., K.G. and D.D.C. designed the experiments. M.L. performed the experiments. M.L. and H.E.M. analyzed the data. All authors contributed to the manuscript writing.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, M., Muñoz, H., Goda, K. et al. Shape-based separation of microalga Euglena gracilis using inertial microfluidics. Sci Rep 7, 10802 (2017). https://doi.org/10.1038/s41598-017-10452-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-10452-5
This article is cited by
-
Acoustofluidic separation of prolate and spherical micro-objects
Microsystems & Nanoengineering (2024)
-
Microfluidic chip-assisted separation process and post-chip microalgae cultivation for carotenoid production
Journal of Applied Phycology (2024)
-
Addition of humic acid accelerates the growth of Euglena pisciformis AEW501 and the accumulation of lipids
Journal of Applied Phycology (2022)
-
Square microchannel enables to focus and orient ellipsoidal Euglena gracilis cells by two-dimensional acoustic standing wave
Microchimica Acta (2022)
-
Application of electrical treatment on Euglena gracilis for increasing paramylon production
Applied Microbiology and Biotechnology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.