Abstract
Corynebacterium striatum is a nosocomial opportunistic pathogen increasingly associated with a wide range of human infections and is often resistant to several antibiotics. We investigated the susceptibility of 63 C. striatum isolated at the Farhat-Hached hospital, Sousse (Tunisia), during the period 2011–2014, to a panel of 16 compounds belonging to the main clinically relevant classes of antimicrobial agents. All strains were susceptible to vancomycin, linezolid, and daptomycin. Amikacin and gentamicin also showed good activity (MICs90 = 1 and 2 mg/L, respectively). High rates of resistance to penicillin (82.5%), clindamycin (79.4%), cefotaxime (60.3%), erythromycin (47.6%), ciprofloxacin (36.5%), moxifloxacin (34.9%), and rifampicin (25.4%) were observed. Fifty-nine (93.7%) out of the 63 isolates showed resistance to at least one compound and 31 (49.2%) were multidrug-resistant. Twenty-nine resistance profiles were distinguished among the 59 resistant C. striatum. Most of the strains resistant to fluoroquinolones showed a double mutation leading to an amino acid change in positions 87 and 91 in the quinolone resistance-determining region of the gyrA gene. The 52 strains resistant to penicillin were positive for the gene bla, encoding a class A β-lactamase. Twenty-two PFGE patterns were identified among the 63 C. striatum, indicating that some clones have spread within the hospital.
Similar content being viewed by others
Introduction
Corynebacterium species are widely distributed in the environment and in the microbiota of humans and animals. Medically relevant Corynebacterium species include Corynebacterium diphtheriae, the primary cause of diphtheria, and the non-diphtherial corynebacteria, which are part of the normal flora of the skin and mucous membranes. Nondiphtherial corynebacteria have been frequently dismissed as a contaminant when isolated from clinical materials. However, the role of these bacteria in clinical disease is now more clearly established. Corynebacterium striatum is recognized as a true pathogen when isolated in several samples from sterile body sites or from indwelling medical devices1, 2. Consideration of whether an isolate represents infection, colonization, or contamination is based upon clinical assessment. A variety of infections have been associated with this bacterium: bacteremia, endocarditis, valvular damage, meningitis, vaginitis and infections of the urinary tract, the respiratory tract, wounds, skin and eye3,4,5,6,7,8,9,10.
Susceptibility testing of C. striatum is necessary to establish a specific therapy. Although initial studies indicated that C. striatum clinical isolates were susceptible to a wide range of antibiotics11, 12, recent reports have shown increased multidrug resistance13, 14. Patients suffering underlying diseases and receiving multiple antibiotic courses are at high risk for developing serious opportunistic infections by drug-resistant C. striatum strains, which can be at the origin of major outbreaks. Thus, several outbreaks of clonal multidrug-resistant C. striatum have been recently reported15,16,17.
C. striatum, considered as an emerging pathogen in different countries, has been the most frequently isolated Corynebacterium species since 2011 at the Farhat-Hached University (FHU) hospital, Sousse, Tunisia. In this study, we aimed to investigate the susceptibility of the 63 C. striatum isolated at the FHU hospital during the period 2011–2014 to 16 agents, representative of the main groups of antibiotics, including the most used compounds to treat infections caused by Corynebacterium spp. as well as second-line and complementary agents. We describe the high incidence of drug resistance in our C. striatum and characterize the molecular mechanisms related to resistance to aminoglycosides, compounds of the MLSB group, fluoroquinolones and β-lactams. The clonal relationships among C. striatum isolates were analysed by Pulsed-field Gel Electrophoresis (PFGE).
Results
Antimicrobial susceptibility testing and resistance profiles
Fifty-nine (93.7%) out of the 63 isolates showed resistance to at least one of the 16 tested compounds, whereas the remaining 4 isolates were susceptible. The MIC50 and MIC90 distributions and the percentage of resistance to the different antibiotics for the 63 C. striatum included in this study are presented in Table 1.
The MIC90 values of vancomycin, daptomycin and linezolid were in the range 0.25–0.5 mg/L, and none of the 63 isolates showed resistance against them. Considering the MIC90 values, the most active compounds were vancomycin and daptomycin (MIC90 = 0.25 mg/L for both compounds) followed by linezolid (MIC90 = 0.5 mg/L). Among the 5 aminoglycosides tested, amikacin and gentamicin were the most active compounds (MIC90 values of 1 and 2 mg/L, respectively). Only three C. striatum were resistant to both compounds (MICs > 64 mg/L). Tobramycin was also very active (56 isolates were susceptible). Kanamycin and streptomycin showed low activity against our C. striatum (MICs90 > 64 mg/L). The compounds of the MLSB group were poorly active against our C. striatum (MICs90 of 8 and >64 mg/L for erythromycin and clindamycin, respectively). Rifampicin was poorly active too (MIC90 = 16 mg/L). Ciprofloxacin and moxifloxacin also showed low activity against our isolates (MIC90 > 16 mg/L for both compounds). A high number of the isolates were resistant to the β-lactams penicillin and cefotaxime (MICs90 = 16 mg/L).
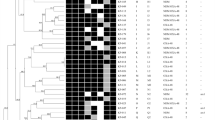
The 59 C. striatum resistant to at least one of the tested compounds were divided in 29 resistance profiles (Fig. 1). The most frequently encountered resistance phenotype was penicillin (12 strains), followed by penicillin-cefotaxime and erythromycin-clindamycin-penicillin (6 strains each). Multidrug resistance (MDR), defined as non-susceptibility to at least one agent in three or more antimicrobial categories (as defined for other microorganisms18), was observed in 31 (49.2%) strains. Among the 31 MDR isolates, 19 resistance profiles could be distinguished. Eight MDR isolates showed resistance to eight or more compounds, belonging to seven main classes of antibiotics (aminoglycosides, ansamycins, macrolides, lincosamides, fluoroquinolones, penicillins and cephalosporins).
Molecular detection of resistance genes
The aph(3′)-Ic gene was detected in the 10 strains resistant to kanamycin (MIC > 64 mg/L). However, fourteen strains susceptible to kanamycin were also aph(3′)-Ic-positive. The genes aph(3″)-Ib and aph(6)-Id were detected in 8 isolates, five of them resistant to streptomycin (MICs > 64 mg/L), and the other three showed MICs of streptomycin in the range 1–4 mg/L. The gene aac(3)-XI was found in 7 isolates. Two of them were susceptible to both gentamicin and tobramycin (MICs < 0,06 mg/L) whereas the other five isolates showed MICs of 8 and 16 mg/L for gentamicin and tobramycin, respectively.
Twenty out of the 24 C. striatum resistant to erythromycin carried the erm(X) gene. Ten out of 20 carried erm(X) plus the erm(B) gene. From the 24 clindamycin-resistant isolates 22 were positive for the gene erm(X) and 10 out of 22 were also erm(B)-positive. The mef(A-E) gene was not found in any strain tested.
The sequences of the QRDR region of the gyrA gene of 21 isolates categorized as resistant or intermediate to fluoroquinolones were compared to that of the quinolone-susceptible C. striatum ATCC 6940 (GenBank accession number AY559038). The relationships between the MICs of ciprofloxacin and moxifloxacin and the mutations in the gyrA QRDRs of the 21 isolates are summarized in Table 2. Eleven strains showing MICs = 16 mg/L for these two compounds carried a double mutation at resistance hotspots Ser-87 and Asp-91 (C. striatum numbering), generating a change from Ser-87 to Phe and another change from Asp-91 to Gly or Ala. In two double mutants showing the same changes the increase of MICs of moxifloxacin was limited to 8 mg/L. Another double mutant with Ala instead of Ser in position 87 and Gly instead of Asp in position 91 showed lower MIC of moxifloxacin (4 mg/L) whereas the MIC of ciprofloxacin was >16 mg/L. Three strains had a mutation at amino acid codon Ser-87 that changes this amino acid for Tyr or Phe. These single mutants showed lower increases of MICs of ciprofloxacin (to 4 or 8 mg/L) and a moderate increase of MICs of moxifloxacin (to 1 mg/L). Two strains showed a replacement in the amino acid codon Asp-91, producing a change from Asp to Gly. The MICs of ciprofloxacin and moxifloxacin for these strains were 2 and 1 mg/L, respectively. Two fluoroquinolone-resistant isolates lacked mutations in their QRDRs.
The 52 isolates showing resistance to penicillin were positive for the gene bla, encoding a class A β-lactamase. The C. striatum bla gene encodes a serine hydrolase belonging to the class A β-lactamase family protein. Forty-six out of the 63 C. striatum were positive for the ampC gene, encoding a class C β-lactamase. Forty-two of the 46 ampC-positive isolates were penicillin-resistant whereas 4 isolates were sensitive. The ampC gene was detected in 22 of the 28 cefotaxime-resistant C. striatum. The 52 penicillin-resistant isolates had MICs of cefotaxime in the range 1–256 mg/L.
Analysis of the strain clonal relationship by PFGE
XbaI digestion of the 63 C. striatum isolates revealed 22 distinct PFGE patterns, which were designated from A to V (Fig. 1). On these, PFGE types D and G could be further classified into 3 subtypes. Pulsotype E was predominant (n = 12 isolates) and it was observed in 11 MDR strains, six of which were isolated in the neonatology ward. The remaining 20 MDR clinical isolates belonged to PFGE patterns A (n = 6), C, D2, G3 (n = 2), H, J, K, N, O (n = 2), P, Q (n = 2) and S, indicating that they were unrelated. Other patterns including a significant number of isolates were A (n = 7), D (n = 7) and G (n = 9). Relationship between PFGE patterns and antibiotic resistance profiles could not be established, although the most prevalent PFGE patterns (E and A) included principally MDR isolates. Two pairs of isolates assigned to pulsotype E showed the same resistance profile.
Discussion
One of the most serious problems related to treatment of the infections caused by C. striatum is the isolation of multidrug-resistant strains from clinical material and the selection of the appropriate antibiotic therapy for a given type of infection. C. striatum infections should be treated according to the results of the susceptibility tests. Multiple drug resistance is caused by the interplay of multiple resistance mechanisms those emerge via the acquisition of extraneous resistance determinants or spontaneous mutations. This work highlights the high prevalence of multi-resistant strains and resistance genes among the C. striatum isolated in a hospital in Tunisia, in particular resistance to aminoglycosides, compounds of the MLSB group, fluoroquinolones, and β-lactams.
The 63 C. striatum analysed in this study were susceptible to vancomycin and linezolid. In an earlier report using the disk diffusion method, Martinez et al.19 showed that 31 C. striatum isolated from clinical samples were susceptible to vancomycin. More recently, Gomila et al.20 reported that their 52 C. striatum isolated from patients with chronic obstructive respiratory disease were susceptible to vancomycin too. Vancomycin is still active today and therefore it represents an adequate option for treatment of severe infections caused by C. striatum. Linezolid has also shown an excellent activity, with MICs routinely below 0.5 mg/L21. Gómez-Garcés et al.22 showed that vancomycin and linezolid were equally active against 30 clinical C. striatum (MICs90 of both compounds = 0.5 mg/L). Linezolid can be considered as an alternative to vancomycin against C. striatum, although its side effects during the long courses of treatment required for hardware or device-associated infections must be pondered. All our C. striatum were susceptible to daptomycin (MIC90 = 0.25 mg/L). Daptomycin has also been proven to be active against C. striatum, alone23 or combined with rifampicin24. Therefore, daptomycin can also be considered as an alternative to vancomycin for treatment of C. striatum infections, although rapid emergence of high-level daptomycin resistance has been recently reported25, 26.
Aminoglycosides are used as complementary antibiotics to treat serious infections caused by diphtheroids. Among aminoglycosides, amikacin and gentamicin showed good activity “in vitro” against our C. striatum (MICs90 = 1 and 2 mg/L, respectively). Aminoglycoside resistance occurs through several mechanisms that can coexist simultaneously in the same cell. Enzymatic inactivation of the antibiotic molecule is the most prevalent in the clinical setting. The aac(3)-XI gene, encoding an aminoglycoside 3-N acetyl transferase conferring resistance to gentamicin and tobramycin in C. striatum 27, was not found in our 3 gentamicin-resistant isolates, suggesting that resistance is mediated by another mechanism. Kanamycin and streptomycin are not used in clinical practice in Tunisia but were tested in this study because they are good markers for detecting the presence of the aph(3′)-Ic, aph(3″)-Ib and aph(6)-Id genes. The aph(3′)-Ic gene, encoding an aminoglycoside-O-phosphotransferase implicated in resistance to kanamycin, neomycin, paromomycin, ribostamycin and lividomycin, is part of a larger DNA region containing the aph(3″)-Ib - aph(6)-Id tandem pair of resistance genes conferring streptomycin resistance in Corynebacterium spp28. Identical aminoglycoside resistance regions were found in the plasmid pTP10 from C. striatum in close vicinity to the erythromycin and chloramphenicol resistance regions29. As expected, the 10 C. striatum resistant to kanamycin carried the aph(3′)-Ic gene. However, this gene was also detected in strains susceptible to kanamycin, probably due to mutations affecting its coding sequence or its promoter. These results confirm that the aph(3′)-Ic gene is widespread in Corynebacterium spp. Resistance to streptomycin in Corynebacterium spp. is related to the presence of the tandem of genes aph(3″)-Ib and aph(6)-Id, encoding for aminoglycoside-3″-phosphotransferase [APH (3″)-Ib] and aminoglycoside-6-phosphotransferase [APH (6)-Id], respectively28. Five of the 8 isolates resistant to streptomycin carried the aph(3″)-Ib and aph(6)-Id genes, and the remaining three had MICs for streptomycin in the range of 1 to 4 mg/L, probably as a consequence of mutations in the above mentioned genes or in their promoter. Other mechanisms such as active efflux of the antimicrobial and reduced intake into the bacterial cell can contribute to streptomycin resistance in these isolates. Amikacin is eventually prescribed in combinative therapy against severe infections caused by C. striatum at the FHU hospital. Our results indicate that amikacin is the preferable aminoglycoside for treatment of C. striatum infections whereas gentamicin could be a valid alternative. However, the occurrence of resistant strains requires continual vigilance.
Erythromycin and clindamycin were inactive against the majority of our C. striatum, in particular clindamycin, with a MIC90 eight times greater than that of erythromycin. This fact confirms the previously reported high prevalence of resistance to compounds of the MLSB group among Corynebacterium spp., including C. striatum 30. MLS resistance in Corynebacterium spp. is most often mediated by two mechanisms: target-site modification mediated by ribosomal RNA methylases codified by the so-called erm genes and active drug-efflux mediated by a membrane efflux pump encoded by the mef(A-E) gene31. Our results confirmed those of previous studies which pointed out that erm(X) is the most important gene implicated in MLS resistance in Corynebacterium spp29, 32. For the first time we have detected the erm(B) gene encoding the ribosomal RNA methylase Erm(B) in C. striatum. The gene erm(B) confers high-level resistance to macrolides in Campylobacter coli 33 and other relevant pathogens but is exceptional in Corynebacterium spp31, 34. Ten of our strains carried the erm(B) and erm(X) genes simultaneously, a characteristic previously reported only in one strain of C. urealyticum 31.
One third of our C. striatum showed intermediate or high-level resistance to ciprofloxacin and moxifloxacin. Fluoroquinolones have been intensively used at the FHU hospital during the last two decades. Upon antibiotic administration, a selective pressure is created in body organs where fluoroquinolones tend to accumulate. Exposure to fluoroquinolones selects for spontaneous mutants in large bacterial populations, including those that colonize the skin and mucous membranes such as corynebacteria. Thus, fluoroquinolone resistance emerged in clinical isolates of C. striatum and C. amycolatum 35. Resistance to fluoroquinolones in Corynebacterium spp. is caused by mutations in the QRDR of the gyrase gene gyrA. In our strains, single amino-acid substitutions in position 87 of the GyrA protein generated ciprofloxacin resistance but double mutations in the gyrA gene leading to changes in positions 87 and 91 were necessary for high level resistance to ciprofloxacin and moxifloxacin. In fourteen of the 21 fluoroquinolone-resistant C. striatum, increases of MICs of ciprofloxacin and moxifloxacin until 16 mg/L were related to a double non-conservative mutation at positions 87 and 91. Sierra et al.35 also reported double mutations at positions 87 and 91 in the gyrA gene of six of their C. striatum, although the MICs of moxifloxacin for their strains were lower (6–8 mg/L). Five of our strains with single mutations at positions 87 or 91 are still resistant to ciprofloxacin although with lower MICs (in the range 2–8 mg/L) whereas the MICs of moxifloxacin remained at 1 mg/L. Single mutations in the residue Ser-87 or in the residue Asp-91 described by Sierra et al.35 increased the MICs of ciprofloxacin to 1–6 mg/L, whereas remaining susceptible to moxifloxacin. The higher level of moxifloxacin resistance in our strains suggest the existence of a resistance mechanism additional to mutations in gyrA. In two strains no changes in their QRDRs were detected, indicating that resistance was mediated by a different mechanism.
β-lactams are the most broadly used class of antimicrobials. Successful treatments of C. striatum infections with penicillin36 or amoxicillin37 have been reported. However, low susceptibility to penicillin and cefotaxime among other β-lactams have been communicated10, 38, although the genetic mechanism of resistance has not been characterized so far. Considering the MICs90 values (16 mg/L), penicillin and cefotaxime showed the same low activity against our C. striatum. The fact that the rate of penicillin-resistant strains is higher than that of cefotaxime-resistant strains is explained because the CLSI susceptibility breakpoint for penicillin has recently been dropped from 1 mg/l to 0.125 mg/L39. Hydrolysis of β-lactam antibiotics by β-lactamases is the most common mechanism of resistance for this class of antibacterial agents in clinically important bacteria. The β-lactamases are classified by protein sequence in four molecular classes, A, B, C, and D, based on conserved and distinguishing amino acid motifs. Fifty-two of our strains were resistant to penicillin and this resistance was related to the presence of a bla gene encoding a class A β-lactamase. The chromosomes of Corynebacterium jeikeium K41140, Corynebacterium urealyticum DSM 710941, and Corynebacterium resistens DSM 4510042, encode the corresponding counterparts of the C. striatum bla gene, although it has not been associated with resistance to β-lactams in these species. The ampC gene, encoding a class C β-lactamase, was detected in 42 of the 52 penicillin-resistant C. striatum. The ampC genes, which are widely distributed among the Enterobacteriaceae, encode enzymes active on both penicillins and cephalosporins43. We show here that two β-lactamase-encoding genes, bla and ampC, are present in β-lactam-resistant C. striatum. Our data revealed high resistance rates to β-lactams and a high prevalence of the bla and the ampC genes among the C. striatum isolated in our hospital. These data are of value to practitioners, discouraging the use of β-lactam compounds for the treatment of infections caused by C. striatum.
PFGE is considered the gold standard in epidemiological studies of pathogenic microorganisms, providing important insights into their population structure44. Our results showed 22 distinct PFGE patterns from 63 C. striatum strains. The high diversity of genotypes among the 63 C. striatum revealed that they are mainly not closely related. Therefore, C. striatum at the FHU hospital may originate from different lineages and sources instead of expansion of a single clonal lineage. This corresponds to the pathogenic condition of C. striatum as an opportunistic pathogen that causes occasional disease in predisposed patients. Some PFGE patterns were more frequently isolated, suggesting the existence of a few more prevalent clones. We consider patterns E and A as high-prevalence pulsotypes. Most of C. striatum assigned to pulsotypes E and A were highly resistant, indicating that the most prevalent clones are highly resistant, as has been previously reported10, 16. The fact that many different antibiotic resistance profiles could be distinguished among the strains belonging to a particular PFGE pattern revealed that there was not a single strain but several closely related clones producing sporadic infections.
In conclusion, this study highlights the relevance of C. striatum as an emerging multidrug-resistant nosocomial pathogen at the FHU hospital. The C. striatum isolates showed 100% susceptibility to vancomycin, linezolid and daptomycin and high rates of resistance to rifampicin, compounds of the MLSB group, fluoroquinolones and β-lactams. Among the several clones of C. striatum circulating at the FHU hospital, the most prevalent were the most resistant. Therefore, surveillance of MDR C. striatum should be continued.
Methods
Bacterial strains and growth conditions
During the period 2011–2014, 90 strains recovered from clinical specimens submitted for routine culture to the microbiology laboratory of the FHU hospital were assigned to the genus Corynebacterium on the basis of colony morphology, Gram staining, and catalase production. They were isolated in pure culture except the 7 specimens from vaginal swabs, where the Corynebacterium were the predominant microorganisms in a poly-microbial culture. In that cases the Corynebacterium were considered of clinical significance since they were associated with a strong leukocyte reaction in Gram staining45. In all cases the Corynebacterium were isolated after two different culture sets. Sixty-three out of the 90 strains were identified as putative C. striatum using API Coryne V2.0 strips (bioMérieux, Marcy l’Etoile, France). C. striatum was differentiated from C. amycolatum by additional phenotypic tests (tyrosine hydrolysis, N-acetylglucosamine assimilation, phenylacetic acid assimilation and susceptibility to the vibriostatic agent O/129). Identification was confirmed by MALDI-TOF using the Vitek MS (bioMérieux) system, in accordance with manufacturer’s instructions. The anatomical sites of specimens from whom the 63 C. striatum were isolated and the clinical diagnosis for the infected patients are shown in Table 3. All strains were grown on blood agar plates at 37 °C and kept frozen at −80 °C in Brain Heart Infusion broth with 20% glycerol until use.
Antimicrobial susceptibility assays
Antimicrobial susceptibilities were determined by micro-dilution in cation adjusted Muller-Hinton broth and interpreted following Clinical and Laboratory Standards Institute (CLSI) guidelines39. Sixteen antimicrobials were tested: amikacin, gentamicin, kanamycin, streptomycin, tobramycin, rifampicin, erythromycin, clindamycin, doxycycline, ciprofloxacin, penicillin, cefotaxime, vancomycin (all of them purchased from Sigma Aldrich, Madrid, Spain), moxifloxacin (Discovery Fine Chemicals, Dorset, United Kingdom), linezolid (Pfizer, Bilbao, Spain) and daptomycin (Cubist, Madrid, Spain). Daptomycin broth was supplemented to 50 mg/L calcium for determinations of susceptibility to that drug. Of note, CLSI susceptible interpretive criteria for penicillin has been recently dropped from 1 mg/l to 0.125 mg/L39. MIC values of the antibiotics not considered in CLSI guidelines for Corynebacterium spp. (amikacin, tobramycin, kanamycin, moxifloxacin and linezolid) were interpreted in accordance to criteria defined by CLSI for Staphylococcus aureus 46. Since the CLSI lacks breakpoints of streptomycin for staphylococci we have considered the MIC breakpoints values proposed by the French Society of Microbiology (http://www.sfmmicrobiologie.org/UserFiles/files/casfm/CASFM2013vjuin.pdf) to classify the C. striatum as susceptible, intermediate or resistant to streptomycin. Escherichia coli ATCC 25922 and Streptococcus pneumoniae ATCC49619 were used as control strains for susceptibility testing assays.
Amplification and sequencing of genes related to resistance
The presence of aminoglycoside modifying enzyme (AME) genes common in Corynebacterium spp. [aph(3′)-Ic, aac(3)-XI, and the tandem of genes aph(3″)-Ib and aph(6)-Id] was investigated by PCR. Resistance to compounds of the MLSB group was investigated by amplification of the erm(X), erm(B) and mef(A-E) genes. Resistance to quinolones in Corynebacterium spp. is related to point mutations in the sequence of the quinolone resistance-determining region (QRDR) of the gyrA gene. Thereby, the QRDR at that gene was amplified and sequenced as previously described35. The 63 C. striatum were analysed for the presence of the bla gene, encoding a class A β-lactamase involved in resistance to penicillins and cephalosporins in Corynebacterium spp. Primers used to amplify the above mentioned genes are listed in Supplementary Table S1. PCR reactions were performed as previously described47. PCR products were purified using the QIAquick PCR Purification kit (Qiagen, Madrid, Spain). Purified DNA was sequenced by Macrogen (Seul, Korea) with the primers outlined in Table S1. Mutations in gyrA were identified by aligning sequences of resistant isolates to the sequence of C. striatum ATCC6940 (GenBank accession number AY559038) using the Clustal W program47.
Pulsed-field Gel Electrophoresis and dendrogram analysis
We obtained XbaI macro-restriction patterns of the 63 C. striatum with a published protocol48 and a CHEF-DRIII variable angle system (Bio-Rad, Hercules, California, USA). The PFGE patterns were analysed with Fingerprinting II v4.5 software (Bio-Rad). Each isolate was compared with all other isolates using the Dice similarity coefficient and the unweighted pair Group method with arithmetic means (UPGMA), with 1% of optimization and tolerance. Isolates were classified as indistinguishable if they showed 100% similarity, as closely related subtypes if they showed 95–99% similarity, and as different strains if they showed <95% similarity.
Ethics statement
This study was performed in accordance with the ethical guidelines of the Declaration of Helsinki (1975). Written informed consent was obtained from each patient from whom samples were taken.
References
Boltin, D. et al. Corynebacterium striatum-A classic pathogen eluding diagnosis. Eur. J. Intern. Med. 20, 49–52 (2009).
Martínez-Martínez, L., Suárez, A. I., Rodríguez-Baño, J., Bernard, K. & Muniáin, M. A. Clinical significance of Corynebacterium striatum isolated from human samples. Clin. Microbiol. Infect. 3, 634–639 (1997).
Ishiwada, N. et al. Clinical and bacteriological analyses of bacteremia due to Corynebacterium striatum. J. Infect. Chemother. 22, 790–793 (2016).
Hong, H. L., Koh, H. I. & Lee, A. J. Native valve endocarditis due to Corynebacterium striatum confirmed by 16S ribosomal RNA sequencing: a case report and literature review. Infect. Chemother. 48, 239–245 (2016).
Chen, F. L., Hsueh, P. R., Teng, S. O., Ou, T. Y. & Lee, W. S. Corynebacterium striatum bacteremia associated with central venous catheter infection. J. Microbiol. Immunol. Infect. 45, 255–258 (2012).
Weiss, K., Labbé, A. C. & Laverdière, M. Corynebacterium striatum meningitis: case report and review of an increasingly important Corynebacterium species. Clin. Infect. Dis. 23, 1246–1248 (1996).
Beteta López, A., Gil Ruiz, M. T., Vega Prado, L. & Fajardo Olivares, M. Cystitis and haematuria due to Corynebacterium striatum. A case report and review. Actas Urol. Esp. 33, 909–912 (2009).
Roig-Rico, P., Safont-Gaso, P., Marín-Tordera, D. & Ortíz de la Tabla, V. Corynebacterium striatum pneumonia in an HIV patient. Enferm. Infecc. Microbiol. Clin. 29, 402 (2011).
Olender, A. & Letowska, I. Wound infections due to opportunistic Corynebacterium species. Med. Dosw. Mikrobiol. 62, 135–140 (2010).
Otsuka, Y. et al. Emergence of multidrug-resistant Corynebacterium striatum as a nosocomial pathogen in long-term hospitalized patients with underlying diseases. Diagn. Microbiol. Infect. Dis. 54, 109–114 (2006).
Soriano, F., Zapardiel, J. & Nieto, E. Antimicrobial susceptibilities of Corynebacterium species and other non-spore-forming gram-positive bacilli to 18 antimicrobial agents. Antimicrob. Agents Chemother. 39, 208–214 (1995).
Martínez-Martínez, L., Pascual, A., Bernard, K. & Suárez, A. I. Antimicrobial susceptibility pattern of Corynebacterium striatum. Antimicrob. Agents Chemother. 40, 2671–2672 (1996).
Yoo, G. et al. Multidrug-resistant Corynebacterium striatum bacteremia: first case in Korea. Ann. Lab. Med. 35, 472–473 (2015).
Hahn, W. O., Werth, B. J., Butler-Wu, S. M. & Rakita, R. M. Multidrug-resistant Corynebacterium striatum associated with increased use of parenteral antimicrobial drugs. Emerg. Infect. Dis. 22, 1908–1914 (2016).
Campanile, F. et al. Clonal multidrug-resistant Corynebacterium striatum strains, Italy. Emerg. Infect. Dis. 15, 75–78 (2009).
Verroken, A. et al. Epidemiological investigation of a nosocomial outbreak of multidrug-resistant Corynebacterium striatum at one Belgian university hospital. Clin. Microbiol. Infect. 20, 44–50 (2014).
Baio, P. V. et al. Clonal multidrug-resistant Corynebacterium striatum within a nosocomial environment, Rio de Janeiro, Brazil. Mem. Inst. Oswaldo Cruz 108, 23–29 (2013).
Magiorakos, A. P. et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 18, 268–281 (2012).
Martínez-Martínez, L., Suárez, A. I., Winstanley, J., Ortega, M. C. & Bernard, K. Phenotypic characteristics of 31 strains of Corynebacterium striatum isolated from clinical samples. J. Clin. Microbiol. 33, 2458–2461 (1995).
Gomila, M. et al. Identification and diversity of multiresistant Corynebacterium striatum clinical isolates by MALDI-TOF mass spectrometry and by a multigene sequencing approach. BMC Microbiol. 12, e52, doi:10.1186/1471-2180-12-52 (2012).
Jones, R. N., Stilwell, M. G., Hogan, P. A. & Sheehan, D. J. Activity of linezolid against 3,251 strains of uncommonly isolated gram-positive organisms: report from the SENTRY antimicrobial surveillance program. Antimicrob. Agents Chemother. 51, 1491–1493 (2007).
Gómez-Garcés, J. L., Alos, J. I. & Tamayo, J. In vitro activity of linezolid and 12 other antimicrobials against coryneform bacteria. Int. J. Antimicrob. Agents 29, 688–692 (2007).
Fernández-Roblas, R. et al. In vitro activity of tigecycline and 10 other antimicrobials against clinical isolates of the genus. Corynebacterium. Int. J. Antimicrob. Agents 33, 453–455 (2009).
Shah, M. & Murillo, J. L. Successful treatment of Corynebacterium striatum endocarditis with daptomycin plus rifampin. Ann. Pharmacother. 39, 1741–1744 (2005).
Werth, B. J., Hahn, W. O., Butler W., S. M. & Rakita, R. M. Emergence of high-level daptomycin resistance in Corynebacterium striatum in two patients with left ventricular assist device infections. Microb. Drug Resist. 22, 233–237 (2016).
McElvania Tekippe, E., Thomas, B. S., Ewald, G. A., Lawrence, S. J. & Burnham, C. A. Rapid emergence of daptomycin resistance in clinical isolates of Corynebacterium striatum, a cautionary tale. Eur. J. Clin. Microbiol. Infect. Dis. 33, 2199–2205 (2014).
Galimand, M. et al. AAC (3)-XI, a new aminoglycoside 3-N-acetyltransferase from Corynebacterium striatum. Antimicrob. Agents Chemother. 59, 5647–5653 (2015).
Ramírez, M. S. & Tolmasky, M. E. Aminoglycoside modifying enzymes. Drug Resist Updat. 13, 151–171 (2010).
Tauch, A., Krieft, S., Kalinowski, J. & Pühler, A. The 51,409-bp R-plasmid pTP10 from the multi-resistant clinical isolate Corynebacterium striatum M82B is composed of DNA segments initially identified in soil bacteria and in plant, animal, and human pathogens. Mol. Gen. Genet. 263, 1–11 (2000).
Ortíz-Pérez, A. et al. High frequency of macrolide resistance mechanisms in clinical isolates of Corynebacterium species. Microb. Drug Resist. 16, 273–277 (2010).
Van Hoek, A. H. et al. Acquired antibiotic resistance genes: an overview. Frontiers in Microbiology 2, e203, doi:10.3389/fmicb.2011.00203 (2011).
Yagüe Guirao, G. et al. Implication of ermX genes in macrolide and telithromycin resistance in Corynebacterium jeikeium and Corynebacterium amycolatum. Rev. Esp. Quimioter. 18, 236–242 (2005).
Qin, S. et al. Report of ribosomal RNA methylase gene erm(B) in multidrug-resistant Campylobacter coli. J. Antimicrob. Chemother. 69, 964–968 (2014).
Luna, V. A. et al. A variety of Gram-positive bacteria carry mobile mef genes. J. Antimicrob. Chemother. 44, 19–25 (1999).
Sierra, J. M., Martínez-Martínez, L., Vázquez, F., Giralt, E. & Vila, J. Relationship between mutations in the gyrA gene and quinolone resistance in clinical isolates of Corynebacterium striatum and Corynebacterium amycolatum. Antimicrob. Agents Chemother. 49, 1714–1719 (2005).
Superti, S. V. et al. Corynebacterium striatum infecting a malignant cutaneous lesion: the emergence of an opportunistic pathogen. Rev. Inst. Med. Trop. S. Paulo 51, 115–116 (2009).
Brandenburg, A. H. et al. Patient-to-patient spread of a single strain of Corynebacterium striatum causing infections in a surgical intensive care unit. J. Clin. Microbiol. 34, 2089–2094 (1996).
Scholle, D. A spontaneous joint infection with Corynebacterium striatum. J. Clin. Microbiol. 45, 656–658 (2007).
CLSI. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of Infrequently Isolated or Fastidious Bacteria. 3rd ed. CLSI guideline M45. Wayne, PA: Clinical and Laboratory Standards Institute (2016).
Tauch, A. et al. Complete genome sequence and analysis of the multiresistant nosocomial pathogen Corynebacterium jeikeium K411, a lipid-requiring bacterium of the human skin flora. J. Bacteriol. 187, 4671–4682 (2005).
Tauch, A. et al. The lifestyle of Corynebacterium urealyticum derived from its complete genome sequence established by pyrosequencing. J. Biotechnol. 136, 11–21 (2008).
Schröder, J. et al. Complete genome sequence, lifestyle, and multi-drug resistance of the human pathogen Corynebacterium resistens DSM 45100 isolated from blood samples of a leukemia patient. BMC Genomics 13, e141, doi:10.1186/1471-2164-13-141 (2012).
Jacoby, G. A. AmpC β-lactamases. Clin. Microbiol. Rev. 22, 161–182 (2009).
Goering, R. V. Pulsed field gel electrophoresis: a review of application and interpretation in the molecular epidemiology of infectious disease. Infect. Genet. Evol. 10, 866–875 (2010).
Funke, G., Von Graevenitz, A., Clarridge, J. E. III & Bernard, K. A. Clinical microbiology of Coryneform bacteria. Clin. Microbiol. Rev. 10, 125–129 (1997).
CLSI. Performance Standards for Antimicrobial Susceptibility Testing. 26th ed. CLSI supplement M100S. Wayne, PA: Clinical and Laboratory Standards Institute (2016).
Thomson, J. D., Higgins, D. G., & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Navas, J., Fernández-Martínez, M., Salas, C., Cano, M. E. & Martínez-Martínez, L. Susceptibility to aminoglycosides and distribution of aph and aac(3)-XI genes among Corynebacterium striatum clinical isolates. PLoS ONE 11, e0167856, doi:10.1371/journal.pone.0167856 (2016).
Acknowledgements
This work was supported by Plan Nacional de I + D + i 2013–2016 and Instituto de Salud Carlos III, Subdirección General de Redes y Centros de Investigación Cooperativa, Ministerio de Economía, Industria y Competitividad, Spanish Network for Research in Infectious Diseases (REIPI RD12/0015) - co-financed by European Development Regional Fund “A way to achieve Europe” Operative program Intelligent Growth 2014–2020, and grants from the Farhat-Hached University Hospital and the University of Carthage (Tunisia).
Author information
Authors and Affiliations
Contributions
S.A., J.B., L.M.M. and J.N. conceived and designed the study, S.A. and A.F. collected and identified the clinical isolates, S.A., M.F.M., M.E.C. and J.N. conducted the experiments, S.A., M.F.M., J.N. and M.E.C. analysed the data, S.A., L.M.M. and J.N. drafted the manuscript. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alibi, S., Ferjani, A., Boukadida, J. et al. Occurrence of Corynebacterium striatum as an emerging antibiotic-resistant nosocomial pathogen in a Tunisian hospital. Sci Rep 7, 9704 (2017). https://doi.org/10.1038/s41598-017-10081-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-10081-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.