Abstract
Rhegmatogenous retinal detachment (RRD) is a serious condition that can lead to blindness; however, it is highly treatable with timely and appropriate treatment. Thus, early diagnosis and treatment of RRD is crucial. In this study, we applied deep learning, a machine-learning technology, to detect RRD using ultra–wide-field fundus images and investigated its performance. In total, 411 images (329 for training and 82 for grading) from 407 RRD patients and 420 images (336 for training and 84 for grading) from 238 non-RRD patients were used in this study. The deep learning model demonstrated a high sensitivity of 97.6% [95% confidence interval (CI), 94.2–100%] and a high specificity of 96.5% (95% CI, 90.2–100%), and the area under the curve was 0.988 (95% CI, 0.981–0.995). This model can improve medical care in remote areas where eye clinics are not available by using ultra–wide-field fundus ophthalmoscopy for the accurate diagnosis of RRD. Early diagnosis of RRD can prevent blindness.
Similar content being viewed by others
Introduction
Rhegmatogenous retinal detachment (RRD) is a highly curable condition if properly treated early1, 2; however, if it is left untreated and develops proliferative changes, it becomes an uncontrollable condition called proliferative vitreoretinopathy (PVR). PVR is a serious condition that can result in blindness regardless of repeated treatments3,4,5. It is important, therefore, for patients to be seen and treated at a vitreoretinal centre at the early RRD stage to preserve visual function. However, establishing such vitreoretinal centres that provide advanced ophthalmological procedures is not practical because of rising social security costs, a problem that is troubling many nations around the world6.
On the other hand, medical equipment has made remarkable advances, and one such advancement is the ultra–wide-field scanning laser ophthalmoscope (Optos 200Tx; Optos PLC, Dunfermline, United Kingdom). The Optos can provide nonmydriatic, noninvasive, wide-field fundus images easily (Fig. 1) and has been used to diagnose or follow-up various fundus conditions and treatment evaluation7. Because pupillary block and elevated intraocular pressure due to dilation can be avoided, examiners who are not qualified to perform ophthalmologic surgeries can safely capture images, which makes it ideal especially for telemedicine applications in areas where ophthalmologists are not available. In recent years, image processing technology using deep learning, a machine-learning technology, has attracted attention for its extremely high classification performance, and there have been a few studies regarding its applications to medical imaging8,9,10,11,12. To the best of our knowledge, there have been no studies on automatic diagnosis of retinal detachment nor studies on deep learning using Optos images.
In this study, we assessed the ability of a deep learning technology to detect RRD using Optos images.
Results
A total of 420 non-RRD images from 238 patients (mean age, 69.8 ± 9.8 years; 103 men and 135 women) and 411 RRD images from 407 patients (mean age, 54.4 ± 15.2 years; 257 men and 150 women) were analysed.
The deep learning model’s sensitivity was 97.6% [95% confidence interval (CI), 94.2–100%] and specificity was 96.5% (95% CI, 90.2–100%), and the area under the curve (AUC) was 0.988 (95% CI, 0.981–0.995). On the other hand, the support vector machine (SVM) model’s sensitivity was 97.5% (95% CI, 94.1–100%), specificity was 89.3% (95% CI, 82.6–96.0%) and AUC was 0.976 (95% CI, 0.957–0.996) (Fig. 2).
Representative receiver operating characteristic curves (AUC) of the deep learning model and support vector machine (SVM) model. The AUC of the deep learning model was 0.988 (95% CI, 0.981–0.995) and AUC of the SVM model was 0.976 (95% CI, 0.957–0.996). The AUC was better for the deep learning model than for the SVM model.
Discussion
Our results showed that the deep learning technology for detecting RRD on the Optos fundus photographs had high sensitivity and high specificity. The results indicate the great potential for RRD diagnosis in areas where ophthalmologists are not available. If a community has Optos, noninvasive wide-field fundus images can be captured easily without pupillary mydriasis and does not cause medical complications. A deep learning technology can provide accurate RRD classification at a high rate and patients classified as RRD by the deep learning technology are at first remotely reviewed by ophthalmologists. The patients diagnosed as having RRD in this way can be encouraged to see a specialist before the condition progresses to difficult-to-treat PVR, which shows the potential of this model for ophthalmic care. Generally, RRD does not immediately deteriorate to PVR; therefore, if air travel is available for patients, this model could be used to cover considerably wide areas. Because RRD is a highly curable condition if patients receive timely appropriate surgical treatment, the telemedicine system using Optos that we suggest here could significantly reduce the risk of blindness in RRD patients who live in areas where ophthalmic care is not available.
In this study, we used a deep learning technology to classify the Optos images based on the presence of retinal detachment. Our analysis showed that the deep learning model using convolutional neural network (CNN) achieved a better AUC than that of the SVM model; however, slight difference was noted. Although the advantage of deep learning was slight, it was considered clinically important for achieving higher accuracy. It is suggested that multiple layers of non-linear processing in a deep learning model can learn features from a broader perspective with greater flexibility relative to what can be achieved by using SVM, which maps data to a feature space and determines identification boundary. Our results demonstrated classification performance that was almost equal to an ophthalmologist’s judging ability.
This study had several limitations. When clarity of the eye is reduced because of severe cataract or dense vitreous haemorrhage, capturing images with Optos becomes challenging; thus, such cases were not included in this study. Additionally, this study only compared the images of normal eyes and eyes with RRD and did not include eyes with any other fundus diseases. We also need to point out that use of deep learning requires a large number of data sets. In the future, further studies using larger samples and including eyes with other fundus diseases are necessary to more broadly assess the performance and versatility of deep learning.
Methods
Data Set
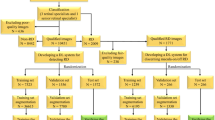
Of the patients diagnosed with RRD by ophthalmologists at Tsukazaki Hospital since December 2011, those who had taken a fundus photograph with Optos were included. In addition, the patients without fundus diseases were extracted from the clinical database of the Ophthalmology Department of Tsukazaki Hospital. These images were reviewed by two ophthalmologists to confirm the presence of RRD and were registered in an analytical database. Eyes that contained vitreous haemorrhage to the extent that ophthalmologists could not determine the area of RRD with Optos image were excluded. Of 831 fundus images, 420 images were from non-RRD patients and 411 images were from RRD patients. According to ophthalmologists who use Optos images, the area of detachment of RRD eyes was less than one quadrant for 93 eyes (22.6%), one to two quadrants for 163 eyes (39.7%), two to three quadrants for 120 eyes (29.2%), and three or more quadrants for 35 eyes (8.5%). In addition, the number of retinal breaks was one for 292 eyes (71.0%), two for 46 eyes (11.2%), and three or more for 14 eyes (3.4%); however, breaks could not be confirmed in 59 eyes (14.4%). Of these, 665 images (336 non-RRD and 329 RRD images) were used for training, and 166 images (84 non-RRD and 82 RRD images) were used for validation of the trained model. The image data set for training was augmented to 11,970 images (6,048 non-RRD and 5,922 RRD images) by performing contrast adjustment for brightness, gamma correction, smoothing, noise imparting and reversal processing on the original images. This research adhered to the Declaration of Helsinki and was approved by the ethics committee of Tsukazaki Hospital. Since this was a study that retrospectively reviewed the Optos images and there were no anonymous issues involved, the Institutional Review Board of Tsukazaki Hospital waived the need for consent.
Deep Learning Model
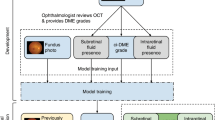
We implemented a deep learning model that uses a CNN shown in Fig. 3 for use in our classification system. This type of multi-layer CNNs is known to automatically learn local features in images and generate a classification model13,14,15. The first layer is called Convolutional Layer 1 (Conv1) and obtains the feature quantities of the target through convolutional filters. Each of the convolutional layers (Conv1, 2 and 3) are followed by rectified linear unit16 and max pooling layers (MP1, 2 and 3) to decrease position sensitivity and allow for more generic recognition. The last two layers (FC1, FC2) are fully connected, remove spatial information from extracted feature quantities and statistically recognise the target from other feature vectors. The last layer is a classification layer, which uses feature vectors of target images acquired in previous layers and the softmax function for binary classification. This network architecture is ideal for learning and recognition of local features of complex image data with individual differences.
Overall architecture of the model. The data set for the retinal fundus images (96 × 96 pixels) is labelled as Input. Each of the convolutional layers (Conv1–3) is followed by an activation function (ReLU) layer, pooling layers (MP1–3) and two fully connected layers (FC1, FC2). The final output layer performs binary classification by using a softmax function.
Training the Deep Convolutional Neural Network
The aspect ratio of the Optos original image is 3900 × 3072 pixels, which is not square; but for analysis, we changed the aspect ratio of all input images and resized them to 96 × 96 pixels. For training, 100 images were mini-batch processed. The initial value of the network weight was randomly given as the Gaussian distribution with a zero mean and a standard deviation of 0.05. To avoid over-fitting during the training, a dropout technique was applied to two fully connected layers (FC1, FC2) to mask out with 70% probability and improve the performance17. The optimisation algorithm called AdaGrad (learning rate = 0.001), one of the stochastic gradient descent methods18, was used to train the network weights. We trained the model by using the GPU of GeForce GTX970M by NVIDIA installed on a commercially available computer.
Support Vector Machine (SVM)
The SVM model was used as a reference to evaluate our deep learning recognition model19. For binary classification SVM, LIBSVM20 from the Scikit–Learn library21 with the Radial Basis Function (RBF) kernel was used. Other feature extraction or feature quantity selection was not performed so that we matched the criteria of the input data used in the deep learning training. The cost parameter (C = 10) and the RBF kernel parameter (γ = 0.001) were used, which were the results of the optimisation by grid search (i.e. C = 1, 10, 100, 1,000, 10,000; gamma = 0.000001, 0.00001, 0.0001, 0.001). Scoring for the grid search was performed on the basis of the mean AUC-receiver operating characteristic obtained by 5-fold cross validation of the grading data. The data set used in the training and validation were the same as the set used in the deep learning model.
Evaluation of Deep Learning and SVM Models
The deep learning and SVM models were operated one hundred times each. Subsequently, the average value and 95% CI for AUC were calculated.
Data availability
The Optos image datasets analysed during the current study are available with the corresponding author on reasonable request.
References
Miki, D., Hida, T., Hotta, K., Shinoda, K. & Hirakata, A. Comparison of scleral buckling and vitrectomy for retinal detachment resulting from flap tears in superior quadrants. Jpn. J. Ophthalmol. 45, 187–191 (2001).
Heussen, N. et al. Scleral buckling versus primary vitrectomy in rhegmatogenous retinal detachment study (SPR Study): predictive factors for functional outcome. Study report no. 6. Graefes Arch. Clin. Exp. Ophthalmol. 249, 1129–1136 (2011).
Lean, J. S. et al. Silicone Study Group. Vitrectomy with silicone oil or sulfur hexafluoride gas in eyes with severe proliferative vitreoretinopathy: results of a randomized clinical trial. Silicone Study Report 1. Arch. Ophthalmol. 110, 770–779 (1992).
AZEN, S. et al. Silicone Study Group. Vitrectomy with silicone oil or sulfur hexafluoride gas in eyes with severe proliferative vitreoretinopathy: results of a randomized clinical trial. Silicone Study Report 2. Arch. Ophthalmol. 110, 780–792 (1992).
Scott, I. U., Flynn, H. W. Jr., Murray, T. G. & Feuer, W. J. Perfluoron study group. Outcomes of surgery for retinal detachment associated with proliferative vitreoretinopathy using perfluoro-n-octane: a multicenter study. Am. J. Ophthalmol. 136, 454–463 (2003).
Mrsnik, M. Global aging 2013: rising to the challenge. Standard & poor’s rating services Network https://www.nact.org/resources/2013_NACT_Global_Aging.pdf (2013).
Nagiel, A., Lalane, R. A., Sadda, S. R. & Schwartz, S. D. Ultra-widefield fundus imaging: a review of clinical applications and future trends. Retina 36, 660–678 (2016).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Liu, S. et al. Multimodal neuroimaging feature learning for multiclass diagnosis of Alzheimer’s disease. IEEE Trans. Biomed. Eng. 62, 1132–1140 (2015).
Litjens, G. et al. Deep learning as a tool for increased accuracy and efficiency of histopathological diagnosis. Sci. Rep. 6, 26286 (2016).
Gulshan, V. et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. JAMA 316, 2402–2410 (2016).
Pinaya, W. H. et al. Using deep belief network modelling to characterize differences in brain morphometry in schizophrenia. Sci. Rep. 6, 38897 (2016).
Deng, J. et al. Imagenet: a large-scale hierarchical image database. Computer Vision and Pattern Recognition 248–255 (2009).
Russakovsky, O. et al. Imagenet large scale visual recognition challenge. Int. J. Comput. Vision 115, 211–252 (2015).
Lee, C. Y., Xie, S., Gallagher, P., Zhang, Z. & Tu, Z. Deeply-supervised nets. InAISTATS 2, 5 (2015).
Glorot, X., Bordes, A. & Bengio, Y. Deep sparse rectifier neural networks. The 14th International Conference on Artificial Intelligence and Statistics. 315–323 (2011).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Duchi, J., Hazan, E. & Singer, Y. Adaptive subgradient methods for online learning and stochastic optimization. J. Mach. Learn. Res. 12, 2121–2159 (2011).
Brereton, R. G. & Lloyd, G. R. Support vector machines for classification and regression. Analyst 135, 230–267 (2010).
Chang, C. C. & Lin, C. J. LIBSVM: a library for support vector machines. ACM Trans. Intell. Syst. Technol. 2, 27 (2011).
Pedregosa, F., Varoquaux, G., Gramfort, A., Michel, V. & Thirion, B. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Acknowledgements
We thank Masayuki Miki and orthoptists of Tsukazaki Hospital for support in data collection.
Author information
Authors and Affiliations
Contributions
H.O. and H.E. wrote the main manuscript text. H.T. and H.E. designed the research. H.O., H.T. and H.E. conducted the research. H.E. undertook the deep learning methods and statistical analysis. N.I. collected the data. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ohsugi, H., Tabuchi, H., Enno, H. et al. Accuracy of deep learning, a machine-learning technology, using ultra–wide-field fundus ophthalmoscopy for detecting rhegmatogenous retinal detachment. Sci Rep 7, 9425 (2017). https://doi.org/10.1038/s41598-017-09891-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09891-x
This article is cited by
-
Developing and validating a clinlabomics-based machine-learning model for early detection of retinal detachment in patients with high myopia
Journal of Translational Medicine (2024)
-
Deep Learning for the Detection of Multiple Fundus Diseases Using Ultra-widefield Images
Ophthalmology and Therapy (2023)
-
Development and validation of a routine blood parameters-based model for screening the occurrence of retinal detachment in high myopia in the context of PPPM
EPMA Journal (2023)
-
Automated detection of retinal exudates and drusen in ultra-widefield fundus images based on deep learning
Eye (2022)
-
Deep learning model for analyzing the relationship between mandibular third molar and inferior alveolar nerve in panoramic radiography
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.