Abstract
Asymmetric cell divisions are of fundamental importance for the development of multicellular organisms, e.g. for the generation of founder cells. Prime examples are asymmetric cell divisions in germline precursors during the early embryogenesis of the transparent roundworm Caenorhabditis elegans, one of the major developmental model organisms. However, due to a lack of quantitative data it has remained unclear how frequent unequal daughter cell sizes emerge in the worm’s early embryogenesis, and whether these originate from sterical or biochemical cues. Using quantitative light-sheet microscopy, we have found that about 40% of all cell divisions in C. elegans until gastrulation generate daughter cells with significantly different volumes. Removing the embryo’s rigid eggshell revealed asymmetric divisions in somatic cells to be primarily induced by steric effects. Division asymmetries in the germline remained unaltered and were correctly reproduced by a model based on a cell-size independent, eccentric displacement of the metaphase plate. Our data suggest that asymmetric cell divisions, imposed by physical determinants, are essential for establishing important cell-cell interactions that eventually fuel a successful embryogenesis.
Similar content being viewed by others
Introduction
Asymmetric cell divisions are of crucial importance for developmental processes, e.g. in the context of tissue or body axis formation1, 2. Many protein species that are involved in asymmetric cell divisions have been shown to be evolutionary conserved (reviewed, for example, in ref. 1), indicating that general mechanisms for asymmetry generation are utilized in different biological systems. Studies on the model organism Caenorhabditis elegans have been instrumental in this context due to its relative simplicity, its susceptibility to modern genetic and molecular-biological tools, and its optical transparency (see www.wormbook.org for an introduction). A plethora of (fluorescence) microscopy-based studies have, for example, revealed detailed insights into the first asymmetric cell division of the zygote (P0) and the concomitant creation of an anterior-posterior body axis2,3,4,5,6,7,8. Also a fair understanding of the associated formation of biochemical gradients, from Turing-like patterns7, 9 to condensation phenomena10, has been possible. Virtually all of these and similar studies have been focusing on the single-cell stage and the first, asymmetric cell division since monitoring dynamic intracellular events in the comparatively large P0 cell is straightforward.
In fact, although C. elegans has been studied as a model organism for several decades by now, cell division asymmetry has remained a rather vaguely defined term as it may describe purely biochemical or geometrical asymmetries, or the combination of both. Defining biochemical asymmetries of daughter cells necessarily requires the quantification of a non-uniform distribution of specific molecular markers and hence virtually all of such reported asymmetries are properly defined (see, for example, ref. 2 for a comprehensive summary on biochemical asymmetries in the zygote). However, geometrical asymmetries, i.e. the emergence of two unequally sized daughter cells, have been studied in much less detail. Frequently utilized techniques like differential interference contrast (DIC) microscopy or even confocal microscopy have method-intrinsic limitations that hamper a thorough three-dimensional quantification, hence requiring simplifying extrapolations to arrive at approximate cell volumes (see ref. 11 for a recent example). Moreover, due to volume-conserving (blastomeric) division cycles, cell sizes in the early C. elegans embryo decrease rapidly, therefore amplifying the uncertainty about actual cell volumes. As a consequence, extrapolated cell volumes are quite error-prone and may not report reliably on geometrical asymmetries in cell division events.
Despite these limitations, it is well established that at least cells of the future germline, the so-called P lineage (cf. the embryo’s early lineage tree in Fig. 1A), undergo geometrically asymmetric divisions2, 12. Yet, a thorough quantification of their (and other cells’) asymmetries has, to the best of our knowledge, not been done. As a consequence, it is neither clear how many geometrically asymmetric cell divisions beyond the P lineage occur until gastrulation nor is it known what causes them. Indeed, one may even ask why C. elegans has geometrically asymmetric cell divisions at all since a biochemical asymmetry might have been sufficient to run the proper molecular-biological developmental program.
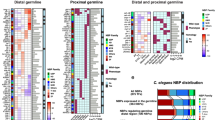
Division asymmetries in unperturbed C. elegans embryos. (A) Lineage tree of early C. elegans embryogenesis (prior to gastrulation). Different lineages are color-coded, the germline is highlighted in red. (B) Representative maximum-intensity projections of image stacks taken on early C. elegans embryos (strain OD95) with the plasma membrane and chromatin stained in red and green, respectively. Scale bar: 10 μm. (C) Single two-dimensional slices taken from the image stacks shown in A. (D) The corresponding membrane segmentation shows how well details of the plasma membrane are identified. Please note: Color-coding of cell boundaries was chosen for best contrast and does not indicate correspondence to specific lineages. (E) Volumetric ratio, VR, of daughter cells emerging from the named mother cell (median of n = 10 embryos with error bars indicating the standard deviation). Color-coding of lineages like in (A). The volume-dependent level of uncertainty for each cell (grey) quantifies the apparent division asymmetry that is attributed solely to segmentation errors (see Materials and Methods for a detailed definition). As a result, cells of the P, MS, and C lineages, but also few cells of the AB lineage show significant division asymmetries that are well beyond the level of uncertainty.
Here we have used selective plane illumination microscopy, SPIM, to address this topic (see, for example, refs 13,14,15 for introductory reviews on SPIM). Due to the gentle illumination via a light sheet, we were able to monitor the development of C. elegans embryos with and without an eggshell in three-dimensional detail up to gastrulation. A custom-made image segmentation approach enabled us to derive volumes and division asymmetries from these raw data. As a result, we observed that about 40% of all cell divisions before gastrulation are significantly asymmetric with many of these events being enhanced by sterical forces from the confining eggshell. For predominantly biochemically governed asymmetric cell divisions, i.e. for the P lineage, we were able to predict the degree of asymmetry via a simple model that relies on a cell-size independent, eccentric displacement of the mitotic spindle.
Results and Discussion
About 40% of all cell divisions until gastrulation are asymmetric
According to the literature, asymmetric cell divisions have been observed for cells of the P lineage (P0, …, P3) and EMS2, 12, 16. These cell divisions coincide with the emergence of so-called founder cells (AB, MS, E, C, D, P4) that establish new lineages (Fig. 1A). All other cell divisions until gastrulation are typically interpreted as being symmetric with respect to daughter cell sizes. In fact, data on volumetric asymmetries of daughter cells are mostly qualitative or extrapolated from two-dimensional imaging techniques rather than reporting faithful three-dimensional quantifications.
In order to obtain more quantitative insights into the amount and degree of volumetric asymmetries in cell divisions during early embryogenesis of C. elegans, we used a custom-made SPIM setup17,18,19 and a custom-written segmentation approach (see Materials and Methods for details). The chosen worm strain (OD95) stably expressed fluorescent markers for histones (H2B::mCherry) and the plasma membrane (PH(PLC1δ1)::GFP), hence facilitating three-dimensional imaging and volume rendering during early embryogenesis. Representative examples of images and segmentation results are shown in Fig. 1B–D.
Using this approach we were able to identify all cell division events and to quantify all cell volumes until the onset of gastrulation via temporally resolved three-dimensional image stacks. Using these data as experimental input, we calculated for each mother cell the volume ratio of daughter cells, VR = V 1 /V 2. For somatic cells we used the more posterior cell for V 2, for germline cells always the (smaller) new germline cell volume was used for V 2. To explore whether a division was significantly asymmetric, i.e. whether the value of VR deviated sufficiently from unity, we defined a level of uncertainty that arises solely from segmentation errors (see Materials and Methods for details). Roughly speaking, relative deviations of daughter cell volumes by 10% or more indicated a significant volumetric asymmetry in the respective cell division. The resulting data and their significance rating are shown in Fig. 1E.
As expected, all cells of the P lineage showed a significant division asymmetry, albeit with markedly different VR values (Fig. 1E). Also EMS was found to divide asymmetrically, although the asymmetry was less than for any cell of the P lineage. Surprisingly, also other cells at this early stage, namely MSa, MSp, Ca, and Cp, showed significant asymmetries with daughter cells differing by 30–60% in volume. In contrast, E, MS, and C divided almost perfectly symmetrical like most cells from the AB lineage. Yet, even some cells from the AB lineage showed a significant but borderline asymmetry, especially ABar.
Thus, our data indicate that altogether about 40% of all cell divisions until gastrulation in C. elegans embryos feature a significant volumetric asymmetry.
Geometrical constraints induce asymmetric divisions
Since, to our knowledge, a biochemical asymmetry in the cell division of MSa, MSp, Ca, and Cp has not been described, we hypothesized that at least some of the observed volumetric asymmetries might arise from geometric constraints during the respective cell divisions rather than being governed by biochemical cues. Indeed, geometrical constraints and physical forces have been seen to have a significant impact on the positioning of cells in the early embryo17, 18 but also in the context of tissue organization and wound healing20, 21, making this hypothesis an attractive option.
In order to test our hypothesis, we sought to decrease possible mechanical constraints by removing the chitin-based eggshell, while leaving the inner, more flexible vitellin layer intact to maintain the embryo’s integrity (see Materials and Methods). As expected, in the absence of the eggshell the intact vitellin layer lead to a rather compact arrangement of the blastomeres that was often, but not always, similar to wild-type embryos (Fig. 2A). In fact, developmental phenotypes became significantly more variable: The stereotypical diamond-shape arrangement of cells in the four-cell state (Fig. 1B, second image) was seen only in about 50% of all embryos without an eggshell, whereas the remaining embryos displayed arrangements that resembled a (partially skewed) T-shape (Fig. 2A, second image). These early deviations from the wild-type phenotype correlated with increasingly severe misarrangements in later stages, hence underlining the supportive role of the eggshell for a robust embryogenesis.
Division asymmetries in C. elegans embryos lacking the eggshell. (A) Representative maximum-intensity projections of early C. elegans embryos after removing the eggshell but leaving the vitellin layer intact (red, green: plasma membrane, chromatin). Scale bar: 10 μm. (B) Volumetric ratio, VR, of daughter cells emerging from the named mother cell (median of n = 11 embryos, error bars indicate the standard deviation) with the volume-dependent level of uncertainty. Color-coding as in Fig. 1E. (C) Comparison of the median values without (red) and with (black) an intact eggshell. Please note that bars for P2 and P3 have been reduced by 0.5 for better visibility. Error bars are equal to those shown in Figs 1E and 2B and have been omitted here for better readability of the plot; please see Fig. S2 for a version including error bars. While asymmetries in the germline are preserved, most somatic cells tend to decrease their level of asymmetry upon removal of the eggshell.
The volume ratios associated with eggshell-free embryos (Fig. 2B) and the direct comparison to untreated embryos (Fig. 2C and Fig. S2) highlighted that the P lineage (P0 to P3) as well as Ca did not change markedly. This suggests that their volumetrically asymmetric divisions are a consequence of an internal biochemical asymmetry. Almost all other volume ratios showed a clear tendency to decrease, indicating that sterical forces induced by the confining eggshell cause at least partially these asymmetries. In particular, EMS and the MS lineage showed markedly reduced asymmetries, making them almost as borderline as the somatic outlier ABar. Cp showed a significantly reduced value of VR while still dividing in significantly asymmetric fashion, whereas Ca was almost unchanged.
Thus, asymmetric divisions in the P and C lineages are well preserved even with softened geometric constraints while the asymmetry in EMS and in the MS lineage seem to rely predominantly on sterical forces imposed at least indirectly by the eggshell.
A cell-size independent displacement of the mitotic spindle quantitatively explains division asymmetries in the germline
Inspired by previous work5, 6 on the first cell division in C. elegans embryos, in which a pronounced shift of the mitotic spindle apparatus along the AP-axis has been identified as major cause for an asymmetric cell division of P0, we hypothesized that also subsequent asymmetric cell divisions in the germline are driven by a displacement of the spindle’s center of mass. In particular, we wondered to which extent a shift of the mitotic spindle could quantitatively explain the experimentally observed volumetric ratios VR of daughter cells in the post-zygote germline. For this analysis, we deliberately excluded P0, since several molecular players that influence an eccentric spindle displacement in P0 from the anterior side22, 23 are segregated into the AB cell during the first cell division. Since gene expression is mostly shut down in the germline during early development24, these molecular players are therefore unlikely to play a major role in subsequent cell divisions in the germline, rendering the first division a special case (see also discussion below). Moreover, we have used data from embryos without eggshell for the subsequent analysis since these display asymmetric divisions without the influence of eggshell-induced cues; we did not observe significant differences when using data from untreated embryos.
For simplicity we assumed the mother cell to be spherical (radius R0), and we asked how the volumetric ratio would be if the daughter cells emerged from a spindle that was displaced by an increment Δx away from the cell center (Fig. 3A, inset). Given that the position of the metaphase plate defines the plane of cytokinesis25, the spherical mother cell would be split approximately into two spherical caps with heights R0 + Δx and R0 − Δx, yielding volumes V1 = π/3·(R0 + Δx)²·(2R0 − Δx) and V2 = π/3·(R0 − Δx)²·(2R0 + Δx), respectively. The mathematically simplest assumption for this division scheme would be a constant shift Δx that does not depend on the size of the mother cell.
Comparison of experimental findings and model predictions. (A) Modeling cells as spherical entities and allowing for an off-center division into spherical caps (cf. inset upper left) predicts well the experimentally observed cell volumes Vexp (main plot). Using the volume of P1 as sole input, volumes of EMS and P2, and from this volumes of P3 and C, and finally P4 and D were deduced assuming a constant shift Δx of the division plane. Predicted values Vtheo matched best the experimentally found ones, when choosing Δx = 1.75 µm (see inset lower right for the relative deviations when varying Δx). (B) Spindle displacements measured relative to the cell center in somatic cells (black dots) follow a normal distribution with a mean of −0.1 µm and standard deviation 0.56 µm (gray dashed line). In contrast, spindle displacements in germline cells (red dots) are grouped around a constant shift of 1.36 µm (dashed vertical line). A Kolmogorov-Smirnov test yielded a significance level of 0.38% that data from somatic and germline cells are from the same distribution, i.e. they can be regarded as different with a high significance. (C) Division asymmetries VR, predicted for P1, P2, and P3 on the basis of the last image stack that shows an unambiguous metaphase (grey bars), follow the experimental results for the daughter cells (black bars) but consistently underestimate the asymmetry. In fact, using these image stacks and dissecting segmented cells into two caps via a division plane through the metaphase plate is consistent with a median spindle shift of only 1.36 µm (see Fig. 3B and main text). Accounting for an additional, unmonitored spindle shift by approximately 450 nm during the lag period between consecutive image stacks (see main text) and repeating the dissection scheme the extrapolated asymmetries (red bars) show a favorable agreement with our experimental data. (D) Spindle displacement by a constant offset Δx can be rationalized by assuming constant forces FA < FP that pull the spindle towards the anterior and posterior end of the cell, respectively. As a result, the spindle is stretched and its center of mass moves into the posterior direction. Stress resistance of the spindle is modeled via a passive Hookean element (spring constant k, resting length L0) until a maximum extension is reached and the spindle ruptures at the onset of anaphase. See main text for details.
To compare this naïve approach with our experimental data, we used the measured volume of P1 from which we extracted the apparent cell radius via R0 = (3 V/4π)1/3. Then, we iteratively predicted from this single experimental input the volumes V theo of all subsequent daughter cells along the lineage tree (P2, EMS, P3, C, P4 and D): Volumes of EMS and P2, i.e. V1 and V2, were derived via the spherical-cap scheme outlined above, using R0 of P1 as input. Assuming P2 to be spherical again, we extracted its apparent radius from the predicted volume and repeated the division scheme until all volumes had been determined. The predictions we got from this procedure showed a remarkably good agreement with the experimentally observed volumes of daughter cells, Vexp, when setting Δx ≈ 1.75μm (Fig. 3A). Using instead a shift Δx that depended on the mother cell size did not capture the experimental data.
Thus, a cell-size independent shift of the mitotic spindle by approximately 1.75 μm can quantitatively explain all experimentally observed volumetric division asymmetries in the post-zygote germline.
Next we sought to obtain experimental support for this simple approach and its prediction of a cell-independent displacement of the mitotic spindle by Δx ≈ 1.75μm during asymmetric division events in cells P1-P3. In contrast to the distinct first cell division, division axes of cells P1-P3 do not necessarily lie in a single imaging plane, which virtually eliminates the possibility to determine the spindles’ shift via very rapid two-dimensional imaging. We therefore utilized our three-dimensional image stacks, acquired with a moderate time resolution, as a proxy.
In particular, we exploited the last image stacks in which P1, P2, and P3 showed an unambiguous metaphase pattern. Combining segmented image stacks and tracking data of chromatin, we determined position and orientation of the metaphase plate in the respective cell. As a reference point, we also calculated the cell center according to a mechanism proposed by Grill and Hyman (see Materials and Methods and Fig. 2c in ref. 26): Astral microtubules, docked to the cell cortex via special sites that are equi-distributed on the plasma membrane, exert pulling forces on the spindle, with the magnitude of force being independent of microtubule length. The equilibrium position determined by this mechanism is equal to the surface’s center of mass for symmetrical cells but can deviate for unsymmetrical or non-convex cells. Assuming the plane of division to coincide with the metaphase plate, we determined the distance of the cell center from the metaphase plate along its surface normal as an estimate for the eccentric displacement of the spindle. Displacements were assigned a direction with negative/positive signs indicating a shift towards the larger/smaller daughter cell.
The analysis outlined above was applied to all eggshell-free embryos (n = 11) from which median values and standard errors for each cell along the lineage tree were determined. As a result, we observed that displacements in somatic cells did not show a clear trend (Fig. 3B). In line with this, a Jarque-Bera test rated these data to be consistent with values drawn randomly from a normal distribution (mean: −0.1 µm; standard deviation: 0.56 µm; p = 0.5). Moreover, a very small correlation coefficient of 0.009 between these spindle displacements and the respective cell division asymmetries further supported the notion that somatic asymmetries are not a consequence of displaced mitotic spindles. This is in stark contrast to consistent spindle displacements in germline cells that were seen to cluster around a median value of 1.36 µm (Fig. 3B, red points). Moreover, a Kolmogorov-Smirnov test revealed a significance of 0.38% that displacement values of germline and somatic cells belong to the same distribution, i.e. it is extremely unlikely that the observed shifts in germline cells are only statistical fluctuations.
The observed shift in germline cells is clearly nonzero but somewhat lower than the predicted value, Δx ≈ 1.75μm. We attributed this difference to the fairly long lag time of ΔT = 30 s between successive image stacks, i.e. even after the last stack with a metaphase phenotype the spindle could still be on the move for up to 30 s. Based on data acquired for P0 a peak velocity in the range of 40 nm/s can be expected for the spindle motion6. Assuming that all spindles in the post-zygote germline move with a somewhat lower, average velocity of 30 nm/s (see below for a justification) and estimating that the onset of cytokinesis happens on average ΔT/2 = 15 s after the image stack has been taken, an unmonitored distance of 15 s × 30 nm/s = 450 nm should be taken into account to extrapolate the typical shift of the spindle in cells P1-P3. The result, a shift by Δx≈1.36μm + 450 nm = 1.81μm, is in favorable agreement with our prediction derived via the division scenario into spherical caps.
This extrapolation for germline percursor cells is further corroborated by the volumetric asymmetries, VR, determined from the very same segmented image stacks: Relying again on the last image stacks in which P1, P2, and P3 showed an unambiguous metaphase pattern, the metaphase plate was used to determine the future division plane. All voxels of cells P1-P3 were sorted into putative daughter cells (see Materials and Methods), i.e. real cells were dissected through the metaphase plate into slightly deformed spherical caps. Values of VR determined via this procedure indeed followed the trend of the experimental data for fully developed daughter cells (Fig. 3C), yet consistently underestimated the asymmetry (consistent with the somewhat too low spindle displacement Δx≈1.36μm determined from these images). Shifting the putative division plane by 450 nm along its surface normal to account for the unmonitored spindle movement between successive image stacks, we obtained a favorable agreement between the estimated and experimentally determined asymmetries (Fig. 3C).
Thus, the experimentally determined displacement of the metaphase plate in cells P1-P3 confirms the reasoning of a cell-size independent eccentric position of the spindle before asymmetric division events, whereas a significant shift of the metaphase plate in the ensemble of somatic cells was not observed.
A simple model explains the constant spindle displacement
The above results raise the question of how cells actually ensure a size-independent spindle displacement to an eccentric position before entering anaphase. To arrive at a meaningful model, we started from experimental observations in the zygote: In P0, the anterior spindle pole is tethered and remains in a fixed position until the spindle is fully assembled27. Upon release of the tethering, the metaphase plate starts to displace to the posterior with a constant velocity27. While the anterior spindle pole is displaced only slightly to the posterior27, the posterior spindle pole is displaced significantly stronger5. Hence, concomitant to its displacement the spindle is stretched along its migration path. Moreover, laser ablation experiments in P0 have revealed that astral microtubules provide the pulling forces for the metaphase plate’s displacement5, 6, 8, 27. These pulling forces are generated at the cell cortex, at which astral microtubules are in contact with force generator complexes that involve Gα, GPR-1/2, LIN-5 and dynein (see ref. 2 for a recent comprehensive review). A net force ratio of ~1.5 towards the posterior is achieved in P0 by locally increasing pulling forces on the posterior cortex5, 6, 8 but also by decreasing forces on the anterior cortex22, 23. Using RNA interference to switch both poles to the same force generator phenotype lead to a vanishing net force on the spindle6, and consequently these embryos lacked the native spindle displacement towards the posterior end. When both poles were forced to assume an ‘anterior’ force generator phenotype, mitosis even was stalled in late metaphase due to a too low absolute force that was insufficient to initiate a rupturing of the spindle.
Combining these observations with our results, we can formulate a simple one-dimensional model for achieving a cell-size independent displacement of the spindle that precedes an asymmetric division (Fig. 3D): Forces acting on the anterior and posterior spindle poles, FA and FP, do not depend on the distance to the cell cortex as microtubules only transmit pulling forces that are created at the cortex. Due to a low residence time of each microtubule at the cortex, about 1–2 s (ref. 28), microtubules also do not contribute a memory-driven restoring force. Therefore, FA and FP can be modeled as constant forces. The spindle does not provide active forces for its displacement but needs to oppose the net stress Fp + FA applied via the spindle poles. For simplicity, we model it as a Hookean element with spring constant k and equilibrium length L 0. Since initial spindle lengths at early metaphase are almost cell-size independent at these early stages of embryogenesis29, we can assume L 0 to be approximately the same for all P cells. Upon stretching this spring beyond a limit L 0 + s max , the spindle is assumed to rupture. This assumption is based on the observation that pulling on both spindle poles with only the anterior force magnitude is insufficient to go from metaphase to anaphase, whereas bidirectional pulling with the posterior force magnitude allows for spindle rupturing and (symmetric) cell division6. Due to the low Reynolds number in cell biology problems30, the motion of the spindle and/or its poles can be described in the overdamped limit, i.e. we can neglect all inertia terms.
For convenience, we express the equations of motion in terms of the distances xA and xP that the anterior and posterior spindle poles assume over time with respect to their initial position. Initially, the two poles are separated by the equilibrium length L 0 of the unstressed spindle, i.e. xA(t = 0) = xP(t = 0) = 0. Upon releasing the tethering at time t = 0, the forces FA < FP act on the spindle poles and the spindle is stretched by a distance s = xP−xA. Hence, the equations of motion read for t > 0:
and
with γ denoting the effective friction coefficient for the spindle poles. Solving these coupled differential equations, one obtains xMP(t) = [xA(t) + xP(t)]/2 = (FP − FA)·t/(2γ) for the position of the metaphase plate, and s(t) = (FP + FA)/(2k)·(1 − exp(−2kt/γ)) for the extension of the stressed spindle. Upon reaching a maximum extension s max , the spindle ruptures and cytokinesis is initiated. The associated instant of time, T, is determined via the equation s max = s(T) = (FP + FA)/(2k)·(1 − exp(−2kT/γ)) from which one can infer the maximum travel distance of the spindle, Δx = xMP(T) = (FP − FA)·T/(2γ). Since neither the forces FA and FP nor the spindle parameters dependent on cell size, this model predicts a constant displacement of the mitotic spindle.
It is worth noting that the spindle displacement explicitly depends on the ratio of anterior and posterior pulling forces, FA/FP. In fact, adapting the aforementioned approach of spherical caps to the ellipsoidal zygote, a displacement of ~3 µm was needed to explain the asymmetry ratio VR for P0, which is significantly larger than the value Δx≈1.75μm for P1-P3. This apparent discrepancy can be rationalized when taking into account that in P0 several molecular agents downstream of the PAR machinery are localized in the anterior domain of the cortex where they contribute to a lowering of pulling forces towards the anterior22, 23. Assuming that the cytoplasmic pool of these proteins is much smaller than the pool localized on the anterior cortex23, the majority of these very proteins would be segregated into the somatic cell AB and hence was lost for the future germline. Since gene expression is mostly shut down in the germline at these early stages of development, it is unlikely that comparable levels of these proteins could be restored soon after the first cell division. As a consequence, these players would not be available to reduce FA in cells P1-P3 as much as in P0, hence increasing the ratio FA/FP in these cells. This leads to a slower spindle displacement in P1-P3 in comparison to P0 (as assumed when estimating the unmonitored spindle shift by 450 nm), whereas the spindle-internal stress builds up more rapidly. Therefore, upon reaching its maximum extension smax, the spindle has travelled a smaller distance. Following this reasoning, cells P1-P3 are predicted to display a smaller displacement Δx than P0, in favorable agreement with experimental observations.
Finally, one may wonder why nature has chosen to equip C. elegans with volumetrically asymmetric cell divisions during early embryogenesis, as biochemical asymmetries could have been fully sufficient. While cell sizes seem to have little influence on the positioning of cells until gastrulation17, 18, the number of cell-cell contact areas certainly depends quite strongly on the surface area of cells. We therefore speculate that distinct division asymmetries cause, or at least support, the formation and/or prevention of cell contact areas to achieve a wiring diagram of cells that can fuel a successful embryogenesis.
Materials and Methods
Sample preparation and imaging
For imaging we used C. elegans strain OD95 in which the plasma membrane and histones are fluorescently labeled (PH(PLC1δ1)::GFP, H2B::mCherry). Worm culture and preparation of untreated embryos for SPIM imaging was done as described before17,18,19. Removal of the eggshell was done similar to previous approaches31: Zygotes with visible pronuclei were chosen from dissected gravid worms. These eggs were placed on a coverslip and immersed in approximately 30 μl of NaOCL solution (3% Na) for two to three minutes. Subsequently they were washed three times with M9 buffer32 to remove all of the NaOCL solution before pipetting 25 μl Chitinase solution onto them. The solution was prepared by dissolving 5 units of Chitinase from Streptomyces (Sigma) in 2 ml of sterile egg buffer32.
Removal of the eggshell took roughly 10 to 15 minutes. The remaining vitellin layer was left intact. When the eggshell was not visible any more, embryos were transferred rapidly to the SPIM setup for immediate imaging (starting in most cases during mitosis of the zygote). Embryos adhered to the plain, untreated glass surface without the need for Poly-L-lysine or other mounting agents. During imaging, unperturbed embryos were immersed in water, embryos without eggshell in M9 buffer. Imaging was performed with a custom-made dual color SPIM setup as described before17,18,19. For long-term imaging of wildtype and eggshell-free embryos, full dual-color stacks, consisting of 50 individual layers with a spacing of \(\sqrt{2}\) μm, were taken every 30 s for a total time of three hours (i.e. 360 stacks).
Segmentation, image analysis, and evaluation
Tracking of nuclei via H2B::mCherry was done as described before17. We generally tracked at least until the embryo consisted of 44 cells and included a manual correction step to account for potential errors.
Three-dimensional segmentation of cell membranes from PH(PLC1δ1)::GFP images required a refined approach to account for SPIM-inherent shadowing effects. These arise from absorption and scattering events at bright structures when being illuminated by the light sheet, i.e. some shadowing is observed behind such structures. As a result, image segmentation of membrane-labeled embryos via global filtering and thresholding operators was not reliable. We have therefore developed a novel segmentation algorithm that is based on growing a seed region in each cell: The goal of the segmentation is a division of a three-dimensional image of an embryo with n cells into n + 1 regions, with each voxel of the image stack being uniquely assigned to one of the cells or to none ( = outside of the embryo). After an initial box-filtering (kernel size 5 × 5 × 1 voxels) the background intensity of the image, i.e. anything outside of the embryo, is set to zero via global thresholding. Then, a seed is placed inside each cell, either manually or by using voxels that have been identified during the tracking of nuclei. During the segmentation process, these n seeds are grown simultaneously and iteratively. Boundaries of each seed are computed by eroding the region of voxels belonging to the seed with the smallest possible kernel (3 × 3 × 3 voxels) and subtracting this result from the original region. Boundary voxels therefore share one face, edge, or corner with voxels outside of this seed’s region. Next, for each of these boundary voxels one neighbor outside the region is chosen randomly and both voxel intensities are compared. If the outside voxel’s intensity is larger than the boundary voxel’s intensity (F out ≥ qF boundary ), the outside voxel is added to the region unless it belongs already to another seed’s region. The multiplier q ≈ 0.97…0.99 is introduced to compensate for noise, and it needs to be chosen carefully for each image or image series. This procedure is carried out for all cells/seed regions prior to the next iteration. Aiming at short processing durations, a total of N = 300/log(n + 1) iterations were performed initially on a downscaled version of the image stack. After upscaling to the original size, 40 additional iterations were performed.
This scheme leads to a local expansion of each seed until it collides with another region or when its boundary arrives at a significant drop in voxel intensity. The latter typically occurs at the cytoplasm-membrane interface, i.e. the boundary of each region becomes a faithful representation of the plasma membrane. Minor artifacts (stray pixels etc.) are removed after the iteration process by opening and closing operations with a kernel of 15 × 15 × 3 voxels. Results were finally controlled manually stack by stack to ensure a high segmentation quality. Results obtained with this algorithm provided us with data of cellular volumes (and shapes) of unprecedented precision.
From the segmentation process, the number of cells, n, and, the number of voxels of each cell, m i (i = 1, …, n), is known for each embryo at each instant of time. From this, each cell’s volume was determined as the product V i = L x L y L z m i with L x = L y = 0.16μm being determined via the objective and the camera sensor, and L z = \(\sqrt{2}\)μm the spacing between two consecutive optical sections within an image stack. Volumes do not show any significant changes during the cell cycle, i.e. a cell’s volume can be assumed constant (data not shown). For our analysis, we used the median value of the obtained time series of cell volumes to suppress few possible outliers in the scheme. As a result, the sum of volumes of the daughter cells generally deviated from the mother cell’s volume by less than 3%, indicating reliable segmentation results throughout the whole image series.
Volume ratios of somatic daughter cells were defined as the volume of the more anterior cell divided by the volume of the more posterior cell, e.g. VR AB = V ABa /V ABp . For germline cells always the smaller volume of the new germline cell was used in the denominator, e.g. VR P3 = V D /V P4 . The level of uncertainty due to segmentation errors, i.e. the minimal ratio VR that reports a significant division asymmetry, was determined as follows: The main source of error during volume determination originates from the decision whether or not an additional layer of voxels around the already detected volume is considered while segmenting a cell, i.e. if the region is expanded even further or not. Assuming all cells to be spherical with radii being determined by the cell volume, \(R={(3V/4\pi )}^{1/3}\), we can express this additional volume ΔV in analytical terms. We first note that due to Lz ≫ Lx we have to separately consider the bottom and top slice in the (sub)stack containing the respective cell, i.e. we have two contributions, ΔV1 and ΔV2, from the inner layers and the two spherical caps in the top and bottom layer, respectively. We reasoned that the true cell boundary will, on average, bisect the thickness of the top and bottom layers (cf. sketch in Fig. S1), i.e. top and bottom caps have a height h = Lz/2 and a squared in-plane radius r2 = R2-(R-h)2 yielding a volume contribution \(\Delta {V}_{2}=2\pi h(3{r}^{2}+{h}^{2})/6=\pi {L}_{z}(R{L}_{z}-\frac{{L}_{z}^{2}}{6})/2\). The contribution from the inner layers, where additional voxels have an average width Lx, is simply the volume of a spherical shell with thickness Lx, reduced by the volume of the two caps that are situated in the top and bottom layer, \(\Delta {V}_{1}=4\pi {R}^{2}{L}_{x}-2\pi R{L}_{z}{L}_{x}\). Since any of these additional volumes will contribute only by chance, adding or not adding these voxels has equal probability 50%, the average volume that is added or not considered amounts to ΔV=(ΔV1 + ΔV2)/2. Hence, even a symmetric division into daughter cells with volume V sym can lead to an apparent asymmetry VR max = (V sym + ΔV)/(V sym − ΔV) > 1 or VR min = (V sym − ΔV)/(V sym + ΔV) < 1, depending on which daughter cell determines the ratio’s (de)nominator. Based on this reasoning, we only deemed measured asymmetries as significant when they exceeded this mother-cell volume-dependent uncertainty range.
To estimate the spindle displacement in late metaphase and to extrapolate the future division plane, we used three consecutive stacks (named S1, …, S3) after the last image stack that showed an unambiguous metaphase phenotype (named S0). From stack S0, the center-of-mass position of the metaphase plate, r 0, is known from tracking the histone stain (H2B::mCherry). The cell center, c, associated with a symmetric spindle position was determined from the plasma membrane segmentation according to an iterative mechanism proposed by Grill and Hyman26: First, we selected an evenly distributed subset of ~3000 voxels on the cell surface and used their coordinates p i as origins of forces. As a starting point for the iteration we used the center of mass of these points, c(0). This position was iterated by calculating the direction of the effective net force, f = Σ(c(j) − p i) / |c(j) − p i|, with the sum running over all voxel indices. Then, c(j + 1) = c(j) + εf with a small, empirical coefficient ε. This step was repeated until the sequence of c(j) converged to a position c that did not change significantly any more. For symmetrical objects, c will be identical to the center of mass of the object’s boundary, while for unsymmetric or non-convex objects like real cells a small but significant deviation remains.
Positions r 1 and r 2 of daughter-cell chromatin assemblies are known for S1, …, S3 from tracking the histone stain (H2B::mCherry). Normalizing and averaging the vector r 2 − r 1 over stacks S1, …, S3 yielded a robust estimate for the surface normal of the future division plane, d. The distance by which the metaphase plate had been shifted to an eccentric position in stack S0 was then determined as the scalar product Δx = d·(c − r 0 ). Then, the position of the metaphase plate was split into two artificial points q 1 and q 2 along the direction of d with a small separation of 10 nm («Lx), i.e. q = r 0 ± d·5 nm. Based on the shortest distance to these points, all voxels of the cell were then classified to belong to q 1 or q 2, and the resulting volumes V1 and V2 were used for Fig. 3A. Indeed, this scheme splits the volume of the mother cell into two parts along a plane (perpendicular to d) that runs through the last metaphase plate position, r 0. When extrapolating the unmonitored spindle movement between stacks S0 and S1, the same approach was used with points q 1 and q 2 being shifted by d·450 nm.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Betschinger, J. & Knoblich, J. A. Dare to be different: asymmetric cell division in Drosophila, C. elegans and vertebrates. Curr Biol 14, R674–685 (2004).
Rose, L. & Gonczy, P. Polarity establishment, asymmetric division and segregation of fate determinants in early C. elegans embryos. WormBook, 1–43 (2014).
Boyd, L., Guo, S., Levitan, D., Stinchcomb, D. T. & Kemphues, K. J. PAR-2 is asymmetrically distributed and promotes association of P granules and PAR-1 with the cortex in C. elegans embryos. Development 122, 3075–3084 (1996).
Watts, J. L. et al. par-6, a gene involved in the establishment of asymmetry in early C. elegans embryos, mediates the asymmetric localization of PAR-3. Development 122, 3133–3140 (1996).
Colombo, K. et al. Translation of polarity cues into asymmetric spindle positioning in Caenorhabditis elegans embryos. Science 300, 1957–1961 (2003).
Grill, S. W., Gonczy, P., Stelzer, E. H. & Hyman, A. A. Polarity controls forces governing asymmetric spindle positioning in the Caenorhabditis elegans embryo. Nature 409, 630–633 (2001).
Goehring, N. W. et al. Polarization of PAR proteins by advective triggering of a pattern-forming system. Science 334, 1137–1141 (2011).
Grill, S. W., Howard, J., Schaffer, E., Stelzer, E. H. & Hyman, A. A. The distribution of active force generators controls mitotic spindle position. Science 301, 518–521 (2003).
Kondo, S. & Miura, T. Reaction-diffusion model as a framework for understanding biological pattern formation. Science 329, 1616–1620 (2010).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Galli, M. & Morgan, D. O. Cell Size Determines the Strength of the Spindle Assembly Checkpoint during Embryonic Development. Dev Cell 36, 344–352 (2016).
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev Biol 100, 64–119 (1983).
Hockendorf, B., Thumberger, T. & Wittbrodt, J. Quantitative analysis of embryogenesis: a perspective for light sheet microscopy. Dev Cell 23, 1111–1120 (2012).
Huisken, J. & Stainier, D. Y. Selective plane illumination microscopy techniques in developmental biology. Development 136, 1963–1975 (2009).
Tomer, R., Khairy, K. & Keller, P. J. Light sheet microscopy in cell biology. Methods Mol Biol 931, 123–137 (2013).
Schierenberg, E. Embryological variation during nematode development. WormBook, 1–13 (2006).
Fickentscher, R., Struntz, P. & Weiss, M. Mechanical cues in the early embryogenesis of Caenorhabditis elegans. Biophys J 105, 1805–1811 (2013).
Fickentscher, R., Struntz, P. & Weiss, M. Setting the Clock for Fail-Safe Early Embryogenesis. Phys Rev Lett 117, 188101 (2016).
Struntz, P. & Weiss, M. Multiplexed measurement of protein diffusion in Caenorhabditis elegans embryos with SPIM-FCS. J Phys D Appl Phys 49, 044002 (2016).
Puliafito, A. et al. Collective and single cell behavior in epithelial contact inhibition. Proc Natl Acad Sci USA 109, 739–744 (2012).
Streichan, S. J., Hoerner, C. R., Schneidt, T., Holzer, D. & Hufnagel, L. Spatial constraints control cell proliferation in tissues. Proc Natl Acad Sci USA 111, 5586–5591 (2014).
Galli, M. et al. aPKC phosphorylates NuMA-related LIN-5 to position the mitotic spindle during asymmetric division. Nat Cell Biol 13, 1132–1138 (2011).
Panbianco, C. et al. A casein kinase 1 and PAR proteins regulate asymmetry of a PIP(2) synthesis enzyme for asymmetric spindle positioning. Dev Cell 15, 198–208 (2008).
Seydoux, G. et al. Repression of gene expression in the embryonic germ lineage of C. elegans. Nature 382, 713–716 (1996).
Rappaport, R. Cytokinesis in animal cells. Int Rev Cytol 31, 169–213 (1971).
Grill, S. W. & Hyman, A. A. Spindle positioning by cortical pulling forces. Dev Cell 8, 461–465 (2005).
Labbe, J. C., McCarthy, E. K. & Goldstein, B. The forces that position a mitotic spindle asymmetrically are tethered until after the time of spindle assembly. J Cell Biol 167, 245–256 (2004).
Kozlowski, C., Srayko, M. & Nedelec, F. Cortical microtubule contacts position the spindle in C. elegans embryos. Cell 129, 499–510 (2007).
Hara, Y. & Kimura, A. Cell-size-dependent spindle elongation in the Caenorhabditis elegans early embryo. Curr Biol 19, 1549–1554 (2009).
Phillips, R. Physical biology of the cell. Second edition, Garland Science (2013).
Edgar, L. G. & McGhee, J. D. DNA synthesis and the control of embryonic gene expression in C. elegans. Cell 53, 589–599 (1988).
Shaham, S. WormBook: Methods in Cell Biology WormBook (2006).
Acknowledgements
Financial support from DFG grant WE4335/3-1 is gratefully acknowledged. Worm strains were provided by the Caenorhabditis Genetics Center, which is funded by the NIH Office of Research Infrastructure Programs (P40 OD010440). This publication was funded by the German Research Foundation (DFG) and the University of Bayreuth in the funding programme Open Access Publishing.
Author information
Authors and Affiliations
Contributions
R.F. performed and analyzed all experiments and wrote all related software; M.W. conceptualized the study and designed experiments; R.F. and M.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fickentscher, R., Weiss, M. Physical determinants of asymmetric cell divisions in the early development of Caenorhabditis elegans . Sci Rep 7, 9369 (2017). https://doi.org/10.1038/s41598-017-09690-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09690-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.