Abstract
Efficient interventions to reduce blood triglycerides are few; newer and more tolerable intervention targets are needed. Understanding the molecular mechanisms underlying blood triglyceride levels variation is key to identifying new therapies. To explore the role of epigenetic mechanisms on triglyceride levels, a blood methylome scan was conducted in 199 individuals from 5 French-Canadian families ascertained on venous thromboembolism, and findings were replicated in 324 French unrelated patients with venous thromboembolism. Genetic context and functional relevance were investigated. Two DNA methylation sites associated with triglyceride levels were identified. The first one, located in the ABCG1 gene, was recently reported, whereas the second one, located in the promoter of the PHGDH gene, is novel. The PHGDH methylation site, cg14476101, was found to be associated with variation in triglyceride levels in a threshold manner: cg14476101 was inversely associated with triglyceride levels only when triglyceride levels were above 1.12 mmol/L (discovery P-value = 8.4 × 10−6; replication P-value = 0.0091). Public databases findings supported a functional role of cg14476101 on PHGDH expression. PHGDH catalyses the first step in the serine biosynthesis pathway. These findings highlight the role of epigenetic regulation of the PHGDH gene in triglyceride metabolism, providing novel insights on putative intervention targets.
Similar content being viewed by others
Introduction
Triglycerides are pivotal players in lipid metabolism in health and disease. They are important sources of energy and transporters of dietary fat. In the blood, triglycerides enable the bidirectional transfer of adipose fat and glucose from the liver. Elevated blood triglyceride levels, however, are associated with several chronic conditions. Two large Mendelian randomization studies employing complementary analytic strategies supported a causal role of triglyceride levels in coronary heart disease1, 2. Further, elevated triglyceride levels is a key feature in several rare and common metabolic abnormalities, including familial hypercholesterolemia, lipodystrophy, diabetes mellitus types I and II, hypertension, metabolic syndrome and obesity, as well as chronic kidney disease3,4,5. Severe hypertriglyceridemia (fasting levels ≥ 500 mg/dL) is further complicated by the life-threatening complication of acute pancreatitis and its chronic consequences5.
Elevated blood triglycerides, defined by fasting levels ≥150 mg/dL (1.7 mmol/L), is present in over 30% of the general adult US population4. Thus, elevated blood triglyceride is a common public health challenge with a broad ranging impact on population health. Managing triglyceride levels is of critical importance for patient’s care and overall population health, but it is challenging. Blood triglyceride levels are largely under the influence of body weight and body fat distribution, lifestyle choices, as well as genetic regulation4. Robust genetic factors for triglyceride variation have been identified but collectively they account for less than 10% of triglyceride level variability in the general population6, 7. On the other hand, over 80% of individuals with high triglyceride levels are either overweight or obese4. Lifestyle interventions associated with decreased Body Mass Index (BMI) have shown to reduce blood triglycerides in some contexts (reviewed in4) but not all8.
Hypertriglyceridemia is undertreated5. Pharmacological interventions to reduce blood triglycerides are scarce, and none have been specifically designed to reduce blood triglyceride levels. Fibrates, which are the most efficient and commonly used drugs to reduce triglyceride levels, are associated with 30–50% decrease in triglyceride levels4. A large proportion of patients, however, do not reach the therapeutic target4. Newer drugs have emerged but their success potential is mitigated by their low tolerability and ineffectiveness in reducing cardiovascular events9. Thus, more efficient and tolerable drugs are needed.
Understanding the molecular mechanisms underlying blood triglyceride variation is key to identifying new intervention targets. Epigenetic marks, such as DNA methylation, are molecular mechanisms that are heritable and reversible, and have been associated with a wide range of environmental stimuli and disease phenotypes10. Others and we have shown that many blood DNA methylation markers can act as surrogates for DNA methylation marks in effector tissues11, 12. Thus, informing function of clinically inaccessible target tissues. This supports the use of DNA methylation profiles in blood as a means to elucidate clinically relevant mechanisms.
DNA methylation marks are highly promising prognostic and therapeutic biomarkers as they are rooted in the mechanisms underlying phenotype expression; thus, more likely to directly guide clinical management of disease13. Emerging studies illustrate the potential impact of DNA methylation biomarkers as risk predictors14, 15 and intervention targets16,17,18.
Others and we previously reported robust association of blood DNA methylation levels in the CPT1A gene with fasting triglyceride levels12, 19,20,21,22,23. In addition to methylation levels at CPT1A gene, other studies found association of methylation levels at the ABCG1 gene with triglyceride levels21,22,23,24,25 and with hypertriglyceridemic waist phenotype26.
To further explore the role of epigenetic mechanisms on fasting triglyceride levels, we conducted a blood methylome study. To reduce environmental and genetic heterogeneity, and increase statistical power, we conducted our discovery analyses in extended French-Canadian families27, and replicated our findings in a dataset of unrelated French individuals28. To better understand the genetic context and functional relevance of our findings, we investigated proximal genetic variations in the same individuals, as well as in silico mRNA expression levels analyses using publicly available datasets. Finally, we explored the role of triglyceride levels in a previously reported association between BMI and methylation levels.
Materials and Methods
Study population
Overall study design
We applied a multi-design strategy that relied on a family-based design and a set of unrelated individuals, for the discovery stage and replicate stage, respectively. The family design allows for better quality control of the genotypic data and control of population stratification, while the use of a second different design replicates effects without replicating confounders by introducing variation in discovery and replication stages; a recommended approach to tackle such bias29, 30.
Discovery Study Sample
The French-Canadian Family study on Factor V Leiden (F5L) Thrombophilia (henceforth the F5L family study) included 369 individuals from 5 extended families27 of which 255 invited subjects consented to participate in the study. The study was originally designed to investigate genetic determinants of quantitative traits related – or potentially related – to venous thromboembolism, and we recently expanded its scope to include the investigation of epigenetic determinants. The families were identified through the Thrombosis Clinic of the Ottawa Hospital between 2005 and 2006, and probands were ascertained on objectively diagnosed idiopathic venous thromboembolism (i.e. venous thromboembolism in the absence of cancer, myeloproliferative disease, pregnancy, puerperium, prolonged immobilization, trauma, surgery, antiphospholipid syndrome, and inherited thrombophilia) and the F5L mutation. All probands self-reported to be of French-Canadian origin and were free of rare genetic risk factors, including anti-thrombin, protein C, and protein S deficiencies, and homozygosity for the F5L and Factor II G20210A mutations. Participants completed an interviewer-administered questionnaire on their personal and medical information. The research ethics boards of the Ottawa Hospital and the University of Toronto approved this study. An informed consent was obtained for all study participants. All methods were performed in accordance with the relevant guidelines and regulations.
Replication Study sample
The MARseille Thrombosis Association (MARTHA) study included 1592 unrelated venous thromboembolism patients of French origin28, recruited between January 1994 and October 2005 from the Thrombophilia Center of La Timone Hospital, Marseille, France. The study was designed to investigate venous thromboembolism and quantitative traits related – or potentially related – to venous thromboembolism. Recruitment occurred at least 3 months after the venous thromboembolism event, which was objectively diagnosed by venography, Doppler ultrasound, angiography and/or ventilation/perfusion lung scan. Study subjects were free of chronic conditions, as well as any well-characterized strong genetic risk factors for venous thromboembolism as described above for the F5L family study. Medical and personal histories were obtained from physician interviews. The research ethics boards of “Ministère de la Recherche et de l’Innovation” approved this study. An informed consent was obtained for all study participants for the purposes of data analysis and publications. All methods were performed in accordance with the relevant guidelines and regulations.
Measurements
DNA methylation
In both the F5L family and MARTHA studies, DNA was extracted from peripheral blood using a salting out procedure adapted from31. Bisulphite conversion and DNA methylation measurements were performed at The Center for Applied Genomics, Toronto, Canada in 227 subjects from the F5L family study and in 350 subjects from the MARTHA study randomly selected from those with whole blood DNA available12. Eleven duplicate samples from both studies were included for quality control. The DNA samples from the F5L family and MARTHA studies were mixed randomly by plate and batch to minimize potential technical biases between the study samples. Bisulphite conversion was performed on 1 µg genomic DNA for each sample using the Qiagen EpiTect 96 Bisulphite Kit and 200 ng of bisulphite-converted DNA at 50 ng/µl was independently amplified, labelled and hybridized to Infinium HumanMethylation450 Bead Chip microarrays. For each sample, the intensities of the methylated and unmethylated signals were measured at 485,577 CpG sites using the Illumina iScan (with default settings). DNA methylation refers to the addition of a methyl group on a CpG site. At a single molecule level, methylation is a then binary mark, i.e. either methylated or unmethylated32 However, the microarray measurements at the tissue level (e.g. blood) reflect the average over all copies of DNA present in the tested sample. As a consequence, DNA methylation measurement at any CpG site is a quantitative signal that can be viewed as the percentage of cells methylated in the sample. Methylation levels at individual CpG sites were reported as β-value, i.e. the ratio of the methylated probe intensity to the overall intensity (sum of methylated and unmethylated probe intensities).
Methylation data from the F5L family and MARTHA studies were merged for quality control and data normalization as previously described11, 33. The data were normalized using SWAN34, Noob35 and a dye bias adjustment method proposed by Illumina. Individual outliers were detected based on a principal component analysis, and four individuals from the F5L family study were excluded. Probe quality was assessed by the detection P-value as defined in the minfi R package, and low detection P-values indicated that the probe signal differed from the background signal. We excluded 6033 probes with detection P-value > 0.05 in more than 5% of the samples, as well as 66,877 polymorphic and 30,969 cross-reactive probes identified by Chen et al.36. The final dataset included 378,594 autosomal probes.
Genotyping
Measurement and procedures for quality control of the genotypic data are described in the Supplemental Data. Briefly, genome-wide genotypes in the F5L family study were typed using the Illumina 660W-Quad Beadchip. SNPs with call rate ≤90% and minor allele count <20 were excluded28. Family structures were validated using genome-wide microsatellite markers and SNP data as described elsewhere27.
MARTHA subjects were genotyped using the Illumina 610 or 660W-Quad Beadchips28. SNPs with a Hardy-Weinberg equilibrium P-value < 10−5, a call rate <99% or a minor allele frequency <1% were excluded from the final dataset.
Lipid-related measures and BMI
Blood samples were collected from study participants after a 12 hour overnight fast, and F5L family study participants had additionally been instructed to abstain from smoking in the previous 12 hours. In the F5L family study, lipoproteins were measured at the Lipid Research Laboratory at St-Michael’s Hospital as described elsewhere37. Blood samples were collected from MARTHA participants after a 12 hours overnight fast at the Department of Haematology of La Timone Hospital. In both studies, serum levels of triglycerides (in mmol/L), very low-density lipoprotein (VLDL, in g/L) and high-density lipoprotein (HDL, in g/L) were assessed by spectrophotometry. The low-density lipoprotein (LDL, in g/L) level was calculated with the Friedewald equation. Triglyceride levels were measured in 239 F5L family study subjects and in 327 subjects of the 350 epigenotyped MARTHA subjects. Subjects self-reported their height and weight, and BMI was calculated as body mass divided by the square of body height.
Cell type proportions
The percentage of cells methylated may vary depending on the tissue - and cell types that composed that tissue - in which it is measured. In the case of whole blood, the DNA methylation measured is that of all leukocyte subtypes together (e.g. neutrophils, monocytes, etc.). Thus, the percentage of cells methylated (i.e. methylation level) in whole blood is an aggregate measure of the proportion from each blood cell types. Inter-individual differences in blood cell type proportions can thus affect blood DNA methylation levels38, and should be accounted for in the analyses. In the MARTHA study, the proportions of lymphocytes, monocytes, neutrophils, eosinophils and basophils were directly measured from whole blood using the ADVIA 120 Haematology System (Siemens Healthcare Diagnostics, Deerfield, IL). Cell type proportions were not measured in the F5L family study. Instead, we used Remove Unwanted Variation (RUV) method39, 40 to capture the cell type proportions as described in Supplemental Data.
Statistical analysis
Due to a highly right-skewed distribution, fasting triglyceride levels were log transformed to improve model fit. DNA methylation β-values were logit transformed into M-values as recommended by Du et al.41.
Methylome association scan
A methylome association scan of triglyceride levels was conducted. A variance components model was fit in the F5L family study to account for familial correlation. Factor components from the Remove Unwanted Variation (RUV) method39, 40 were included in the model to capture cell type proportions and batch effects, and the model was additionally adjusted for age (years) and sex. Triglyceride level was treated as independent variable, and methylation level at each CpG site (M-value) was regressed on triglycerides and potential confounding factors. The statistical significance of the association of triglycerides with DNA methylation levels was tested using the likelihood ratio test. Full details of the statistical methods are provided in the Supplemental Data.
CpG sites that met the genome-wide significance threshold of FDR q-value ≤ 0.05 were further tested for association with triglyceride levels in the MARTHA study. Models were adjusted for age, sex and proportions of measured lymphocytes, monocytes, basophils and eosinophils. P-values ≤ 0.05 after Holm-Bonferroni correction42 were considered statistically significant.
Technical validation and Sensitivity analyses
To assess the robustness of our novel findings, technical validation and sensitivity analyses were performed. Details are provided in the Supplementary materials. Briefly, technical validation of the novel DNA methylation findings was performed by qRT-PCR, and various models to account for cellular heterogeneity and other potential confounders were tested and compared in both studies. Approaches to account for cellular heterogeneity include the Remove Unwanted Variation (RUV) method39, 40 components and adjustments for the estimated or the true cell type proportions. The influence of additional adjustments for potential confounders (e.g. lipid lowering medication, oral contraceptive use and current smoking) was also examined. Finally, we used the gaphunting approach43 to assess whether the CpG sites showed a clustered distribution.
Follow-up analyses
For the novel identified triglyceride-DNA methylation locus, a difference in the strength of association between our discovery and replication studies was observed. To understand the source of this variability, further in-depth analyses were conducted. First, a non-linear association between methylation and triglyceride levels was explored. A Generalized Additive Model (GAM)44 with same covariates adjustment as in the discovery model was fitted. In the F5L family study, the family clusters were treated as random effect to adjust for inter-individual correlation. Upon confirmation of the non-linear association, to quantify the associations, further analyses were conducted using piecewise linear regression models with a pre-determined breakpoint value. The piecewise regression model approach assumes linear relationship within each defined segment on the independent variable’s scale, and estimates linear associations within defined intervals separately45. Full details of the method are described in the Supplemental Data.
A recent methylome study showed an inverse association between BMI and DNA methylation levels46. Since triglycerides and BMI are positively correlated, additional analyses to clarify the interrelationship among BMI, triglycerides and DNA methylation levels at the discovered locus were conducted. First, the strength of the association between BMI and methylation levels at the locus was estimated applying a linear regression model adjusted for same set of covariates as described above. Then, triglyceride level was added as a covariate in the model. The regression coefficient estimates of BMI between these two models were compared. A reduction in the magnitude of the coefficient toward the null when adjusting for triglyceride levels suggests that triglyceride levels lay in the pathway linking BMI to methylation levels. Similarly, the BMI-triglyceride-methylation relationship was investigated by comparing models for triglycerides–methylation association, with or without adjustment for BMI. The direct effect of BMI on methylation levels was also estimated using Structural Equation Modelling (SEM). Triglyceride levels were assumed to lie in the pathway linking BMI and methylation levels. The models were adjusted for age, sex and cell type composition in both studies.
Genetic association analysis
Genetic analysis was carried out to assess whether the observed associations between methylation and triglyceride levels were due to underlying common SNPs33, 47 (Fig. 1a). SNPs within 1 Mb of the CpG sites were first tested for association with methylation levels in the F5L family study, and those with FDR q-value ≤ 0.05 were tested for validation in the MARTHA study. SNPs that were associated with DNA methylation in the MARTHA study after Holm-Bonferroni correction were then tested for association with triglyceride levels across both study samples. These SNP-triglycerides associations were further investigated in silico in the Global Lipids Genetics Consortium meta-analysis results, which involved more than 90,000 individuals48. Finally, the identified SNPs were used as instrumental variables in a Mendelian randomization framework49, 50 to examine a putative causal effect of DNA methylation at the CpG site on triglyceride levels (Fig. 1b). The possibility of reverse causation (Fig. 1c) was examined using the 32 SNPs reportedly associated with triglyceride levels, individually and as a genetic score as described in the Supplemental Data.
Possible relationships between a SNP, a CpG site and triglyceride levels. The SNP is assumed to act on the methylation levels at the CpG site: (a) The SNP can cause variation independently in the level of methylation and triglycerides (common cause); (b) The CpG site can be on the pathway between the SNP and triglyceride levels. In this case, the SNP is expected to be associated with triglyceride levels; (c) The methylation change can be the consequence of the triglyceride levels variation (reverse causation). Reverse causation can be inferred with a genetic variant (SNPtg) associated with triglyceride levels.
All analyses were conducted using R version 3.0.1. Variance components models were fitted with the pedigreemm package, GAM models with the mgcv package, and SEM models with the Lavaan package51.
Results
Study population characteristics
Data on triglyceride levels, genotypes, and DNA methylation levels were available from 199 individuals in the F5L family study and from 324 individuals in MARTHA (Table 1). The two study populations differed in a few aspects: compared to the MARTHA study participants, who are all venous thromboembolism patients, only 11 participants in the F5L family study had a venous thromboembolism. The F5L family participants had, on average, a higher BMI (26.8 kg/m2 vs 24.2 kg/m2; P-value = 1.6 × 10−4) and higher triglyceride levels (1.5 mmol/L vs 1.0 mmol/L; P-value = 9.7 × 10−10); and were on average 4 years younger. Triglyceride levels in the F5L family study were systematically shifted toward higher values compared to the levels observed in the MARTHA study (Fig. 2). The MARTHA study was mainly composed of females (77.5%), whereas the F5L family study had an equal proportion of males and females. Participants in the two studies were similar with respect to LDL and HDL levels. Lipid-lowering drug use was low in both studies (7% and 11%; P-value = 0.15).
Methylome association scan
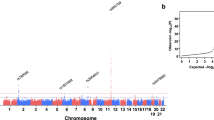
The methylome-wide association scan of triglyceride levels in the discovery data set (i.e. the F5L family study) identified two CpG sites reaching genome-wide significance level. These results are shown in Manhattan plot of the Supplementary Fig. S1; the associated Q-Q plot is shown in Supplementary Fig. S2. DNA methylation levels at these two CpG sites (cg06500161 and cg14476101) were found associated with triglyceride levels in the F5L family study and replicated in the MARTHA study (Table 2). The first CpG site, cg06500161, is located in the ABCG1 gene. DNA methylation at this site was recently reported to be positively associated with triglyceride levels in a population-based cohort25 and in female patients with familial hypercholesterolemia24. We detected a similar strength of the association with respect to magnitude and sign in the MARTHA study (β = 0.11, P-value = 1.92 × 10−8) as in the F5L family study (β = 0.11, P-value = 1.5 × 10−7). The second locus is novel, and maps to the PHGDH gene. DNA methylation at cg14476101 was inversely associated with triglyceride levels in the F5L family study (β = −0.21, P-value = 2.3 × 10−7, 95% confidence interval (CI) (−0.29, −0.13)) and in the MARTHA study (β = −0.08, P-value = 4.8 × 10−2, 95% CI (−0.16, −6.7 × 10−3). Variation in triglyceride levels explained 13% and 6.4% of the methylation level variance at this CpG in the F5L family study and in the MARTHA study, respectively. The effect of triglyceride levels on DNA methylation was independent of the effects of age, sex and cell type proportions. Additional adjustments for lipid lowering medications, oral contraceptive use or current smoking did not alter the association of triglyceride levels with methylation levels at cg14476101 (Supplementary Table S1). In addition, cg14476101 is not annotated as SNP probe in the Illumina provided manifest. Based on the concordance between whole-genome bisulphite sequencing data and the Illumina assay52, the signal provided by cg14476101 is not affected by a SNP. In addition, the cg14476101 did not show clustered distribution in either study. Technical validation of cg14476101 by bisulfite qRT-PCR compared to H450M quantification showed high correlation, 0.865 (Supplementary materials Supplementary Fig. S11).
Hereafter, the focus for the remaining analyses is on the novel cg14476101 PHGDH CpG site. A regional plot of the association of triglyceride levels with DNA methylation levels in the vicinity of cg14476101 is shown in Supplementary Fig. S3.
Non-linear triglyceride-CpG relationship
Although the association between methylation levels at PHGDH cg14476101 and triglyceride levels was replicated in MARTHA, the strength of the association was slightly different across both samples. To investigate the underlying cause of this disparity, a non-linear relationship between methylation and triglyceride levels that may not be captured by a linear model was considered. Under the generalized additive model (GAM) framework, the estimated degrees of freedom for triglyceride levels were similar in both studies: 1.91 (P-value = 1.3 × 10−6) in the F5L family study and 1.97 (P-value = 0.085) in the MARTHA study. These results suggest a non-linear association between triglycerides and methylation levels. A visual inspection of the GAM plots showing the cg14476101 methylation-triglyceride levels relationship (Fig. 3) indicated a similar trend in both studies. A monotonic inverse association between methylation and triglyceride levels was observed. Methylation levels rapidly decreased when triglyceride levels were above a threshold of approximately 1.1 mmol/L in the MARTHA study and 1.6 mmol/L in the F5L family study. This trend was more apparent in the F5L family study. The different threshold and slope observed between these two study samples likely reflect the difference in their triglyceride level distributions observed between the two studies: the distribution was shifted to the left in the MARTHA study compared to the F5L study. Estimation of the non-linear effects separately in each study showed wider confidence intervals at higher values in the MARTHA study compared to confidence intervals at lower values; and confidence intervals in the lower range of the distribution in the F5L family were larger than those observed in the MARTHA study (Fig. 3). Consequently, the strength of associations was estimated with higher uncertainty for lower triglyceride levels in the F5L family study compared to the MARTHA study; and in the MARTHA study, the level of uncertainty was higher in the higher value range of the distribution. That may impact the point estimate of the threshold in both studies.
Partial residual plots from the GAM model showing the relationship between triglyceride and the PHGDH cg14476101 methylation levels in the F5L family and MARTHA studies. The non-linear relationship was modelled with a GAM model adjusted on age, sex and cell type proportions (RUV component in the F5L family study). A random effect was added to the model for the analysis of the F5L family study data to adjust for the relatedness among the family members.
To compare and quantify the observed threshold effect, a piecewise linear regression with a triglyceride levels breakpoint at 1.12 mmol/L was fitted (Table 3). This breakpoint value was selected based on the American Heart Association guidelines for triglyceride levels management4, and aligned with the observed thresholds in this study. Triglyceride levels below the breakpoint were not statistically associated with methylation levels in either dataset, while triglyceride levels above the breakpoint were statistically significant and of similar strength in both the F5L family study (β = −0.26, P-value = 8.4 × 10−6, 95% CI (−0.37, −0.14)) and the MARTHA study (β = −0.25, P-value = 9.1 × 10−3, 95% CI (−0.44, −0.062)).
Genetic association analysis
Of the 217 genotyped SNPs within 1 Mb of cg14476101, six were associated with methylation levels at cg14476101 in both the F5L family and MARTHA studies (Supplementary Table S2). However, none of these SNPs were associated with triglyceride levels in either dataset, nor were they significant in the Global Lipids Genetics Consortium meta-analysis dataset (Supplementary Table S3). Thus, the relationship between cg14476101 methylation and triglyceride levels appears to be independent of proximal genetic variants. These results can be interpreted within a Mendelian Randomization framework. The six genetic variants were assumed to influence methylation levels at cg14476101, and the absence of association between these SNPs and triglyceride levels indicates that variation in cg14476101 methylation levels is unlikely a causal factor in triglyceride level variation. On the other hand, the triglyceride-associated genetic score showed consistent effect size and direction across datasets with cg14476101 methylation levels (β = −0.089; P-value = 0.48 in the F5L family study and β = −0.13; P-value = 0.11 in the MARTHA study, Supplementary Table S4). Thus, these results are consistent with reverse causation but are inconclusive.
Triglycerides-CpG relationship with BMI
An association between PHGDH cg14476101 methylation levels and BMI was recently reported46. Given the well-established relationship between triglycerides and BMI, the potential role of triglyceride levels in the reported BMI-cg14476101 association was investigated (as described in Supplementary Fig. S5).
First, the effect of BMI on cg14476101 methylation levels was estimated. The association was marginal within the F5L family (β = −7.1 × 10−3, P-value = 0.040) and MARTHA (β = −6.9 × 10−3, P-value = 0.11) studies as compared to the one observed for triglycerides. However, the strengths of the associations were similar and directionally concordant with those previously reported46. The strengths of the association of BMI on cg14476101 methylation levels were then compared using a linear model with and without triglycerides as a covariate. BMI was not associated with methylation levels in either study (Table 4). In addition, adjusting for triglyceride levels in the model reduced the effect of BMI on cg14476101 methylation levels by 54% and 29%, respectively, in the F5L family study and in the MARTHA study. These results suggest that triglyceride levels variation likely plays an intermediate role in the BMI-cg14476101 relationship. Conversely, BMI did not significantly attenuate the association between cg14476101 methylation and triglyceride levels in piecewise linear models (3.1% and 3.6% effect estimate reduction, respectively, in the F5L family and MARTHA studies), suggesting that BMI is neither a confounder nor a mediator in the DNA methylation-triglyceride levels relationship (Supplementary Fig. S5; Table 5). We further conducted mediation analyses using structural equation modelling in the F5L and MARTHA datasets assuming the pathway to be: BMI → triglyceride levels → methylation levels (Supplementary Fig. S6). Results from both datasets indicate that the BMI effect on methylation level at the cg14476101 CpG site were mediated through the triglyceride levels (Table 6).
DNA methylation in the ABCG1 and CPT1A genes
As previously reported, associations of methylation levels at the ABCG1 (cg06500161) and CPT1A (cg00574958) genes with triglyceride levels were observed (Supplementary Table S5). Thus, providing external validity to our findings, and highlighting the value of our small but well-characterized study samples for epigenomic investigations. To assess whether non-linearity was also a characteristic of the CPT1A and ABCG1 DNA methylation-triglyceride relationships, a GAM model was applied. None of the associations showed a non-linear trend, and results confirmed the linear relationship (P-value = 4.5 × 10−6 with a degree of freedom estimated at 1 for cg06500161; P-value = 4.0 × 10−4 with a degree of freedom estimated at 1 for cg00574958). Finally, to assess whether methylation levels at the triglyceride-associated PHGDH CpG site were influenced by methylation levels at the ABCG1 and CPT1A sites, additional models adjusting for these sites were tested. Adjustments for methylation levels at the ABCG1 and CPT1A CpG sites did not affect the association between triglyceride levels and the PHGDH CpG site, suggesting that their effects are independent (Supplementary Table S6).
Gene expression association
The cg14476101 probe tags a CpG site located in the first intron of the PHGDH gene on chromosome 1. In order to assess the potential functional relevance of the PHGDH gene and triglyceride levels variation, we looked at the association of the PHGDH expression with the methylation levels at the PHGDH CpG site and with the triglyceride levels. Since gene expression levels were not measured in F5L family and MARTHA studies, we turned to two public datasets: eMS53 and GHS Express54. eMS is a database of associations between DNA methylation levels and gene expression levels in purified peripheral monocytes from the Multi-Ethnic Study of Atherosclerosis55. GHS Express is a public database of expression quantitative trait loci, which includes results of associations between triglyceride levels and monocyte gene expression levels. In the eMS dataset, increased methylation levels at cg14476101 were significantly associated with lower PHGDH expression (β = −0.42, P-value = 2.2 × 10−41, N = 1264). In the GHS Express dataset, the expression of PHGDH was positively associated with triglyceride levels (β = 0.19, P-value = 1.5 × 10−9, N = 1490). Taken together, these results support the association between triglyceride and methylation levels at cg14476101, and suggest a novel role of the PHGDH gene in triglyceride metabolism. For comparison, gene expressions of CPT1A and ABCG1 were also positively associated with triglyceride levels in the GHS Express dataset (β = 0.036, P-value = 5.4 × 10−3 and β = −0.11, P-value = 2.1 × 10−13, respectively). Methylation levels in the CPT1A and ABCG1 genes (cg00574958 and cg06500161) were not available in the eMS dataset.
Discussion
Applying a multi-design strategy and two data sets that have previously been shown to be efficient for the discovery and replication of genetic and epigenetic variants associated with cardiovascular traits, this work reports associations between fasting triglyceride and blood methylation levels at two CpG sites in two genes: PHGDH and ABCG1. Although both datasets used in this study have a common underlying ascertainment scheme favouring the investigation of venous thrombosis, our DNA methylation-triglyceride associations are likely generalizability to other populations, as previously shown in Gagnon et al. 2014.
DNA methylation levels at PHGDH cg14476101 were inversely associated with blood triglycerides. Mendelian randomization and genetic score analysis findings were consistent with triglyceride levels most likely affecting DNA methylation at these sites, rather than vice-versa. In addition, the observed association between DNA methylation at the ABCG1 cg06500161 and blood triglycerides provided robustness to the above novel findings as the cg06500161-triglycerides association had previously been reported24, 25.
The cg14476101 site is located in the first intron of the PHGDH gene. According to the Roadmap Epigenome browser56, the site is located in the liver in a region enriched for the posttranslational histone modification H3K27Ac (acetylation of lysine 27 of the H3 histone protein) that is indicative of active enhancers and in H3K4me3 (Histone 3 lysine 4 trimethylation) a modification that is indicative of proximal promoter regions (Supplementary Fig. S7). In the peripheral blood mononuclear cells, the CpG site lies in close proximity to a region enriched in H3K4me4. These results suggest that the PHGDH CpG site is located in or close to a promoter region. Methylation in a promoter region is often associated with repression of gene expression57. This is consistent with the observed expression results in two public datasets: methylation levels at cg14476106 were inversely associated with PHGDH expression. In addition, an inverse association between the expression of PHGDH and fasting triglycerides was observed in circulating monocytes. The results show that the PHGDH gene is likely involved in triglyceride metabolism. But our analysis is limited to the monocytes. Further studies are needed to understand the relationship between the methylation levels at cg14476101, the expression of PHGDH and fasting triglycerides and in different blood cell types.
While results from the GHS Express and eMS indicated that PHGDH is expressed in monocytes, the Roadmap Epigenome browser showed that PHGDH was barely expressed or completely repressed in peripheral blood mononuclear cells (Supplementary Fig. S7). This disparity may be due to the fact that monocytes only make up 2 to 8% of white blood cells58. Thus, the effect on monocytes might be mitigated by other white blood cell types that have low or no expression of PHGDH. Under the ChromHMM prediction model59, the region that contained cg14476101 is an active transcription starting site and is close to a weak transcription region. These lines of evidence indicate that, under normal conditions, PHGDH is expressed in monocytes and is less likely to be actively transcribed. This is consistent with the study findings: only after reaching a clinically relevant triglyceride levels threshold did methylation levels decrease. This implies that when triglyceride levels are below the threshold, PHGDH is less likely to be expressed; above the threshold, PHGDH expression is likely triggered.
The PHGDH gene encodes 3-phosphoglycerate dehydrogenase, an enzyme that catalyses 3-phosphoglycerate into 3-phosphohydroxypyruvate in the biosynthesis pathway of L-serine60. L-serine is a key enzyme for de-novo sphingolipid synthesis. Sphingolipids are a family of membrane lipids that are involved in diverse functions including lipid metabolism61. Sphingolipids can be found in most lipoproteins62. Increased intake of dietary sphingolipids have been shown to reduce triglyceride levels in mice63 but not in humans64, 65. Additionally, the lipid-lowering drug statin has been shown to lower subtypes of plasma sphingolipids in males with metabolomics syndrome66. Studies investigating the PHGDH gene relationship with sphingolipid levels in humans are needed.
Mutations in the PHGDH gene have been associated with phosphoglycerate dehydrogenase deficiency67 and Neu Laxova syndrome68. No variations in the gene have been associated with triglyceride levels.
The question on the nature of the relationship between blood triglycerides and methylation levels described in this paper is of broad interest: does variation in methylation levels cause variation in triglyceride levels, or vice-versa? In this study, several proximal SNPs associated with methylation levels at cg14476106 were identified. These SNPs can be used as proxies for methylation levels to infer causation through Mendelian randomization approaches49, 50, 69. Intuitively, if variation in methylation levels is causing variation in triglyceride levels, then these SNPs should be associated with triglyceride levels. However, there were no strong evidence to show these SNPs being associated with triglyceride levels in our datasets. Additionally, despite the sufficiently large sample size of the Global Lipids Genetics Consortium study, little or marginal effects of SNPs in the neighbourhood of the methylation site were observed on the triglyceride levels (Supplementary Table S3). These results suggest that variation in methylation levels at cg14476106 do not cause variation in fasting triglyceride levels. Thus, it is unlikely that the triglyceride-associated PHGDH methylation accounts for the reported triglyceride level missing heritability. However, it is plausible that elevated fasting triglyceride levels decrease methylation levels at this site (reverse causation) promoting the expression of the PHGDH gene. Reverse causation can be tested using triglyceride-related SNPs in a Mendelian randomization framework. However, SNP effects on triglyceride levels variation are known to be small, and the datasets available for this study were too small to detect such effects. In a tentative to increase statistical power, a genetic score was developed and tested for association with methylation levels at cg14476106. The genetic risk score explained only a small proportion of triglyceride level variation, which leaves the main findings remain the same. Thus, suggesting that the study datasets were still underpowered or that association between the genetic risk score and methylation levels at cg14476106 is truly absent. Well-powered studies are needed to unequivocally exclude either possibility and to address potential violation of the underlying Mendelian Randomization assumptions.
BMI, a proximate measure for adiposity, is moderately correlated with fasting triglyceride levels. The association between fasting triglycerides and methylation levels may be confounded by BMI. Recently, an association between BMI and methylation levels at cg14476101 was reported46. However, triglyceride levels were not considered in that study. In the present study, evidence for association between BMI and methylation levels at cg14476101 was weak. Instead, the results suggested that blood triglycerides lay on the pathway between BMI and PHGDH methylation. Thus, the previously reported PHGDH methylation associations with BMI may reflect the association with triglycerides.
Using a standard linear model, a difference in effect sizes for the PHGDH cg14476101-triglyceride relationship was observed between the discovery and replication datasets. Triglyceride levels, however, were systematically lower in the MARTHA study compared to the F5L family study. Follow-up analyses showed that the differential effect estimates could be explained by the non-linear association between triglyceride levels and DNA methylation at cg14476101. When triglyceride levels were below 1.12 mmol/L, the relationship was flat in the MARTHA study, and mildly declining in the F5L family study. Conversely, above this threshold, the association declined dramatically in both datasets. The threshold effect implies that methylation levels may change (i.e. the CpG site is less methylated) when triglyceride levels are above a certain value.
An alternative explanation for the difference in the strength of association between cg14476101 methylation and triglyceride levels in the MARTHA and F5L family studies is confounding by an unknown factor that affects both variables. However, we used the RUV method to capture cellular heterogeneity as well as other potential unmeasured confounders, and performed sensitivity analyses. Modelling for additional potential confounders (e.g. lipid lowering medication, oral contraceptive use and current smoking) and for SNPs in the vicinity of cg14476101 did not alter the association between triglyceride levels and methylation levels at cg14476101. Thus, the observed difference between the two studies is unlikely due to the above factors.
Other studies have investigated epigenetic variation with triglyceride levels. In a candidate gene study, pyrosequencing of selected genomic targets revealed associations with CpG sites mapped to ABCG1 and LIPC genes in familial hypercholesterolemia24. In a genome-wide study using the Infinium HumanMethylation450 BeadChip array in CD4+T cells, an association with CpG sites in the CPT1A gene were detected19, and this association was robustly replicated in independent datasets, including ours12, 20. Using the same array on whole blood, another group found associations with CpG sites in ABCG1 and SREBF1 genes25. Those associations were confirmed in a recent epigenome-wide study between lipid profiles and DNA methylation23. The CpG sites in the SREBF1 gene did not meet the genome-wide significance in our data. The reported strength of the association was small (β = 0.043), suggesting that we likely had insufficient statistical power to detect such small effects (Supplementary Fig. S8). But the statistical power was enough to detect the associations with ABCG1 and PHGDH methylation sites. In addition, the F5L family and MARTHA projects were designed to study, respectively, haemostatic proteins and venous thromboembolism. The ascertainment scheme of these studies may have impacted the observed associations with methylation levels. In particular, methylation levels in the MARTHA study were measured only in patients who developed venous thromboembolism. Nevertheless, these study samples identified the previously reported triglycerides-methylation associations in the ABCG1 and CPT1A genes, supporting the appropriateness of these datasets for epigenetic analyses of lipid-related phenotypes. Recently, a Mendelian Randomization analysis21 on 3396 individuals from six studies was performed to investigate the causal relationship between lipid profiles and DNA methylation levels. Results suggest that variations in methylation levels at CPT1A and ABCG1 were a consequence of inter-individual variation of triglyceride levels. Finally, methylation levels at the TXNIP, SLC7A11, and MYLIP were also associated with triglyceride levels22. Both the studies were also based on the Infinium HumanMethylation450 BeadChip array.
No association with PHGDH DNA methylation was reported in previous triglyceride methylome studies19, 20, 22, 23, 25. However, association of PHGDH DNA methylation with triglyceride levels was mentioned - but was not followed up - in the Mendelian Randomization study21. The threshold effect we observed for the PHGDH DNA methylation can affect the effect size estimate, and reduce statistical power in a linear model. Depending on the triglyceride level distribution, the linear model may not have been adequate to detect this association in previous methylome studies.
In summary, a novel association linking increased blood methylation levels at the PHGDH cg14476106 with decreased fasting triglyceride levels was discovered. This association may be non-linear with a clinically relevant threshold of approximately 1.12 mmol/L. These results suggest that the decline in methylation levels at cg14476101 may be a consequence, as opposed to a cause, of high triglyceride levels. These findings may bring new insights into the molecular mechanisms underlying triglyceride level variation.
References
Do, R. et al. Common variants associated with plasma triglycerides and risk for coronary artery disease. Nat Genet 45, 1345–1352, doi:10.1038/ng.2795 (2013).
The Emerging Risk Factors, C. Diabetes mellitus, fasting blood glucose concentration, and risk of vascular disease: A collaborative meta-analysis of 102 prospective studies. Lancet 375, 2215–2222, doi:10.1016/S0140-6736(10)60484-9 (2010).
Upadhyay, R. K. Emerging risk biomarkers in cardiovascular diseases and disorders. Journal of Lipids 2015, 50, doi:10.1155/2015/971453 (2015).
Miller, M. et al. Triglycerides and cardiovascular disease: A scientific statement from the american heart association. Circulation 123, 2292–2333, doi:10.1161/CIR.0b013e3182160726 (2011).
Toth, P. P., Grabner, M., Ramey, N. & Higuchi, K. Clinical and economic outcomes in a real-world population of patients with elevated triglyceride levels. Atherosclerosis 237, 790–797, doi:10.1016/j.atherosclerosis.2014.09.029.
Surakka, I. et al. The impact of low-frequency and rare variants on lipid levels. Nature genetics 47, 589–597, doi:10.1038/ng.3300 (2015).
Teslovich, T. M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Belalcazar, L. M. et al. Lifestyle intervention for weight loss and cardiometabolic changes in the setting of glucokinase regulatory protein inhibition: Glucokinase regulatory protein-Leu446Pro variant in look ahead. Circulation: Cardiovascular Genetics 9, 71–78, doi:10.1161/circgenetics.115.001192 (2016).
Wierzbicki, A. S., Hardman, T. C. & Viljoen, A. New lipid-lowering drugs: An update. International Journal of Clinical Practice 66, 270–280, doi:10.1111/j.1742-1241.2011.02867.x (2012).
Cortessis, V. K. et al. Environmental epigenetics: Prospects for studying epigenetic mediation of exposure–response relationships. Human Genetics 131, 1565–1589, doi:10.1007/s00439-012-1189-8 (2012).
Dick, K. J. et al. DNA methylation and body-mass index: A genome-wide analysis. The Lancet 383, 1990–1998, doi:10.1016/S0140-6736(13)62674-4 (2014).
Gagnon, F., Aïssi, D., Carrié, A., Morange, P.-E. & Trégouët, D.-A. Robust validation of methylation levels association at CPT1A locus with lipid plasma levels. Journal of Lipid Research 55, 1189–1191, doi:10.1194/jlr.E051276 (2014).
Robinson, W. H., Lindstrom, T. M., Cheung, R. K. & Sokolove, J. Mechanistic biomarkers for clinical decision making in rheumatic diseases. Nat Rev Rheumatol 9, 267–276, doi:10.1038/nrrheum.2013.14 (2013).
Reis, I. M., Ramachandran, K., Speer, C., Gordian, E. & Singal, R. Serum GADD45A methylation is a useful biomarker to distinguish benign vs malignant prostate disease. Br J Cancer 113, 460–468, doi:10.1038/bjc.2015.240 (2015).
Chambers, J. C. et al. Epigenome-wide association of DNA methylation markers in peripheral blood from indian asians and europeans with incident type 2 diabetes: A nested case-control study. Lancet Diabetes Endocrinol. 3, doi:10.1016/s2213-8587(15)00127-8 (2015).
Kelly, T. K., De Carvalho, D. D. & Jones, P. A. Epigenetic modifications as therapeutic targets. Nat Biotech 28, 1069–1078, doi:10.1038/nbt.1678 (2010).
Milagro, F. I. et al. A dual epigenomic approach for the search of obesity biomarkers: DNA methylation in relation to diet-induced weight loss. The FASEB Journal 25, 1378–1389, doi:10.1096/fj.10-170365 (2011).
Crider, K. S. et al. Genomic DNA methylation changes in response to folic acid supplementation in a population-based intervention study among women of reproductive age. PLoS ONE 6, e28144, doi:10.1371/journal.pone.0028144 (2011).
Frazier-Wood, A. C. et al. Methylation at CPT1A locus is associated with lipoprotein subfraction profiles. Journal of Lipid Research 55, 1324–1330, doi:10.1194/jlr.M048504 (2014).
Irvin, M. R. et al. Epigenome-wide association study of fasting blood lipids in the genetics of lipid-lowering drugs and diet network study. Circulation 130, doi:10.1161/circulationaha.114.009158 (2014).
Dekkers, K. F. et al. Blood lipids influence DNA methylation in circulating cells. Genome Biology 17, 138, doi:10.1186/s13059-016-1000-6 (2016).
Sayols-Baixeras, S. et al. Identification and validation of seven new loci showing differential DNA methylation related to serum lipid profile: An epigenome-wide approach. The regicor study. Human Molecular Genetics ddw285 (2016).
Braun, K. V. E. et al. Epigenome-wide association study (ewas) on lipids: The rotterdam study. Clinical Epigenetics 9, 15, doi:10.1186/s13148-016-0304-4 (2017).
Guay, S.-P., Brisson, D., Lamarche, B., Gaudet, D. & Bouchard, L. Epipolymorphisms within lipoprotein genes contribute independently to plasma lipid levels in familial hypercholesterolemia. Epigenetics 9, 718–729, doi:10.4161/epi.27981 (2014).
Pfeiffer, L. et al. DNA methylation of lipid-related genes affects blood lipid levels. Circulation: Cardiovascular Genetics. doi:10.1161/circgenetics.114.000804 (2015).
Mamtani, M. et al. Genome- and epigenome-wide association study of hypertriglyceridemic waist in mexican american families. Clinical Epigenetics 8, 1–14, doi:10.1186/s13148-016-0173-x (2016).
Antoni, G. et al. A multi-stage multi-design strategy provides strong evidence that the BAI3 locus is associated with early-onset venous thromboembolism. Journal of Thrombosis and Haemostasis 8, 2671–2679, doi:10.1111/j.1538-7836.2010.04092.x (2010).
Antoni, G. et al. Combined analysis of three genome-wide association studies on VWF and FVIII plasma levels. BMC Medical Genetics 12, 102 (2011).
Rosenbaum, P. R. Replicating effects and biases. The American Statistician 55, 223–227, doi:10.1198/000313001317098220 (2001).
Kraft, P., Zeggini, E. & Ioannidis, J. P. A. Replication in genome-wide association studies. Statistical science: a review journal of the Institute of Mathematical Statistics 24, 561–573, doi:10.1214/09-STS290 (2009).
Miller, S., Dykes, D. & Polesky, H. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic acids research 16, 1215 (1988).
Michels, K. B. et al. Recommendations for the design and analysis of epigenome-wide association studies. Nat Meth 10, 949–955 (2013).
Aïssi, D. et al. Genome-wide investigation of DNA methylation marks associated with FV Leiden mutation. PLoS ONE 9, e108087, doi:10.1371/journal.pone.0108087 (2014).
Maksimovic, J., Gordon, L. & Oshlack, A. Swan: Subset-quantile within array normalization for illumina infinium humanmethylation450 beadchips. Genome Biology 13, R44 (2012).
Triche, T. J. & Weisenberger, D. J. Van Den Berg, D., Laird, P. W. & Siegmund, K. D. Low-level processing of illumina infinium DNA methylation beadarrays. Nucleic Acids Research 41, e90–e90, doi:10.1093/nar/gkt090 (2013).
Chen, Y.-A. et al. Discovery of cross-reactive probes and polymorphic cpgs in the illumina infinium humanmethylation450 microarray. Epigenetics 8, 203–209, doi:10.4161/epi.23470 (2013).
Martin, L. J. et al. Serum lipids, lipoproteins, and risk of breast cancer: A nested case-control study using multiple time points. Journal of the National Cancer Institute 107, djv032 (2015).
Reinius, L. E. et al. Differential DNA methylation in purified human blood cells: Implications for cell lineage and studies on disease susceptibility. PLoS ONE 7, e41361, doi:10.1371/journal.pone.0041361 (2012).
Gagnon-Bartsch, J. A. & Speed, T. P. Using control genes to correct for unwanted variation in microarray data. Biostatistics 13, 539–552, doi:10.1093/biostatistics/kxr034 (2012).
Jaffe, A. & Irizarry, R. Accounting for cellular heterogeneity is critical in epigenome-wide association studies. Genome Biology 15, R31 (2014).
Du, P. et al. Comparison of beta-value and m-value methods for quantifying methylation levels by microarray analysis. BMC bioinformatics 11, 587 (2010).
Holm, S. A simple sequentially rejective multiple test procedure. Scandinavian Journal of Statistics 6, 65–70, doi:10.2307/4615733 (1979).
Andrews, S. V., Ladd-Acosta, C., Feinberg, A. P., Hansen, K. D. & Fallin, M. D. “Gap hunting” to characterize clustered probe signals in illumina methylation array data. Epigenetics & Chromatin 9, 56, doi:10.1186/s13072-016-0107-z (2016).
Wood, S. N. Fast stable restricted maximum likelihood and marginal likelihood estimation of semiparametric generalized linear models. Journal of the Royal Statistical Society: Series B (Statistical Methodology) 73, 3–36, doi:10.1111/j.1467-9868.2010.00749.x (2011).
Von Eye, A. & Schuster, C. Regression analysis for social sciences. (Elsevier Science Publishing Co Inc, 1998).
Aslibekyan, S. et al. Epigenome-wide study identifies novel methylation loci associated with body mass index and waist circumference. Obesity 23, 1493–1501, doi:10.1002/oby.21111 (2015).
Bell, J. T. et al. DNA methylation patterns associate with genetic and gene expression variation in hapmap cell lines. Genome Biol 12, R10 (2011).
Willer, C. J. et al. Discovery and refinement of loci associated with lipid levels. Nature genetics 45, doi:10.1038/ng.2797 (2013).
Brion, M.-J., Benyamin, B., Visscher, P. & Smith, G. Beyond the single snp: Emerging developments in mendelian randomization in the “omics” era. Curr Epidemiol Rep 1, 228–236, doi:10.1007/s40471-014-0024-2 (2014).
Relton, C. L. & Davey Smith, G. Two-step epigenetic mendelian randomization: A strategy for establishing the causal role of epigenetic processes in pathways to disease. International Journal of Epidemiology 41, 161–176, doi:10.1093/ije/dyr233 (2012).
Rosseel, Y. Lavaan: An r package for structural equation modeling. 2012 48, 36, doi:10.18637/jss.v048.i02 (2012).
Naeem, H. et al. Reducing the risk of false discovery enabling identification of biologically significant genome-wide methylation status using the humanmethylation450 array. BMC Genomics 15, 51, doi:10.1186/1471-2164-15-51 (2014).
Liu, Y. et al. Methylomics of gene expression in human monocytes. Human Molecular Genetics 22, 5065–5074, doi:10.1093/hmg/ddt356 (2013).
Zeller, T. et al. Genetics and beyond – the transcriptome of human monocytes and disease susceptibility. PLoS ONE 5, e10693, doi:10.1371/journal.pone.0010693 (2010).
Bild, D. E. et al. Multi-ethnic study of atherosclerosis: Objectives and design. American Journal of Epidemiology 156, 871–881, doi:10.1093/aje/kwf113 (2002).
Bernstein, B. E. et al. The nih roadmap epigenomics mapping consortium. Nat Biotech 28, 1045–1048, doi:10.1038/nbt1010-1045 (2010).
Jones, P. A. Functions of DNA methylation: Islands, start sites, gene bodies and beyond. Nature Reviews Genetics 13, 484–492 (2012).
McPherson, R. A., Pincus, M. R. & Henry, J. B. Henry’s clinical diagnosis and management by laboratory methods. (Saunders Elsevier, 2007).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Yoshida, K. et al. Targeted disruption of the mouse 3-phosphoglycerate dehydrogenase gene causes severe neurodevelopmental defects and results in embryonic lethality. Journal of Biological Chemistry 279, 3573–3577 (2004).
Worgall, T. S. In Sphingolipids and metabolic disease 139–148 (Springer, 2011).
Hammad, S. M. In Sphingolipids and metabolic disease 57–66 (Springer, 2011).
Duivenvoorden, I. et al. Dietary sphingolipids lower plasma cholesterol and triacylglycerol and prevent liver steatosis in APOE* 3 Leiden mice. The American journal of clinical nutrition 84, 312–321 (2006).
Ohlsson, L., Burling, H., Duan, R.-D. & Nilsson, Å. Effects of a sphingolipid-enriched dairy formulation on postprandial lipid concentrations. European journal of clinical nutrition 64, 1344–1349 (2010).
Ramprasath, V. R., Jones, P. J., Buckley, D. D., Woollett, L. A. & Heubi, J. E. Effect of dietary sphingomyelin on absorption and fractional synthetic rate of cholesterol and serum lipid profile in humans. Lipids in health and disease 12, 1 (2013).
Ng, T. W. et al. Dose-dependent effects of rosuvastatin on the plasma sphingolipidome and phospholipidome in the metabolic syndrome. The Journal of Clinical Endocrinology & Metabolism 99, E2335–E2340 (2014).
Tabatabaie, L. et al. Novel mutations in 3-phosphoglycerate dehydrogenase (PHGDH) are distributed throughout the protein and result in altered enzyme kinetics. Human Mutation 30, 749–756, doi:10.1002/humu.20934 (2009).
Acuna-Hidalgo, R. et al. Neu-laxova syndrome is a heterogeneous metabolic disorder caused by defects in enzymes of the l-serine biosynthesis pathway. The American Journal of Human Genetics 95, 285–293, doi:10.1016/j.ajhg.2014.07.012 (2014).
VanderWeele, T. J., Tchetgen Tchetgen, E. J., Cornelis, M. & Kraft, P. Methodological challenges in mendelian randomization. Epidemiology (Cambridge, Mass.) 25, 427–435, doi:10.1097/EDE.0000000000000081 (2014).
Acknowledgements
F.G. and M.D.W. hold Canada Research Chairs. J.D. is a Vanier Canada Graduate Scholar. J.D. and N.Z. are Fellows in the Canadian Institutes of Health Research Strategic Training for Advanced Genetic Epidemiology (CIHR STAGE) program. D.A. was supported by a Ph D grant from the Région Ile de France (CORDDIM) and the Nouvelle Société Française d’Athérosclérose (NSFA). This work was partially supported by the Canadian Institutes of Health Research, the Heart and Stroke Foundation of Canada and by the ICAN Institute for Cardiometabolism and Nutrition (ANR-10-IAHU-05). The authors are thankful to the individuals who participated in the study, and to Karl Everett and Michael Lerro for reviewing the manuscript.
Author information
Authors and Affiliations
Contributions
F.G., P.W., D.A.T. and P.E.M. designed the F5L Family and MARTHA studies and collected the data. J.D. and D.A. pre-processed the data. V.T. led the data analysis with support from S.H. F.G., V.T. and S.H. participated to the manuscript preparation. V.T., S.H., J.D., M.L., N.Z., D.A., I.K., P.W., P.E.M., M.W., D.A.T., F.G. contributed to the revision process. C.P. performed the technical validation of DNA methylation at cg14476101 by relative quantitative PCR with specific TaqMan probes. All the authors approved the manuscript for publication.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Truong, V., Huang, S., Dennis, J. et al. Blood triglyceride levels are associated with DNA methylation at the serine metabolism gene PHGDH . Sci Rep 7, 11207 (2017). https://doi.org/10.1038/s41598-017-09552-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09552-z
This article is cited by
-
Obesity-driven mitochondrial dysfunction in human adipose tissue-derived mesenchymal stem/stromal cells involves epigenetic changes
Cell Death & Disease (2024)
-
Transcription factor NKX2–1 drives serine and glycine synthesis addiction in cancer
British Journal of Cancer (2023)
-
Fallopian tube lesions as potential precursors of early ovarian cancer: a comprehensive proteomic analysis
Cell Death & Disease (2023)
-
DNA methylation trajectories and accelerated epigenetic aging in incident type 2 diabetes
GeroScience (2022)
-
ABCA1 and ABCG1 DNA methylation in epicardial adipose tissue of patients with coronary artery disease
BMC Cardiovascular Disorders (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.