Abstract
In this study, we categorize the microbial community in mangrove sediment samples from four different locations within a vast mangrove system in Kerala, India. We compared this data to other samples taken from the other known mangrove data, a tropical rainforest, and ocean sediment. An examination of the microbial communities from a large mangrove forest that stretches across southwestern India showed strong similarities across the higher taxonomic levels. When ocean sediment and a single isolate from a tropical rain forest were included in the analysis, a strong pattern emerged with Bacteria from the phylum Proteobacteria being the prominent taxon among the forest samples. The ocean samples were predominantly Archaea, with Euryarchaeota as the dominant phylum. Principal component and functional analyses grouped the samples isolated from forests, including those from disparate mangrove forests and the tropical rain forest, from the ocean. Our findings show similar patterns in samples were isolated from forests, and these were distinct from the ocean sediment isolates. The taxonomic structure was maintained to the level of class, and functional analysis of the genes present also displayed these similarities. Our report for the first time shows the richness of microbial diversity in the Kerala coast and its differences with tropical rain forest and ocean microbiome.
Similar content being viewed by others
Introduction
The mangrove ecosystem plays a crucial role by acting as a buffer zone between land and sea, maintaining the sea level and protecting the coast1. Mangroves are a crucial component of the food chain in the saline coastal biome of the tropics and subtropics. Mangrove trees convert solar energy into organic matter via photosynthesis, with their leaves and branches serving as a source of energy and providing a habitat for a variety of aquatic organisms, which in turn, support a higher level in the food chain. This ecosystem is an enormous food web, supplying a myriad of microorganisms with both protection and nutrients2,3. It is considered to be one of the most critical in tropical regions, and also one of the most vulnerable to global climate change4.
The complexity of the mangrove microbial communities has generated deep interest among microbial ecologists. The dynamic environment of the mangrove ecosystem, brought about by the regular tidal variations, pH, temperature, salinity, light, rainfall and nutrient availability provides an excellent environment for a wide range of organisms with diversified functional roles5. Studies have shown that microbial communities play a vital role in this ecosystem, being essential for biogeochemical cycles and biocycling of most nutrients, including nitrogen6,7. Except for absorption of nutrients, transformation processes of nutrients can also be important for avoiding eutrophication8.
The impact that mangrove forests and their supporting ecosystem extends beyond their local ecological impact. Explorations of this ecosystem have revealed some interesting bioactive compounds that could have a potential impact on human health. These include some of the novel compounds with cytotxoic activity9,10, some p-aminoacetophenonic acids that may have antibiotic properties11,12,13, some indolesesquiterpenes that appear to have selective anti-HIV activity14,15 and an anti-fibrotic compound9.
Mangroves cover up to 152,000 km2 globally8, but their range is decreasing due to pollution, urbanization and other human activities. Conservation and research on mangroves are imperative due to their dominant role in the marine food chain and the recognition that they may play a role in human health by their potential ecological, industrial and pharmaceutical impacts5. There is an increasing urgency to understand the structural and functional architecture that underlies the mangrove ecosystem, and the microbial community is an important part that remains unexplored. With more than 98% of microbes currently unculturable16, examination of environmental samples by metagenomic analysis is now the only avenue available for exploring these communities. Until now, the few metagenomic studies in mangroves have been concentrated South America17. The South American mangrove ecosystems are important, but constitute only 11% of the range of mangroves across the globe18. In this study, we carried out the first detailed analysis of microbial communities within the mangrove ecosystems from four different locations in mangrove forests across Kerala, India, and compare them to a similar Brazilian ecosystem, a tropical rainforest, and samples taken from ocean sediment.
Results
Sequencing, quality control and annotation of proteins
A total of 64413433 sequencing reads were obtained from the four samples from Kerala with an average length of 254 ± 12.75 bases. Following quality trimming, a total of 54833910 (85.13% of the total) remained, including for 7809729 reads for PGD (average read length of 408 ± 138 bp), 7303470 reads for MAL (average read length of 408 ± 138 bp), 6414174 reads for VL1 (average read length of 200 ± 68 bp), and 5920270 reads for PYN (average read length of 193 ± 64 bp). A total of 18.6 GB (64 million reads) were obtained across all four of the Kerala mangrove samples, and 54 million reads (85.13%) were retained for further analysis following quality control. The sequences for these reads are publicly available from the MG-RAST server under the following IDs: 4671371, 4671370, 467136 and 4671368. Of the sequences that passed quality control, each had a certain percentage that mapped to proteins of known (PGD= 39%, MAL= 41.2%, VL1= 35.4%, and PYN= 35.4%) and also to proteins of unknown function (PGD= 7.5%, MAL= 41.2%, VL1= 44.5%, and PYN= 44.1%). Asymptotic rarefaction curves were generated for each sample indicating that the majority of taxonomic diversity was covered in the samples (data not shown).
Taxonomic Diversity across the Kerala samples
The taxonomic classifications of genes were assigned to the RefSeq annotation source19 using the Best Hit Classification algorithm of MG-RAST. A total of 13269130 representative sequences were assigned to different taxonomies using the RefSeq database from all the four datasets accounting for more than 99% of the reads for PYN, MAL, VL1 and PGD samples respectively (Table 1). There were also a number of sequences that could not be assigned to the highest taxonomy levels, with an average of 15.5% of the reads either unassigned, unclassified, or identified as other. Collectively, bacterial sequences dominated the overall reads accounting for 96.81% of the total assigned reads, with Eukaryota and Archaea assigned 1.8% and 1.32% respectively. Less than 1% of the reads mapped to viruses. Sequences from all the datasets were assigned to 28 different bacterial, 5 archaeal, and 34 eukaryotic phyla. All four samples shared Proteobacteria as the phyla with the most reads assigned, having an average of 65.70% (Supplementary Table 1). Similar rich dominance of Proteobacteria was found in Brazilian oil contaminated mangroves20. Other dominant bacterial phyla included Bacteriodes (11.83%), Firmicutes (5.56%) and Actinobacteria (3.61%), but there was some variation in the ranking of each of these among the Kerala samples. Although all samples shared Proteobacteria as the predominant phylum, there was diversity in the percentage of the reads assigned the classes within that taxon (Fig. 1 and Table 2), with PYN and MAL both sharing Gammaproteobacteria as the dominant class, and VL1 and PGD with the majority of their Proteobacteria reads mapped to Betaproteobacteria. A similar lack of structure is seen within the phylum Bacteriodes. Both PYN and MAL have Flavobacteriia as the dominant class, but VL1 has most of its Bacteriodes reads map to Bacteroidia, and in PGD it is Flavobacteria. The classes within Firmicutes and Actinobacteria phyla have similar diversity and percentage of reads assigned to their top classes across all of the Kerala samples.
A majority of the reads assigned to Archaea (1.32 ± 0.44% of total reads) mapped to the phylum Euryarchaeota (81.77% ± 7.02) across all four samples (Fig. 2 and Table 2). The remainder of the reads in Archaea mapped predominantly to Crenarchaeota (10.43% ± 2.16) and Thaumarchaeota (6.43 ± 8.47), although the order and the percentage varied across the four Kerala locations. Of the 1.32% of the assigned reads that mapped to Eukaryota, 90% of those reads were found across 8 phyla (Fig. 3 and Table 2), with the majority (18.04% ± 3.55) mapping to the phylum Streptophyta, closely followed by Ascomycota (16.4% ± 1.78) in all of the samples except PGD. Unlike what was seen in Bacteria and Archaea, a large number of the Eukaryota reads (14.94% ± 0.88) could not be classified to a specific phylum.
An analysis of the genera found in these sediment samples showed unique differences at each sampling location, with some similarities. The reads that mapped to each genus were calculated, and only those genera with more than 1% of the total mapped reads were noted (Fig. 4). Burkholderia and Geobacter were the only genera that were found in each of the sampling locations. Each location had a different predominant genus, and while three locations had most of the same dominant genera (PYN, MAL and VL1), PGD was quite different in both the number and members identified.VL1 had Sulfuricurvum, which at 6% had almost double the reads as the next genus (Sulfurimonas). Burkholderia was the most prominent in PGD, with Acidovorax also prominent. Geobacter, Gramella and Shewanella were the dominant genera found at the PYN location, and Marinobacter was the most prominent in MAL.
Geographic locations and diversity in the sampling sites in the mangrove ecosystem of Kerala, India. Genera with more than 1% of the mapped reads are listed, with their rank and indications of their abundance are shown, and organisms shared across sampling locations are indicated by shading or boxes. (Image source: http://www.d-maps.com/carte.php?num_car=24853&lang=en andInkscapev0.92 was used for sample site labelling).
Comparison of metagenomes from different soil samples
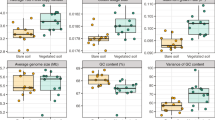
Sediment samples from three different ecosystems were compared to the Kerala mangrove samples. These included four samples from a Brazilian mangrove forest21, sediment from a tropical rain forest in Puerto Rico22, and four ocean sediment samples from the South China Sea23. A Principal Component Analysis (PCA) of all the samples clearly grouped the forest isolates together on one axis, clearly separating them from the ocean samples (Fig. 5).
Principal Component Analysis and geographic location of samples compared across the globe. A. PCA analysis comparing mangrove forests, a rain forest, and four ocean sediment samples. B. Geographic locations of the samples that were compared in this analysis. (Image source- Link:https://commons.wikimedia.org/wiki/File:BlankMap-World-noborders.pngand Inkscape v0.92 was used for labelling).
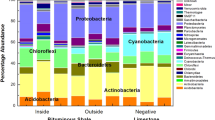
MG-RAST’s organism tree tool was used to examine patterns across the different sediment types. At the highest taxonomic level (Super Kingdom) clear differences were seen between the ocean sediment samples, and those isolated from the two types of forests. The four ocean samples had an average of 868769.5 (±123583) reads, with an average of more than 99% assigned to a taxonomic category (Table 1 and Fig. 6). In contrast, the majority of the reads isolated from any of the forest samples mapped to Bacteria, with Archaea being less than 5% of the assigned total in any sample (Table 1 and Fig. 6). All of the sediment samples had approximately 1.5% of the reads assigned to Eukaryota. Very few virus reads were found.
A more detailed look at the two taxonomic levels with the majority of the reads assigned (Archaea and Bacteria) showed some striking differences between the assigned phyla within these two groups. Most notably, the ocean sediment samples had the highest archaeal reads, while all of the arboreal samples are predominantly bacterial (Table 1). An average 9. 3% of the ocean reads mapped to Archaea, and more than 95% of the arboreal reads, both mangrove and rain forest, are bacterial. Those bacterial reads, regardless of the ecology of the samples, map to Proteobacteria across all samples (Fig. 6), but that similarity does not extend beyond the phylum. Within this phylum, the Kerala mangrove samples have either Gamma- or Betaproteobacteria as the class that has the most reads assigned to it, with the Brazilian mangrove and Puerto Rican forest samples both having Gammaproteobacteria as the dominant class (Supplementary Table 4). The ocean samples have either Beta- or Alphaproteobacteria as the most dominant class. The lack of consensus, both within and between the groups, is also seen with classes within the Bacteriodes and Firmicutes phyla (Supplementary Table 4). The only bacterial phylum where all groups shared a similar structure was the Actinobacteria, with Actinomycetales as the dominant class (Supplementary Table 4).
The ocean sediments had a high abundance of reads that map to the archaeal kingdom and within that, Thaumarchaeota as the dominant phylum (Fig. 6). Archaea reads were less than 5% of the total from the samples isolated from forests, but across all of them, Euryarchaeota was the dominant phylum, followed by Crenarchaeota. All of the samples, both from the mangrove and rain forests, had Methanomicrobia as the dominant class in Euryarchaeota. They also shared Thermoprotei as the most prevalent Crenarchaeota class. Reads that mapped to these particular classes were not even found (or barely registered) in any of the ocean samples, which instead had Thermococci and Methanococci as the most prevalent classes among the ocean reads that mapped to the Euryarchaeota phyla.
Less than 2% of the total reads from any sample mapped to Eukaryota. Most of these reads mapped to Streptophyta in the Indian and Brazilian samples, but the majority of these could not be classified to any particular class (Supplementary Table 4). The dominant class in the sample from the rain forest was Ascomycota. Streptophyta was also an important part of the ocean eukaryotic reads, but the Bacillariophyta, which contains the diatoms, was the dominant phylum. There was no clear consensus in a dominant class across the groups.
Functional analysis
A functional comparison that mapped the abundance profiles to different categories across the Subsystems24 was performed for each of the Indian mangroves samples. All four samples had similar distributions and abundance of reads that were mapped at the highest levels of subsystem categorization (Supplementary Table 2).
A single sample was chosen from each of the groups for a functional comparison (India mangrove = PGD/4671368.3, Brazil mangrove = BrMgv2/4451034.3, Puerto Rico Forest = 4446153.3, South China Sea = E201-2/4487295.3). The numbers of hits for each of the functional levels were summed, and the percentages of hits per each level compared to the total number were compared (Supplementary Table 3 and Fig. 7). The forest samples had similar profiles, the exception being Protein Metabolism, where the India mangrove sample (PGD) had many fewer reads (Supplementary Table 2), and the ocean sample with a clear majority of mapping reads. The subsystems that had the highest Z score when comparing the average across the three forest samples to the ocean sample were Carbohydrates, Clustering-based subsystems, Protein metabolism and Amino acids derivatives (Supplementary Table 3 and Fig. 7).
Discussion
Similarities and differences seen across the Kerala mangrove samples
All of the Kerala mangrove samples shared certain microbial community structures. Bacteria were the predominant organisms, averaging 81.72% of the total reads across all four locations. Metagenomic analysis of mangrove at Cardoso Island State Park, Brazil, through 16S rRNA pyrosequencing showed similar dominance of Proteobacteria (88% of overall sequence) irrespective of soil depth through 16S rRNA pyrosequence25. Within the Bacteria, Proteobacteria was the dominant phylum, followed by Bacteriodes, Firmicutes and then Actinobacteria. Similar bacterial phylum dominancy was found using PCR-Clone based metagenomic library screening26 as well as 16S rRNA ribo-typing27. The similarities within Kerala datasets seen across the higher taxonomic levels (Kingdom and Phylum) did not continue at the class level, where the composition of the bacteria varied at each location (Fig. 1), especially among the classes within the Proteobacteria and Bacteriodes phyla. Archaea followed next with the most reads, but with an average of 1.8%, they were not considered a dominant member of the Kerala microbial community. Despite their small numbers, they had a remarkable consistency in their taxonomic structure, maintaining the same divisions even to the class level (Fig. 2). Previous whole metagenome study in Brazilian mangroves sediments by Andreote et al.21 have found similar abundance (0–3.4%) of archaea. Furthermore, mangrove soil sediment from Saudi Arabia28 has also showed similar percentage abundance (3.5%). The consistent level of archaea in mangrove samples could denote its importance in the ecosystem such as N cycle by Thaumarchaeota and the favorable conditions for Methanogens29. Eukaryota had almost as many reads assigned to it as the Archaea, with 1.32% of the total, and all four samples had similar distributions at the taxon class level (Fig. 3).
Genera that had more than 1% of the reads mapped to them were examined at each sampling location, revealing some similarities and differences across the Kerala isolates (Fig. 4). Only two genera, Burkholderia and Geobacter, had more than one percent of the sequences map to them in each of the four samples. The mangrove samples from Kerala (India) were affected by anthropogenic activities, hence, it is very likely that the high dominance of Burkholderia could be due to its role involved in degradation of various compounds such as polycyclic aromatic hydrocarbons (PAHs), diesel, kerosene, naphthalene, and phenol30,31. Burkholderia sp. was also abundant in Okinawa (Japan) oil contaminated mangrove sediments32. Three of the locations (MAL, PYN and VL1) shared most of the genera that had more than 1% of the total reads, but those found in PGD at similar levels were mostly unique. The two predominant genera found in the VL1 were Sulfuricurvum and Sulfurimonas, with 6.2% and 3.66% of the total reads mapping to them. Both of these genera have been associated with sulfur oxidation33,34,35,36,37. Sulfuricurvum is in high concentration only in VL1, but Sulfurimonas is also a predominant organism in PYN, with 1.19% of the reads mapping to this genus in this sample. MAL and PYN were in close proximity compared to the other samples, and they shared Marinobacter as a dominant genus (with 3.93% and 0.79% of the reads, respectively). Marinobacter has been described as being ubiquitous across the global oceans38, and is known to degrade hydrocarbons39 and fix nitrogen40. This genus is also of interest as it has been shown to be one of the few bacterial genera known to “bloom” when oil or oil constituents are introduced into seawater41, and its predominant presence in the co-located MAL and PYN, as compared to VL1 and PGD could suggest some pollution in that environment when the samples were taken.
Although there were differences in the bacterial membership at certain taxonomic levels, this was not repeated when the functional capacity of the four samples was compared. The reads across all four samples had similar abundance patterns that were assigned to the 28 different functional categories used by Subsystems22. No significant differences were seen in sulfur metabolism, despite VL1 having the two most abundant genera being sulfur oxidizers.
Similarities and differences across global samples
To look for similarities that might be shared across similar ecosystems, the Indian mangrove samples were compared with four samples isolated from mangrove forests in Brazil21, a tropical forest in Puerto Rico22 and four samples collected sediment from the South China sea23. A principal component analysis clustered the samples taken from forests, which includes both the mangrove samples and the tropical rain forest isolate, closer to the y-axis than those samples collected in the ocean (Fig. 5). A taxonomic analysis revealed supported this finding. At the highest taxonomic levels, all of the forest samples had similar taxonomic patterns (Fig. 6), with the ocean samples being distinctly different. Bacteria were the dominant kingdom in all samples taken from forests with Archaea less than 5%, however, the ocean isolates have almost 12.9% archaeal reads. Previously, more than 87% of the microbial biomass was seen to be dominated by Archaea in deep subsurface sediments42. In addition, Antarctic circumpolar continental shelf waters have been shown to be highly dominated by Archaea43. Dominance of archaea in deep ocean subsurface and bacteria in arboreal samples could be explain according to the theory proposed by Valentine44 which states that bacteria can adapt to the changing environment while archaea can sustain in the nutrient limited environment. At the phylum taxon, all forest samples were predominantly Proteobacteria. This was also true of the ocean samples, but as they accounted for less than 10% of the total reads they were not the dominant ocean phylum. Nevertheless, Proteobacteria was found to be the most dominant in all the ecosystems within the bacterial kingdom. Saline environments have shown to harbor Proteobacteria in the past45,46. The mangrove samples from Brazil and India had a mixture of Bacteriodes, Firmicutes and Actinobacteria as a significant part of the bacterial phyla diversity, which corroborates to mangroves data generated from other part of India47, but the Puerto Rican rainforest had two additional phyla (Acidobacteria and Planctomycetes) that were only a small part of the diversity in the mangroves. Acidobacteria have been found to be dominant in rain forest sample48,49. Ocean samples, which also included reads that mapped to Bacteriodes, Firmicutes and Actinobacteria.
Similarities were also seen in the structure of the Archaea across the four ecosystems examined. Of the Archaea reads that were present, the forest samples contained mainly Euryarchaeota. The ocean samples were primarily archaeal and mostly of the phylum Thaumarchaeota. The finding is in corroboration with the work of Quaiser et al.50, which showed the dominance of archaea by Group-I Thaumarchaeota in the sediment of Marmara Sea. They found that the archaeal amo (ammonia mono-oxygenase) genes were highly abundant in Marmara Sea suggestion the dominance of Thaumarchaeota in ammonia oxidation. Marine Group-I (MG) group were also found to be the most dominated group among the archaeal kingdom in deep Mediterranean Sea51. Studies have shown that the seasonal variation affects the archaeal diversity wherein Marine Group-I (MG) and Euryarchaeota MG II.b dominates during winter and Euryarchaeota MG II.b during summer52. Archaeal diversity can also be influenced by the difference in zones (depth) of the sea/ocean; the oxic/anoxic interface zone featured high dominance of Marine Group-I (MG) archaea. However, significant reduction was exhibited in sulfatemethane transition zone and methanic zone53. Similar patterns were also seen in the Eukaryota, which were not in the majority in any sample. Across both the forest and ocean samples the Streptophyta, which include the green plants, were a predominant phylum found in the eukaryotic reads. The forest samples also had reads that mapped to the phyla Ascomycota (fungi) in significant numbers and a large number in Chordata (vertebrates), Cnidaria, and Arthropoda (insects and arachnids). These same phyla were also present in the ocean. The mangrove and ocean samples had a number of reads mapping to the phylum Bacillariophyta (data not shown), which include the diatoms. Bacillariophyta play a crucial role in generation of organic carbon soluble in organic compounds in the ocean bottom and also produces exopolysaccharides (EPS) which stabilizes the sedimentary materials54.
A similar pattern that groups the forest samples distinctly from the ocean isolates was also seen in the functional analysis. While all samples had similar functional profiles in some of the subsystems, the forest samples were clearly distinct from the ocean samples in several of the subsystems examined. Interestingly, the ocean samples examined had more reads that mapped to genes active in the subsystems defined as Carbohydrates, Amino Acids and Derivatives, Protein Metabolism, Respiration, and RNA Metabolism (Fig. 7), but had fewer reads that mapped to the Virulence, Disease and Defense.
Conclusions
A metagenomic analysis of isolates from soil sediment in four different locations across the Kerala, India mangrove forest ecosystem showed strong similarities across higher taxonomic divisions extending to the level of phylum. Comparisons of the Indian mangrove isolates to samples from mangrove in Brazil, and to a tropical rain forest in North America showed similar patterns, with most of the reads mapping to phylum Proteobacteria within the Bacteria kingdom. Fewer reads were found that mapped to Archaea, but those that were present predominantly to the phylum Euryarchaeota. Similar numbers of reads in each of the isolates from mangrove or rainforest, mapped to Eukaryota, which showed comparable divisions across the phyla Streptophyta (green plants), Ascomycota (fungi), Chordata (vertebrates), Cnidaria and Arthropoda (insects and arachnids). However, differences were noted, when these samples were compared to isolates taken from ocean sediments. The ocean samples had, on average, larger number of Archaea, with almost all the reads mapping to the phylum Thaumarchaeota. Like the forest samples, the predominant bacterial phylum in the ocean samples mapped to the phylum Proteobacteria.
This finding shows strong patterns in metagenomics structure across the samples taken from the two types of forest ecosystems. It shows distinct patterns that unite the forest samples and differentiates them from an ocean sediment ecosystem.
Methods
Soil samples
Approximately 250 g of soil were collected at 20 cm in depth from the surface in four different locations within Kerala, India and included Pyannur (PYN, Coordinates: 12.1050687, 75.2058), Panangod (PGD, Cord: 9.8959941, 76.326094), Vallarpadam(VL1, Cord: 9.9994138, 76.253705) and Madakal (MAL, Cord: 9.9091896, 76.30629) (Fig. 4B). Physical characteristics of the samples sites are provided (Supplementary Table 4). Following collection, each soil sample was preserved at −20 °C prior to DNA isolation.
Genomic DNA extraction and Sequencing
Genomic DNA was extracted using MoBio Powersoil DNA isolation Kit (MO BIO Laboratories, Inc., California) as per manufacturer’s instructions. Sample quality was verified by gel electrophoresis. Paired end read sequencing was performed on an Illumina HiSeq platform 2500 at SciGenome Labs Pvt Ltd, Cochin (India). The raw fastQ files were uploaded in NCBI SRA database with accession numbers SRR2844600, SRR2844601, SRR2844602 and SRR2844616.
Quality Control and Annotation Pipeline
The paired end fastQ read files of all the samples were uploaded to the Metagenome Rapid Annotation using Subsystem Technology (MG-RAST) server (http://metagenomics.anl.gov/)55 and processed following their standard protocol. Briefly, the mate-pairs were joined with overlap setting of 8 base pairs (bp) and a maximum difference of 10% and were then processed for quality control. Low-quality regions were trimmed off using SolexaQA56, de-replicated, and analyzed the artificially duplicated reads (ADRs)57 using Duplicate Read Inferred Sequencing Error Estimation (DRISEE)58. The near-exact matches against model organisms including fly, mouse, cow and human were removed using Bowtie59. Coding regions in DNA sequences of 75 bp and longer were predicted using FragGeneScan60. Protein clusters with 90% identity were built using the UCLUSTimplementation61 in Quantitative Insights into Microbial Ecology (QIIME). A representative of each cluster is subjected to similarity analysis using BLAST-like alignment tool (BLAT)62. Sequence similarity searches to identify proteins and mapped annotations are computed against a MG-RAST protein databases M5NR19, Genbank63,64, SEED23, Integrated Microbial Genomes & Microbiomes (IMG)65, Universal Protein Resource (UniProt)40, Kyoto Encyclopedia of Genes and Genomes (KEGG)66 and Evolutionary genealogy of genes: Non-supervised Orthologous Groups (eggNOG)67.
Taxonomic and Functional Analysis
Taxonomic assignments were carried out against the RefSeq protein database, Metagenomics Rapid Annotation using Subsystem Technology (MG-RAST)55. This database is an integration of many sequence databases into one single, searchable database. Cut-offs included a maximum E-value of 1 × 10−5, a minimum percentage identity of 60%, and a minimum alignment length of 15 were used. The subsystems platform annotation that assigns genes to functional roles68,69, was used for functional analysis comparisons, with the same cut-off values used for the taxonomic assignments.
Comparison of metagenomes from different soil samples
Different sediment samples isolated from mangroves in Brazil21, a tropical forest in Puerto Rico22, and different sediment samples collected from the South China Sea23 were compared to the Kerala, India soil samples (Table 1), which were publically available at MG-RAST. We selected samples that could be linked to public data. A PCA provided by MG-RAST was used to examine dimensions of maximal variation, and an examination of the taxonomic diversity across all samples was conducted using the Organism tree tool. Taxonomic assignments used M5NR with the same cut-offs used for the Kerala samples, and an average was made across all the samples from a similar location/ecosystem for final comparison. The subsystems platform was used to assigned functional roles, with settings similar to those described above. When functional comparisons were made across metagenomes from geographic and ecological variants, a single isolate that was chosen based on the number of reads to represent each of the geographic groups (India-PGD-4671368.3, Brazil-BrMgv2-4451034.3, Puerto Rico Rain forest- 4446153.3, South China Sea-E201-1-4487294.3).
Availability of data and materials
The raw fastQ files for the Kerala India samples were uploaded in NCBI SRA database with accession numbers SRR2844600, SRR2844601, SRR2844602 and SRR2844616. In addition, the sequences for these reads are publicly available from the MG-RAST server under the following IDs: 4671371, 4671370, 467136 and 4671368.
Change history
04 January 2018
A correction to this article has been published and is linked from the HTML version of this paper. The error has been fixed in the paper.
References
Basak, P., Pramanik, A., Roy, R., Chattopadhyay, D. & Bhattacharyya, M. Cataloguing the bacterial diversity of the Sundarbans mangrove, India in the light of metagenomics. Genomics data 4, 90–92 (2015).
Parks, D. H. & Beiko, R. G. Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26, 715–721 (2010).
Jayadev, A., RajikaLekshman, V. & Parvathy, V. S. Activity Screening of Mangrove Microbes through a Metagenomic Approach. Int. J. Pure App. Biosci 3, 202–207 (2015).
Alongi, D. M., Christoffersen, P. & Tirendi, F. The influence of forest type on microbial-nutrient relationships in tropical mangrove sediments. Journal of Experimental Marine Biology and Ecology 171, 201–223 (1993).
Holguin, G., Vazquez, P. & Bashan, Y. The role of sediment microorganisms in the productivity, conservation, and rehabilitation of mangrove ecosystems: an overview. Biology and fertility of soils 33, 265–278 (2001).
Holguin, G. et al. La microbiologia de losmanglares, bosquesen la frontera entre el mar y la tierra. Ciencia Desarrollo 144, 26–35 (1999).
Kathiresan, K. & Bingham, B. L. Biology of mangroves and mangrove ecosystems. Advances in marine biology 40, 81–251 (2001).
Hou, Lijun et al. Anaerobic ammonium oxidation (anammox) bacterial diversity, abundance, and activity in marsh sediments of the Yangtze Estuary. Journal of Geophysical Research: Biogeosciences 118, 1237–1246 (2013).
Arumugam, M. et al. Isolation of an unusual metabolite 2-allyloxyphenol from a marine actinobacterium, its biological activities and applications. Applied microbiology and biotechnology 86, 109–117 (2010).
Li, X. B. et al. A new staurosporine analog from Actinomycetes Streptomyces sp.(172614). Journal of Asian natural products research 13, 765–769 (2011).
Li, H. et al. Identification and characterization of xiamycin A and oxiamycin gene cluster reveals an oxidative cyclization strategy tailoring indolosesquiterpene biosynthesis. Journal of the American Chemical Society 134, 8996–9005 (2012).
Wang, F., Xu, M., Li, Q., Sattler, I. & Lin, W. p-Aminoacetophenonic acids produced by a mangrove endophyte Streptomyces sp.(strain HK10552). Molecules 15, 2782–2790 (2010).
Hong, K. [Actinomycetes from mangrove and their secondary metabolites]. Wei sheng wuxuebao = ActamicrobiologicaSinica 53, 1131–1141 (2013).
Ding, L., Maier, A., Fiebig, H. H., Lin, W. H. & Hertweck, C. A family of multicyclic indolosesquiterpenes from a bacterial endophyte. Org. Biomol. Chem 9, 4029–4031 (2011).
Ding, L. et al. Hertweck C. Xiamycin, a pentacyclicindolosesquiterpene with selective anti-HIV activity from a bacterial mangrove endophyte. Bioorganic & medicinal chemistry letters 20, 6685–6687 (2010).
Stewart, E. J. Growing unculturable bacteria. Journal of bacteriology 194, 4151–4160 (2012).
Basak, P. et al. Spatiotemporal analysis of bacterial diversity in sediments of Sundarbans using parallel 16S rRNA gene tag sequencing. Microbial ecology 69, 500–511 (2015).
Giri, C. et al. Status and distribution of mangrove forests of the world using earth observation satellite data. Global Ecology and Biogeography 20, 154–159 (2011).
Wilke, A. et al. The M5nr: a novel non-redundant database containing protein sequences and annotations from multiple sources and associated tools. BMC bioinformatics 13, 141 (2012).
Dos Santos, H. F. et al. Mangrove bacterial diversity and the impact of oil contamination revealed by pyrosequencing: bacterial proxies for oil pollution. PLoS One 6(3), e16943 (2011).
Andreote, F. D. et al. The microbiome of Brazilian mangrove sediments as revealed by metagenomics. PLoS One 7, e38600 (2012).
DeAngelis, K. M. et al. Strategies for Enhancing the Effectiveness of Metagenomic-based Enzyme Discovery in Lignocellulolytic Microbial Communities. Bio Energy Research 3, 146–158 (2010).
Chen, B. et al. Metagenomic profiles of antibiotic resistance genes (ARGs) between human impacted estuary and deep ocean sediments. Environmental science &technology 47, 12753–12760 (2013).
Overbeek, R. et al. The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic acids research 33, 5691–5702 (2005).
Mendes, L. W. & Tsai, S. M. Variations of Bacterial Community Structure and Composition in Mangrove Sediment at Different Depths in Southeastern Brazil. Diversity 6(4), 827–843 (2014).
Ghosh, A. et al. Culture independent molecular analysis of bacterial communities in the mangrove sediment of Sundarban, India. Saline systems 6(1), 1 (2010).
Wu, P. et al. Bacterial Communities in the Rhizospheres of Three Mangrove Tree Species from Beilun Estuary China. PloS one 11(10), e0164082 (2016).
Alzubaidy, H. et al. Rhizosphere microbiome metagenomics of gray mangroves (Avicennia marina) in the Red Sea. Gene 576(2), 626–636 (2016).
Lüke, C., Speth, D. R., Kox, M. A., Villanueva, L. & Jetten, M. S. Metagenomic analysis of nitrogen and methane cycling in the Arabian Sea oxygen minimum zone. Peer J 4, e1924 (2016).
Gomes, N. C. M. et al. Diversity of ndo genes in mangrove sediments exposed to different sources of polycyclic aromatic hydrocarbon pollution. Applied and environmental microbiology 73(22), 7392–7399 (2007).
Revathy, T., Jayasri, M. A., &Suthindhiran, K. Biodegradation of PAHs by Burkholderia sp. VITRSB1 Isolated from Marine Sediments. Scientifica, Vol. 2015 Article ID 867586, 9 pages (2015).
Bacosa, H. P., Suto, K. & Inoue, C. Degradation potential and microbial community structure of heavy oil-enriched microbial consortia from mangrove sediments in Okinawa, Japan. Journal of Environmental Science and Health, Part A 48(8), 835–846 (2013).
Kodama, Y. & Watanabe, K. Sulfuricurvumkujiense gen. nov., sp. nov., a facultatively anaerobic, chemolithoautotrophic, sulfur-oxidizing bacterium isolated from an underground crude-oil storage cavity. Int J Syst Evol Microbiol 54, 2297–2300 (2004).
Han, C. et al. Complete genome sequence of the sulfur compounds oxidizing chemolithoautotrophSulfuricurvumkujiense type strain (YK-1(T)). Stand Genomic Sci 6, 94–103 (2012).
Valdes, F. et al. Sulfide-oxidizing bacteria establishment in an innovative microaerobic reactor with an internal silicone membrane for sulfur recovery from wastewater. Biodegradation 27, 119–130 (2016).
Takai, K. et al. Sulfurimonasparalvinellae sp. nov., a novel mesophilic, hydrogen- and sulfur-oxidizing chemolithoautotroph within the EpsilonProteobacteria isolated from a deep-sea hydrothermal vent polychaete nest, reclassification of Thiomicrospiradenitrificans as Sulfurimonasdenitrificans comb. nov.and emended description of the genus Sulfurimonas. Int J Syst Evol Microbiol 56, 1725–1733 (2006).
Inagaki, F., Takai, K., Kobayashi, H., Nealson, K. H. & Horikoshi, K. Sulfurimonasautotrophica gen. nov., sp. nov., a novel sulfur-oxidizing epsilon-proteobacterium isolated from hydrothermal sediments in the Mid-Okinawa Trough. Int J Syst Evol Microbiol 53, 1801–1805 (2003).
Singer, E. et al. Genomic potential of Marinobacteraquaeolei, a biogeochemical “opportunitroph”. Appl Environ Microbiol 77, 2763–2771 (2011).
Fathepure, B. Z. Recent studies in microbial degradation of petroleum hydrocarbons in hypersaline environments. Front Microbiol 5, 173 (2014).
UniProt Consortium. The universal protein resource (UniProt). Nucleic acids research 36, D190–D195 (2006).
Yakimov, M. M., Timmis, K. N. & Golyshin, P. N. Obligate oil-degrading marine bacteria. Curr Opin Biotechnol 18, 257–266 (2007).
Lipp, J. S., Morono, Y., Inagaki, F. & Hinrichs, K. U. Significant contribution of Archaea to extant biomass in marine subsurface sediments. Nature 454(7207), 991–994 (2008).
Alonso‐Sáez, L., Andersson, A., Heinrich, F. & Bertilsson, S. High archaeal diversity in Antarctic circumpolar deep waters. Environmental microbiology reports 3(6), 689–697 (2011).
Valentine, D. L. Adaptations to energy stress dictate the ecology and evolution of the Archaea. Nature Reviews Microbiology 5(4), 316–323 (2007).
Holmes, A. J. et al. Phylogenetic structure of unusual aquatic microbial formations in Nullarbor caves, Australia. Environmental microbiology 3(4), 256–264 (2001).
Bowman, J. P. & McCuaig, R. D. Biodiversity, community structural shifts and biogeography of prokaryotes within Antarctic continental shelf sediment. Applied and Environmental Microbiology 69(5), 2463–2483 (2003).
Basak, P., Pramanik, A., Roy, R., Chattopadhyay, D. & Bhattacharyya, M. Cataloguing the bacterial diversity of the Sundarbans mangrove, India in the light of metagenomics. Genomics data 4, 90–92 (2015).
Lima-Perim, J. E. et al. Linking the composition of bacterial and archaeal communities to characteristics of soil and flora composition in the Atlantic Rainforest. PloS one 11(1), e0146566 (2016).
Tripathi, B. M. et al. Distinctive Tropical Forest Variants Have Unique Soil Microbial Communities, But Not Always Low Microbial Diversity. Frontiers in microbiology, Vol. 7, article 376, https://doi.org/10.3389/fmicb.2016.00376 (2016).
Quaiser, A., Zivanovic, Y., Moreira, D. & López-García, P. Comparative metagenomics of bathypelagic plankton and bottom sediment from the Sea of Marmara. The ISME journal 5(2), 285–304 (2011).
Zaballos, M. et al. Comparison of prokaryotic diversity at offshore oceanic locations reveals a different microbiota in the Mediterranean Sea. FEMS Microbiol Ecol 56, 389–405 (2006).
Hugoni, M. et al. Structure of the rare archaeal biosphere and seasonal dynamics of active ecotypes in surface coastal waters. Proc Natl AcadSci 110, 6004–6009 (2013).
Rasigraf, O., Schmitt, J., Jetten, M. S., & Lüke, C. Metagenomic potential for and diversity of N‐cycle driving microorganisms in the Bothnian Sea sediment. Microbiology Open e475, https://doi.org/10.1002/mbo3.475 (2017).
Shnyukova, E. I. & Zolotareva, E. K. Ecological Role of Exopolysaccharides of Bacillariophyta: A Review. International Journal on Algae 19(1), 56 (2017).
Meyer, F. et al. The metagenomics RAST server–a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC bioinformatics 9, 386 (2008).
Cox, M. P., Peterson, D. A. & Biggs, P. J. SolexaQA: At-a-glance quality assessment of Illumina second-generation sequencing data. BMC bioinformatics 11, 1 (2010).
Gomez-Alvarez, V., Teal, T. K. & Schmidt, T. M. Systematic artifacts in metagenomes from complex microbial communities. The ISME journal 3, 1314–1317 (2009).
Keegan, K. P. et al. platform-independent method for detecting errors in metagenomic sequencing data: DRISEE. PloS Comput Biol 8, e1002541 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome biol 10, R25 (2009).
Rho, M., Tang, H. & Ye, Y. FragGeneScan: predicting genes in short and error-prone reads. Nucleic acids research 38, e191–e191 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome research 12, 656–664 (2002).
Benson, D. A. et al. GenBank. Nucleic acids research 41, D36–42 (2013).
Benson, D. A. et al. GenBank. Nucleic acids research 43, D30–35 (2015).
Markowitz, V. M. et al. The integrated microbial genomes (IMG) system. Nucleic acids research 34, D344–D348 (2006).
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic acids research 36, D480–D484 (2008).
Muller, J. et al. eggNOG v2. 0: extending the evolutionary genealogy of genes with enhanced non-supervised orthologous groups, species and functional annotations. Nucleic acids research 38, D190–D195 (2010).
Overbeek, R. et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic acids research 42, D206–214 (2014).
Alfaro-Espinoza, G. & Ullrich, M. S. Marinobacteriummangrovicola sp. nov., a marine nitrogen-fixing bacterium isolated from mangrove roots of Rhizophora mangle. International journal of systematic and evolutionary microbiology 64(12), 3988–3993 (2014).
Acknowledgements
The authors would like to acknowledge the gracious access to research datasets made publicly available by Andreote, Jimenez et al. 2012; DeAngelis, Gladden et al. 2010; Chen, Yang et al. 2013 and to the MG-RAST team for the free availability of the pipeline and server resources. RK thanks to DST-SERB (Young Scientist Award & EEQ project) and also UGC-BSR startup grants for partial financial assistance and the research facilities supported by Central University of Kerala. DB would like to thank twas- cnpq for the PDF fellowship.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: R.K., D.B., and A.R.W. Analyzed the data: A.R.W., M.V., I.M., R.K., Contributed reagents/materials/analysis tools: R.K. Wrote the paper: A.R.W., R.K., D.B., V.A., and P.G.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Change History: A correction to this article has been published and is linked from the HTML version of this paper. The error has been fixed in the paper.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41598-017-18550-0.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Imchen, M., Kumavath, R., Barh, D. et al. Searching for signatures across microbial communities: Metagenomic analysis of soil samples from mangrove and other ecosystems. Sci Rep 7, 8859 (2017). https://doi.org/10.1038/s41598-017-09254-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09254-6
This article is cited by
-
Comparative metagenomic analysis from Sundarbans ecosystems advances our understanding of microbial communities and their functional roles
Scientific Reports (2024)
-
Bacterial diversity in the aquatic system in India based on metagenome analysis—a critical review
Environmental Science and Pollution Research (2023)
-
Metagenomics and metatranscriptomics as potential driving forces for the exploration of diversity and functions of micro-eukaryotes in soil
3 Biotech (2023)
-
Effect of salinity on genes involved in the stress response in mangrove soils
Antonie van Leeuwenhoek (2023)
-
Metatranscriptomic insight into the possible role of clay microbiome in skin disease management
International Journal of Biometeorology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.