Abstract
The conformational behaviour of three phosphate-bridged dimannosides was studied by means of NMR and computational molecular modelling. First, the conformations of the phosphodiester linker were determined by quantum chemistry methods using dimethyl phosphate as a model. Then, a series of conformations was constructed for each of the studied molecules. Preliminary molecular dynamics (MD) simulations revealed that the inclusion of a cation had a drastic influence on the obtained results. Additionally, triethylammonium had the same effect as sodium as the counter-ion. After that, another series of MD simulations was run. The resulting MD trajectories were used to define the conformations responsible for the observed nuclear Overhauser effects and inter-nuclear coupling.
Similar content being viewed by others
Introduction
Phosphodiester bridges represent an important functional fragment widely encountered in natural polysaccharides including capsular bacterial polysaccharides (examples of producers: Streptococcus pneumonia 1,2,3,4,5,6,7,8,9, Haemophilus influenzae 10,11,12,13,14, Escherichia coli 15,16,17) and others (see review paper18). The polysaccharides bearing phosphodiester bridges (i.e., structurally phosphoglycans) are of ongoing interest as models for immunological investigations, vaccine design, diagnostics development and other purposes. For example, an exocapsular phosphomannan from Pichia holstii 19 was used as a raw material for the synthesis of anti-cancer drug-candidate PI-8819, which was produced by hydrolysis and partial sulfation of the phosphomannan.
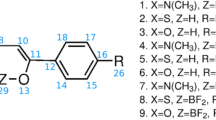
Meanwhile, the conformational properties of oligosaccharide fragments around phosphodiester bridges have not been systematically studied until now. The studies to date are limited to model ab initio calculations of a tetrahydropyranyl phosphate derivative20 and investigation of the molecular dynamics of glucosyl nucleotides21. To fill this gap, we conduct systematic conformational studies of oligo- and polysaccharides bearing phosphodiester bridges. Herein, we report the results of theoretical (density functional theory (DFT) and molecular mechanics calculations) and NMR-based conformational studies of a series of three isomeric glycosyl phosphosaccharide compounds (Fig. 1) in which the α-D-mannosylphosphate unit is connected to the methyl α-D-mannoside section at O-3 (1), O-4 (2), and O-6 (3). Fragments related to compound 3 have been discovered in the chains of cell walls and extracellular phosphomannas of yeast22,23,24,25,26,27,28,29,30. Phosphodiesters 1 and 2 have been prepared as model compounds, since those types of intersaccharidic linkage [(1 → 3) and (1 → 4)] are common in natural bacterial polysaccharides18, but between different type of monosaccharides (not for α-D-Manp). The synthesis of compounds 1–3 was described previously31, 32. Their 1H and 13C NMR data are given in Tables 1 and 2. The assignment of the chemical shifts was performed by the use of 2D COSY, HSQC and HMBC techniques.
Results and Discussion
Conformational analysis of the dimethyl phosphate as a model of the phosphodiester bridge
Prior to performing conformational analysis of disaccharides 1-3, the general behaviour of the phosphodiester bridge was investigated by using the example of dimethyl phosphate (4) as a simplified model. The conformational state of compound 4 is determined by two C-O-P-O torsional angles, which can have either gauche or trans orientation (Fig. 2), and gauche conformers can be either gauche+ (g+) or gauche− (g−). All possible combinations of these rotamers were constructed as starting conformations. Of course, in the case of such a simple compound, some of them form “pseudo-enantiomeric” pairs, but keeping in mind that these fragments would be used further as linkers for asymmetric fragments such as mannoside residues, each of the “pseudo-enantiomers” should have been optimized. These starting structures were subjected to geometry optimization by DFT using the B3LYP functional and 6–311 G(2d,2p) basis set. Only one conformation, namely, g + g-, could be excluded from consideration since it transformed to one of the trans-conformers during the optimization. This result is probably explained by the significant repulsion between two oxygen atoms in such a conformation. The five remaining conformers of the phosphodiester bridge model 4 are shown in Fig. 2.
Building and modelling the phosphodiester-linked dimannosides 1–3
Using the five conformations of the model phosphodiester 4 described above, several conformations for each of the studied phosphodisaccharides 1-3 were built. The methylated oxygen atoms of the linker’s conformers, after deletion of the methyl groups, were used to connect reducing and non-reducing sugar residues. This gave seven possible conformations: Man-(g + g +)-Man, Man-(g−g−)-Man, Man-(g + t)-Man, Man-(tg +)-Man, Man-(g − t)-Man, Man-(tg−)-Man and Man-(tt)-Man. Additionally, each bond involving atoms forming a glycosidic linkage could have its own conformational preferences. For example, when a non-reducing, or glycosylating, mannose residue was concerned, the torsional angle O5′-C1′-O1′-P was expected to exist in a gauche rather than a trans orientation due to the exo-anomeric effect33. Hence, during the construction of the starting phosphodisaccharide models, the value of this torsion was set to +60°.
As for the residue at the reducing end, or glycosyl phosphorylated residue (i.e., Man-OMe), they are known to have greater rotational freedom. The latter can be described, for example, by the P-Ox-Cx-Hx (x = 3, 4 or 6) torsion angle, which can lie in the region of ±60° or 180°. All three orientations were used, yielding 21 starting conformations for each phosphodisaccharide. These were subjected to molecular dynamics (MD) simulations using an MM3 force field and the Solvent Accessible Surface Area (SASA34) approach for water consideration.
In our initial experiments, no counter-ions were included in the simulation. However, analysis of the produced MD trajectories showed that soon after the start of the simulations, all O5′-C1′-O1′-P torsions determining the conformation of the glycosyl phosphate residues changed their orientation to trans (sample illustrations of MD trajectories are given in Fig. 3). Since that contradicted the expectations arising from the well-known exo-anomeric effect33, 35, a literature search was performed. Tvaroska et al.20 observed a similar effect during ab initio modelling of tetrahydropyranyl methyl phosphate. In particular, when the calculations were performed in the absence of a cation, the conformation with the lower energy was also that having O5′-C1′-O1′-P torsion in the trans orientation (reverse exo-anomeric effect36). When a sodium atom was included, the expected situation (result of the exo-anomeric effect) was restored with the O5′-C1′-O1′-P torsion having a gauche orientation. Tvaroska20 speculated that this was probably due to the ability of Na+ not only to shield the negative charge on the phosphate bridge but also to coordinate with the intra-ring oxygen O5′.
However, (1 → 3)- and (1 → 4)-linked compounds 1 and 2 were primarily investigated in their triethylammonium forms. It was unclear using general considerations to deduce whether the NHEt3 + cation could have the same influence on the conformation of the O5′-C1′-O1′-P torsion as the Na+ cation. The situation had additional complications such as hydroxyl groups from the other mannoside residue, which could also interact in some fashion with the NHEt3 + or Na+ moieties. Hence, another series of DFT calculations was performed for (1 → 3)- (1) and (1 → 4)-linked (2) compounds with the NHEt3 + cation and for the (1 → 6)-linked disaccharide 3 with Na+ as the counter-ion. This time, two different starting conformations were used for O5′-C1′-O1′-P torsion of the glycosylating mannose, either gauche or trans. The conformations of the rotatable bonds in the phosphate bridge were the same as described above.
In all of the performed DFT calculations, it was found that the NHEt3 + cation indeed produced the same conformational effect as Na+, i.e., in all cases, structures with the gauche orientation of the O5′-C1′-O1′-P torsion in accordance with the exo-anomeric effect were found to have lower energies. Thus, the quantum mechanical calculation results led to a conclusion that shielding the negative charge on the phosphate oxygens played a greater role than any plausible additional interactions with O5′ or other fragments of the studied molecules.
With these results obtained, MD simulations were performed again on all the conformers with inclusion of the counter-ions. Additionally, restraints on O5′-C1′-O1′-P torsion stemming from DFT calculations were applied to hold it in the gauche conformation determined by the exo-anomeric effect. The resulting ensembles of the conformers were analysed to measure their average inter-atomic distances and torsional angles of interest to compute NOE and coupling constants.
Nuclear Overhauser effect (NOE) analysis
To estimate the theoretical NOE values, corresponding inter-unit proton-proton distances were used. They were measured in each of the snapshot conformations and then averaged. Although averaged values of r−6 are required for the quantitative calculations of NOE, some assumptions about the presence or absence of the effect for certain proton pairs can be based on the fact that the NOE is measurable only in case when the inter-proton distance is less than 4.5 Å.
Experimental data were obtained by means of 1D NOESY. During the analysis of the MD snapshots, it was found that while the glycosylating section was initially restrained from rotation around the C1′-O1′ bond, the glycosyl phosphorylated section (i.e., Man-OMe) demonstrated a high degree of conformational flexibility. However, in the case of the (1 → 3)-linked structure 1 and the (1 → 4)-linked structure 2, this flexibility led to the possibility of the studied molecules adopting conformations in which the respective inter-ring H1′-H3 and H1′-H4 distances became lower than 4 Å and their averaged values were in the range of 3–3.5 Å, signifying the possibility of NOE (Table 3, Fig. 4). In case of (1 → 4)-linked compound 2, the distances H1′-H6a and H1′-H6b were also small enough to suggest NOE. In addition, in this case some NOE was indeed observed on H6 protons (Fig. 5b). Similarly, NOE was observed on the H2 proton of (1 → 3)-linked compound 1 (Fig. 5a and Table 3). This explained the appearance of inter-ring NOEs in the case of compounds 1 and 2 (Figs 4 and 5).
Quantitative analysis of the observed NOE values was attempted. The H1′-H2′ distance that was measured to be approximately 2.6 Å was used as a reference. Considering that relative NOEs are proportional to the reciprocal sixth power of inter-proton distances, this suggested that in compounds 1 and 2, the H1′-H2′ NOE should have values 3 to 6 times larger than those of the inter-unit effects. From Fig. 5a,b it can be seen that these proportions are all 5x. This means that no exact quantitative correlation could be established in this case.
Unlike compounds 1 and 2, for (1 → 6)-linked molecule 3, the presence of an extra C6-C5 bond near the phosphodiester bridge leads to a spatial separation of the two pyranoside residues that could not be overcome by the enhanced conformational flexibility of the P-O6-C6-C5 fragment. The averaged distances in compound 3 between H1′ and the protons H6 of the pyranoside section in the Man-OMe unit exceeded 4 Å (Table 3). In fact, no reliable NOE effects were observed experimentally in this instance (Fig. 5c).
Additionally, the temperature dependence of the NOE effects was studied for compounds 1 and 2 (as their ammonium salts) to gain further insight into their conformational equilibrium. 1D NOESY experiments were performed for these substances at temperatures of 263, 283, 303 and 323 K in methanol-d4 solutions. At the lowest temperature (263 K), no NOE were observed, most likely because of the total decrease in molecular motion. 1D ROESY spectroscopy was attempted in this case, but since the quantitative integration of ROE signals may give significant errors, these values are not shown; the corresponding spectra are presented in Supplementary Information.
At higher temperatures, different tendencies were found for the investigated structures (Table 4). For (1 → 4)-linked compound 2, no detectable changes were observed in H1′-H4 NOE with increasing temperature, while the strength of the H1′-H6a,b signal increased. Some slight changes occurred in (1 → 3)-linked molecule 1: a relative decrease in H1′-H2 NOE along with a simultaneous slight increase in H1′-H3. This confirms, in our opinion, that the conformational distribution in the studied structures has a rather complex nature due to the relative flexibility of the phosphate linkage.
Analysis of three-bond 13C-31P and H-31P coupling constants (3 J C,P and 3 J H,P)
3 J C,P as well as other three-bond coupling constants can be calculated empirically using a specially parametrized Karplus-type equation. Equation (1), given in work37, was used in the present study:
where φ defines the corresponding torsional angle C-C-O-P. These torsions and subsequent coupling constants were calculated for each of the snapshot conformations to evaluate theoretical 3 J C,P values. The averaged values of these constants were than compared to the experimental ones that are readily measured from 1D 13C NMR spectra (Table 5).
A generally good coincidence can be observed between the theoretical and the calculated data. However, the deviation sometimes reaches 1 Hz. This can be explained by the already mentioned fact that the studied molecules demonstrate high conformational flexibility, especially at the glycosyl phosphorylated end, which leads to a greater error in the computed constants.
Since no reliable NOE were observed for (1 → 6)-linked phosphodimannoside 3, an additional investigation that consisted of the measurement of 1H-31P three-bond couplings was performed. For their theoretical calculation, another Karplus-type equation (2) was employed, taken from work38:
Table 6 shows the experimentally measured 3JH,P constants in comparison with those calculated as averages of the ensemble of conformers as described above for the 3JC,P. Unlike the P-C constants that were generally underestimated in calculations, P-H couplings tended to be largely (more than 1 Hz) overestimated. We believe this can result from the used Karplus-type equation (2) having been originally derived for nucleotides such that it may give larger errors in the case of phosphodimannosides, although the general trend is reproduced.
Conclusions
The conformations of numerous di- and trisaccharides (i.e., conformations around various glycoside bonds) have been thoroughly examined over the last 30 years. In contrast, the conformational properties of phosphodiester-linked disaccharides have not been studied to date. To fill this gap, we conducted the first conformational study of oligosaccharides bearing phosphodiester bridges.
The conformational behaviour of a series of phosphodiester-bridged dimannosides was studied by means of NMR spectroscopy, DFT calculations and MD simulations using a continuum solvation model and the MM3 force field. When the presence of a counter-ion was neglected, the reverse exo-anomeric effect occurred at the glycosylating part of the molecules, most likely due to the repulsion between the negative charges on the phosphate group and the saccharide oxygen atoms. When either sodium or triethylammonium cations were added, the standard conformation determined by the exo-anomeric effect was restored. In general, the studied molecules demonstrated a high degree of conformational flexibility, which in the case of (1 → 3)- and (1 → 4)-linked structures led to the spatial proximity of H1′ and Hx atoms resembling (in some extent) that one in common disaccharides linked by a glycosidic bond.
Methods
The NMR spectra were recorded on a Bruker Avance 600 spectrometer for solutions in CD3OD at 30 °C or other temperatures when specified. TSP (δH 0.0 and δC −1.6 ppm) was used as the internal standard for 1H and 13C spectra, and 85% H3PO4 (δP 0.0) was used as an external standard for 31P spectra. The 2D NMR spectra were recorded and treated according to the standard Bruker software. A mixing time of 60 ms was used in TOCSY experiments. 1D ROESY spectra were registered with cw spinlock for mixing using refocusing with a shaped pulse. The spinlock time was 0.2 s. 1D NOESY spectra were registered using selective refocusing with a shaped pulse. The spinlock time was 0.7 s.
DFT calculations were performed using DALTON201539 software. Optimizations were performed in vacuo until the gradient reached values less than 10−4. Molecular mechanics calculations were performed using the TINKER 5.040 software package with the MM3 force field. The Solvent Accessible Surface Area (SASA) continuum solvation model was used in all cases. Each MD trajectory length was 50 ns with the snapshot conformations being written each 2 ps, thus producing 25000 structures for each simulation. Geometrical constraints were applied using the standard algorithm provided by the software using force constants of 5.
1D ROESY and 1D NOESY spectra recorded at different temperatures are available in the Supplementary file. Other experimental data used in this study are available from the authors upon request.
References
Jannson, P.-E., Lindberg, J., Wimalasiri, K. M. S. & Henrichsen, J. The structure of the capsular polysaccharide from Streptococcus pneumoniae type 7B. Carbohydr. Res. 217, 171–180 (1991).
Jennings, H. J., Rosell, K.-G. & Carlo, D. J. Structural determination of the capsular polysaccharide of Streptococcus pneumoniae type-19 (19F). Can. J. Chem. 58, 1069–1074 (1980).
Lee, C.-J. & Fraser, B. A. The structures of the cross-reactive types 19 (19F) and 57 (19A) pneumococcal capsular polysaccharides. J. Biol. Chem. 255, 6847–6853 (1980).
Ohno, N., Yadomae, T. & Miyazaki, T. The structure of the type-specific polysaccharide of Pneumococcus type XIX. Carbohydr. Res. 80, 297–304 (1980).
Katzenellenbogen, E. & Jennings, H. J. Structural determination of the capsular polysaccharide of Streptococcus pneumoniae type 19A (57). Carbohydr. Res. 124, 235–245 (1983).
Beynon, L. M., Richards, J. C., Perry, M. B. & Kniskern, P. J. Antigenic and structural relationships within group 19 Streptococcus pneumoniae: chemical characterization of the specific capsular polysaccharides of types 19B and 19C. Can. J. Chem. 70, 218–232 (1992).
Richards, J. C., Perry, M. B. & Carlo, D. J. The specific capsular polysaccharide of Streptococcus pneumoniae type 20. Can. J. Biochem. Cell Biol. 61, 178–190 (1983).
Beynon, L. M., Richards, J. C. & Perry, M. B. Identification of the common antigenic determinant shared by Streptococcus pneumoniae serotypes 35A and 20 capsular polysaccharides — structural analysis of the Streptococcus pneumoniae serotype 35A capsular polysaccharide. Eur. J. Biochem. 250, 163–167 (1997).
Karlsson, C., Jannson, P.-E. & Sorensen, U. B. The chemical structures of the capsular polysaccharides from Streptococcus pneumoniae types 32F and 32A. Eur. J. Biochem. 255, 296–302 (1998).
Branefors-Helander, P., Classon, B., Kenne, L. & Lindberg, B. Structural studies of the capsular antigen of Haemophilus infuenzae type C. Carbohydr. Res. 76, 197–202 (1979).
Branefors-Helander, P., Kenne, L. & Lindqvist, B. Structural studies of the capsular antigen from Haemophilus influenzae type F. Carbohydr. Res. 79, 308–312 (1980).
Egan, W., Fai-Po, T., Climenson, P. A. & Schneerson, R. Structural and immunological studies of the Haemophilus influenzae type C capsular polysaccharide. Carbohydr. Res. 80, 305–316 (1980).
Egan, W., Fai-Po, T. & Schneerson, R. Structural studies of the Haemophilus influenzae type F capsular polysaccharide. Carbohydr. Res. 79, 271–277 (1980).
Egan, W., Schneerson, R., Werner, K. E. & Zon, G. Structural studies and chemistry of bacterial capsular polysaccharides. Investigations of phosphodiester-linked capsular polysaccharides isolated from Haemophilus influenzae types A, B, C, and F: NMR spectroscopic identification and chemical modification of end groups and the nature of base-catalyzed hydrolytic depolymerization. J. Am. Chem. Soc. 104, 2898–2910 (1982).
Rodriguez, M.-L., Jann, B. & Jann, K. Structure and serological properties of the capsular K11 antigen of Escherichia coli O13:K11:H11. Carbohydr. Res. 196, 101–109 (1990).
Jann, B., Dengler, T. & Jann, K. The capsular (K51) antigen of Escherichia coli 01:K51:H−, an O-acetylated poly-N-acetylglucosamine phosphate. FEMS Microbiol. Lett. 29, 257–261 (1985).
Hofmann, P., Jann, B. & Jann, K. Structure of the fructose‐containing K52 capsular polysaccharide of uropathogenic Escherichia coli O4:K52:H−. Eur. J. Biochem. 147, 601–609 (1985).
Nikolaev, A. V., Botvinko, I. V. & Ross, A. J. Natural phosphoglycans containing glycosyl phosphate units: structural diversity and chemical synthesis. Carbohydr. Res. 342, 297–344 (2007).
Ferro, V., Fewings, K., Palermo, M. C. & Li, C. Large-scale preparation of the oligosaccharide phosphate fraction of Pichia holstii NRRL Y-2448 phosphomannan for use in the manufacture of PI-88. Carbohydr. Res. 332, 183–189 (2001).
Tvaroška, I., André, I. & Carver, J. P. Ab initio molecular orbital study of the conformational behavior of the sugar-phosphate linkage. Toward an understanding of the catalytic mechanism of glycosyltransferases. J. Phys. Chem. B. 103, 2560–2569 (1999).
Petrová, P., Koča, J. & Imberty, A. Effect of cation concentration on molecular dynamics simulations of UDP-glucose. Mol. Simulat. 24, 325–340 (2000).
Gorin, P. A. J. & Mazurek, M. Structure of a phosphonomannan, as determined by the effect of lanthanide ions on its carbon-13 magnetic resonance spectrum. Can. J. Chem. 52, 3070–3076 (1974).
Bretthauer, R. K., Kaczorowski, B. J. & Weise, M. J. Characterization of a phosphorylated pentasaccharide isolated from Hansenula holstii NRRL Y-2448 phosphomannan. Biochemistry 12, 1251–1256 (1973).
Parolis, L. A. S., Duus, J. Ø., Parolis, H., Meldal, M. & Bock, K. The extracellular polysaccharide of Pichia (Hansenula) holstii NRRL Y-2448: The structure of the phosphomannan backbone. Carbohydr. Res. 293, 101–117 (1996).
Parolis, L. A. S., Parolis, H., Kenne, L., Meldal, M. & Bock, K. The extracellular polysaccharide of Pichia (Hansenula) holstii NRRL Y-2448: The phosphorylated side chains. Carbohydr. Res. 309, 77–87 (1998).
Thieme, T. R. & Ballou, C. E. Nature of the phosphodiester linkage of the phosphomannan from the yeast Kloeckera brevis. Biochemistry 10, 4121–4129 (1971).
Raschke, W. C. & Ballou, C. E. Immunochemistry of the phosphomannan of the yeast Koeckera brevis. Biochemistry 10, 4130 (1971).
Cawley, T. N. & Ballou, C. E. Identification of two Saccharomyces cerevisiae cell wall mannan chemotypes. J. Bacteriol. 111, 690 (1972).
Antalis, C., Fogel, S. & Ballou, C. E. Genetic control of yeast mannan structure: mapping the first gene concerned with mannan biosynthesis. J. Biol. Chem. 248, 4655–4659 (1973).
Kardošová, A., Daniš, J. & Alföldi, J. Extracellular phosphogalactomannan of candida-mucifera CCY-29-170-1 strain. Chem. Papers 48, 189–194 (1994).
Nikolaev, A. V., Ivanova, I. A., Shibaev, V. N. & Kochetkov, N. K. Application of the hydrogenphosphonate approach in the synthesis of glycosyl phosphosugars linked through secondary hydroxyl groups. Carbohydr. Res. 204, 65–78 (1990).
Nikolaev, A. V., Ryabtseva, E. V., Shibaev, V. N. & Kochetkov, N. K. Fragments of biopolymers containing glycosyl phosphate residues: use of hydrogenphosphonate approach for synthesis of glycosyl phosphosugars, immunodominant fragments of yeast phosphoglycans. Sov. J. Bioorg. Chem. 15, 897 (1989).
Lemieux, R. U. & Koto, S. The conformational properties of glycosidic linkages. Tetrahedron 30, 1933–1944 (1974).
Ooi, T., Oobatake, M., Némethy, G. & Scheraga, H. A. Accessible surface areas as a measure of the thermodynamic parameters of hydration of peptides. Proc. Natl. Acad. Sci. USA 84, 3086–3090 (1987).
Alonso, E. R., Peña, I., Cabezas, C. & Alonso, J. L. Structural expression of exo-anomeric effect. J. Phys. Chem. Lett. 7, 845–850 (2016).
Randell, K. D., Johnston, B. D., Green, D. F. & Mario-Pinto, B. Is there a generalized reverse anomeric effect? Substituent and solvent effects on the configurational equilibria of neutral and protonated-arylglucopyranosylamines and N-aryl-5-thioglucopyranosylamines. J. Org. Chem. 65, 220–226 (2000).
Coxon, B. Developments in the Karplus equation as they relate to the NMR coupling constants of carbohydrates. Adv. Carbohydr. Chem. Biochem. 62, 17–82 (2009).
Lankhorst, P. P., Haasnoot, C. A. G., Erkelens, C. & Altona, C. Nucleic-acid constituents C-13 NMR in conformational-analysis of nucleic-acid fragments. A reparametrization of the Karplus equation for vicinal NMR coupling-constants in CCOP and HCOP fragments. J. Biomol. Struct. Dyn. 1, 1387–1405 (1984).
Aidas, K. et al. The Dalton quantum chemistry program system. WIREs Comput. Mol. Sci. 4, 269–284 (2014).
J P Lab TINKER: Software Tools for Molecular Design. Department of Chemistry, Washington University, Saint Louis, Missouri, USA, http://dasher.wustl.edu/tinker (2009).
Acknowledgements
This investigation was supported by the Russian Scientific Foundation (grant 14-50-00126).
Author information
Authors and Affiliations
Contributions
N.E.N. conceived the work. A.V.N. and D.V.Y. performed synthetic operations. A.G.G. conducted computer simulations. A.S.S. and A.S.D. performed NMR experiments. All the authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gerbst, A.G., Nikolaev, A.V., Yashunsky, D.V. et al. Theoretical and NMR-based Conformational Analysis of Phosphodiester-linked Disaccharides. Sci Rep 7, 8934 (2017). https://doi.org/10.1038/s41598-017-09055-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09055-x
This article is cited by
-
Theoretical DFT interpretation of infrared spectra of biologically active arabinogalactan sulphated derivatives
Chemical Papers (2020)
-
Rapid Solution-Phase Hydrogen/Deuterium Exchange for Metabolite Compound Identification
Journal of the American Society for Mass Spectrometry (2019)
-
Classical and interdisciplinary approaches to the design of organic and hybrid molecular systems
Russian Chemical Bulletin (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.