Abstract
Liver X receptorβ (LXRβ) is a promising therapeutic target for lipid disorders, atherosclerosis, chronic inflammation, autoimmunity, cancer and neurodegenerative diseases. Druggable LXRβ agonists have been explored over the past decades. However, the pocket of LXRβ ligand-binding domain (LBD) is too large to predict LXRβ agonists with novel scaffolds based on either receptor or agonist structures. In this paper, we report a de novo algorithm which drives privileged LXRβ agonist fragments by starting with individual chemical bonds (de novo) from every molecule in a LXRβ agonist library, growing the bonds into substructures based on the agonist structures with isomorphic and homomorphic restrictions, and electing the privileged fragments from the substructures with a popularity threshold and background chemical and biological knowledge. Using these privileged fragments as queries, we were able to figure out the rules to reconstruct LXRβ agonist molecules from the fragments. The privileged fragments were validated by building regularized logistic regression (RLR) and supporting vector machine (SVM) models as descriptors to predict a LXRβ agonist activities.
Similar content being viewed by others
Introduction
Liver X receptorβ (LXRβ, also known as NR1H2) is a nuclear receptor, which is considered as the core of modern pharmacology, and the promising therapeutic target for lipid disorders, atherosclerosis, chronic inflammation, autoimmunity, cancer and neurodegenerative diseases1, 2. But, LXRβ ligand-binding domains (LBDs) have a big binding pocket, which tolerates diverse sizes and shapes of ligands. This makes difficult to predict LXRβ ligand structures with novel scaffolds based upon known receptor or ligand structures3. Thousands of natural or synthetic LXR agonists have been reported. Conventional approaches were also tried to predict LXR agonists4,5,6,7. In current studies, we were motivated to figure out privileged LXRβ agonist fragments from the known LXRβ agonists to guide fragment-based8 LXRβ agonist design and discovery.
There are many ways to define or derive structural fragments (substructures) from a chemical structure library, such as, maximal common substructure (MCSS) algorithm9, fingerprint algorithms10, scaffold-based classification approach (SCA)11, atom center fragments12, 13, etc. These approaches were based upon empirically or algorithmically pre-defined rules14, 15 and, the resulting substructures could be subjective. To build predictive SAR models, we need substructures that are statistically representative in a chemical structure library and related to the concerned activity.
Over the last decade, subgraph mining algorithms were developed and applied in QSAR modeling. Dehaspe and colleagues16 used a subgraph discovery algorithm to predict the toxicity of a compound based upon its chemical structure. Yan and Han developed the gSpan program for subgraph mining17. Huan and colleagues addressed the isomorphism problem in the subgraph mining process18. Kuramochi and coworkers also developed a subgraph discovery program19. Borgelt and colleagues developed MoSS for subgraph mining20, 21. Meinl and co-workers developed the ParMol package for subgraph mining22. Wang and colleagues paralleled a subgraph mining algorithm with the CUDA technology23. Most recently, Khashan and co-workers used the subgraph mining approach in QSAR Modeling to predict compound toxicity24. Shao and colleagues used a subgraph mining technology to identify common functional groups to predict drug adverse effects25.
These algorithms were tested on smaller data sets ranging from 10 K to 100 K compounds, some of them were tested on artificially generated data17, 19, 26, 27. Nowadays, chemical structure data (such as ZINC, one of the largest databases for medicinal chemistry, contains approximately 21 million compounds) grow rapidly28. Our studies revealed that a conventional subgraph (substructure) mining algorithm would encounter a huge computational challenge when it was tested on a million-compounds database due demanding huge memory for the isomorphism checking (more than 128 GB). Those subgraph mining algorithms elected substructures based upon a minimal support threshold, which was determined by trial-and-errors. Raising the threshold would be at the risk of losing substructures, which were related to the activity. Lowering the threshold would be at the risk of introducing too many trivial substructures, which reduced the prediction accuracy for lowering signal-to-noise ratio. Consequently, the classification accuracies were around 70%24. Moreover, most of the previous subgraph mining approaches did not interpret the subgraph chemistry, which should be of interest to chemists. Khashan and colleagues did study the relationship of their substructures and toxiphores. But, these fragments were derived without considering the chemical integrity (such as, an aromatic ring was broken in the middle of the ring).
In order to solve these problems, we propose a new de novo substructure generation algorithm (DSGA), which discovers substructures from a chemical structure library with improved substructure mining strategies:
-
(1)
To avoid generating too many trivial substructures and reducing the memory requirements for the isomorphism checking, we coded growing subgraphs with linear notations (subIDs, see Fig. 1). The advantage of the subID linear notation is that the isomorphism checking can be done by a substring search instead of a subgraph search, which demands memory and computing resource.
-
(2)
When substructures were generated with a depth-first search strategy, the computing complexity could grow exponentially. Therefore, we developed a strategy to prune the depth-first search tree to converge the results. The algorithm only grows the nodes with the maximal substructure on the search tree, other branches in the tree will be pruned. To examine if a substructure is a maximal substructure, the GMA algorithm9 was employed to exclude isomorphic or homomorphic substructures.
-
(3)
The further integrity checking was applied to ensure the chemical relevant of the maximal substructures.
Conventionally, the library under investigation is used to find substructures as a structural descriptor vector for a molecule in a compound library. However, electing substructures only based on the minimal support threshold may include too many trivial substructures or missed the under-supported substructures that are still strongly related to the activity. This problem can be resolved by taking chemical and biological background knowledge (chemical functional groups or synthetic feasibility, and biological activities) into account. We ran the de novo algorithm on the ZINC database (the commonly recognized database in medicinal chemistry) to gain maximal substructures as the background knowledge to produce substructures as privileged fragments for LXRβ ligands. The knowledge is in the form of a standard substructure dictionary (SSD). With the SSD, a substructure can still be elected as a privileged fragment even if its population is below the “minimal support threshold”; a substructure can still be excluded even if its population is much higher than the “minimal support threshold”. The gold criterion is the relation between the substructure and the concerned property.
With DSGA, a compound library can be converted into a set of substructure descriptor vectors or an m x n matrix (m is the number of maximal frequent substructures, and n is the number of compounds in the library). If the matrix is associated with activities, a regularized logistic regression (RLR) model29 or other machine learning models can be constructed to predict the activity for a new compound based on its chemical structure.
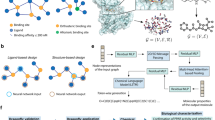
Comparing with previous studies, we emphasize more on gaining new chemical insights from the substructure mining algorithm. In fragment-based drug discovery (FBDD)30, key questions to be answered are what are the fragments for a drug lead, and what are the rules to combine these fragments. In this work, we present an example on how one can answer these questions by applying a de novo substructure generation algorithm (Fig. 1). This work can be applied for analyzing privileged fragments for the ligands against other biological targets.
Results and Discussion
De novo substructure generation process with a pruning strategy
The pruning strategy significantly reduced the number of substructures discovered from the three testing libraries(LXRβ, PPARα, and VR libraries). The ratios of total-substructures/pruned-substructures are 113, 105, and 114 for the LXRβ library (634 compounds), PPARα library (606 compounds), and VR library (619 compounds), respectively. This means the pruning strategy improves the performance more than one hundred times. The pruning strategy is particularly important when de novo substructure generation algorithm (DSGA) is used in a big compound library (such as a library with more than 100 K compounds). In our studies, the LXRβ library has only 161 frequent substructures, a program without pruning strategy has to check 18,170 substructures; for the VR library (83 frequent substructures) checking 9,439 substructures; and for the PPARα library (114 frequent substructures) checking 12,021 substructures. This costs not only computing time, but exhausts so much memory that an algorithm cannot continue the calculation due to no enough memory.
With the pruning strategy, we, for the first time, are able to generate substructures from the ZINC database31, which has approximately 9.1 million drug-like compounds. The algorithm discovered 51,770 substructures from the ZINC database. The number of substructures increased exponentially before the first 10 K structures of the ZINC database were scanned, and the number significantly slowed down because most of the maximal substructures had been discovered. This suggested that the structural diversity of substructures is limited in the currently explored chemical space (Figure S1).
By using the frequent substructuresthat were generated from the ZINC library as descriptors, we were able to discriminate three focused compound libraries associated with three different biological targets (LXRβ, PPARα, and VR) with principal component analyses (PCA) as depicted in Fig. 2.
Substructures used for predicting activities with the RLR approach
One way to examine the quality of DSGA is to study the relations between the substructures and bioactivities. 51,170 substructures were derived from the ZINC database (9,107,119 compounds), and used as the SSD for building RLR classification models. Again, three compound libraries for LXRβ, PPARα, and VR, were studied for SSD-based RLR classification modeling to predict the activities against LXRβ, PPARα, and VR. Figure 3 demonstrates the prediction capacities of the SVM models.
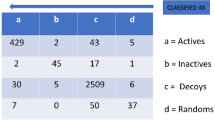
The RLR classification model performances are summarized in Table 1. The ratio of splits between train and validation data is 2:1.
These results demonstrated that the substructures discovered by DSGA are objective structural descriptors for RLR classifications.
Substructures used for predicting activities with SVM regression
Regression modeling requires reducing the number of descriptors in order to avoid high computational costs. A subset of the SSD was derived based upon the population tuning points for a specific compound library. In Fig. 5, the X-axis stands for the substructure, the Y-axis stands for the frequency of the corresponding substructure in a compound library. This plot demonstrates the distributions of the substructures in the VR, LXRβ, and PPARα libraries. The curves begin to flatten at the frequency of 40, where the LXRβ and PPARα libraries can adopt 3,000 substructures, and the VR library can adopt 4,375 substructures as their structural descriptors.
The SVM regression models were built for the LXRβ, PPARα, and VR libraries with 277, 484, and 495 training compounds. The predictive models were validated with 5-fold validation processes. The performances were measured using the average mean square errors (MSE) and Pearson Correlations as listed in Table 2.
These results conclude that the substructures are highly related to the bioactivities. The prediction accuracies of regression models were not very high due to the paradox of predictivity versus diversity (that is, the greater the chemical diversity of the investigated compounds, the smaller the chance that SAR models exist and can be uncovered)15. The limit of this approach is that it is difficult for a common structure fragment descriptor to distinguish tiny structural differences among molecules. However, the advantage of this approach is that privileged structural fragments can be derived from these models.
Identifying privileged fragments for privileged scaffold exploration
The substructures used in SVM models were scored with a privileged fragment index (PFI) as the following,
where, f i is the population of the ith substructure appearing in a given compound library, T is the total number of compounds in the library, and a i is the number of active compounds in the library. All the substructures used in the SVM models for the LXRβ, PPARα, and VR libraries were sorted in descending order of the PFIs (Supplementary Materials).Privileged fragments for the LXRβ ligands found by DSGA are listed in Table 3.
Rules to construct LXRβ agonists from the privileged fragment
Rule 1
Fragments D and F are linkers connecting A and B. There were 36 LXRβ ligands made through this rule. There were 23 such ligands linked through the MF substructure D, other 13 ligands were linked through the MF substructure F (details can be found in the supplementary material SM Table 1). The linker MF substructure D can produce more active ligands. The schema of Rule 1 is depicted in Fig. 4.
Rule 2
The MF substructures A and C connect through direct covalent binding to make LXRβ ligands. The Fragment C is modified to allow any heavy atom at the position of the nitrogen atom. Thus, we got 31 LXRβ agonists based upon this rule. Fragment C has two classes of bioisosteres (the hetero atom linker can be nitrogen or oxygen), which do not significantly change the binding affinity. It seams that Fragment A cannot be simplified, and it is critical to maintain an acidic polar group at the terminal of Fragment C (details can be found in supplementary material SM Table 2). The schema of Rule 2 is demonstrated in Fig. 5.
Rule 3
Fragments B and C can be linked to form an LXRβ agonist. This combination can also be viewed as Fragment F merges with Fragments B and C. Although, only 5 LXRβ agonists were discovered, there are many opportunities to explore (Fig. 6).
Rule 4
Typical LXRβ agonist constructing cases are demonstrated in Figs 7 and 8. By inspecting the data set, we recognize that A and D have bioisosteres. Therefore, we define Fragments A′ and D′ as shown in Fig. 9. LXRβ agonists can be created by merging Fragments A’, B, and D’. Fragment A’ connects to Fragment D’, and Fragment D’ merges with B at the aromatic rings. This results in 66 LXRβ ligands with EC50 values ranging between 0.011 and 3.40 μM (details can be found in supplementary material SM Table 3 (Rule 4)). In this case, Fragment D’ is a linker to connect Fragments A’ and B.
Rule 5
Fragment C itself can be an LXRβ agonist scaffold. It can also be merged with Fragment F. The typical agonists are listed in supplementary material (details can be found in supplementary material SM Table 4 (Rule 5)). Typical ligands and their activities are depicted in Fig. 9.
Rule 6
Fragment I itself can form a star-shaped scaffold with a pentagon for an LXRβ agonist. It may merge with Fragments F and B, or C. 66 LXRβ agonists were constructed with this rule as shown in Fig. 10 (SM Table 5: Rule 6).
Rule 7
Fragments J and D’ can form a new scaffold by a methylene linker as shown in Fig. 11. These ligands are listed in supplementary material (SM Table 6: Rule 7).
Rule 8
Fragment H forms a scaffold without connected with any other fragments reported in Table 3. These ligands are listed in supplementary material (SM Table 7: Rule 8). Typical examples are depicted in Fig. 12.
With the frequent fragment descriptors derived from ZINC database, the compounds in LXRβ library are depicted in three-dimensional space by means of PCA as shown in Fig. 13.
Figure 13 demonstrates that the privileged fragments (Table 3) and their combinations are capable at discriminating compounds with similar scaffolds.
Experimental results
Based upon the above-mentioned rules, we selected compounds from our in-house compounds library for biological assays. Six compounds are found active against LXRβ in cell-based LXRβ agonistic assays. The compounds are listed in Fig. 14.
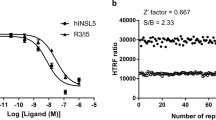
The activities of the confirmed LXRβ agonistic compounds are depicted in Fig. 15. GW3965 is for positive control. The compound 2 activated LXRβ significantly, the EC50 of which is 2.66 µM.
The activities of the confirmed LXRβ agonistic compounds and their fragment combination patterns. The letters above the bars represent the privileged fragments discovered by our algorithm. The structure of GW3965 is depicted in Fig. 11.
Discussion
Over the past decades, many substructure generation approaches have been reported, such as empirical search keys32, algorithm-based atom center fragments13, 33, 34, fingerprints (http://www.daylight.com/)15, 35. De novo substructures are derived by algorithms with a given minimal support threshold (popularity threshold). It is difficult to determine the threshold. The lower threshold results in too many trivial substructures, and the higher threshold results in potentially losing substructures that have strong relations with the activity. Another concern is that these substructure mining algorithms produce partial substructures (incomplete rings or aromaticity). In essence, these frequent substructures need to be refined with chemical and biological knowledge. Our approach is developed to resolve these problems. The features of our algorithm are summarized as follows:
-
(1)
We introduce a linear notation to encode growing substructures into strings which are used to filter out most of isomorphic substructures. This technique converted the atom-by-atom isomorphism checking process to a string comparison, dramatically reducing the computing complexity, and allowed us to run the frequent substructure discovery algorithm on “big” data (over ten million compounds level).
-
(2)
Thus, we have derived the standard MF substructure dictionary (SMFSD) for selecting substructures for a small compound library to keep relevant chemical and biological substructures and exclude trivial substructures. We proved that this method improved the accuracies of the predictions (Table 1).
-
(3)
Most of the previous substructure mining algorithms did not interpret the chemistry of the frequent substructures. Based on our method, we can derive privileged structures for a focused compound library, and figure out the rules to assembly these substructures (or building blocks for a drug lead). These rules can be used to guide a medicinal chemist in synthetic design for a drug target.
-
(4)
Regarding the comparison of our approach with the matched molecular pairs (MMP) approach36, MMP method focuses on identifying every pair of molecules that differ only by a particular, well-defined, structural transformation. Our method, however, focuses on gaining new chemical insights from the substructure mining algorithm without predefined chemical substructures.
Methods and Materials
Molecular graph
A compound is represented in a molecular graph (MG). MG is an object consisting of an atom list, a bond-list, and a molecular attribute list. Each atom in the atom list is an object containing atomic attributes, such as, atom ID, atomic number, mass, charge status, binding adjacency etc. Each bond in the bond list is an object containing chemical bond attributes, such as, bond ID, bond types, two binding atom IDs, and stereo description, etc. The molecular attribute list holds data including molecular ID, weight, name, activities, and other properties. The MG external representation is MOL format. A compound library consists of a number of small molecules represented in MGs. In graph theory, a compound library is a molecular graph database.
Maximal substructure tree
The tree is generated by a restricted depth-first search process, which only grows the node with the maximal substructure on the tree, other branches in the tree will be pruned. The tree starts with a single-edge fragments (for example, an edge with two carbon atoms connected in a single bond) called root fragments. Each substructure is expanded from a root fragment and is assigned with a subID (substructure identifier) vector. An element in the subID vector encodes the information of its parent molecule and root fragment. The tree grows by expanding root fragments through including adjacent edges (bonds). As shown in Fig. 16, the tree started with a root fragment of two carbon atoms with a single bond (subID = {An Bm Ch De Dk}). A subID consists of molecular IDs (denoted with capital letters) and bond IDs (denoted with lowercase letters). If a subID is generated from more than one molecule, the corresponding fragment is expanded and new nodes are added to the tree. In Fig. 1 at the root node of the tree, its subID consists of four members (popularity = 4). By expanding the fragment in the root node, two more nodes (Node-11 and Node-12) were added into the tree, and new subIDs were generated. The process was repeated till all successor nodes were undividable (subID consists only one member).
An example of a substructure generation tree. The tree started with a four compounds library with a C-C root fragment with a subID vector containing {An Bm Ch De Dk}. The popularity of this root fragment is 4. The root node produced two successor nodes (Node-11 and Node-12) by generating two new substructures. The process is repeated till all successor nodes are undividable. The substructures in the thick boxes are all possible fragment substructures created from a C-C root fragment. Other types of root fragments will be used to generate more substructure generation trees.
Pruning a substructure generation tree
The tree can grow rapidly, and cause a serious “combinatorial explosion”, because a MG can have \({2}^{n}-1\) possible substructures, where n is the number of edges (chemical bonds, the chemical bonds with hydrogens are omitted). These substructures contain huge amount of redundant information that can be pruned to significantly reduce computing complexity and simplify substructure trees37. As shown in Fig. 16, The tree was generated from node 1, and searched from the left branch (Node 1.1). Since 1.1 was a leaf node, the algorithm kept searching on right branch (node 1.2) until reached node 1.2.1.1.1.1.1, which was termed as potential reporting node (PRN). A PRN node is defined as follows:
Let P be the subID component set for a parent node, C 1 , C 2 , …C n be the subID component sets for the children nodes of the parent node (n is the number of the children nodes for PRN node).
Then, the parent node will be recognized as PRN node if (2) is satisfied:
For example, 1.2.1.1.1.1.1 (green box) was considered as a PRN because it had four children nodes and no child had the same as the subID of current node’s subID.
Each PRN had a popularity, which was the molecular counts encoded in subID. A substructure in PRN would be reported as a substructure if the PRN popularity was great than a designated threshold (t > 1), and the corresponding subID was recorded as well. Thus, a substructure library (containing subIDs and substructures) was generated and expanded when the tree was growing. When a new node was searched on the tree, its subID would be retrieved against the library. If the subID was found in the library, it would be pruned (red boxes in Fig. 17). Consequently, the successors of the pruned node would not be searched. The redundant information was avoided, and the computing complexity was significantly reduced.
Usually, a substructure, for example, the fragment in 1.2.1.1.1.1.1 node (Fig. 18), was the maximal substructure fragment (MSF) in a depth-first search path. After the MFS library was generated, the subIDs were converted into frequent fragment IDs (FFIDs), which came from subIDs by removing bondIDs. FFIDs encoded the information regarding their parents and popularities. Some FFIDs were assigned to unique fragments. Other FFIDs could have multiple fragments.
FFID and frequent substructures. Case 1: One FFID could have more than one fragments, one of the fragments was the substructure of another. The larger fragment was kept in this case. Case 2: One FFID could have more than one fragments, they were not topologically included to each other. Both fragments would be assigned to the same FFID.
As shown in Fig. 18, if one FFID was assigned with two fragments, and if one is the substructure of another (checked by the substructure match algorithm9), then the smaller fragment was removed (Fig. 18 Case 1). The detailed implementation of this tree pruning strategy can be found in the supplementary material.
Substructures as descriptors for a compound
QSAR study requires a descriptor vector for a compound. Each component in the descriptor vector is the count of a designated substructure that appeared in the compound. The designated substructures for the vector can be empirical (such as MDL 166 search keys or 960 extended search keys32, Daylight fingerprints38, or atom center fragments13, 39). In this work, we select the designated substructures for the vector based upon statistics. First, a SSD was derived from the ZINC database31, which contains more than 9.1 million chemical structures, to ensure the library covers known chemical diversity space. Let SSD have n substructures, a compound can be represented by a binary vector BV with n components, each component BV[i] (i ∈ 1..n) has a value 0 or 1 for SSD[i] being absent or present in the compound structure, for further QSAR or classification studies.
Data sets for classification and regression models using SSD
To examine the performance of the QSAR models using SSD, three data sets, LXRβ (Liver X receptor β), PPARα (peroxisome proliferator-activated receptor α), and VR (vasopressin receptor) libraries with chemical structures and bioactivities (IC50 values), were extracted from the BindingDB40. Duplicated structures in the libraries were filtered. Salt moieties in the connection tables were removed.
The activity data were pre-processed differently. For classification, the IC50 values were converted to zeros (in-actives) if they were greater than 10,000 nM, otherwise ones (actives). For regression, the records with the IC50 values, which were greater than 10,000 nM, were removed. Then, the IC50 values were converted to pIC50 values. This resulted in 717 and 634 LXRβ records, 784 and 621 PPARα records, and 619 and 491 VR records for classifications and regressions respectively.
The structural descriptors were selected from the SSD based upon their appearances in the corresponding data set. The frequency of a structural descriptor in a data set less than 5% was not selected.
Regularized logistic regression (RLR) method for compound classification
A compound library with known bioactivity results is represented in a matrix L[1..n, 1..m], where n is the number of the substructures selected from the SSD, and m is the number of compound structures in the library. The bioactivity data of the library is represented with A[1..m]. RLR41 will figure out parameter vector W[1..m] in (3).
L j is the descriptors for the jth compound, A j is the predicted activity (0 or 1) for the jth compound. Let F stand for SSD, X stand for the structures in the compound library L, S [1..m] stand for the scores of compounds being active, then, L[i, j], an element of L is defined in (4),
where SMFSD[i] is the ith MFS in SMFSD, and X[j] is the jth compound in L.
According to RLR approach41, the bioactive probability of the ith compound can be calculated in (5),
and the non-bioactive probability of the ith compound can be calculated in (6),
where S[j] is the activity prediction for the jth compound in L.
Machine learning process is to figure out W by optimizing (7) and (8) through logic regressions. For the jth compound,
where σ is standard deviation, C is a constant.
For all compounds,
We obtain optimized values for W through Newton iteration method29, because the second gradient of L(W) is always greater than zero.
Evaluating SSD-based classification results
ROC and following parameters were calculated to evaluate the MFS-based classification approach42.
SSD-based models were validated with the random sub-sampling cross validation method43. Initially, each experimental data set was randomly divided into 3 subsets; randomly selected 2 subsets to train the models, and the remaining subset was used for validating the models. The validating parameters were calculated and averaged over each batch of validations.
Predicting activities with support vector machines (SVM) using the substructure discriptors
In a SVM regression model44, the IC50 was converted to pIC50 (−log(IC50)), which is proportional to the bioactivity. pIC50 is the function of the descriptors, f(L x ), and was calculated as the following:
where L i is the descriptors (FS) of the ith compound; n is the number of the subsets (support vectors) from a training set; j is a compound in the subset; α j is the regression parameter for the jth compound; b is a regression constant to be determined by SVM regression process; k is a RBF kernel function defined in (10),
where σ is the standard deviation.
SVM model evaluation method
The SVM models were cross-validated through the average mean square error (MSE) and Pearson correlation of predicted and experimental pIC50 values with a k-fold cross-validation approach.
Method for selecting privileged substructure from SVM models
A privileged substructure is the one that is responsible for desired activities. Privileged substructures were derived from SVM models by ranking them with their p-values(p). If a MF substructure were used in a SVM model, its p-value (the function of the observed sample results that is used for testing a statistical hypothesis) was calculated with one-tailed test Fisher’s exact test45. Let A and B be the numbers of matched and unmatched substructures for the ith substructure in an active molecule from a training set; let C and D be the numbers of matched and unmatched substructures for ith substructures in a molecule from a background set (in our case, it is ZINC compound library). The p-value (p i ) is calculated as the following:
where a and b are numbers of matched and unmatched substructures for the ith substructure in an active molecule from a training set; and c and d are numbers of matched and unmatched substructures for ith MF substructures in a molecule from a background set. In an active data set. And, \(\frac{a}{c} > \frac{A}{C}\), a + b = A + B, c + d = C + D. The p-values were adjusted with false-discovery rate (FDR) approach46.
The substructures were sorted in the ascending orders of p-values. The significant substructures are with p-values < 0.05. Privileged substructures were elected by using high-scored substructures as substructure queries searching against the targeted compound library. The hits with a high number of active compounds were identified as the privileged substructures of the focused library.
Deriving the rules of combining fragments
Let A = {a[0], a[1], … a[x], …a[M − 1]} as a privileged substructure list derived from a compound library; B = {b[0], b[1], … b[y], … b[N − 1]} as the compound list. The rules for combining fragments for FBDD study can be discovered in the following pseudo-code:
References
Hong, C. & Tontonoz, P. Liver X receptors in lipid metabolism: opportunities for drug discovery. Nature reviews. Drug discovery 13, 433–444 (2014).
Lin, C.-Y., Vedin, L.-L. & Steffensen, K. R. The emerging roles of liver X receptors and their ligands in cancer. Expert Opinion on Therapeutic Targets 20, 61–71 (2016).
Zhao, W. et al. Three-dimensional pharmacophore modeling of liver-X receptor agonists. Journal of chemical information and modeling 51, 2147–2155 (2011).
Li, Y. et al. Predicting selective liver X receptor b agonists using multiple machine learning methods. Molecular BioSystems 11, 1241–1250 (2015).
Temml, V., Voss, C. V., Dirsch, V. M. & Schuster, D. Discovery of New Liver X Receptor Agonists by Pharmacophore Modeling and Shape-Based Virtual Screening. Journal of chemical information and modeling 54, 367–371 (2014).
von Grafenstein, S. et al. Identification of Novel Liver X Receptor Activators by Structure-Based Modeling. Journal of chemical information and modeling 52, 1391–1400 (2012).
Lagarde, N., Delahaye, S., Zagury, J.-F. & Montes, M. Discriminating agonist and antagonist ligands of the nuclear receptors using 3D-pharmacophores. Journal of cheminformatics 8, 43 (2016).
Keserű, G. M. et al. Design Principles for Fragment Libraries: Maximizing the Value of Learnings from Pharma Fragment-Based Drug Discovery (FBDD) Programs for Use in Academia. J Med Chem 59, 8189–8206 (2016).
Xu, J. GMA: a generic match algorithm for structural homomorphism, isomorphism, and maximal common substructure match and its applications. Journal of chemical information and computer sciences 36, 25–34 (1996).
Willett, P., Barnard, J. M. & Downs, G. M. Chemical similarity searching. Journal of chemical information and computer sciences 38, 983–996 (1998).
Xu, J. A New Approach to Finding Natural Chemical Structure Classes. J Med Chem 45, 5311–5320 (2002).
Batista, J., Tan, L. & Bajorath, J. Atom-centered interacting fragments and similarity search applications. Journal of chemical information and modeling 50, 79–86 (2010).
Xu, J. 13C NMR Spectral Prediction by Means of Generalized Atom Center Fragment Method. Molecules 2, 114 (1997).
Xu, J. & Stevenson, J. Drug-like Index: A New Approach To Measure Drug-like Compounds and Their Diversity. Journal of Chemical Information and Computer Sciences 40, 1177–1187 (2000).
Xu, J. & Hagler, A. Chemoinformatics and drug discovery. Molecules 7, 566–600 (2002).
Dehaspe, L., Celestijnenlaan, A., Toivonen, H., King, R. D. & Ceredigion, P. A. Finding frequent substructures in chemical compounds. Proceedings of KDD-98, 30–36 (1998).
Yan, X. F. & Han, J. W. gSpan: Graph-based substructure pattern mining. 2002 Ieee International Conference on Data Mining, Proceedings, 721–724 (2002).
Huan, J., Wang, W. & Prins, J. In IEEE International Conference on Data Mining 549–552 (2003).
Kuramochi, M. & Karypis, G. An efficient algorithm for discovering frequent subgraphs. Ieee T Knowl Data En 16, 1038–1051 (2004).
Borgelt, C., Meinl, T. & Berthold, M. MoSS: a program for molecular substructure mining. Osdm’05 Proceedings of International Workshop on Open Source Data Mining, 6–15 (2005).
Borgelt, C. & Meinl, T. Full Perfect Extension Pruning for Frequent Graph Mining. (Springer Berlin Heidelberg, 2009).
Meinl, T., Wörlein, M., Urzova, O., Fischer, I. & Philippsen, M. The ParMol package for frequent subgraph mining. Electronic Communications of the Easst (2007).
Wang, F., Dong, J. Q. & Yuan, B. Graph-Based Substructure Pattern Mining Using CUDA Dynamic Parallelism. Lect Notes Comput Sc 8206, 342–349 (2013).
Khashan, R., Zheng, W. & Tropsha, A. The Development of Novel Chemical Fragment-Based Descriptors Using Frequent Common Subgraph Mining Approach and Their Application in QSAR Modeling. Molecular Informatics 33, 201–215 (2014).
Shao, Z., Hirayama, Y., Yamanishi, Y. & Saigo, H. Mining Discriminative Patterns from Graph Data with Multiple Labels and Its Application to Quantitative Structure-Activity Relationship (QSAR) Models. Journal of chemical information and modeling 55, 2519–2527 (2015).
Yan, X. & Han, J. In ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Washington, Dc, Usa, August 286–295 (2003).
Kuramochi, M. & Karypis, G. In IEEE International Conference on Data Mining 313–320 (2001).
Takigawa, I. & Mamitsuka, H. Graph mining: procedure, application to drug discovery and recent advances. Drug Discovery Today 18, 50–57 (2013).
Lin, C. J., Weng, R. C. & Keerthi, S. S. Trust Region Newton Method for Large-Scale Logistic Regression. Journal of Machine Learning Research 9, 627–650 (2008).
Jhoti, H., Williams, G., Rees, D. C. & Murray, C. W. The ‘rule of three’ for fragment-based drug discovery: where are we now? Nature Reviews Drug Discovery 12, 644–645 (2013).
Irwin, J. J. & Shoichet, B. K. ZINC-a free database of commercially available compounds for virtual screening. Journal of chemical information and modeling 45, 177–182 (2005).
Durant, J. L., Leland, B. A., Henry, D. R. & Nourse, J. G. Reoptimization of MDL Keys for Use in Drug Discovery. Journal of chemical information and modeling 42, 1273–1280 (2002).
Liu, Z. et al. ChemStable: a web server for rule-embedded naive Bayesian learning approach to predict compound stability. Journal of computer-aided molecular design 28, 941–950 (2014).
Wang, L. et al. Predicting mTOR inhibitors with a classifier using recursive partitioning and Naive Bayesian approaches. Plos One 9, e95221 (2014).
Awale, M. & Reymond, J.-L. Atom Pair 2D-Fingerprints Perceive 3D-Molecular Shape and Pharmacophores for Very Fast Virtual Screening of ZINC and GDB-17. Journal of chemical information and modeling 54, 1892–1907 (2014).
Leach, A. G. et al. Matched Molecular Pairs as a Guide in the Optimization of Pharmaceutical Properties; a Study of Aqueous Solubility, Plasma Protein Binding and Oral Exposure. J Med Chem 49, 6672–6682 (2006).
Yan, X. & Han, J. In Proceedings of the ninth ACM SIGKDD international conference on Knowledge discovery and data mining 286–295 (ACM, 2003).
Butina, D. Unsupervised data base clustering based on Daylight’s fingerprint and Tanimoto similarity: A fast and automated way to cluster small and large data sets. Journal of Chemical Information and Computer Sciences 39, 747–750 (1999).
Yan, X., Gu, Q., Lu, F., Li, J. & Xu, J. GSA: a GPU-accelerated structure similarity algorithm and its application in progressive virtual screening. Molecular diversity 16, 759–769 (2012).
Chen, X., Liu, M. & Gilson, M. K. BindingDB: a web-accessible molecular recognition database. Combinatorial chemistry & high throughput screening 4, 719–725 (2001).
Fan, R.-E., Chang, K.-W., Hsieh, C.-J., Wang, X.-R. & Lin, C.-J. LIBLINEAR: A Library for Large Linear Classification. J. Mach. Learn. Res. 9, 1871–1874 (2008).
Fawcett, T. An introduction to ROC analysis. Pattern recognition letters 27, 861–874 (2006).
Kohavi, R. A study of cross-validation and bootstrap for accuracy esti-mation and model selection, IJCAI 14, 1137–1145 (1995).
Javed, F. et al. In Engineering in Medicine and Biology Society, 2009. EMBC 2009. Annual International Conference of the IEEE 4352–4355 (IEEE, 2009).
Fisher, R. A. On the interpretation of χ2 from contingency tables, and the calculation of P. Journal of the Royal Statistical Society, 87–94 (1922).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. Series B (Methodological), 289–300 (1995).
Acknowledgements
This work was supported by the National Science Foundation of China (81473138), Guangdong Province Frontier and Key Technology Innovation Program (2015B010109004), Guangdong National Science Foundation (2016A030310228), Guangdong Provincial Key Laboratory of Construction Foundation (2011A060901014), Guangdong NSF (2016A030310228), and the Fundamental Research Funds for the Central Universities (2013HGCH0015). We also thank Professor Johann Gasteiger for his advice and proof-reading the manuscript.
Author information
Authors and Affiliations
Contributions
Ideas and experiment design: J.X. and H.P. Computational development and design: J.X., H.P. and Z.L. Analyses and data interpretation: H.P., J.X. and X.Y. Write manuscript: J.X. and H.P. Read and revised the manuscript: J.X. Study supervision: J.X. and J.R.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peng, H., Liu, Z., Yan, X. et al. A de novo substructure generation algorithm for identifying the privileged chemical fragments of liver X receptorβ agonists. Sci Rep 7, 11121 (2017). https://doi.org/10.1038/s41598-017-08848-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-08848-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.