Abstract
Genome duplication and repeat multiplication contribute to genome evolution in plants. Our previous work identified a recent allotetraploidization event and five high-copy LTR retrotransposon (LTR-RT) families PgDel, PgTat, PgAthila, PgTork, and PgOryco in Panax ginseng. Here, using whole-genome sequences, we quantified major repeats in five Panax species and investigated their role in genome evolution. The diploids P. japonicus, P. vietnamensis, and P. notoginseng and the tetraploids P. ginseng and P. quinquefolius were analyzed alongside their relative Aralia elata. These species possess 0.8–4.9 Gb haploid genomes. The PgDel, PgTat, PgAthila, and PgTork LTR-RT superfamilies accounted for 39–52% of the Panax species genomes and 17% of the A. elata genome. PgDel included six subfamily members, each with a distinct genome distribution. In particular, the PgDel1 subfamily occupied 23–35% of the Panax genomes and accounted for much of their genome size variation. PgDel1 occupied 22.6% (0.8 Gb of 3.6 Gb) and 34.5% (1.7 Gb of 4.9 Gb) of the P. ginseng and P. quinquefolius genomes, respectively. Our findings indicate that the P. quinquefolius genome may have expanded due to rapid PgDel1 amplification over the last million years as a result of environmental adaptation following migration from Asia to North America.
Similar content being viewed by others
Introduction
Nuclear genome sizes in flowering plants are diverse, and can vary over 2,400-fold, ranging from 63 Mb in Genlisea margaretae to 149 Gb in Paris japonica 1. This dramatic genome size variation is attributed to both whole-genome duplication and accumulation of repeated sequences, or repeats2,3,4. During the diploidization process following genome duplication, euchromatic DNA is usually reduced by deletion of unnecessary paralogous regions5, 6 while heterochromatic DNA is often expanded by species-specific multiplication of repeats7. Repeats are categorized into two major types: tandem repeats (TRs) and transposable elements (TEs)8. TRs exist in a head-to-tail arrangement in distinct chromosomal regions, generally found at centromeric, subtelomeric, and telomeric regions7, 9. By contrast, TEs are dispersed throughout the genome. TEs are classified based on their transposition mechanisms as class I (copy-and-paste) or class II (cut-and-paste). Class I TEs include the class I.1 LTR-retrotransposons (LTR-RTs) and the class I.2 non-LTR retrotransposons, whereas class II TEs include DNA transposons10. Repeats play important roles in gene regulation, evolution, and adaptation11,12,13.
The family Araliaceae is composed of approximately 55 genera and 1,500 species, which include many valuable medicinal and ornamental plants14. Within this family, the genus Panax contains economically important medicinal plants including the diploids P. japonicus, P. vietnamensis, and P. notoginseng (2n = 2x = 24), and the tetraploids P. quinquefolius and P. ginseng (2n = 4x = 48). These five species are perennial and absolute shade plants that have been used for medicinal purposes in Asia and North America because of their beneficial effects on human health15. Although Panax species display relatively limited morphological diversity, their genome sizes vary from 2.02 Gb (P. vietnamensis) to 4.9 Gb (P. quinquefolius)16, 17. Several genomic studies have been conducted to elucidate the genome structure, function, and evolution of genomes in the Panax genus18,19,20,21,22,23,24,25,26.
Recently, we described the evolution of five Panax species by comparative analysis of complete chloroplast genome sequences and ribosomal DNA25. We also characterized the major repeats that occupied more than 35% of the P. ginseng genome, namely five high-copy LTR-RT families26, 27. In this study, we aimed to explore the role of major repeats in the evolution of the Panax genus, which shows large genome size variation. Accordingly, we established a reliable quantification method for major repeats within a genome using low-coverage whole-genome sequences and quantified each of these LTR-RTs in the genomes of five Panax species. Our comparative analysis revealed dynamic impacts of these major repeats on genome size variation, speciation, and evolution in the Panax genus.
Results
Whole genome sequence (WGS)-based quantification of major repeats in P. ginseng
In P. ginseng, we recently reported 11 LTR-RT subfamilies contained within five superfamilies, namely PgDel1–6, PgTat1 and 2, PgAthila, PgTork, and PgOryco, and two tandem repeat sequences, namely Pg167TR and 45 S rDNA (Supplementary Table S1). These 13 repeats are high-copy, major repeats and are estimated to occupy more than 41% of the P. ginseng genome (Supplementary Table S2, S3, and Fig. S1)24, 26, 27. Here, we aimed to quantify these major repeats in the WGS datasets of five Panax species. We determined the amount of each repeat by calculating its genomic proportion (GP) in each WGS, via quantification of homologous nucleotides in each WGS based on repeat masking using RepeatMasker28. We validated RepeatMasker-based GP (R-GP) estimation and the quantification of each major repeat using various WGS data sets. We then compared repeat quantification in WGS data sets with different genome coverages (0.00005–10x), as well as in WGS data sets from different libraries using P. ginseng cv. ‘Chunpoong’ and in WGS data sets from different ginseng cultivars (Table 1, Supplementary Table S2, S4, and Fig. S1). The reproducibility of R-GP estimation for each of major repeats was evaluated in each WGS.
The R-GP of each repeat displayed little variation in datasets of the same WGS that represented nine different genome coverages, and low variation in datasets from four different WGS libraries created using the same ginseng cultivar (Table 1 and Supplementary Table S4). Furthermore, low R-GP variation for repeats was observed across WGS datasets of 11 ginseng cultivars, with the 13 repeats displaying a R-GP of 41.0–46.3% (Supplementary Table S2 and Fig. S1). High-copy LTR-RTs showed little variation, while low-copy LTR-RTs occupying less than 1% GP and tandem repeat units such as Pg167TR and 45 S rDNA showed relatively high variation (Table 1 and Supplementary Table S4). The R-GP of PgDel1 was 23–26% among 11 cultivars (Supplementary Table S2 and Fig. S1).
Genomic quantification of major repeats in five Panax species
We used the above WGS-based R-GP estimation to quantify the major repeats in the genomes of five Panax species alongside a species from a related genus (Table 2). The quantification of repeats using PE reads corresponding to 0.3–1.5x haploid genome equivalents for each species revealed a R-GP of 46%, 45%, 50%, 41%, 53%, and 17% in P. japonicus, P. vietnamensis, P. notoginseng, P. ginseng, P. quinquefolius, and Aralia elata, respectively (Fig. 1). Each individual major repeat possessed a similar R-GP in the five Panax species, whereas in A. elata, the R-GP was comparatively low. The Ty3/Gypsy-type LTR-RT families, such as PgDel1–6, PgTat1–2, and PgAthila, covered approximately 37.7–47.5% of the genomes. Among these, PgDel1 had 22.6–34.5% R-GP in five Panax species but approximately 1% R-GP in A. elata. In particular, the larger genome of the Panax species in P. quinquefolius had a high amount of PgDel1 elements, with a R-GP of 34.5% (Fig. 1).
The R-GP of PgDel2, PgDel5, PgDel6, and PgTork displayed large variation among the five Panax species (Fig. 1). PgDel2 had 2.6–3.0% R-GP in the three diploid Panax species, and 1.5% and 1.4% R-GP in the two tetraploids P. ginseng and P. quinquefolius, respectively, which was approximately half of that measured in the diploids. PgDel5 was more abundant in P. notoginseng and A. elata compared to that in other species. PgDel6 had 4.3% and 5% R-GP in the two diploids P. japonicus and P. vietnamensis, respectively, whereas it had 1.5–2.4% R-GP in the remaining three Panax species. The R-GP of PgTork varied dynamically between Panax species (Fig. 1).
Dynamics of the PgDel1 subfamily members in Panax species
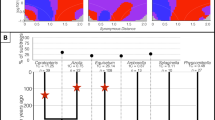
We analyzed the structural dynamics of PgDel1 subfamily members in the Panax species. Five PgDel1 subfamily members (PgDel1_1–5) were identified from three complete BAC clone sequences (GenBank accession nos. KF357943, KF357944, and KF357942)27. These five members displayed relatively complete structures including both LTRs and an inner sequence, although there were nested insertions caused by other repeats or subsequent deletion events. Inspection of the complete unit of these repeats, which was 7.7–10.1 kb, revealed an overall similarity in the large structural variations in the LTR regions. To estimate the distribution of PgDel1 members in the P. ginseng genome, we mapped the 1x genome coverage Chunpoong WGS data onto the representative PgDel1_1 element because of the well-conserved LTR domains of PgDel1. Mapping depth had a range of 111–157,407 with an average of 50,952 (mode and median values were 48,399 and 47,503, respectively) (Fig. 2).
Structural characteristic of five PgDel1 subfamily members. (A) Representation of the distribution of 1x WGS data of P. ginseng cv. CP. (B) Horizontal schematic diagrams of PgDel1 subfamily members 1–5. Boxed orange triangles indicate LTR regions of PgDel1. Yellow boxes indicate the internal LTR-RTs domains of PgDel1 detected in each subfamily member. (AP: aspartic protease, CH: chromodomain, GAG: capsid protein, INT: integrase, RH: RNase H, RT: reverse-transcriptase, and Zn: zinc knuckle). Homologous sequence were indicated as grey panels.
Cytogenomic mapping of PgDel1 and PgDel2 in three Panax species
To validate the R-GP variation identified via in silico analysis, we analyzed the distribution patterns of PgDel1 and PgDel2 by fluorescence in situ hybridization (FISH) using somatic metaphase chromosomes of three Panax species: P. notoginseng, as a representative of the three diploid Panax species, and the two tetraploids P. ginseng and P. quinquefolius. The PgDel1 elements displayed high-density FISH signals throughout the chromosomes in all three Panax species (Fig. 3A,B and C). The intensive FISH signal of PgDel1 throughout the chromosome regardless of the ploidy level of the species it originated from supported our in silico analysis results, which estimated 23–35% R-GP for PgDel1 in the Panax species (Figs 1 and 3A–C).
Fluorescence in situ hybridization (FISH) analysis of PgDel1 and PgDel2 distribution in P. ginseng, P. quinquefolius, and P. notoginseng chromosomes. The PgDel1 FISH signals in somatic metaphase chromosomes of (A) P. notoginseng (purple), (B) P. ginseng, and (C) P. quinquefolius. The PgDel2 FISH signals in somatic metaphase chromosomes of (D) P. notoginseng, (E) P. ginseng (blue), and (F) P. quinquefolius (blue). Bar = 10 μm.
PgDel2 had nearly two-fold greater R-GP values in the three diploid Panax species compared to the two tetraploids (Fig. 1). Consistent with this result, FISH analysis revealed different distribution patterns of PgDel2 in diploid and tetraploid Panax species. PgDel2 signal was localized to pericentromeric regions in all 24 chromosomes of the diploid P. notoginseng, whereas strong PgDel2 signal was detected in half of the 48 chromosomes of both tetraploid Panax species (Fig. 3D,E and F). In these tetraploids, PgDel2 distribution was concentrated to the pericentromeric regions in P. ginseng chromosomes but was more broadly located in P. quinquefolius chromosomes (Fig. 3E and F).
Contribution of major repeats to genome size variation
We investigated the contribution of the four most abundant LTR-RT families, PgDel, PgAthila, PgTat, and PgTork, to the overall genome contents. Each family was present in varied proportions in the six analyzed species (Figs 1 and 4). Combined, the four LTR-RTs had a 39–52% R-GP in each of five species, corresponding to 0.9–2.6 Gb (Fig. 4). Of these repeats, PgDel occupied 30–41% of R-GP, accounting for 0.7–2.0 Gb. PgTat had a 5–7% R-GP, corresponding to 97–285 Mb. The estimated quantity of PgTork was 241 Mb in P. quinquefolius, whereas it was 24–57 Mb in the other four Panax species. Interestingly, PgTork was the most abundant LTR-RT in the A. elata genome, occupying 7% R-GP (56 Mb) (Fig. 4).
Comparison between proportions of four major repeats in five Panax species and A. elata. (A) Phylogenetic relationships based on chloroplast sequences modified from Kim et al.25. Estimated time since divergence (MYA) is indicated at the root of branch divisions. (B) The predicted genome size of Panax species and A. elata are depicted as bar charts with the estimated amounts of PgDel, PgTat, PgAthila, and PgTork families contained in each genome represented by blue, green, yellow, and brown regions, respectively. Genome contents not containing these repeats are represented by the grey region. Blue letters above bars indicate GP of PgDel alone (left), GP of four LTR-RT families (middle), and total GP of the genome (right). Red letters below bars indicate estimated amount in Gb of PgDel contents alone (left), contents of four LTR-RT families (middle), and total genome size (right). For A. elata, blue letters above bars indicate GP of four LTR-RT families (left) and total GP of the genome (right), whereas red letters below bars indicate estimated amount in Gb of four LTR-RT families (left) and total genome size (right). Total genome size of P. japonicus was estimated in the present study.
Discussion
In this work, we used low-coverage WGS sequences to calculate the GP of major repeats. We estimated the prevalence of each repeat by determining GP using various WGS data sets, based on the calculation of masked homologous sequence in WGS reads by RepeatMasker28. GP can also be calculated using clustered WGS reads or mapped WGS reads29, 30. Mapping-based GP (M-GP) and clustering-based GP (C-GP) calculations are based on numbers of homologous WGS reads, whereas R-GP calculation is based on real amounts of homologous sequences in WGS reads. We compared the ability of R-GP and M-GP methods to estimate PgDel1 GP using different WGS sets, which resulted in a consistent pattern whereby R-GP calculations estimated 3–4% more GP than M-GP calculations (Supplementary Fig. S3 and S4). This variation may be attributed to the difference in how homologous sequences are counted in both methods, namely the number of homologous reads and the number of homologous nucleotides for M-GP and R-GP, respectively.
The estimated GPs were highly reproducible for the major high-copy LTR-RTs, although we observed relatively high CV values (25–65%) for GP estimation of tandem repeats using different genome coverage and different WGS libraries (Table 1 and Supplementary Table S4). The number of tandem repeat reads might be uneven because of biased fragmentation during WGS library construction (Table 1 and Supplementary Table S4). Overall, though, the coefficients of variation (CVs) of the high copy PgDel1 and the low copy PgOryco were 3.23% and 13.53%, respectively, when we estimated GP using various levels of genome coverage in the data sets, i.e., 0.0001−10x genome coverage for WGS reads of P. ginseng (Table 1). We observed slightly increased variation when we reduced the genome coverage below 0.0001x, but all the data showed similarly low CVs when we utilized over 0.0001x coverage WGS reads. As WGS data can be produced at low cost by high-throughput NGS processes and over 1 Mbp of WGS reads produced reproducible GP estimation, we conclude that genome coverage in WGS data is not a critical constraint limiting the application of this approach for analysis.
Although there is some variation, the GPs calculated here for the major repeats were reproducible and are thus representative of the abundance of each repeat in the genomes of the different species. However, it is possible that the true GP for each major repeat is higher than the GP estimate presented here because, in our analysis, only a single representative structure was used for each repeat and other structural variations were not considered31. For example, five PgDel1 elements displayed large structural variation in the LTR region and a large bias in WGS read mapping for the representative PgDel1 family member in the P. ginseng genome (Fig. 2).
Our results point to tetraploidization and four LTR-RTs as the primary reasons for genome size variation in the genus Panax. Divergence of a common ancestor into the genera Panax and Aralia is predicted to have occurred approximately eight MYA25. A. elata was estimated to have an approximate haploid genome equivalent size of 0.8 Gb on 12 chromosome pairs (Supplementary Fig. S2). However, the genome sizes of Panax species (2.0–4.9 Gb) are much larger than that of A. elata. We propose that the multiplication of some major repeats influenced the genome size in the Panax lineage. In particular, a large proportion of the increased genome size can be explained by multiplication of the four LTR-RTs investigated here, which occupied 0.9–2.6 Gb in the Panax lineage (Fig. 4). The GP of the four LTR-RTs was 39% (1.4 Gb) and 52% (2.6 Gb) in two tetraploids, P. ginseng and P. quinquefolius, respectively, and 44–49% (0.9–1.2 Gb) in the three diploid Panax species. Among them, PgDel was the predominant repeat with a GP of 30–41%, which corresponds to 0.7–2.0 Gb in the five Panax species.
LTR-RTs make up a large proportion of the genomes of many higher plants32,33,34. The repeats can play an important role as promoters of genomic diversification and speciation35. It is possible that, even in the same genus, a rapid burst of retrotransposition can induce genome size variance with different evolutionary effects, as observed for Oryza, Nicotiana, and Genlisea 36,37,38. Here we investigated abundant, high-copy LTR-RTs and performed a comparative analysis of these repeats in Panax species to understand their influence on genome evolution. The presence of these repeats in five Panax species and a further related species suggests that they likely existed in the genome of a common ancestor39. However, extensive multiplication of LTR-RTs occurred only in the Panax genus and appears to have a decisive effect on the expansion of the genome sizes in Panax species (Figs 1 and 4). This finding suggests that the repeat amplification occurred concomitantly with or following divergence in the five Panax species during the last eight million years.
We identified six PgDel subfamilies based on LTR sequences from P. ginseng BAC clone sequences26, 27. Among them, PgDel1 was highly abundant in each Panax species. The abundance, sequence diversity, and cytogenetic distribution of PgDel1 LTR-RTs indicated that considerable multiplication and transposition may have occurred across the five Panax species genomes (Figs 1, 2, 3 and 4). We found a positive correlation (coefficient of 0.6 with a p-value of 0.40) between the R-GP values for PgDel1 and the genome size of each Panax species (Table 2 and Fig. 4). This correlation indicates that the accumulation of PgDel1 elements has greatly contributed to the increased genome sizes in the genus Panax. In this regard, we speculate that the genome size of P. japonicus might be below 2.0 Gb, based on the relatively small PgDel1 GP we found in the diploid Panax species (Figs 1 and 4).
Correlation between PgDel1 abundance and genome size in Panax species could explain the expansive genome of P. quinquefolius, which is the largest within the Panax genus. The two tetraploids P. ginseng and P. quinquefolius were reported to exhibit a difference of 1.3 Gb. Based on divergence of orthologous gene pairs, we estimated that these species diverged approximately one MYA, following the recent allotetraploidization two MYA40. The considerable disparity in genome size that has evolved between P. ginseng and P. quinquefolius is largely explained by the different amount of PgDel1 in each genome, which is 0.8 and 1.7 Gb, respectively, indicating that PgDel1 was exclusively amplified in P. quinquefolius during last one MY.
The difference between PgDel1 GP in P. ginseng and P. quinquefolius can be explained by two hypotheses concerning TE dynamics. The first hypothesis is that there was a considerable loss of PgDel1 GP in P. ginseng after speciation. Polyploidization often results in genome downsizing via expulsion of genomic DNA, mostly repetitive DNA sequence, for stable meiotic rebuilding in nascent polyploids37, 41, 42. The second hypothesis is that there was a sizeable expansion of PgDel1 GP in P. quinquefolius after speciation. We believe that the second hypothesis holds more merit than the first. Drastic environmental change could have triggered epigenetic restructuring42,43,44, resulting in the unusual accumulation of LTR-RTs in P. quinquefolius 45. P. quinquefolius is said to have migrated from Asia to America through the Bering land bridge during glacial and interglacial cycles one MYA40. Consequently, P. quinquefolius would have been exposed to extreme abiotic stress during the process of migration and adaptation to new habitats. The influence of PgDel1 amplification in genome organization and gene function accordingly might play an important role in the interspecific genomic barriers between species.
PgDel1 made up a large proportion of the genome in all five Panax species analyzed in this study. In addition, other PgDel subfamily members also had notable genome distributions in the Panax species. PgDel2 occupied approximately 1.4% GP in the two tetraploids and 2.8% GP in the three diploid Panax species (Fig. 1). This variation in PgDel2 GP between diploids and tetraploids was confirmed by cytogenetic analysis using FISH. In the tetraploids P. ginseng and P. quinquefolius, PgDel2 signals were observed in half of the chromosomes whereas all chromosomes of the diploid P. notoginseng displayed PgDel2 signals (Fig. 3D,E, and F). PgDel5 and PgDel6 showed large differences among three Panax species. P. notoginseng had approximately twice the amount of PgDel5 than the other Panax species, and P. japonicus and P. vietnamensis had more abundant PgDel6 compared to the other species. These findings highlight the likely importance of the PgDel subfamily contribution to diversification of Panax species.
Materials and Methods
Plant materials, genomic DNA isolation, and Illumina sequencing
Eleven P. ginseng cultivars as well as P. quinquefolius, P. notoginseng, P. japonicus, P. vietnamensis and A. elata were used for genomic DNA preparation and sequencing (Table 2 and Supplementary Table S3). P. ginseng cv. Chunpoong was used as a representative for GP estimation in the current analysis. Leaf tissue for the above species, apart for P. notoginseng, P. japonicus, P. vietnamensis, and A. elata, was obtained from the ginseng farms of Seoul National University and Korean Ginseng Corporation (http://www.kgc.or.kr). A. elata and P. vietnamensis leaf tissue was collected from Susinogapy Corporation (http://www.susinogapy.com), Korea, and Da Lat City, Tay Nguyen Institute of Scientific Research, Vietnam, respectively. P. notoginseng and P. japonicus leaf tissue was collected from Dafang Country, Guizhou province, and Enshi County, Hubei province, China, respectively.
Genomic DNA was extracted using a modified cetyltrimethylammonium bromide (CTAB) method46. All genomic libraries were prepared according to the recommended Illumina paired-end standard protocol (http://www.illumina.com). The whole genomes of those plants listed in Table 2 and Supplementary Table S3 were sequenced using an Illumina genome analyzer at the National Instrumentation Center for Environmental Management (NICEM: http://nature.snu.ac.kr/kr.php) and LabGenomics (www.labgenomics.co.kr/), South Korea. All sequence data were uploaded to the National Agricultural Biotechnology Information Center (http://nabic.rda.go.kr) (Table 2 and Supplementary Table S3)47.
Major repeat sequences of Panax ginseng
In our previous study, we described the major repeats of P. ginseng including 11 LTR-RTs and two tandem repeat sequences (Pg167TR and 45 S rDNA), which occupy more than one third of the genome24, 26, 27. These reference sequences were used as queries to estimate their abundance in Panax and Aralia genomes. Most of LTR-RTs of major repeats analyzed in this work have a complete structure that includes both flanking LTRs and inter-LTR domains, with the exception of PgAthila that has one LTR27. The 45 S rDNA of P. ginseng was used as a representative rDNA sequence for all Panax species (Supplementary Table S1 and Data S1).
Quantification of major repeats using WGS
The GP of each repeat was quantified by masking nucleotides of WGS reads into the representative repeat sequence using RepeatMasker (ver. 4.0.6)28. WGS reads were trimmed based on their quality score (minimum quality score: ≥20) using the software Trimmomatic ver. 0.3348. WGS reads were directly surveyed for homology to each repeat using RepeatMasker, using the slow search parameters and option that does not mask low complexity DNA or simple repeats (applying ‘-s –no low’). Homologous nucleotides were masked and the amounts of masked nucleotides were counted to calculate GP for each repeat. The RepeatMasker-based genomic proportion (R-GP) was calculated as the proportion that masked nucleotides make of total nucleotides in each data set: R-GP (%) = (masked read length/total read length) × 100. The actual amounts of each repeat in the genome was estimated based on the R-GP and the size of the genome: Repeat amount = (R-GP/100) × genome size) (Fig. 4B). The mapping-based GP (M-GP) and copy number of PgDel1_1 LTR-RTs were estimated using CLC mapper ver. 4.21.104315 (CLC Inc, Aarhus, Denmark) with the parameters of minimum 50% fraction of the read and 80% similarity (Fig. 2A, Supplementary Fig. S3 and S4).
Fluorescence in situ hybridization (FISH) analysis
Preparation of P. notoginseng, P. ginseng, and P. quinquefolius chromosome spreads and FISH procedures were performed according to dual-color FISH analysis protocols49. Briefly, root tips were treated with 2 mM 8-hyroxyquinoline, fixed with Carnoy’s solution, and were enzymatically digested with pectolytic enzyme solution (2% Cellulase R-10 (C224, Phytotechnology Laboratories) and 1% Pectolyase Y-23 (P8004.0001, Duchefa)) in 100 mM citrate buffer) for 1 h. Root tips were then squashed onto slides pre-cleaned with 70% ethanol. Air-dried slides were fixed in 2% formaldehyde for 5 min and dehydrated with a series of ethanol treatments (70%, 90%, and 100%)50. PgDel1 and PgDel2 probes were obtained by PCR amplification using P. ginseng genomic DNA and primers detailed in our previous study27. PgDel1 was labeled with Cy5-dUTP (Jena Bioscence), whereas PgDel2 was labeled with Diethyl amino coumarin-5-dUTP (NEL455001EA, Perkin Elmer) or Alexa Fluor 488-5-dUTP (C11397, Life Technologies). Images were captured using an Olympus BX53 epifluorescence microscope equipped with a Leica DFC365 FS CCD camera, and processed using Cytovision version 7.2 (Leica Microsystems, Germany). Further image enhancements were performed using Adobe Photoshop CS6.
References
Pellicer, J., Fay, M. F. & Leitch, I. J. The largest eukaryotic genome of them all? Biol. J. Linn. Soc. Lond. 164, 10–15 (2010).
SanMiguel, P. et al. Nested retrotransposons in the intergenic regions of the maize genome. Science 274, 765–768 (1996).
SanMiguel, P., Gaut, B. S., Tikhonov, A., Nakajima, Y. & Bennetzen, J. L. The paleontology of intergene retrotransposons of maize. Nat. Genet. 20, 43–45 (1998).
Wendel, J. F. Genome evolution in polyploids. Plant Mol. Biol 42, 225–249 (2000).
Leitch, I. J. & Bennett, M. D. Genome downsizing in polyploid plants. Biol. J. Linn. Soc. Lond. 82, 651–663 (2004).
Yang, T. J. et al. Sequence-level analysis of the diploidization process in the triplicated FLOWERING LOCUS C region of Brassica rapa. Plant Cell 18, 1339–1347 (2006).
Lim, K. B. et al. Characterization of the centromere and peri‐centromere retrotransposons in Brassica rapa and their distribution in related Brassica species. Plant J. 49, 173–183 (2007).
Richard, G. F., Kerrest, A. & Dujon, B. Comparative genomics and molecular dynamics of DNA repeats in eukaryotes. Microbiol Mol. Biol. Rev. 72, 686–727 (2008).
Csink, A. K. & Henikoff, S. Something from nothing: the evolution and utility of satellite repeats. Trends Genet. 14, 200–204 (1998).
Piégu, B., Bire, S., Arensburger, P. & Bigot, Y. A survey of transposable element classification systems–a call for a fundamental update to meet the challenge of their diversity and complexity. Mol. Phylogenet. Evol. 86, 90–109 (2015).
Volff, J. N. Turning junk into gold: domestication of transposable elements and the creation of new genes in eukaryotes. Bioessays 28, 913–922 (2006).
Feschotte, C. & Pritham, E. J. DNA transposons and the evolution of eukaryotic genomes. Annu. Rev. Genet. 41, 331–368 (2007).
Oliver, K. R., McComb, J. A. & Greene, W. K. Transposable elements: powerful contributors to angiosperm evolution and diversity. Genome Biol. Evol. 5, 1886–1901 (2013).
Wen, J., Plunkett, G. M., Mitchell, A. D. & Wagstaff, S. J. The evolution of Araliaceae: a phylogenetic analysis based on ITS sequences of nuclear ribosomal DNA. Syst. Bot. 26, 144–167 (2001).
Yun, T. K. Brief introduction of Panax ginseng CA Meyer. J. Korean Med. Sci. 16, S3–5 (2001).
Obae, S. G. & West, T. P. Nuclear DNA content and genome size of American ginseng. J. Med. Plants Res. 6, 4719–4723 (2012).
Pan, Y. Z., Zhang, Y. C., Gong, X. & Li, F. S. Estimation of Genome Size of Four Panax Species by Flow Cytometry. Plant Diversity Resour. 36, 233–236 (2014).
Jayakodi, M. et al. Transcriptome profiling and comparative analysis of Panax ginseng adventitious roots. J. Ginseng Res. 38, 278–288 (2014).
Bai, D., Brandle, J. & Reeleder, R. Genetic diversity in North American ginseng (Panax quinquefolius L.) grown in Ontario detected by RAPD analysis. Genome 40, 111–115 (1997).
Ho, I. S. & Leung, F. C. Isolation and characterization of repetitive DNA sequences from Panax ginseng. Mol. Genet. Genomics 266, 951–961 (2002).
Hong, C. P. et al. Construction of a BAC library of Korean ginseng and initial analysis of BAC-end sequences. Mol. Genet. Genomics 271, 709–716 (2004).
Kim, N. H., Choi, H. I., Ahn, I. O. & Yang, T. J. EST-SSR marker sets for practical authentication of all nine registered ginseng cultivars in Korea. J. Ginseng Res. 36, 298–307 (2012).
Jayakodi, M. et al. Comprehensive analysis of Panax ginseng root transcriptomes. BMC Plant Biol. 15, 138 (2015).
Kim, K. et al. Comprehensive Survey of Genetic Diversity in Chloroplast Genomes and 45S nrDNAs within Panax ginseng Species. PloS one 10, e0117159 (2015).
Kim, K. et al. Evolution of the Araliaceae family inferred from complete chloroplast genomes and 45S nrDNAs of 10 Panax-related species. Sci. Rep. 7, 4917 (2017).
Jang, W. et al. A glimpse of Panax ginseng genome structure revealed from ten BAC clone sequences obtained by SMRT sequencing platform. Plant Breed. Biotech. 5, 25–35 (2017).
Choi, H. I. et al. Major repeat components covering one-third of the ginseng (Panax ginseng C.A. Meyer) genome and evidence for allotetraploidy. Plant J. 77, 906–916 (2014).
Smit, A., Hubley, R. & Green, P. RepeatMasker Open-4.0. http://www.repeatmasker.org. (2013–2015).
Macas, J., Neumann, P. & Navrátilová, A. Repetitive DNA in the pea (Pisum sativum L.) genome: comprehensive characterization using 454 sequencing and comparison to soybean and Medicago truncatula. Bmc Genomics 8, 427 (2007).
Waminal, N. E., Perumal, S., Lee, J., Kim, H. H. & Yang, T. J. Repeat Evolution in Brassica rapa (AA), B. oleracea (CC), and B. napus (AACC) Genomes. Plant Breed. Biotech. 4, 107–122 (2017).
Devos, K. M., Brown, J. K. & Bennetzen, J. L. Genome size reduction through illegitimate recombination counteracts genome expansion in Arabidopsis. Genome Res. 12, 1075–1079 (2002).
Neumann, P., Koblizkova, A., Navratilova, A. & Macas, J. Significant expansion of Vicia pannonica genome size mediated by amplification of a single type of giant retroelement. Genetics 173, 1047–1056 (2006).
Hawkins, J. S., Proulx, S. R., Rapp, R. A. & Wendel, J. F. Rapid DNA loss as a counterbalance to genome expansion through retrotransposon proliferation in plants. Proc. Natl. Acad. Sci. USA 106, 17811–17816 (2009).
Gill, N. et al. Dynamic Oryza genomes: repetitive DNA sequences as genome modeling agents. Rice 3, 251–269 (2010).
Levy, A. A. Transposons in Plant Speciation (ed. Fedoroff, N. V.) ch9 (John Wiley & Sons, Inc., 2013).
Piegu, B. et al. Doubling genome size without polyploidization: dynamics of retrotransposition-driven genomic expansions in Oryza australiensis, a wild relative of rice. Genome Res. 16, 1262–1269 (2006).
Renny-Byfield, S. et al. Diploidization and genome size change in allopolyploids is associated with differential dynamics of low‐and high‐copy sequences. Plant J. 74, 829–839 (2013).
Vu, G. T. H. et al. Comparative genome analysis reveals divergent genome size evolution in a carnivorous plant genus. Plant Genome 8 (2015).
Fry, K. & Salser, W. Nucleotide sequences of HS-alpha satellite DNA from kangaroo rat Dipodomys ordii and characterization of similar sequences in other rodents. Cell 12, 1069–1084 (1977).
Choi, H. I. et al. Evolutionary relationship of Panax ginseng and P. quinquefolius inferred from sequencing and comparative analysis of expressed sequence tags. Genet. Resour. Crop Evol. 60, 1377–1387 (2013).
Renny-Byfield, S. et al. Next generation sequencing reveals genome downsizing in allotetraploid Nicotiana tabacum, predominantly through the elimination of paternally derived repetitive DNAs. Mol. Biol. Evol. 28, 2843–2854 (2011).
Fedoroff, N. V. Transposable elements, epigenetics, and genome evolution. Science 338, 758–767 (2012).
Kalendar, R., Tanskanen, J., Immonen, S., Nevo, E. & Schulman, A. H. Genome evolution of wild barley (Hordeum spontaneum) by BARE-1 retrotransposon dynamics in response to sharp microclimatic divergence. Proc. Natl. Acad. Sci. USA 97, 6603–6007 (2000).
Tank, D. C. et al. Nested radiations and the pulse of angiosperm diversification: increased diversification rates often follow whole genome duplications. New Phytol. 207, 454–467 (2015).
Alzohairy, A. M. et al. Environmental stress activation of plant long-terminal repeat retrotransposons. Funct. Plant Biol. 41, 557–567 (2014).
Allen, G. C., Flores-Vergara, M. A., Krasynanski, S., Kumar, S. & Thompson, W. F. A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat. Protoc. 1, 2320–2325 (2006).
Seol, Y. J., Lee, T. H., Park, D. S. & Kim, C. K. NABIC: A New Access Portal to Search, Visualize, and Share Agricultural Genomics Data. Evol. Bioinform. Online 12, 51–58 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Waminal, N. E. & Kim, H. H. Dual-color FISH karyotype and rDNA distribution analyses on four Cucurbitaceae species. Hort. Environ. Biotechnol. 53, 49–56 (2012).
Vrana, J., Simkova, H., Kubalakova, M., Cihalikova, J. & Dolezel, J. Flow cytometric chromosome sorting in plants: the next generation. Methods 57, 331–337 (2012).
Bai, C., Alverson, W. S., Follansbee, A. & Waller, D. M. New reports of nuclear DNA content for 407 vascular plant taxa from the United States. Ann. Bot. 110, 1623–1629 (2012).
Acknowledgements
We would like to thank members in the Lab. of Functional Plants for preparation of plant material. This research was supported by the “Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ01100801)”, Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Contributions
J.L. and T.J.Y. conceived and designed the experiments. J.L., N.E.W., V.B.N., W.J. and N.H.K. prepared the samples and performed the experiments. J.L., N.E.W., H.I.C. and T.J.Y. analyzed the data. J.L., N.E.W., S.P., S.C.L., L.G. and T.J.Y. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, J., Waminal, N.E., Choi, HI. et al. Rapid amplification of four retrotransposon families promoted speciation and genome size expansion in the genus Panax . Sci Rep 7, 9045 (2017). https://doi.org/10.1038/s41598-017-08194-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-08194-5
This article is cited by
-
Genome structure and diversity among Cynanchum wilfordii accessions
BMC Plant Biology (2022)
-
A high-quality genome assembly highlights rye genomic characteristics and agronomically important genes
Nature Genetics (2021)
-
Comparative analyses of DNA repeats and identification of a novel Fesreba centromeric element in fescues and ryegrasses
BMC Plant Biology (2020)
-
Five-color fluorescence in situ hybridization system for karyotyping of Panax ginseng
Horticulture, Environment, and Biotechnology (2020)
-
Study of VIPER and TATE in kinetoplastids and the evolution of tyrosine recombinase retrotransposons
Mobile DNA (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.