Abstract
Obsessive-compulsive disorder (OCD) is a common psychiatric disorder with a lifetime prevalence of 2–3%. Recently, brain activity in the resting state is gathering attention for exploring altered functional connectivity in psychiatric disorders. Although previous resting-state functional magnetic resonance imaging studies investigated the neurobiological abnormalities of patients with OCD, there are concerns that should be addressed. One concern is the validity of the hypothesis employed. Most studies used seed-based analysis of the fronto-striatal circuit, despite the potential for abnormalities in other regions. A hypothesis-free study is a promising approach in such a case, while it requires researchers to handle a dataset with large dimensions. Another concern is the reliability of biomarkers derived from a single dataset, which may be influenced by cohort-specific features. Here, our machine learning algorithm identified an OCD biomarker that achieves high accuracy for an internal dataset (AUC = 0.81; N = 108) and demonstrates generalizability to an external dataset (AUC = 0.70; N = 28). Our biomarker was unaffected by medication status, and the functional networks contributing to the biomarker were distributed widely, including the frontoparietal and default mode networks. Our biomarker has the potential to deepen our understanding of OCD and to be applied clinically.
Similar content being viewed by others
Introduction
Obsessive-compulsive disorder (OCD) is a common psychiatric disorder with a lifetime prevalence of 2–3%1 that is characterized by obsessions (recurrent intrusive thoughts with excessive anxiety) and compulsions (excessive repetitive actions for reducing obsession-induced anxiety). Previous neuroimaging studies using structural and task-based functional magnetic resonance imaging (fMRI) have revealed neurobiological dysfunctions in OCD, most notably in the fronto-striatal circuit2,3,4,5,6. A meta-analysis of task-based fMRI studies using the symptom provocation paradigm revealed consistent increased activation within fronto-striatal regions7. Likewise, a multicenter voxel-based morphometric study revealed altered fronto-striatal gray and white matter volumes in patients with OCD8. Structural- or functional-MRI OCD classifiers constructed based on these findings have been reported9,10,11,12. Furthermore, methods for modulating the neural activity of the brain regions within the fronto-striatal circuit, such as deep brain stimulation (DBS), have been applied as clinical therapy for OCD13.
Besides structural and task-based fMRI studies, resting-state fMRI (rs-fMRI) is gathering attention as a new means of exploring altered functional connectivity (FC) in OCD14. Several studies have reported that rs-fMRI can detect differences in FC between healthy controls (HCs) and patients with OCD15,16,17,18, and find correlations with treatment response to medication19 and behavioral therapy20, 21. Furthermore, DBS reduced excessive FC within the fronto-striatal circuit, and the DBS-induced changes in FC and changes in symptom severity were correlated22.
There are two types of rs-fMRI studies in OCD: hypothesis-driven, seed-based analyses and hypothesis-free, data-driven analyses. Many of the OCD rs-fMRI studies have used seed-based FC analyses with a focus on the hypothesis of local abnormalities, especially within the fronto-striatal circuit15, 16. Recently, other data-driven studies have revealed more global abnormalities, involving a more complex combination of activity throughout the brain23, 24. The latter approach does not require an a priori hypothesis; therefore, it has the potential to quantitatively evaluate the contribution of the fronto-striatal circuit relative to other brain regions2.
Although previous rs-fMRI studies revealed neurobiological abnormalities in patients with OCD, the generalizability of these findings is still elusive. In fact, even for the most promising fronto-striatal circuit hypothesis, the findings have been inconsistent15, 16, 24,25,26; that is, the fronto-striatal circuit in OCD was hypoconnected in some studies25, 26 and hyperconnected in others15, 16, 24. Although these inconsistencies might be due to the clinical characteristics of each dataset25, 26, other studies suggested that the fronto-striatal circuit is not the only target should be considered. In other words, studies have suggested that there were abnormalities in addition to those observed in the fronto-striatal circuit, including the frontoparietal and default mode networks5, 27, 28. No study has quantitatively evaluated the relative importance of the fronto-striatal circuit relative to the whole brain. Thus, it is worth constructing a reliable FC-based “biomarker” that, with a subset of relevant FCs, allows the automatic distinction between patients with OCD and HCs. Such a biomarker may provide a novel framework in which psychiatric disorders, including OCD, are redefined biologically29. Only when such a biomarker is constructed through a fully data-driven approach, and when its validity is confirmed using independent cohorts of patients, will we be in a position to evaluate the relative importance of the fronto-striatal and other networks in the pathophysiology of OCD. Previously, an rs-fMRI study23 attempted to predict the diagnosis of OCD in a data-driven and cross-validated manner, but the generalizability of the result was not verified using an external dataset. Indeed, it is quite challenging to construct a classifier with sufficient generalizability because of two major methodological difficulties30. First, the number of subjects in an rs-fMRI dataset is usually small relative to the high dimensionality of FCs. It is a well-known problem that applying a naïve machine learning classifier to such a dataset leads to over-fitting31. Second, findings obtained from a single dataset are heavily influenced by cohort-specific features such as sex, gender, and medication, that is, nuisance variables (NVs), which may lead to catastrophic over-fitting. The status of medication such as antidepressants and anxiolytics should also be included as NVs23, so that the performance of the HC-OCD classifier is confirmed irrespective of medication status.
Here, to overcome the aforementioned issues, we aimed to construct a reliable whole-brain rs-fMRI-based biomarker using a data-driven approach. The recently developed machine learning algorithm31 employed a cascade of two statistical models: L1-norm regularized sparse canonical correlation analysis (L1-SCCA) and sparse logistic regression (SLR)32, 33. SLR can train a logistic regression model while objectively pruning features that are not useful for classifying OCD. For efficient training of SLR, however, the input dimension must be optimally reduced and the effects of NVs must be maximally removed. Therefore, before training SLR, feature selection was performed using L1-SCCA, which can identify small sets of features that contribute to only a specific variable among the NVs. For example, we can identify a small set of features that are relevant for only the diagnosis label, but not relevant for the NVs such as age or medication use. By adopting a cascade of the sparse estimation methods, our procedure leads to sparse parameters with higher generalizability, while at the same time excluding features correlating with NVs. We hypothesized that our method could be used to distinguish patients with OCD from HCs, even in an external dataset. Furthermore, its predictions were unaffected by NVs such as medication status. Finally, we quantitatively evaluated the contribution of the fronto-striatal circuit relative to other brain regions for the classification of OCD.
Results
Constructing an rs-FC-based classifier
All rs-fMRI data (N = 108) were collected at Kyoto Prefectural University of Medicine (KPUM), Kyoto, Japan. Fifty-six patients with OCD and 52 HCs were included. Table 1 summarizes the demographic data of the participants. There were 16 participants using medication (16 using antidepressants, 4 using antipsychotics, and 2 using anxiolytics). All patients were surveyed for obsessive symptoms using the Yale-Brown Obsessive-Compulsive Scale (Y-BOCS)34.
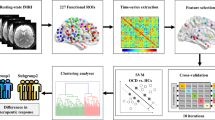
Figure 1 shows the overview of the analysis. Pairwise, interregional FC was evaluated for each participant after standard preprocessing among 140 regions of interests (ROIs) covering the entire brain. The time courses of the voxels in each ROI were averaged to extract its time course. Then, for each participant, a matrix of FC between all ROIs was calculated by evaluating pairwise temporal Pearson correlations of the time course of blood oxygenation level-dependent signals. Further, to avoid multicollinearity between the input features, we used principal component analysis (PCA) and kept all obtained principal components (PCs). This procedure enabled us to reduce the dimensionality of the input feature space from nearly 10,000 to the number of participants, thereby allowing the classifier to learn more stably. It should be noted that PCA was conducted using the whole training dataset; that is, an external dataset was not used to obtain the transformation matrix.
Schematic diagram of the procedure for selecting FCs as an OCD biomarker and assessing their predictive power. Rs-FC matrices were processed through the cascading feature selection procedure. Left-out participants and all participants in the external validation dataset were classified based on the classifier derived from the rs-FC matrix from the other participants.
To avoid problems of over-fitting due to small sample size or irrelevant NVs, we applied the method developed in our previous study (see Methods)31. We constructed the classifier by combining two machine learning algorithms: SLR and L1-SCCA32, 33. In SLR, the probability distribution of the parameter vector is estimated by hierarchical Bayesian estimation, in which the prior distribution of each element of the parameter vector is represented as a Gaussian distribution. Based on the automatic relevance determination prior in the hierarchical Bayesian estimation method, irrelevant features are not used in the classification because the respective Gaussian prior distributions have a sharp peak at zero. In L1-SCCA, latent relationships are identified between PCs and various attributes of each participant, including the diagnostic label and available demographic information. By selecting PCs connected to a canonical variable related to only the “Diagnosis” label and not to NVs, we aimed to reduce the interference of NVs. Here, we defined age, sex, handedness, and medication use (anxiolytics, antidepressants, or antipsychotics) as NVs. Our method avoids the problem of over-fitting by adopting a cascade of the sparse estimation method, a well-known approach for handling small sample sizes. Furthermore, it also avoids extracting irrelevant cohort-specific OCD features or NVs.
Reliable classifier for OCD in the training set
Leave-one-out cross-validation (LOOCV) was used to assess classification accuracy (see Methods). Participants with OCD could be separated from HCs with 73% accuracy and an area under the curve (AUC) of 0.81 (1,000-repetition permutation test, P < 0.001). Thus, the discriminatory ability of the classifier was high. The weighted linear summation (WLS or linear discriminant function) of the identified PC values of the classifier predicted the diagnostic label of each participant. Participants with a positive WLS were classified as OCD patients and those with a negative WLS as HCs. Figure 2a shows that the WLS distributions of the OCD and HC participants were separated to the right and left, respectively.
Distribution of WLSs of functional connections used for the classification of the OCD and HC populations. (a) The number of HC (white) and OCD (black) participants in the internal dataset in a specific WLS interval of width 5 is shown as a histogram. (b) WLS for the validation dataset in a specific WLS interval of width 2 is shown as a histogram.
Generalization of the classifier for the external dataset
The generalizability of the classifier was tested by using an external dataset (N = 28) collected on a different MRI scanner from that used to collect the training dataset (see Methods). We used the same dataset as Sakai et al.16. The patients were recruited at KPUM. None of the participants had been taking any kind of psychotropic medication for at least 8 weeks. Fifteen participants were entered into both experiments. In such a case, we used them in the training dataset and excluded them from the external dataset. Finally, 28 participants, including 10 patients with OCD, were used as the external dataset. Thus, there was no overlap between the training and external datasets. For this external dataset, the present classifier, trained with a different MRI scanner, performed well, with an AUC of 0.70 (1,000 repetitions permutation test, P = 0.049) (Fig. 2b). Notably, the external dataset was not involved in any part of classifier training. Therefore, the successful classification of the external dataset indicates that the developed biomarker has the ability to be generalized to a totally independent dataset.
Effects of NVs and symptom severity
Next, we investigated the effects of medication on classification accuracy. For the training set, the accuracy of LOOCV was 75% (12 of 16) and the AUC was 0.87 for patients on medication, and the accuracy of LOOCV was 67.5% (27 of 40) and the AUC was 0.79 for patients not on medication. Classification accuracy was not significantly different between the two populations (chi-squared test, P = 0.581). None of the patients in the external cohort were on medication. We also examined whether there were significant differences in age, sex, and symptom severity (Y-BOCS) between correctly and incorrectly classified patients. No significant difference was found in either the internal or external dataset in terms of age (two-sample t-test, P > 0.05), sex (chi-squared, P > 0.05), or Y-BOCS (two-sample t-test, P > 0.05).
Contribution to the WLSs of each FC
To understand how each FC contributed individually to the WLSs, their individual contributions to the WLSs through the selected PCs were calculated. As both PCA and the classifier are linear methods, the contribution of each FC can be calculated by examining the transformation matrix of PCA and the weight of the classifier. We considered 200 FCs that contributed the most to the WLSs. Figure 3a shows the spatial distribution of these 200 FCs that were identified from the dataset for the reliable classification of OCD and HC participants.
Functional connections used in the classification of the OCD and HC populations. (a) The 200 most contributing FCs from the left (left top), posterior (left bottom), and top (right) to the WLSs are visualized. (b) Matrices for the most contributing 200 FCs in 18 macroscale regions that were functionally defined in a previous study35. Diagonal and non-diagonal elements show within- and between-network FCs, respectively. The blue box highlights the corresponding area in the matrix discussed in the main text, i.e., FC between the orbitofrontal and basal ganglia-thalamus networks. The color bar indicates the number of FCs included between two networks. (c) Circle plot of the 200 most contributing 200 FCs in 18 macroscale regions. The number of FCs in each of the 2 macroscale regions is presented as the thickness of the connection lines (edges). The connections within the same network are colored blue, and connections between two different networks are colored red.
Next, to interpret their contributions in macroscale regions, all ROIs were grouped into 18 macroscale brain regions that were defined functionally in a previous study35 (e.g., the default mode network36) and examined the number of FCs between each pair of regions in each network. The networks were defined according to the datasets in “BrainMap ICA,” which identified intrinsic connectivity networks by applying independent component analysis (ICA) to BrainMap, a large-scale database of neuroimaging studies. Figure 3b shows the matrices for the 200 FCs in the macroscale regions. Diagonal and non-diagonal elements show within- and between-network FCs, respectively. Figure 3c shows a circle plot of the 200 FCs in the macroscale regions. The number of FCs in each of the two macroscale regions is presented as the thickness of the connection lines. Some trends were observed, for example, the right-lateral frontoparietal network contributed strongly relative to the other regions. However, the FCs were distributed widely rather than constrained locally. As for the FCs between the bilateral basal ganglia-thalamus and orbitofrontal cortex, only 2 FCs between the thalamus and orbitofrontal cortex were included among the 200 most contributing FCs (highlighted by the blue box in Fig. 3b). It is noteworthy that no FC between the orbitofrontal cortex and striatum was included in the 200 most contributing FCs. Note that, because this threshold (200 FCs) was arbitrary and not determined from the perspective of classification performance, we varied the threshold (100, 200, 400, and 600), and confirmed that the contribution of the FCs between the orbitofrontal cortex and striatum was still small. That is, only 0, 0, 1, and 1 FCs between the orbitofrontal cortex and striatum were selected for each respective threshold.
Discussion
A reliable neuroimaging-based classifier for OCD was developed in this study by investigating whole-brain FC patterns using rs-fMRI data. This classifier incorporated the PCs of FCs distributed across the brain, and achieved a high AUC of 0.81 with an accuracy of 73%. Further, the classifier could be generalized to an external dataset (AUC of 0.70). To our knowledge, no neuroimaging-based classifier for OCD has been shown to be generalizable using an external, independent dataset. By interpreting the classifier, we evaluated for the first time the relative contribution of the fronto-striatal and other networks in the successful classification of OCD.
We found that the FCs contributing to the classification were distributed widely rather than being locally constrained. Specifically, many of the FCs were involved in the frontoparietal or default mode network. It is noteworthy that there were relatively fewer investigations that focused on the frontoparietal and default mode networks rather than the fronto-striatal circuit. However, both seed-based17, 27, 28 and data-driven studies21, 23, 37 have reported abnormalities of these networks besides the fronto-striatal circuit. Intriguingly, frontoparietal abnormalities driving OCD pathophysiology were also suggested by a previous study38. Although this previous study found DBS-induced changes in the fronto-striatal circuit, our result suggests that DBS could trigger broad changes in the FC patterns of the whole brain. Our findings suggest that, although previous studies, including our laboratory’s, have often reported abnormalities in the fronto-striatal circuits16, other networks should also be examined in the investigation of OCD.
Although the successful construction of a structural- or functional-MRI OCD classifier has been reported previously9,10,11,12, the present study is the first to classify OCD across internal and external datasets. This was achieved because our analysis pipeline was fully data-driven and cross-validated, instead of using the seed-based analysis employed in most of the previous studies. Furthermore, we employed a cascade of sparse estimation methods by using L1-SCCA and SLR31. We were able to avoid the over-fitting problem by extracting optimal PCs that were relevant only to the core OCD characteristics. At the same time, we could eliminate the effects of NVs such as age, sex, and medication in the feature selection process. Specifically, we did not observe a clear difference in classification accuracy between patients with and without medication. Medication reportedly significantly affects rs-FC patterns39, and a naïve algorithm might over-fit the difference induced by the medication use, which leads to a reduction of generalization accuracy for non-medicated OCD patients in validated data.
The output of the OCD classifier might provide a reliable measure of an individual’s “OCD-ness” along one of the biological dimensions in psychiatric disorders, because our OCD classifier was successfully generalized to an external dataset for the first time. In the field of psychiatry, we have been unable to find any neuroscientific evidence to support the breakdown of complex psychiatric disorders into separate categories. Therefore, the hypothesis of a multiple psychiatric disorder spectrum is gaining attention40. According to this view, psychiatric disorders are the product of shared risk factors, or dimensions, that lead to abnormalities. Although the findings from brain imaging31, 41 and genetic studies42 support this idea, this hypothesis is still premature because of the scarcity of reliable dimensions. Our OCD classifier provides a biologically defined continuous index of OCD, and it can reliably separate HCs from patients with OCD. Therefore, it can be considered an objective, reliable dimension for the spectrum. Although the outputs of the biomarker were not related to a conventional clinical severity-index, our machine learning-based biomarker can be an objective and reliable complementary measure, given that the current diagnosis for this psychiatric disorder is based on a subjective report from an individual. Further studies evaluating the relationship between the classifiers of multiple psychiatric disorders are needed for a deeper understanding of psychiatric disorders and for clinical application.
A limitation of the present study is that we cannot directly compare our finding with previous studies investigating local brain regions15, 16. This is because we employed the PCs of FCs, and they represent a linear combination of whole brain FCs. This is the conventional approach in the field of machine learning to avoid the over-fitting problem when using a dataset with a small sample size and high dimensionality. In addition, unlike our previous study that employed a multi-site dataset31, all participants in the training dataset were scanned in the same site. It might also lead to difficulties with generalization at another site without PCA because of the presence of uncontrolled site-specific NVs. A future study with a much larger sample size assessed at multiple sites will investigate the contribution of each FC independently. Furthermore, in removing artifacts, we applied motion correction, NV regression, and scrubbing. Although recent studies have proposed alternative procedures for denoising (e.g., Kundu et al.43), there is still debate about the optimal denoising procedure. The development of a methodologically more appropriate alternative may lead to further improvement of classification accuracy. In addition, although classification accuracy was not significantly different (chi-squared test, P = 0.100) between the internal and external datasets, our biomarker worked better for LOOCV in the internal dataset than for the generalization to the external dataset. This is reasonable because, usually, generalization to a different dataset is more difficult than to the same dataset due to the distribution of samples or measurement noise being different from those of the training dataset. Finally, it should be noted that the sample size of the external dataset was not large (N = 28). Even though our biomarker was also verified in a fully cross-validated manner with the much larger internal dataset (N = 108), its generalizability might not be estimated with high accuracy.
In summary, we have developed the first generalizable rs-fMRI-based classifier for OCD. It reliably distinguished participants with OCD from HCs even in an independent validation dataset. Our whole-brain biomarker may shed light on the neural substrates of OCD in the form of the abnormal FC pattern across the whole brain.
Methods
Training dataset used for the construction of the OCD classifier
Participants. All resting state fMRI data (N = 108) were collected at KPUM; 69 of these participants were also included in the study of Abe et al.44. Although a few studies have tried to construct a biomarker in a fully cross-validated manner using machine learning, our sample size is larger than that of a previous study of an OCD biomarker23 (N = 46). The demographic data for all experiments are shown in Table 1. Patients with OCD were recruited at KPUM. Trained, experienced clinical psychiatrists and psychologists assessed all participants. All patients were primarily diagnosed using the Structured Clinical Interview for DSM-IV Axis I Disorders-Patient Edition (SCID)45. Exclusion criteria were 1) cardiac pacemaker or other metallic implants or artifacts; 2) significant disease, including neurological diseases, disorders of the pulmonary, cardiac, renal, hepatic, or endocrine systems, or metabolic disorders; 3) prior psychosurgery; 4) DSM-IV diagnosis of mental retardation and pervasive developmental disorders based on a clinical interview and psychosocial history; and 5) pregnancy. We excluded patients with a current DSM-IV Axis I diagnosis of any significant psychiatric illness except OCD as much as possible, and only 1 patient with trichotillomania, 1 patient with tic disorder and specific phobia, and 1 patient with bulimia nervosa were included as patients with a comorbid condition. There was no history of psychiatric illness in the control group as determined by the SCID-Non-Patient Edition46. In addition, they reported no history of psychiatric treatment in any of their first-degree relatives. Handedness was classified based on a modified 25-item version of the Edinburgh Inventory. The Medical Committee on Human Studies at KPUM approved all procedures in this study. All participants gave written, informed consent after receiving a complete description of the study. All methods were carried out in accordance with the approved guidelines and regulations.
Image acquisition. A whole-body 3-T MR system (Achieva 3.0 T Quasar Gyroscan Intera; Philips Medical Systems, Best, The Netherlands) with an 8-channel phased-array head coil at the Kajiicho Medical Imaging Center was used to generate magnetic resonance images. Functional data were collected using gradient echo planar imaging (EPI) sequences (echo time/repetition time, 30/2000 ms; flip angle, 80°; field of view, 192 mm2; imaging matrix, 64 × 64, 39 slices; slice thickness, 3.0 mm, no gaps). High-resolution (1.0 × 1.0 × 1.0 mm) T1-weighted magnetization-prepared rapid gradient echo images were also acquired before scanning the functional data. The first 6 (additional) images were discarded to allow magnetization to reach equilibrium. All participants underwent an approximately 6 min and 40 s resting-state scan, resulting in a total of 200 volumes. They were instructed simply to keep their eyes closed, not to think of anything, and not to fall asleep.
External validation dataset
Participants
We used the same dataset as Sakai et al.16. Fifteen participants were also included in the training dataset; therefore, they were excluded from the validation dataset. Finally, 28 participants were used as the external validation dataset. Thus, there was no overlap between the training and external validation datasets. Patients with a current DSM-IV Axis I diagnosis of any significant psychiatric illness except OCD were excluded. The other settings were the same as for the training dataset. The Medical Committee on Human Studies at KPUM approved all procedures in the study. All participants gave written, informed consent after receiving a complete description of the study. All methods were carried out in accordance with the approved guidelines and regulations.
Image acquisition
A whole-body 1.5-T MR system (Gyroscan Intera; Philips Medical Systems, Best, The Netherlands) with a 6-channel phased-array head coil was used to generate magnetic resonance images. Foam pads were used to reduce head motion and scanner noise. Functional data were collected using gradient EPI sequences (echo time/repetition time, 40/2411 ms; flip angle, 80°; field of view, 192 mm2; imaging matrix, 64 × 64, 35 slices; slice thickness, 3.6 mm, no gaps). High-resolution (1 × 1 × 1.5 mm) T1-weighted magnetization-prepared rapid gradient echo images were acquired before each resting image. All participants underwent an approximately 8 min resting-state scan, resulting in a total of 200 volumes. The experimental settings for the resting-state scan were the same as for the training dataset.
Preprocessing
We used a preprocessing method similar to that of Yahata et al.31 for both the training and external datasets. We used Statistical Parametric Mapping 8 (Wellcome Trust Centre for Neuroimaging, London, UK; http://www.fil.ion.ucl.ac.uk/spm/software/) in MATLAB (The MathWorks, Inc., Natick, MA) for preprocessing and statistical analyses. First, head motion was compensated for by collecting raw functional images for slice-timing and realigning them to the mean image of that sequence. Second, the structural images were co-registered to the mean functional image and segmented into 3 tissue classes in Montreal Neurological Institute (MNI) space. Using associated parameters, we normalized the functional images and resampled them in a 2 × 2 × 2 mm grid. Third, the images were smoothed by a Gaussian function with a full width at half-maximum of 6 mm. To avoid the effects of motion artifacts, the pre-processed sequence of functional images was examined as follows. First, the mean relative displacement in each of the 6 motion parameters (translation along and rotation with respect to the x, y, and z axes) was evaluated by calculating the mean of the absolute frame-to-frame relative changes in each parameter through a given time series (namely, the mean of |Δ p (i)| = |p i + 1 − p i | across the time series, where p is one of the 6 motion parameters and i specifies the time point). In both the training and external datasets, no statistically significant difference between the groups was noted in this measure for the 6 motion parameters (two-sample t-test, P > 0.05 for all parameters in both datasets). Next, frame displacement (FD) was calculated for each participant at each time point by summing all 6 parameters. Using this FD, we used the “scrubbing” procedure to identify and exclude any frame affected by excessive head motion47. Specifically, a frame was flagged and removed, along with the previous and two subsequent frames, from correlation analysis, if the associated FD exceeded 0.5 mm. For both datasets, there was no difference in the number of frames that passed this procedure between the HC and OCD populations (two-sample t-test, P > 0.05).
Interregional correlation analysis
Pairwise, interregional FC was evaluated for each participant among 140 ROIs covering the entire brain. Each region’s spatial extent was defined anatomically according to the digital atlas of the BrainVISA Sulci Atlas (BSA)48. As this atlas does not include the cerebellum, the 3 subregions of the cerebellum were appended to it based on the anatomical automatic labeling (AAL) package49. This BSA-AAL composite atlas was resampled in 2 × 2 × 2 mm grid MNI space. The time course of the voxels in each region was averaged to extract its representative time course. Further, we excluded ROIs with zero-variance in at least 1 participant. The time course sets were band-pass filtered (0.008–0.1 Hz) prior to the following regression procedure. The filtered time courses were linearly regressed by the temporal fluctuations of the white matter, cerebrospinal fluid, and entire brain as well as the 6 head motion parameters. The fluctuation in each tissue class was determined from the average time course of the voxels within a mask created by the segmentation procedure of the T1 image. The mask for the white matter was eroded by 1 voxel to consider a partial volume effect. These extracted time courses were band-pass filtered (0.008–0.1 Hz) before linear regression, as was performed for the regional time courses. Then, for each participant, a matrix of FCs between all ROIs was calculated while discarding flagged frames, if any, in the previous procedure (scrubbing). The scrubbing procedure removed any frames exhibiting abrupt head movements that could be the source of high-frequency fluctuations in the filtered time course50. The FC matrices are symmetric, so values on only one side of the diagonal were kept, resulting in the number of samples × number of FC matrices. Further, to reduce the dimensionality of the matrix from nearly 10,000 to the number of participants, we used PCA and kept all obtained PCs for the following classification analyses, resulting in the number of samples × number of PC matrices. This procedure allowed the classifier to avoid multicollinearity between the input features and to learn in a stable manner. PCA was conducted using the whole training dataset.
Selecting FCs as the OCD classifier
To avoid the problems of over-fitting because of small sample size or irrelevant NVs, we applied the method developed by Yahata et al.31. The procedure for selecting relevant PCs, training the predictive model, and assessing its generalization ability was performed as a sequential process of nested feature-selection and LOOCV. In each LOOCV fold, all-but-one participant was used to train the SLR classifier, while the remaining participants were used for evaluation. SLR can train a logistic regression model while objectively pruning PCs that are not useful for classifying OCD. For efficient training of SLR, however, the input dimension must be optimally reduced and the effects of NVs must be maximally removed. Therefore, before LOOCV, nested feature selection was performed using L1-SCCA.
Prediction of the diagnostic label
Logistic regression analysis was used as the classifier to diagnostically label from the identified PCs. A logistic function was used to define the probability of a participant belonging to the OCD class:
here, y is the diagnosis class label (OCD, y = 1; HC, y = 0), \(\hat{{\bf{z}}}\) = [z T,1]T ∈ ℝm+1 is a feature vector with an augmented input, where the feature vector z is the PCs of a participant’s rs-fMRI sample. Using the augmented input “1” is a standard approach to introduce constant (bias) input for the classifier. w ∈ ℝm+1 is the weight vector of the logistic function. To decrease the dimension of the feature vector further, which was already reduced by L1-SCCA according to the equation, we used an SLR method. SLR automatically selects OCD-classification-related features as input for the logistic function.
References
Ruscio, A., Stein, D., Chiu, W. & Kessler, R. The epidemiology of obsessive-compulsive disorder in the National Comorbidity Survey Replication. Mol. Psychiatry 15, 53–63 (2008).
Menzies, L. et al. Integrating evidence from neuroimaging and neuropsychological studies of obsessive-compulsive disorder: the orbitofronto-striatal model revisited. Neurosci. Biobehav. Rev. 32, 525–49 (2008).
Admon, R. et al. Functional and structural neural indices of risk aversion in obsessive-compulsive disorder (OCD). Psychiatry Res. 203, 207–213 (2012).
Nakamae, T. et al. Altered fronto-striatal fiber topography and connectivity in obsessive-compulsive disorder. PLoS One 9 (2014).
Hou, J. et al. Morphologic and functional connectivity alterations of corticostriatal and default mode network in treatment-naïve patients with obsessive-compulsive disorder. PLoS One 8 (2013).
Jung, W. H. et al. Abnormal corticostriatal-limbic functional connectivity in obsessive-compulsive disorder during reward processing and resting-state. NeuroImage Clin. 3, 27–38 (2013).
Rotge, J. Y. et al. Provocation of obsessive-compulsive symptoms: a quantitative voxel-based meta-analysis of functional neuroimaging studies. J. Psychiatry Neurosci. 33, 405–12 (2008).
De Wit, S. J. et al. Multicenter voxel-based morphometry mega-analysis of structural brain scans in obsessive-compulsive disorder. Am. J. Psychiatry 171, 340–349 (2014).
Soriano-Mas, C. et al. Identifying patients with obsessive-compulsive disorder using whole-brain anatomy. Neuroimage 35, 1028–1037 (2007).
Weygandt, M. et al. FMRI pattern recognition in obsessive-compulsive disorder. Neuroimage 60, 1186–1193 (2012).
Li, F. et al. Multivariate pattern analysis of DTI reveals differential white matter in individuals with obsessive-compulsive disorder. Hum. Brain Mapp. 35, 2643–2651 (2014).
Hu, X. et al. Multivariate pattern analysis of obsessive-compulsive disorder using structural neuroanatomy. Eur. Neuropsychopharmacol. 26, 246–254 (2016).
Denys, D. & Mantione, M. Deep brain stimulation in obsessive-compulsive disorder. Prog. Brain. Res. 175, 419–27 (2009).
Fox, M. D. & Greicius, M. Clinical applications of resting state functional connectivity. Front. Syst. Neurosci. 4, 19 (2010).
Harrison, B. J. et al. Altered corticostriatal functional connectivity in obsessive-compulsive disorder. Arch. Gen. Psychiatry 66, 1189–1200 (2009).
Sakai, Y. et al. Corticostriatal functional connectivity in non-medicated patients with obsessive-compulsive disorder. Eur. Psychiatry 26, 463–469 (2011).
Jang, J. H. et al. Functional connectivity in fronto-subcortical circuitry during the resting state in obsessive-compulsive disorder. Neurosci. Lett. 474, 158–162 (2010).
Vaghi, M. M. et al. Specific fronto-striatal circuits for impaired cognitive flexibility and goal-directed planning in Obsessive-Compulsive Disorder: Evidence from resting-state functional connectivity. Biol. Psychiatry 81, 708–11 (2017).
Shin, D. J. et al. The effects of pharmacological treatment on functional brain connectome in obsessive-compulsive disorder. Biol. Psychiatry 75, 606–14 (2014).
Göttlich, M., Krämer, U. M., Kordon, A., Hohagen, F. & Zurowski, B. Resting-state connectivity of the amygdala predicts response to cognitive behavioral therapy in obsessive compulsive disorder. Biol. Psychol. 111, 100–109 (2015).
Yang, X. Y. et al. Regional homogeneity of spontaneous brain activity in adult patients with obsessive-compulsive disorder before and after cognitive behavioural therapy. J. Affect. Disord. 188, 243–251 (2015).
Figee, M. et al. Deep brain stimulation restores frontostriatal network activity in obsessive-compulsive disorder. Nat. Neurosci. 16, 386–7 (2013).
Gruner, P. et al. Independent component analysis of resting state activity in pediatric obsessive-compulsive disorder. Hum. Brain Mapp. 35, 5306–15 (2014).
Beucke, J. C. et al. Abnormally high degree connectivity of the orbitofrontal cortex in obsessive-compulsive disorder. JAMA psychiatry 70, 619–29 (2013).
Posner, J. et al. Reduced functional connectivity within the limbic cortico-striato-thalamo-cortical loop in unmedicated adults with obsessive-compulsive disorder. Hum. Brain Mapp. 35, 2852–2860 (2014).
Bernstein, G. A. et al. Abnormal striatal resting-state functional connectivity in adolescents with obsessive-compulsive disorder. Psychiatry Res. 247, 49–56 (2016).
Stern, E. R., Fitzgerald, K. D., Welsh, R. C., Abelson, J. L. & Taylor, S. F. Resting-state functional connectivity between fronto-parietal and default mode networks in obsessive-compulsive disorder. PLoS One 7 (2012).
Beucke, J. C. et al. Default mode network subsystem alterations in obsessive-compulsive disorder. Br. J. Psychiatry 205, 376–382 (2014).
Yamada, T. et al. Resting-state functional connectivity-based biomarkers and functional MRI-based neurofeedback for psychiatric disorders: a challenge for developing theranostic biomarkers. arXiv:1704.01350 [q-bio.NC] 1–46 (2017).
Whelan, R. & Garavan, H. When optimism hurts: Inflated predictions in psychiatric neuroimaging. Biol. Psychiatry 75, 746–748 (2014).
Yahata, N. et al. A small number of abnormal brain connections predicts adult autism spectrum disorder. Nat. Commun. 7 (2016).
Witten, D. M., Tibshirani, R. & Hastie, T. A penalized matrix decomposition, with applications to sparse principal components and canonical correlation analysis. Biostatistics 10, 515–34 (2009).
Yamashita, O., Sato, M., Yoshioka, T., Tong, F. & Kamitani, Y. Sparse estimation automatically selects voxels relevant for the decoding of fMRI activity patterns. Neuroimage 42, 1414–29 (2008).
Goodman, W. K. et al. The Yale-Brown obsessive compulsive scale: I. Development, use, and reliability. Arch. Gen. Psychiatry 46, 1006–1011 (1989).
Laird, A. R. et al. Behavioral interpretations of intrinsic connectivity networks. J. Cogn. Neurosci. 23, 4022–37 (2011).
Fox, M. D. et al. The human brain is intrinsically organized into dynamic, anticorrelated functional networks. Proc. Natl. Acad. Sci. USA 102, 9673–8 (2005).
Tian, L. et al. Abnormal functional connectivity of brain network hubs associated with symptom severity in treatment-naive patients with obsessive-compulsive disorder: A resting-state functional MRI study. Prog. Neuro-Psychopharmacology Biol. Psychiatry 66, 104–111 (2016).
Menzies, L. et al. Neurocognitive endophenotypes of obsessive-compulsive disorder. Brain 130, 3223–3236 (2007).
Schaefer, A. et al. Serotonergic modulation of intrinsic functional connectivity. Curr. Biol. 24, 2314–2318 (2014).
Adam, D. On the spectrum. Nature 496, 6–8 (2013).
Dichter, G. S., Damiano, C. A. & Allen, J. A. Reward circuitry dysfunction in psychiatric and neurodevelopmental disorders and genetic syndromes: animal models and clinical findings. J. Neurodev. Disord. 4, 19 (2012).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: A genome-wide analysis. Lancet 381, 1371–1379 (2013).
Kundu, P. et al. Multi-Echo fMRI: A Review of Applications in fMRI Denoising and Analysis of BOLD Signals. Neuroimage 154, 59–80 (2017).
Abe, Y. et al. Hyper-influence of the orbitofrontal cortex over the ventral striatum in obsessive-compulsive disorder. Eur. Neuropsychopharmacol. 25, 1898–1905 (2015).
First, M. B., Spitzer, R. L., Miriam, G. & Williams, J. B. W. Structured Clinical Interview for Axis I DSM-IV Disorders-Patient Edition (SCID-I/P). New York: Biometrics Research Department. (New York State Psychiatric Institute, 1994).
First, M. B., Spitzer, R. L., Gibbon, M. & Williams, J. B. W. Structured Clinical Interview for DSM-IV-TR Axis I Disorders, Research Version, Non-patient Edition. (SCID-I/NP). New York: Biometrics Research Department. (New York State Psychiatric Institute, 2002).
Power, J. D., Barnes, K. A., Snyder, A. Z., Schlaggar, B. L. & Petersen, S. E. Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. Neuroimage 59, 2142–2154 (2012).
Perrot, M., Rivière, D. & Mangin, J. F. Cortical sulci recognition and spatial normalization. Med. Image Anal. 15, 529–550 (2011).
Tzourio-Mazoyer, N. et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 15, 273–89 (2002).
Griffanti, L. et al. ICA-based artefact removal and accelerated fMRI acquisition for improved resting state network imaging. Neuroimage 95, 232–247 (2014).
Acknowledgements
This research is conducted as the “Application of DecNef for development of diagnostic and cure system for mental disorders and construction of clinical application bases” of the Strategic Research Program for Brain Sciences from Japan Agency for Medical Research and development, AMED. We thank T. Okada, H. Ito, and technical engineers for their assistance in MRI data acquisition. We thank K. Tamura, S. Kimura, and K. Inoue for their assistance in the assessment of the patients. We thank the Joint Usage/Research Center at ISER, Osaka University for financial support.
Author information
Authors and Affiliations
Contributions
Y.T., Y.S., M.K., J.N., and S.C.T. conceived and designed the study; Y.S., Y.A., S.N., and T.N. recruited the participants of the study and collected their clinical and imaging data; Y.T., Y.S., L.G., N.Y., and J.M. analyzed the data; Y.T., Y.S., G.L., N.Y., Y.A., S.N., T.N., J.M., M.K., J.N., and S.C.T. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Takagi, Y., Sakai, Y., Lisi, G. et al. A Neural Marker of Obsessive-Compulsive Disorder from Whole-Brain Functional Connectivity. Sci Rep 7, 7538 (2017). https://doi.org/10.1038/s41598-017-07792-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-07792-7
This article is cited by
-
Sampling inequalities affect generalization of neuroimaging-based diagnostic classifiers in psychiatry
BMC Medicine (2023)
-
Landscape and future directions of machine learning applications in closed-loop brain stimulation
npj Digital Medicine (2023)
-
Dissecting Psychiatric Heterogeneity and Comorbidity with Core Region-Based Machine Learning
Neuroscience Bulletin (2023)
-
Depressive symptoms reduce when dorsolateral prefrontal cortex-precuneus connectivity normalizes after functional connectivity neurofeedback
Scientific Reports (2022)
-
Relationship between nuclei-specific amygdala connectivity and mental health dimensions in humans
Nature Human Behaviour (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.