Abstract
Prions are unconventional agents composed of misfolded prion protein that cause fatal neurodegenerative diseases in mammals. Prion strains induce specific neuropathological changes in selected brain areas. The mechanism of strain-specific cell tropism is unknown. We hypothesised that prion strains rely on different endocytic routes to invade and replicate within their target cells. Using prion permissive cells, we determined how impairment of endocytosis affects productive infection by prion strains 22L and RML. We demonstrate that early and late stages of prion infection are differentially sensitive to perturbation of clathrin- and caveolin-mediated endocytosis. Manipulation of canonical endocytic pathways only slightly influenced prion uptake. However, blocking the same routes had drastic strain-specific consequences on the establishment of infection. Our data argue that prion strains use different endocytic pathways for infection and suggest that cell type-dependent differences in prion uptake could contribute to host cell tropism.
Similar content being viewed by others
Introduction
Prions are proteinacious infectious agents that cause transmissible spongiform encephalopathies, fatal neurological disorders of mammals1. A misfolded, aggregated isoform (PrPSc) of the cellular prion protein (PrPC) is the major if not sole component of the infectious agent2, 3. In the central nervous system, prion deposition has been observed in association with neurons, astrocytes and microglia as well as ependymal or endothelial cells4,5,6. PrPSc molecules replicate by binding to PrPC and templating its conversion into an infectious isoform. PrPC is a conserved cell surface glycoprotein that resides within cholesterol- and sphingolipid-enriched cell surface microdomains, such as caveolae or lipid rafts7,8,9,10. PrPSc formation occurs on the cell surface and/or within intracellular vesicles following internalisation of prion particles11,12,13,14,15,16,17. The exact site(s) of prion replication and the cellular events that lead to productive infection have not been resolved.
Prions exist as strains with specific biological and biochemical properties18. In rodents with experimental prion disease, strains can be discriminated by incubation times, clinical signs and neuropathological features18. Prion strains preferentially target specific brain regions and cause characteristic lesion profiles and PrPSc deposition patterns. Prion strains differ in their cell tropism, with some strains exhibiting high tropism for astrocytes and low tropism for neurons and vice versa19. As prions lack coding nucleid acid, strain-specific information cannot be encoded within genes. Instead, differences in the high-order structure of PrPSc multimers are proposed to encipher heritable strain information20. How exactly the conformational diversity of PrPSc multimers associated with different strains relates to different disease phenotypes is unknown.
One possible explanation for cell tropism could be that strains use different cellular receptors or require different cofactors for efficient replication. While several putative prion receptors have been proposed, their roles as general prion receptors are unclear21,22,23,24. Likewise, elegant studies over the last years have identified endogenous cofactors such as phospholipids or polyanions that promote replication of certain prion strains in vitro 25, 26. It has been speculated that prion strains depend on different sets of so far unknown cofactors27. Curiously, certain cell lines can faithfully propagate different prion strains, while other cell lines are permissive to only one strain28, 29. Thus, it is possible that some cell lines lack cellular factors required by certain prion strains. Alternatively, cofactors might be confined to cellular sites to which the specific prion strain has only limited access30. Restricted access to certain subcellular compartments could be the result of different uptake mechanisms for different prion strains. It has been shown previously that the route of internalisation can have drastic consequences on the fate of a pathogen and its ability to establish infection31, 32. We therefore hypothesised that prion strains use different cellular trafficking pathways for cell invasion and productive infection.
Unfortunately, studying cellular aspects of prion replication has been notoriously difficult due to the low number of permissive cell lines and their restricted susceptibility to only certain prion strains (for a review, see ref. 33). Even in susceptible cell populations, only a limited number of cells become infected34. The difficulty to discriminate PrP isoforms as well as exogenous from de novo generated abnormal PrP further restricts detailed microscopic analysis of prion replication. Substantial efforts over the last years led to the isolation of cell sublines with increased susceptibility24, 35,36,37. Some of the most widely used cell lines in prion research to date are of non-neuronal origin, including a growing number of fibroblast cell lines28, 29, 36,37,38,39,40 and primary fibroblasts41. Importantly, recent evidence accumulates that fibroblast-like cells constitute targets of prions in vivo 42, 43. The peculiar susceptibility of fibroblasts to different prion strains makes them particularly suitable for elucidating basic principles of prion strain replication that so far have been unexplored.
In the present study, we aimed to answer if prion strains that are able to replicate in the same cell population might use different internalization pathways to establish persistent infection. Different requirements for specific endocytosis pathways could indicate that the subcellular compartments involved in establishing productive infections differ for different strains. To this end we examined the role of canonical endocytic pathways implicated in prion uptake and establishment of chronic prion infection. A L929 mouse fibroblast cell subline was chosen based on its high susceptibility to prion strains 22L and RML24, 28. Our data reveal 1) that blockage of main endocytic routes does not drastically affect prion entry, 2) that manipulation of endocytosis differentially affects the establishment of infection by different prion strains and 3) that endocytic pathways play different roles during early and late stages of prion infection. The fact that prion strains are differentially sensitive to perturbations in endocytosis during the infection process argues that prion strains rely on different endocytic routes for productive infection. This finding supports the hypothesis that selective trafficking of prion strains through specific subcellular compartments controls access to the cofactors or microenvironments required for prion replication.
Results
PrPC in prion-permissive L929 cells localizes to the cell surface, associates with detergent-resistant membranes and is endocytosed via clathrin-mediated endocytosis
Aim of this study was to test the hypothesis that different prion strains depend on different internalisation routes and might therefore differ in their requirements for subcellular compartments involved in the establishment of infection. To this end, a cell line highly susceptible to different prion strains that is amenable to manipulation of endocytosis pathways was required. We chose to perform our analysis using a murine fibroblast cell line susceptible to different prion strains. L929 cells are a well established cell culture model in prion research and have been fundamental in elucidating basic principles in prion biology24, 29, 40, 44. A clone (L929 - 15.9) of L929 mouse fibroblasts44 was selected based on its high susceptibility to mouse-adapted prion strains 22L and RML (Supplementary Table). Previous studies have demonstrated that caveolae or lipid rafts as well as clathrin-mediated endocytosis (CME) are involved in the internalization of PrPC and might also play a role in prion replication8, 45,46,47. Caveolae are a special class of rafts highly enriched in caveolins, integral membrane proteins that regulate trafficking and sorting of caveolae. Caveolin-1 (Cav-1) expression is required for caveolar biogenesis48, while clathrin-coated pits are characterised by the coat protein clathrin and mediate uptake of a diverse set of molecules from the extracellular space49. Both clathrin- and caveolin-mediated endocytosis pathways have been successfully manipulated in L929 cells to study uptake of viruses50.
In several cell lines, PrPC localizes to rafts or caveolae prior to its internalization by CME23, 51, 52. We first determined the endocytic route of PrPC in L929 cells. In uninfected L929 cells, PrPC was located on the cell surface and within small puncta in the cytosol (Fig. 1a). PrPC partially segregated with sucrose gradient fractions harboring Cav-1, but not with fractions containing clathrin heavy chain (CHC) (Fig. 1b), suggesting that in L929 fibroblasts, PrPC is localized in caveolae. Small interfering RNA (siRNA) was used to disrupt either CME or caveolae-mediated endocytosis. As expected, silencing CHC or Cav-1 expression (Fig. 1c) functionally impaired CME and caveolae-mediated endocytosis of fluorescently labelled transferrin or choleratoxin, respectively (Supplementary Fig. S1). Knock-down of CHC also significantly decreased the uptake of biotinylated cell surface PrPC (Fig. 1d, e, lower panel). No decrease in internalized PrPC was observed when Cav-1 was downregulated (Fig. 1d, e, upper panel). Thus, PrPC in L929 - 15.9 cells partially localizes in membrane environments enriched in Cav-1 and is preferentially endocytosed via clathrin-coated pits. Interestingly, both Cav-1 and CHC knock-down slightly increased total and cell surface expression levels of PrPC (Fig. 1f, g). This suggests that caveolae and clathrin might cooperate in internalizing PrPC 45, 53.
Localization and internalization of cellular PrP in L929 fibroblast cells. (a) Confocal microscopy analysis of non-permeabilized (−Triton) and permeabilized (+Triton) L929 cells. PrPC was detected using monoclonal anti-PrP antibody 4H11 (green), nuclei were stained with Hoechst (blue). Scale bar = 5 μm. (b) PrPC and Cav-1 partially co-localize in detergent-resistant membrane fractions. Lysates from L929 - 15.9 cells were applied to sucrose gradients and fractions were analyzed by western blot. (c) Knock-down of CHC and Cav-1 was determined by western blot analysis of cell lysates harvested 72 h post transfection. GAPDH serves as a loading control. (d) Cell surface biotinylation of L929 cells 72 h post transfection with Cav-1 or CHC siRNAs. Cells were chased at 37 °C for 60 min to allow PrPC internalization and either incubated with Trypsin (+) to remove cell surface proteins or left untreated (−) for evaluation of total biotinylated PrPC. PrPC in cell lysates was immunoprecipitated with mAb 4H11 and biotinylated PrP was detected using horseradish peroxidase- (HRP-) conjugated streptavidin. Bands originate from the same western blot (same acquisition settings). Additional lanes were excized for presentation purposes. (e) Quantification of the biotinylation assay displayed in (d). The amount of biotinylated internalized PrPC (+Trypsin in d) was expressed as the percentage of total biotinylated PrPC detected in control cells not treated with Trypsin (−Trypsin in d). The mean of the control cells was set to 100%. Bars represent mean values ± SD. Signals of three independent transfections/biotinylations were quantified. Statistics were performed using unpaired two-tailed t test. (***p < 0.001, ns = not significant). (f) Flow cytometry analysis of PrPC levels after siRNA-mediated knock-down of Cav-1 or CHC 72 h post transfection. Cells were either permeabilized to detect total PrP (right panels) or left intact for detection of cell surface PrP (left panels). (g) Measured mean fluorescence intensities (MFIs) in (f) were normalized to the mean of control cells set to 100%. Three independently transfected cell populations for each siRNA were measured. Bars represent mean values ± SD. Statistical analysis was performed using the unpaired two-tailed t test. Significant changes are indicated by asterisks (*p < 0.05; **p < 0.01; ***p < 0.001).
PrPSc accumulation in persistently infected cells depends on Cav-1 expression
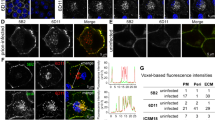
Next, we studied the role of caveolae-mediated endocytosis and CME on prion accumulation in chronically infected cells44. Subclone L929 15.9 persistently propagates prions when infected with prion strains 22L or RML (Suppl. Table). To test if PrPSc produced by cells chronically infected with either 22L (L92922L) or RML (L929RML) partitioned into similar detergent rich membranes like PrPC, we performed sucrose gradient density fractionation (Fig. 2a). Independent of the scrapie strain, PrPSc was recovered from low buoyant density fractions (fractions 4–6) that partially overlapped with Cav-1. Some overlap was also observed with CHC. Interestingly, RML PrPSc but not 22L PrPSc also segregated with very high density fractions. The difference in PrPSc partitioning with different sucrose gradient fractions suggests that the intracellular distribution of 22L and RML PrPSc slightly differs in L929 cells.
Caveolae are required for persistent PrPSc accumulation in L929 cells. (a) Sucrose gradient fractionation of cleared cell lysates of L92922L and L929RML cells. Proteins in gradient fractions were subjected to proteinase K(PK) (50 μg/ml, 45 min, 37 °C) and subsequently analyzed by western blot using mAb 4H11. Shown are representative blots of three fractionated lysates per cell line. (b) Experimental setup for transient siRNA transfection of persistently infected L929 cells. L929 cells chronically infected with 22L or RML prions were transfected with either non-silencing siRNA or with two siRNAs against Cav-1 (Cav-1.1, Cav-1.2) or CHC (CHC.1 and CHC.2). (c) L92922L or L929RML cells transfected with siRNAs against Cav-1 or non-silencing control siRNA (Ctrl) were tested for PrPSc content (+PK) after PK treatment by western blot 72 h post transfection (p.t.). GAPDH protein levels demonstrate comparable protein levels in the non-PK treated samples. For presentation purposes additional lanes were excised. (d) Same experimental setup as shown in (c) but with siRNA against CHC. (e,f) Statistical analysis of PrPSc signals 72 h post transfection with siRNA against Cav-1 (e) or CHC (f). The mean signal value of the control was set to 100%. Bars represent mean values ± SD. Statistical analysis was performed using one-way ANOVA with Dunnett’s multiple comparisons test. For each siRNA, three independent transfections were performed (*p < 0.05, **p < 0.01, ***p < 0.001). Experiments were repeated at least twice with similar results. (g,h) Statistical analysis of PrPSc signals passage 1 post transfection with siRNA against Cav-1 (g) or CHC (h) performed as above.
To study the influence of caveolae-mediated endocytosis or CME on the accumulation of PrPSc in persistently infected cells, levels of PrPSc were determined 72 h post transfection with siRNAs targeting both pathways (Fig. 2b). We chose two independent siRNAs for CHC and Cav-1 that resulted in knock-down efficiencies of at least 69.7% depending on the siRNA (Supplementary Fig. S2). Knock-down of Cav-1 expression significantly decreased PrPSc levels in both 22L or RML infected cells 72 h post transfection (Fig. 2c, e). Likewise, treatment of L92922L or L929RML cells with tyrosine kinase inhibitor Genistein, known to disrupt caveolae-dependent endocytosis, decreased the amount of PrPSc in 22L or RML infected cells in a dose-dependent manner (Supplementary Fig. S3). Knock-down of CHC had the opposite effect on PrPSc accumulation, leading to a slight increase of PrPSc 72 h post transfection (Fig. 2d, f). Several days post transfection, the effects of Cav-1 and CHC siRNA on PrPSc were even more pronounced (Fig. 2g, h). In conclusion, caveolae rather than CME were required for efficient PrPSc accumulation in chronically infected L929 cells.
Caveolin-1-, clathrin- and dynamin-2-independent uptake of PrPSc
Early events in prion biogenesis are binding of exogenous PrPSc to the cell surface and subsequent PrPSc uptake into intracellular compartments24, 54. Association of PrPSc with the cells was first assessed by western blot analysis 18 h post exposure to prion strains (Fig. 3a–d). The PrPSc signal detected in the scrapie brain homogenate treated cell populations likely represents PrPSc originating from the inoculum, as it markedly differed in size and glycosylation pattern from that of cells persistently infected with prion strains 22L or RML (Fig. 3b,c). Control experiments using L929 cells expressing 3F4 antibody epitope-tagged mouse PrP confirmed that 18 h post exposure, de novo generated PrPSc was not detectable (Supplementary Fig. S4). Substantial amounts of PrPSc were associated with cells in which Cav-1 or CHC were downregulated by siRNA (Fig. 3c). Silencing of CHC expression during 22L prion exposure led to an almost two-fold increase in cell-associated PrPSc, while no significant increase in PrPSc was observed when cells were silenced for Cav-1 (Fig. 3d, left panels). By contrast, knock-down of Cav-1 or CHC did not significantly affect the amount of cell-associated PrPSc when cells were exposed to RML prions (Fig. 3d, right panels).
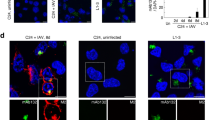
PrPSc is taken up by clathrin-, caveolin- and dynamin-independent routes. (a) Graphical illustration of the experimental setup. BH: Brain homogenate, WB: Western blot, IF: Immunofluorescence staining. (b) Comparison of brain homogenate PrPSc and PrPSc propagating in persistently infected L929 cells. Additional lanes were excised for presentation purposes. Sample L92922L was run on the same gel but had a different exposure time to avoid oversaturation. BH: Brain homogenate. Anti-PrP antibody 4H11. (c) PrPSc bound to or internalized by siRNA-transfected L929 cells 18 h post exposure to 22L or RML prions. As controls, cell lysates of L929 cells chronically infected with either 22L or RML were loaded, demonstrating different sizes and glycosylation patterns characteristic for cell culture derived PrPSc. For presentation purposes additional lanes were excised (dashed lines). PK: Proteinase K. (d) Statistical analysis of western blots shown in (c) was performed using one-way ANOVA with Dunnett’s multiple comparisons test (n = 3 biological replicates; ***p < 0.001, ns = not significant). (e) Uptake of 22L or RML PrPSc in siRNA transfected cells was monitored by immunoflourescence staining 18 h post exposure to prions. PrPSc was detected with mAb 4H11 (red) after denaturation with GdnHCl. Cav-1 and CHC were detected with the appropriate pAbs (green). Nuclei were stained with Hoechst (blue). Cells exposed to Mock brain homogenate served as controls. Every sample group was imaged with identical settings. Scale bar: 10 µm. (f) Analysis of cells with PrPSc signal. A total number of at least 100 cells was scored. (g,h) Box plot demonstrating number and size of PrPSc puncta in cells exposed to 22L (blue) and RML (red) prions upon Cav-1 or CHC siRNA transfection. Plotted is the median with whiskers displaying the 5–95% percentile. Statistical analysis was performed using the non-parametric Mann-Whitney test. Asterisks indicate significant differences between cells treated with non-silencing siRNA and siRNA directed against Cav-1 or CHC (*p < 0.05, ***p < 0.001, ns = not significant).
We performed antigen denaturation55, 56 and indirect immunofluorescence to assess the amount of internalized PrPSc following exposure to prion brain homogenate. As a control, we first tested if PrPC might serve as an uptake receptor for PrPSc. In line with previous studies33, 54, 57, 58, expression of PrPC was not required for PrPSc internalization, suggesting that prion uptake was independent of PrPC (Supplementary Fig. S5). By contrast, upon knock-down of Cav-1, L929 cells exhibited increased internal 22L PrPSc staining compared to cells transfected with non-silencing siRNA (Fig. 3e). High levels of intracellular 22L PrPSc staining were also apparent when cells were transfected with siRNA directed against CHC (Fig. 3e). Similarly, substantial amounts of RML PrPSc were taken up by cells silenced for Cav-1 or CHC expression, arguing that these endocytic pathways are not essential for PrPSc uptake (Fig. 3e). Silencing of Cav-1 slightly reduced the number of cells that had taken up RML PrPSc, while the same treatment left the uptake of 22L PrPSc unaffected (Fig. 3f). Still, PrPSc nternalization was generally highly efficient, leading to more than 70% of positive cells (Fig. 3f). Impairment of endocytic pathways influenced cellular distribution of PrPSc puncta (Fig. 3g, h). Quantitative image analysis revealed that Cav-1 and CHC siRNA transfection increased both number and size of 22L PrPSc puncta compared to control cells (Fig. 3g, h, blue box plots). By contrast, Cav-1 silencing slightly reduced the number of RML PrPSc puncta but left the size of puncta unaffected (Fig. 3f, g, red box plots, left panels). CHC silencing increased the number but not size of RML PrPSc puncta (Fig. 3g, h, red box plots, right panels).
The foregoing experiment suggested that 22L and RML prions were taken up predominately by clathrin- and caveolin-independent pathways, or that the two strains can be internalized by either one of the pathways, dependent on accessibility. Attempts to downregulate Cav-1 and CHC simultaneously were unsuccessful due to toxic effects of the siRNA treatments (data not shown). Therefore, we tested if functional impairment of dynamin-2 downstream of clathrin and Cav-1 prevents PrPSc internalization (Supplementary Fig. S6). Expression of the dominant negative dynamin-2 (Dyn K44A), a mutant that selectively and effectively inhibits dynamin-dependent endocytic pathways including both CME and caveolin-dependent uptake59, 60 (Supplementary Fig. S6a, b), was unable to block PrPSc uptake (Supplementary Fig. S6c, d). Thus, PrPSc predominantly enters L929 cells by pathways independent of clathrin-, caveolin-1 and dynamin-2, or is able to bypass these routes when impaired. The finding that different strains were differentially sensitive to manipulations of endocytic routes suggests that slight differences in the uptake mechanisms of different strains exist.
A fraction of PrPSc is internalized by macropinocytosis
The fact that CHC silencing increased PrPSc uptake prompted us to test if alternative internalization routes might be upregulated by this treatment. Macropinocytosis is a relatively non-selective, clathrin- and caveolin-independent uptake route utilised by several pathogens49, 61 and potentially also prions in some cellular models62. Control experiments confirmed that chemical inhibition of macropinocytosis by EIPA reduced the uptake of macropinocytosis reporter FITC-dextran by 50% (Supplementary Fig. S7a, b). Interestingly, siRNA-mediated knock-down of CHC significantly increased the uptake of FITC-dextran, suggesting that internalization by macropinocytosis is enhanced when CME is impaired (Supplementary Fig. S7b). Thus, the increased uptake of 22L PrPSc upon CHC siRNA transfection could be due to increased macropinocytosis rates. By contrast, knock-down of Cav-1 had no significant effect on FITC-dextran uptake (Supplementary Fig. S7b). To study if macropinocytosis is involved in PrPSc uptake, L929 cells were treated with macropinocytosis inhibitor EIPA prior to exposure to brain homogenates (Fig. 4a). EIPA treatment slightly inhibited the association of RML PrPSc with L929 cells as assessed by western blot (Fig. 4b,c). No significant difference was observed for 22L. Internalized PrPSc was subsequently detected by immunofluorescence staining (Fig. 4d). EIPA treatment only slightly affected the number of cells that had taken up 22L or RML PrPSc (Fig. 4e), but significantly reduced number (Fig. 4f) and size (Fig. 4g) of 22L PrPSc puncta per cell. The same treatment had no effect on the number of RML PrPSc puncta per cell (Fig. 4f), but slightly reduced the size of RML PrPSc puncta (Fig. 4g). These results suggest that the increased uptake of 22L PrPSc upon CHC knock-down is likely due to increased macropinocytosis. As CHC downregulation and macropinocytosis inhibitor EIPA had only small effects on RML PrPSc uptake, macropinocytosis is not essential for cell invasion by the RML scrapie strain. Importantly, however, the fact that inhibition of three major uptake routes did not drastically impair PrPSc uptake strongly argues that prions can enter cells by multiple pathways, as has been documented for other pathogens63. Alternatively, additional endocytic routes might be involved in prion uptake.
Effect of macropinocytosis inhibition on PrPSc uptake. (a) Experimental setup to study the effect of macropinocytosis inhibition on the uptake of PrPSc. L929 cells were treated with EIPA for 1 h prior to exposure to brain homogenates (BH). Note that due to toxic effects of the inhibitor upon prolonged incubation, PrPSc was detected 5 h post exposure to BH. (b) PrPSc bound to or internalized by EIPA treated L929 cells 5 h post exposure to 22L or RML prions. As controls, BH from 22L or RML infected mice were loaded. For presentation purposes, additional lanes were excized (dashed lines). PK: Proteinase K. (c) Statistical analyses of western blots shown in (b) were performed using t test (n = 3 biological replicates; *p < 0.05, ns = not significant). (d) PrPSc was detected by immunofluorescence staining using mAb 4H11 (red). Cells were analyzed by confocal microscopy with identical imaging settings. Under the conditions used, PrPC was not detected in Mock treated cells (insets). Nuclei were stained with Hoechst (blue). Scale bar: 10 µm. (e) Quantification of the relative percentages of cells that internalized 22L (blue) or RML (red) PrPSc when macropinocytosis was inhibited. Numbers were normalized to the mean of PrPSc positive cells in the siRNA controls set to 100%. Displayed are mean values ± SD (n = 3 biological replicates). (f,g) Quantification of the number and sizes of intracellular PrPSc puncta with and without EIPA treatment. Shown are box plots of the number (f) or sizes (g) of 22L (blue) or RML (red) PrPSc puncta. Plotted is the median with whiskers displaying the 5–95% percentile. Asterisks indicate significant changes between control and EIPA treated cells (*p < 0.05, ***p < 0.001, ns = not significant).
The establishment of productive prion infections involves different routes of endocytosis
The foregoing experiments showed that prion invasion could not be blocked by inhibition of the major endocytic routes. We reasoned that blockage of certain endocytic routes does not inhibit prion uptake but could still traffic PrPSc to cellular compartments that do not support prion replication. To assess this, knock-down of CHC or Cav-1 was performed at the time point of prion exposure and successful infection was monitored two to three passages post exposure to prion strains (Fig. 5a). At this time point, residual PrPSc originating from the brain homogenate is diluted out, and PrPSc detected by western blot represents de novo generated abnormal PrP (Supplementary Fig. S8). Interestingly, knock-down of Cav-1 or CHC differentially affected productive prion infections. Both 22L and RML acute infections were insensitive to Cav-1 depletion, as the de novo PrPSc levels were comparable to PrPSc levels in the non-silencing controls (Fig. 5b, c). By contrast, knock-down of CHC nearly doubled overall levels of newly formed PrPSc when cells were exposed to 22L (Fig. 5b, c). Importantly, the same treatment led to a strong and significant reduction of PrPSc in cells infected with RML (Fig. 5b, c). These results suggest that CME is not essential for the establishment of a productive 22L prion infection. By contrast, CME plays a role during the establishment of productive RML infections in L929 cells. Thus, we conclude that the establishment of productive RML and 22L prion infections depends on different endocytic routes.
Knock-down of Cav-1 and CHC during acute prion exposure differentially affects establishment of infection. (a) Experimental setup. L929 cells were transfected with Cav-1 or CHC siRNAs or non-silencing (Ctrl) siRNA and subsequently exposed to brain homogenates (BH). Cells were cultured for several cell doublings in the absence of BH and tested for PrPSc content by western blot. (b) Efficient knock-down at the time point of infection was confirmed in parallel transfections. Please note that these cells were lysed at 72 h post transfection and not used for the infection experiment. (c) Western blot analysis of PrPSc levels in cells that had been siRNA transfected and passaged 2–3 times post exposure to prions. PrPSc was detected by mAb 4H11 after PK treatment (+PK), GAPDH protein levels were detected as loading controls in −PK blots. Representative results of one of three independent experiments are shown. For presentation purposes, additional lanes were excized. (d) Statistical analysis of PrPSc signals detected by western blot shown in (c). The mean signal value in the sample group transfected with ctrl siRNA was set to 100%. Bars represent mean values ± SD. Statistical analysis was performed using one-way ANOVA with Dunnett’s multiple comparisons test. Signals from four independent transfections/infections were quantified (*p < 0.05, ***p < 0.001, ns = not significant). Experiments were repeated twice with similar results.
Discussion
Prion replication is believed to occur on the cell surface or along the endocytic pathway11,12,13. The role of canonical endocytic routes for the establishment of productive infections by different prion strains is unknown. Here we investigated how manipulation of major endocytic routes during prion uptake, the first days post infection or during chronic infection affect replication of mouse-adapted prion strains 22L and RML in L929 cells. We specifically focused on the role of caveolae-dependent endocytosis and CME for the prion infection process, as both caveolae/rafts and CME are involved in PrPC trafficking and potentially PrPSc formation23, 62. Our comparative analysis of sequential steps of cellular prion infection provides evidence that PrPSc formation, induced by different prion strains, has different requirements for endocytic trafficking. Specifically, different infection stages appear to be differentially sensitive to disruption of certain endocytic pathways. We cannot exclude that endocytic pathways active in fibroblasts differ from those of susceptible neuronal subsets in vivo. It is well established that pathogens exploit different entry pathways in different cell types and cell lines (for review see ref. 63). It is thus possible that prion strains use different cell entry pathways to establish persistent infections in different cell populations. Unfortunately, repetition of these studies in primary neurons is difficult to achieve, as these cultures are hard to transfect, sensitive to environmental perturbations64 and also contain a glial subpopulation that increases over time65. However, our finding that distinct prion strains depend on different endocytosis routes for productive infection of a given target cell population provides proof-of-concept that the subcellular compartments involved in establishment of infection can fundamentally differ for different strains.
Prion infection in vitro includes sequential steps of cell surface attachment, internalization, establishment of productive infection and sustained infection over multiple cell divisions. In analogy to other studies, we find that prions are taken up independent of PrPC expression33, 54, 57, 58. The exact mechanism of prion uptake is unknown. In our hands, blockage of caveolae-mediated endocytosis or CME only slightly affected prion internalization. Substantial amounts of PrPSc were also found internalized by cells with blocked dynamin-dependent endocytosis. These findings suggest that clathrin-, caveolin- and dynamin-dependent pathways are either dispensable for prion uptake by L929 cells or can be bypassed when blocked. Multiple entry routes have already been suggested for prions15, 66. The slight but significant differences in strain-specific PrPSc uptake indicate subtle differences in the entry routes of different strains. Similar subtle differences were recently identified to exist in the uptake of prion strains 22L and 87V in primary neurons67. Our data demonstrate that macropinocytosis played a more important role for the uptake of 22L prions compared to RML prions in L929 cells. Interestingly, uptake of 22L prions by primary neurons was shown to be more vulnerable to macropinocytosis inhibitors compared to uptake of 87V67. Macropinocytosis has been proposed as a dynamin-independent uptake mechanism for RML prions in another cellular model62. It is thus possible that the entry pathway depends on both prion strain and target cell. Macropinocytosis has recently also been reported as an uptake mechanism for other exogenous protein aggregates associated with neurodegenerative diseases68, 69. Consistent with our data, macropinocytosis inhibitors reduced superoxide dismutase aggregate uptake by only 20%, arguing that additional pathways might generally participate in the uptake of protein aggregates68.
Most studies on prion uptake have not assessed the establishment of infections22, 70,71,72,73,74. Importantly, the uptake of PrPSc does not necessarily lead to persistent infection. Acute PrPSc formation is even initiated in non-permissive cells or with strains that do not persistently propagate in cell culture28. The plasma membrane has been proposed to be the first site of prion conversion11, 12, 15,16,17, 75. While our data do not negate a possible role of the plasma membrane for initial PrPSc formation, they do point to an involvement of specific endocytic routes for the establishment of productive infections. It is thus possible that initial PrPSc formation on the cell surface is followed by strain-specific endocytosis events that route the agent through cellular compartments that enable the establishment of an infection. Importantly, different endocytic pathways appear to be required for productive infection by different prion strains. In our hands, caveolae-dependent endocytosis does not play a crucial role for the establishment of 22L or RML prion infections in L929 cells. By contrast, the establishment of an infection with RML prions depends on CME. Surprisingly, impairment of CME significantly increased infection by strain 22L, potentially by increasing the uptake of 22L prions by macropinocytosis. Due to the fact that the majority of cells still take up significant amounts of prions when specific endocytic pathways are blocked, manipulation of endocytosis likely affects downstream events in the establishment of infection. The different infection efficiencies might be the result of prions being routed to subcellular compartments that are more or less favorable for establishment of infection by a particular prion strain. The hypothesis that different prion strains preferentially propagate in different subcompartments is supported by our previous in vitro studies30 and the here identified strain-specific differences in Optiprep fractionation profiles of PrPSc propagated in L929 cells. These cellular compartments could for example contain or lack strain-specific cofactors or comprise cellular milieus required for the establishment of productive infections26, 30, 54, 76. As prion strains also vary in their resistance to proteolytic degradation77, 78, routing to the degradative pathway could have different consequences for different prion strains. However, the fact that inhibition of one endocytic pathway can increase or decrease prion infection depending on the strain argues against the idea that manipulation of endocytic pathways simply increases prion transport for lysosomal degradation. Another possibility is that differences in subcellular trafficking restrict the interaction of prions with potential strain-specific cofactors that are available in different subcellular vesicles. Phospholipids, for example, have been shown to act as cofactors for strain-specific PrPSc amplification in vitro and also vary in relative abundance along the endocytic pathway79. Which cellular cofactors catalyse prion strain replication in vivo remains to be elucidated.
Curiously, we found that acute and persistent prion infections differed drastically in their dependence on specific endocytic pathways. While caveolae-dependent endocytosis was dispensable during the first days of prion infection, downregulation of this pathway signficantly decreased PrPSc accumulation in persistently infected cells. This is in line with previous studies that demonstrated that overexpression of Cav-1 in N2a cells persistently infected with RML scrapie led to a more stable infection10. Remarkably, no strain-dependent differences were observed for chronic 22L or RML infections. The fact that manipulation of endocytic routes during acute and chronic infections has divergent effects on prion accumulation suggests that the cellular compartments for acute and persistent prion replication differ.
Our data have important implications for the observed strain tropism in vivo. Prion strains are characterised by preferential targeting of specific cell types or neuronal subpopulations in the central nervous system80. It is possible that uptake by specific entry routes directs prion particles to subcellular compartments that are favorable or less favorable for productive infection by a specific strain. But why would prions be internalized by different routes? Although several putative prion receptors have been described, their role for the establishment of prion infection and their engagement with different prion strains is unclear22, 23, 73. Importantly, PrPSc molecules associated with different strains can be distinguished mainly by their conformation20 and not by their association with different cellular proteins81. Consequently, conformational differences must impact particle association with cellular receptors or membranes and subsequent intracellular sorting. Endocytic sorting due to pleimorphic morphology of particles has recently been reported for influenza virus82. Thus, an interplay between the host cell and the physicochemical properties of prion aggregates could determine the predominant internalization pathway and access to the preferred intracellular compartments. Interestingly, immortalized brain cells have been shown to preferentially internalize PrPSc subpopulations with comparable sedimentation properties, independent of the strain’s ability to infect cells83. Thus, differences in infectivity cannot necessarily be attributed to different PrPSc subpopulations separable by conventional methods such as sucrose density gradient centrifugation. Rather, the finding that biophysically more stable PrPSc aggregates associated with a non-productive strain are more efficiently disaggregated intracellularly83 is in line with the hypothesis that prion strains are differentially trafficked through the endolysosomal system post internalization. Identification of the subcellular compartments involved in prion strain propagation will be of considerable importance for understanding the pathogenesis associated with distinct prion strains and to unravel potential targets for prion disease intervention.
Materials and Methods
Ethics statement
Use of animals in this study was performed in accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. The protocol was approved by the Rocky Mountain Laboratories Animal Care and Use Committee (protocol no. 2009-36). The Rocky Mountain Laboratories are fully accredited by the American Association for Laboratory Animal Care. In Germany, animal care was conducted according to German Law. The protocol was approved by the “Landesamt für Landwirtschaft, Lebensmittelsicherheit und Fischerei Mecklenburg-Vorpommern” (protocol no. LALLF M-V/TSD/7221.3-2.1-012/03).
Mouse bioassay
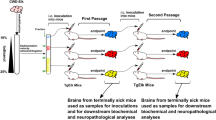
A mouse bioassay was performed with inocula prepared from L929 cell cultures exposed to mouse-adapted scrapie strains RML or 22L. Following exposure to prions, cells were split at a ratio of 1:8 every three to four days. After eleven passages cells were subsequently pelleted by centrifugation. Control cells that were exposed to uninfected Mock brain homogenate were collected the same way. Cell pellets were subjected to five freeze-thaw cycles, then adjusted to 1 × 107 cells/ml in Dulbecco’s modified Eagle’s medium (DMEM) (Invitrogen, Karlsruhe, Germany) and passaged through 20 and 24 gauge needles. Additionally, control groups were inoculated with 1% 22L, RML or Mock brain homogenate. A volume of 30 μl of cell extracts (approximately 3 × 105 cells) or brain homogenates were intracerebrally inoculated into C57BL/6 mice (group size: 15). Mice were monitored for TSE-specific clinical symptoms and euthanised upon signs of clinical disease. Incubation times were calculated as the time between the inoculation and the euthanization of the animal.
Antibodies and siRNA
Mouse monoclonal anti-PrP antibody 4H11 has been described previously84. Rabbit polyclonal anti-caveolin-1 (Cav-1) antibody was purchased from Santa Cruz Biotechnologies (Heidelberg, Germany). Mouse monoclonal antibody anti-GAPDH was obtained from Merck Millipore (Darmstadt, Germany) and rabbit polyclonal antibody anti-clathrin heavy chain (CHC) was obtained from Abcam (Cambrige, UK). GFP was indirectly stained with rabbit polyclonal anti-GFP serum purchased from Life Technologies (Darmstadt, Germany).
Cell culture
Mouse fibroblast cell line L929 (ECACC; L929 (NCTC); #85103115) was purchased from Sigma-Aldrich (Taufkirchen, Germany). Cells were maintained in DMEM containing GlutaMAX and supplemented with 10% fetal calf serum (FCS) and antibiotics. The L929 subclone 15.9, which is highly susceptible to the mouse-adapted scrapie strains 22L and RML, was used for all experiments24. This clone was derived by two rounds of limiting dilution cloning and subsequent determination of susceptibility to mouse-adapted prion strains 22L and RML. The selection of susceptible cell clones is a common and necessary method in prion research to achieve higher infection rates in vitro 29, 34,35,36.
Scrapie strains and preparation of brain homogenates
Mouse-adapted scrapie strains 22L and RML were passaged into C57BL/6 mice or the transgenic mouse line Tg(WT-E1) which overexpresses mouse PrPC containing the epitope for the anti-PrP mouse monoclonal antibody 3F485. Mice were specifically infected to produce stocks at the German TSE reference centre. For this purpose, mice inoculated with the particular strain were checked routinely for earliest signs of clinical disease. Animals were sacrificed after reaching a defined humane end point. Brains of the sacrificed mice were routinely checked biochemically to verify the quality and quantity of the deposited PrPSc. Standard operating procedures were implemented and strictly followed for preparation of brain homogenates. Brain homogenates (10% weight/volume) were prepared in OptiMEM medium by use of a dounce homogenizer by 20 strokes and cell debris was removed by low speed centrifugation (870 × g, 4 °C, 5 min). Mock control brain homogenate was prepared from uninfected C57BL/6 mice. Aliquots of brain homogenate were stored at −80 °C. Brain homogenate was chosen as the source for infectious prions as it exhibits increased activity compared to more purified PrPSc preparations in vitro 54, 86 and PrPSc is internalized faster54, 57. For uptake and de novo infection experiments, brain homogenates from at least three independent brain homogenate preparations were used, yielding similar results.
siRNA and plasmid transfections
To inhibit CME or caveolae-dependent endocytosis, small interfering RNAs (siRNAs) targeting the clathrin heavy chain (CHC) or Caveolin-1 (Cav-1) were used. SiRNAs for knock-down of Cav-1 (SI00942228 and SI02731813), CHC (SI00953617 and SI00953624) and negative control non-silencing siRNA (AllStars neg. control siRNA, 1027281) were purchased from Qiagen (Hilden, Germany). Knock-down of CHC or Cav-1 was confirmed in every experiment. For perturbation of dynamin-dependent endocytosis, L929 cells were transfected with the dominant negative Dyn2 K44A-GFP construct59, 60 or vehicle control pEGFP-N1. Transient transfections were carried out using Lipofectamine 2000 (Invitrogen, Karlsruhe, Germany) and the respective siRNAs according to the manufacturer’s instructions under RNAse free conditions.
De novo prion infection
Prion infections were performed 72 h post transfection. Cells in 24 well plates were exposed to 1% 22L, RML or Mock brain homogenates in DMEM/10% FCS for 5 h, then the brain homogenate was diluted 1:3 with growth medium. The next day the brain homogenate was discarded and the cells were subsequently cultured in fresh medium. Confluent cell monolayers were expanded (passage 1) and cells at passage 2 were tested for PrPSc content. To control for Cav-1 and CHC knock-down at the time point of infection, additional wells of L929 cells were transfected with siRNA, lysed at the time of infection and analyzed by western blot. Results were confirmed in at least three independent experiments. For binding and uptake experiments after transfection with siRNA or plasmids, cells were extensively rinsed in PBS 18 h post infection and either lysed for western blot analysis or subjected to immunofluorescence staining unless otherwise stated. For inhibition of macropinocytosis, the inhibitor EIPA (80 μM) or DMSO as a solvent control were added to the cells at 37 °C for 1 h prior to addition of brain homogenate.
Western blot analysis and PK treatment
Cells were lysed and the lysate digested with 20 µg/ml proteinase K (PK) at 37 °C for 30 min. Proteolysis was stopped by adding 0.5 mM Pefabloc. The samples were then precipitated with methanol at −20 °C and PrPSc detected by an immunoblot assay using antibody 4H11 as described previously13. For detection of PrPC, Cav-1 and CHC, lysates were not treated with PK but were supplemented with 0.5 mM Pefabloc and precipitated with methanol. Aliquots were analyzed on 12.5% SDS-PAGE gels. PrPC blots were stripped with Re-blot solution (Merck Millipore, Darmstadt, Germany) and re-probed with an antibody directed against GAPDH. For detection of CHC, samples without PK treatment were treated the same way, but analyzed on 7.5% gels. Photodensitometric analysis was performed using the ImageQuant TL (GE Healthcare, Freiburg, Germany) or AIDA Image Analyzer (Raytest, Staubenhardt, Germany). Specific protein bands were normalized to GAPDH bands of the same sample (no PK treatment) and the relative protein levels after knock-down compared to the protein signals in cells treated with non-silencing RNA.
Flow cytometry analysis
SiRNA-transfected L929 cells were subjected to flow cytometry analysis 72 h post transfection. Cells were detached from dishes using 1 mM EDTA and centrifuged at 320 × g for 2 min. Cells were resuspended in FACS buffer (PBS supplemented with 2.5% FCS and 0.05% sodium azide). For detection of cell surface PrPC all steps were performed on ice. Cells were incubated with anti-PrP monoclonal antibody 4H11 for 45 min, washed and incubated with Cy2-labelled secondary antibody (Dianova, Hamburg, Germany) for 45 min. Cells were then washed and 7-AAD dye (Beckmann Coulter, Krefeld, Germany) was added to visualize dead cells. To detect total (surface and intracellular) PrPC levels, cells were fixed with 4% Roti-Histofix (Roth, Karlsruhe, Germany) at room temperature for 10 min and stained as above at room temperature using FACS buffer supplemented with 0.1% saponin. Data acquisition was performed with a Gallios Flow Cytometer (Beckmann Coulter, Krefeld, Germany) and data were analyzed using the FlowJo software (Tree Star, Inc., Ashland, Oregon, USA). Mean fluorescence intensities (MFIs) of CHC and Cav-1 siRNA treated cells were normalized to MFIs of cells transfected with non-silencing siRNA.
Flotation assay for detergent-resistant microdomains (DRM) or rafts
Cells were washed twice with MBS pH 6.5 (25 mM MES pH 6.5, 150 mM NaCl) and lysed in 2 ml MES buffer supplemented with 1% Triton-X 100 and complete protease inhibitor cocktail on ice for 30 min. Cell suspensions were homogenized by 20 strokes of a dounce homogenizer and cell debris was pelleted. The supernatant was adjusted to a final concentration of 5% sucrose in MBS. A 5–40% discontinuous sucrose gradient was formed and overlaid with the sample. The samples were ultracentrifuged (189.000 × g, 4 °C) for 18 h in a Beckman type SW41Ti rotor. Fractions (1 ml) were collected from the top to the bottom of the ultracentrifuge tube and proteins were precipitated with 5 volumes of methanol. After centrifugation at 2.600 × g for 25 min, pellets were resuspended and equal volumes were analyzed by western blot analysis. For PrPSc detection, methanol precipitated proteins were resuspended in PBS and subjected to 50 μg/ml proteinase K for 45 min at 37 °C. Proteolysis was terminated by addtion of Pefabloc and proteins were separated as above.
Surface biotinylation assay
L929 cells were plated and transiently transfected with siRNA as described previously. At 72 h post transfection, cells were rinsed with ice cold PBS and biotinylated with 250 mg/ml Sulfo-NHS-LC-Biotin (Thermo Scientific, Schwerte, Germany) on ice for 20 min. Cells were subsequently washed with cold PBS, incubated with 20 mM Glycin/50 mM NH4Cl on ice to quench the biotinylation reaction and rinsed again with ice cold PBS. Cultures were chased at 37 °C for 60 min to allow internalization of PrPC. Cells were subsequently incubated with or without trypsin on ice for 10 min. Proteolysis was terminated by addition of soybean trypsin inhibitor. After lysis of cells, PrPC was immunoprecipitated using the monoclonal antibody 4H11 at 4 °C over night and subjected to SDS-PAGE followed by immunoblot. Biotinylated PrPC was detected using horseradish peroxidase- (HRP-) conjugated streptavidin and signals were analyzed densitometrically. The amount of internalized PrPC (+Trypsin) was expressed as a percentage of the amount of total biotinlytated PrPC detected in control cells without Trypsin treatment (−Trypsin). The mean signal value in this sample group was set to 100%.
Transferrin and cholera toxin uptake assays
L929 cells were plated on glass bottom dishes (35 mm, MatTek In Vitro Life Science Laboratories, Bratislava, Slovak Republic) and transfected with siRNA as described previously. Uptake experiments were performed 72 h post transfection. For transferrin (Tfn) uptake experiments cells were rinsed once with PBS and incubated in serum-free DMEM containing 40 µg/ml transferrin conjugated to Alexa Fluor 488 (Tfn-AF488, Molecular Probes, Thermo Fisher, Darmstadt, Germany) at 37 °C for 30 min. After incubation, cells were washed twice with PBS and living cells were imaged by confocal microscopy. To follow the uptake of cholera toxin subunit B (CtxB) upon Cav-1 silencing, cells were washed in PBS supplemented with 10 mM HEPES and incubated with 7.5 µg/ml Alexa Fluor 647-labelled cholera toxin subunit B (CtxB-AF647, Molecular Probes) at 10 °C for 30 min. Buffer was replaced with fresh HEPES-PBS and cells were further incubated at 37 °C for 40 min prior to a second extensive rinsing. Cells were subsequently imaged by confocal microscopy for CtxB uptake. Fluorescence intensities in at least 50 cells per dish were analyzed using ImageJ software (http://imagej.nih.gov/ij/). The relative uptake of Tfn and CtxB was expressed as a percentage of the signal intensities in non-silencing siRNA transfected control cells.
Fluid-phase uptake
The fluid-phase uptake assay was performed 72 h post transfection except for infection experiments where, after inhibition of macropinocytosis, cells were analyzed for FITC-dextran uptake at the time point of infection. Cells were incubated with 0.5 mg/ml 70 kDa FITC-dextran for 30 min, then placed at 4 °C and washed twice with ice cold PBS and once with 0.1 M sodium acetate (pH 5.5)/0.05 M NaCl to remove surface-bound dextran. Cells were detached with 0.25% Trypsin-EDTA on ice for 25 min and transferred into PBS containing 7% FCS to inhibit proteolysis. After low-speed centrifugation (320 × g, 2 min) cell pellets were resuspended in 3.7% paraformaldehyde (PFA) (pH 7.4), fixed on ice for 20 min and washed again. Cells in PBS were analyzed by flow cytometry. Cells treated with the macropinocytosis inhibitor EIPA (80 μM) or DMSO at 37 °C for 1 h prior to addition of FITC-dextran were used as controls for impaired fluid-phase endocytosis. As a control for the complete removal of membrane-bound FITC-dextran, additional cultures were kept at 4 °C and analyzed in parallel in every experiment.
Confocal laser scanning microscopy
Cells were prepared for immunofluorescence analysis as described previously78, with slight modifications. Briefly, cells plated on coverslips were fixed using 3.7% PFA (pH 7.4), permeabilized (0.1% Triton X-100) and proteins were denatured using 6 M guanidinium hydrochloride. Samples were rinsed with PBS, blocked in 0.2% gelatine and incubated for 2 h with primary antibody diluted in blocking solution. After three washing steps in PBS, cells were incubated for 1 h with Alexa Fluor 488- or Cy3-conjugated secondary antisera (Life Technolgies, Darmstadt, Germany; Dianova, Hamburg, Germany) and nuclei stained with Hoechst DNA staining dye (Sigma, Taufkirchen, Germany). When number and size of intracellular PrPSc puncta were analyzed, cells were also stained with HCS CellMask Blue stain (Life Technolgies, Darmstadt, Germany) according to the manufacturer’s protocol. After washing, slides were mounted in Aqua-Poly/Mount (Polysciences, Eppelheim, Germany). Confocal laser scanning microscopy was performed using a LSM 700 laser scanning microscope (Zeiss, Jena, Germany). All samples within one experiment were imaged with the same acquisition and filter settings.
Quantification of intracellular puncta after exposure to scrapie brain homogenates
Confocal images were acquired using the LSM 700 laser scanning microscope with a 63x objective. An image analysis routine was developed using the CellProfiler cell image analysis software87 to determine the number and size of intracellular PrPSc puncta per cell. Segmentation of images was performed by adjustment of CellProfiler’s object identification modules. Nuclei of single cells were identified as primary objects and the algorithm was adjusted via morphology characteristics and intensity levels of Hoechst. To determine cell boundaries, differences in the intensity levels of CellMask Blue stain were detected in the same channel as Hoechst. Cell boundaries and corresponding nuclei were used to identify single cells. Cells that were only partly imaged (cells at the boundaries of the image) were excluded from the analysis. PrPSc puncta were identified as secondary objects by Cy3 fluorescence determination. The module was carefully adjusted to morphology and intensity levels and identified puncta were linked to host cells. Mock infected cells processed in parallel were used as controls for the specific detection of PrPSc and to exclude PrPC background staining. The number of cells harboring intracellular puncta and cells without puncta were determined, and cells were grouped as PrPSc positive or PrPSc negative. The percentage of cells with intracellular puncta to total number of cells was calculated using Excel. The size of puncta was measured as the pixel area per identified object and consecutively translated in μm². On average 100 cells per group were analyzed in RNAi and macropinocytosis experiments. In uptake experiments following inhibition of dynamin, at least 40 cells per group were analyzed.
Statistical analysis
Results obtained from image analysis were tested for normal distribution and comparisons were made with non-parametric Mann-Whitney test or the unpaired two-tailed Student’s t test. Significant differences between individual biological replicates in western blots were determined using the unpaired two-tailed Student’s t-test for single comparisons or one-way ANOVA with Dunnett’s multiple comparison test using Graph Pad Prism software. P values < 0.05 were considered significant. Experiments were repeated at least twice with consistent results.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information files).
References
Aguzzi, A. & Polymenidou, M. Mammalian prion biology: one century of evolving concepts. Cell 116, 313–327 (2004).
Griffith, J. S. Self-replication and scrapie. Nature 215, 1043–1044 (1967).
Prusiner, S. B. Novel proteinaceous infectious particles cause scrapie. Science 216, 136–144 (1982).
Jeffrey, M., Goodsir, C. M., Bruce, M. E., McBride, P. A. & Scott, J. R. Infection-specific prion protein (PrP) accumulates on neuronal plasmalemma in scrapie-infected mice. Ann N Y Acad Sci 724, 327–330 (1994).
Jeffrey, M., Goodsir, C. M., Race, R. E. & Chesebro, B. Scrapie-specific neuronal lesions are independent of neuronal PrP expression. Ann Neurol 55, 781–792 (2004).
Fournier, J. G., Adjou, K., Grigoriev, V. & Deslys, J. P. Ultrastructural evidence that ependymal cells are infected in experimental scrapie. Acta Neuropathol 115, 643–650 (2008).
Taraboulos, A. et al. Cholesterol depletion and modification of COOH-terminal targeting sequence of the prion protein inhibit formation of the scrapie isoform. J Cell Biol 129, 121–132 (1995).
Vey, M. et al. Subcellular colocalization of the cellular and scrapie prion proteins in caveolae-like membranous domains. Proc Natl Acad Sci USA 93, 14945–14949 (1996).
Madore, N. et al. Functionally different GPI proteins are organized in different domains on the neuronal surface. EMBO J 18, 6917–6926 (1999).
Naslavsky, N. et al. Sphingolipid depletion increases formation of the scrapie prion protein in neuroblastoma cells infected with prions. J Biol Chem 274, 20763–20771 (1999).
Caughey, B. & Raymond, G. J. The scrapie-associated form of PrP is made from a cell surface precursor that is both protease- and phospholipase-sensitive. J Biol Chem 266, 18217–18223 (1991).
Borchelt, D. R., Taraboulos, A. & Prusiner, S. B. Evidence for synthesis of scrapie prion proteins in the endocytic pathway. J Biol Chem 267, 16188–16199 (1992).
Gilch, S. et al. Intracellular re-routing of prion protein prevents propagation of PrP(Sc) and delays onset of prion disease. EMBO J 20, 3957–3966 (2001).
Marijanovic, Z., Caputo, A., Campana, V. & Zurzolo, C. Identification of an intracellular site of prion conversion. PLoS Pathog 5, e1000426 (2009).
Goold, R. et al. Rapid cell-surface prion protein conversion revealed using a novel cell system. Nat Commun 2, 281 (2011).
Goold, R. et al. Alternative fates of newly formed PrPSc upon prion conversion on the plasma membrane. J Cell Sci 126, 3552–3562 (2013).
Rouvinski, A. et al. Live imaging of prions reveals nascent PrPSc in cell-surface, raft-associated amyloid strings and webs. J Cell Biol 204, 423–441 (2014).
Bruce, M. E., McConnell, I., Fraser, H. & Dickinson, A. G. The disease characteristics of different strains of scrapie in Sinc congenic mouse lines: implications for the nature of the agent and host control of pathogenesis. J Gen Virol 72(Pt 3), 595–603 (1991).
Gonzalez, L., Martin, S. & Jeffrey, M. Distinct profiles of PrP(d) immunoreactivity in the brain of scrapie- and BSE-infected sheep: implications for differential cell targeting and PrP processing. J Gen Virol 84, 1339–1350 (2003).
Bessen, R. A. & Marsh, R. F. Distinct PrP properties suggest the molecular basis of strain variation in transmissible mink encephalopathy. J Virol 68, 7859–7868 (1994).
Leucht, C. et al. The 37 kDa/67 kDa laminin receptor is required for PrP(Sc) propagation in scrapie-infected neuronal cells. EMBO Rep 4, 290–295 (2003).
Horonchik, L. et al. Heparan sulfate is a cellular receptor for purified infectious prions. J Biol Chem 280, 17062–17067 (2005).
Jen, A. et al. Neuronal low-density lipoprotein receptor-related protein 1 binds and endocytoses prion fibrils via receptor cluster 4. J Cell Sci 123, 246–255 (2010).
Wolf, H. et al. Modulation of Glycosaminoglycans Affects PrPSc Metabolism but Does Not Block PrPSc Uptake. J Virol 89, 9853–9864 (2015).
Deleault, N. R., Lucassen, R. W. & Supattapone, S. RNA molecules stimulate prion protein conversion. Nature 425, 717–720 (2003).
Deleault, N. R., Harris, B. T., Rees, J. R. & Supattapone, S. Formation of native prions from minimal components in vitro. Proc Natl Acad Sci USA 104, 9741–9746 (2007).
Supattapone, S. Synthesis of high titer infectious prions with cofactor molecules. J Biol Chem 289, 19850–19854 (2014).
Vorberg, I., Raines, A. & Priola, S. A. Acute formation of protease-resistant prion protein does not always lead to persistent scrapie infection in vitro. J Biol Chem 279, 29218–29225 (2004).
Mahal, S. P. et al. Prion strain discrimination in cell culture: the cell panel assay. Proc Natl Acad Sci USA 104, 20908–20913 (2007).
Vorberg, I. & Priola, S. A. Molecular basis of scrapie strain glycoform variation. J Biol Chem 277, 36775–36781 (2002).
Narayan, S., Barnard, R. J. & Young, J. A. Two retroviral entry pathways distinguished by lipid raft association of the viral receptor and differences in viral infectivity. J Virol 77, 1977–1983 (2003).
Padilla-Parra, S., Marin, M., Kondo, N. & Melikyan, G. B. Synchronized retrovirus fusion in cells expressing alternative receptor isoforms releases the viral core into distinct sub-cellular compartments. PLoS Pathog 8, e1002694 (2012).
Grassmann, A., Wolf, H., Hofmann, J., Graham, J. & Vorberg, I. Cellular aspects of prion replication in vitro. Viruses 5, 374–405 (2013).
Butler, D. A. et al. Scrapie-infected murine neuroblastoma cells produce protease-resistant prion proteins. J Virol 62, 1558–1564 (1988).
Klohn, P. C., Stoltze, L., Flechsig, E., Enari, M. & Weissmann, C. A quantitative, highly sensitive cell-based infectivity assay for mouse scrapie prions. Proc Natl Acad Sci USA 100, 11666–11671 (2003).
Raymond, G. J. et al. Inhibition of protease-resistant prion protein formation in a transformed deer cell line infected with chronic wasting disease. J Virol 80, 596–604 (2006).
Bian, J. et al. Cell-based quantification of chronic wasting disease prions. J Virol 84, 8322–8326 (2010).
Courageot, M. P. et al. A cell line infectible by prion strains from different species. J Gen Virol 89, 341–347 (2008).
Leblanc, P. et al. Co-infection with the friend retrovirus and mouse scrapie does not alter prion disease pathogenesis in susceptible mice. PLoS One 7, e30872 (2012).
Li, J., Browning, S., Mahal, S. P., Oelschlegel, A. M. & Weissmann, C. Darwinian evolution of prions in cell culture. Science 327, 869–872 (2010).
Heiseke, A., Aguib, Y. & Schatzl, H. M. Autophagy, prion infection and their mutual interactions. Curr Issues Mol Biol 12, 87–97 (2010).
Heikenwalder, M. et al. Lymphotoxin-dependent prion replication in inflammatory stromal cells of granulomas. Immunity 29, 998–1008 (2008).
Okada, H., Iwamaru, Y., Yokoyama, T. & Mohri, S. Immunohistochemical detection of disease-associated prion protein in the peripheral nervous system in experimental H-type bovine spongiform encephalopathy. Vet Pathol 50, 659–663 (2013).
Vorberg, I., Raines, A., Story, B. & Priola, S. A. Susceptibility of common fibroblast cell lines to transmissible spongiform encephalopathy agents. J Infect Dis 189, 431–439 (2004).
Sunyach, C. et al. The mechanism of internalization of glycosylphosphatidylinositol-anchored prion protein. EMBO J 22, 3591–3601 (2003).
Shyng, S. L., Lehmann, S., Moulder, K. L. & Harris, D. A. Sulfated glycans stimulate endocytosis of the cellular isoform of the prion protein, PrPC, in cultured cells. J Biol Chem 270, 30221–30229 (1995).
Ballmer, B.A. et al. Modifiers of prion protein biogenesis and recycling identified by a highly-parallel endocytosis kinetics assay. J Biol Chem (2017).
Lipardi, C. et al. Caveolin transfection results in caveolae formation but not apical sorting of glycosylphosphatidylinositol (GPI)-anchored proteins in epithelial cells. J Cell Biol 140, 617–626 (1998).
Pelkmans, L. & Helenius, A. Insider information: what viruses tell us about endocytosis. Curr Opin Cell Biol 15, 414–422 (2003).
Lin, H. Y. et al. Caveolar endocytosis is required for human PSGL-1-mediated enterovirus 71 infection. J Virol 87, 9064–9076 (2013).
Shyng, S. L., Heuser, J. E. & Harris, D. A. A glycolipid-anchored prion protein is endocytosed via clathrin-coated pits. J Cell Biol 125, 1239–1250 (1994).
Parkyn, C. J. et al. LRP1 controls biosynthetic and endocytic trafficking of neuronal prion protein. J Cell Sci 121, 773–783 (2008).
Sarnataro, D. et al. Lipid rafts and clathrin cooperate in the internalization of PrP in epithelial FRT cells. PLoS One 4, e5829 (2009).
Greil, C. S. et al. Acute cellular uptake of abnormal prion protein is cell type and scrapie-strain independent. Virology 379, 284–293 (2008).
Veith, N. M., Plattner, H., Stuermer, C. A., Schulz-Schaeffer, W. J. & Burkle, A. Immunolocalisation of PrPSc in scrapie-infected N2a mouse neuroblastoma cells by light and electron microscopy. Eur J Cell Biol 88, 45–63 (2009).
McKinley, M. P. et al. Ultrastructural localization of scrapie prion proteins in cytoplasmic vesicles of infected cultured cells. Lab Invest 65, 622–630 (1991).
Magalhaes, A. C. et al. Uptake and neuritic transport of scrapie prion protein coincident with infection of neuronal cells. J Neurosci 25, 5207–5216 (2005).
Paquet, S. et al. PrPc does not mediate internalization of PrPSc but is required at an early stage for de novo prion infection of Rov cells. J Virol 81, 10786–10791 (2007).
Cao, H., Thompson, H. M., Krueger, E. W. & McNiven, M. A. Disruption of Golgi structure and function in mammalian cells expressing a mutant dynamin. J Cell Sci 113(Pt 11), 1993–2002 (2000).
Ochoa, G. C. et al. A functional link between dynamin and the actin cytoskeleton at podosomes. J Cell Biol 150, 377–389 (2000).
Swanson, J. A. Shaping cups into phagosomes and macropinosomes. Nat Rev Mol Cell Biol 9, 639–649 (2008).
Wadia, J. S., Schaller, M., Williamson, R. A. & Dowdy, S. F. Pathologic prion protein infects cells by lipid-raft dependent macropinocytosis. PLoS One 3, e3314 (2008).
Cossart, P. & Helenius, A. Endocytosis of viruses and bacteria. Cold Spring Harb Perspect Biol 6 (2014).
Karra, D. & Dahm, R. Transfection techniques for neuronal cells. J Neurosci 30, 6171–6177 (2010).
Cronier, S., Laude, H. & Peyrin, J. M. Prions can infect primary cultured neurons and astrocytes and promote neuronal cell death. Proc Natl Acad Sci USA 101, 12271–12276 (2004).
Taylor, D. R. & Hooper, N. M. The prion protein and lipid rafts. Mol Membr Biol 23, 89–99 (2006).
Bett, C. et al. Enhanced neuroinvasion by smaller, soluble prions. Acta Neuropathol Commun 5, 32 (2017).
Munch, C., O’Brien, J. & Bertolotti, A. Prion-like propagation of mutant superoxide dismutase-1 misfolding in neuronal cells. Proc Natl Acad Sci USA 108, 3548–3553 (2011).
Wu, J. W. et al. Small misfolded Tau species are internalized via bulk endocytosis and anterogradely and retrogradely transported in neurons. J Biol Chem 288, 1856–1870 (2013).
Hijazi, N., Kariv-Inbal, Z., Gasset, M. & Gabizon, R. PrPSc incorporation to cells requires endogenous glycosaminoglycan expression. J Biol Chem 280, 17057–17061 (2005).
Schonberger, O. et al. Novel heparan mimetics potently inhibit the scrapie prion protein and its endocytosis. Biochem Biophys Res Commun 312, 473–479 (2003).
Gauczynski, S. et al. The 37-kDa/67-kDa laminin receptor acts as the cell-surface receptor for the cellular prion protein. EMBO J 20, 5863–5875 (2001).
Gauczynski, S. et al. The 37-kDa/67-kDa laminin receptor acts as a receptor for infectious prions and is inhibited by polysulfated glycanes. J Infect Dis 194, 702–709 (2006).
Morel, E. et al. Bovine prion is endocytosed by human enterocytes via the 37 kDa/67 kDa laminin receptor. Am J Pathol 167, 1033–1042 (2005).
Taraboulos, A., Raeber, A. J., Borchelt, D. R., Serban, D. & Prusiner, S. B. Synthesis and trafficking of prion proteins in cultured cells. Mol Biol Cell 3, 851–863 (1992).
Abid, K., Morales, R. & Soto, C. Cellular factors implicated in prion replication. FEBS Lett 584, 2409–2414 (2010).
Kuczius, T. & Groschup, M. H. Differences in proteinase K resistance and neuronal deposition of abnormal prion proteins characterize bovine spongiform encephalopathy (BSE) and scrapie strains. Mol Med 5, 406–418 (1999).
Gilch, S. et al. CpG and LPS can interfere negatively with prion clearance in macrophage and microglial cells. FEBS J 274, 5834–5844 (2007).
Bohdanowicz, M. & Grinstein, S. Role of phospholipids in endocytosis, phagocytosis, and macropinocytosis. Physiol Rev 93, 69–106 (2013).
Hecker, R. et al. Replication of distinct scrapie prion isolates is region specific in brains of transgenic mice and hamsters. Genes Dev 6, 1213–1228 (1992).
Moore, R. A., Timmes, A., Wilmarth, P. A. & Priola, S. A. Comparative profiling of highly enriched 22L and Chandler mouse scrapie prion protein preparations. Proteomics 10, 2858–2869 (2010).
Rossman, J. S., Leser, G. P. & Lamb, R. A. Filamentous influenza virus enters cells via macropinocytosis. J Virol 86, 10950–10960 (2012).
Choi, Y. P. & Priola, S. A. A specific population of abnormal prion protein aggregates is preferentially taken up by cells and disaggregated in a strain-dependent manner. J Virol 87, 11552–11561 (2013).
Ertmer, A. et al. The tyrosine kinase inhibitor STI571 induces cellular clearance of PrPSc in prion-infected cells. J Biol Chem 279, 41918–41927 (2004).
Chiesa, R., Piccardo, P., Ghetti, B. & Harris, D. A. Neurological illness in transgenic mice expressing a prion protein with an insertional mutation. Neuron 21, 1339–1351 (1998).
Baron, G. S., Magalhaes, A. C., Prado, M. A. & Caughey, B. Mouse-adapted scrapie infection of SN56 cells: greater efficiency with microsome-associated versus purified PrP-res. J Virol 80, 2106–2117 (2006).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol 7, R100 (2006).
Acknowledgements
We thank Daniele Bano, Walker Jackson and Eugenio Fava for critical reading of the manuscript. This work was supported by the DFG grant IV 1277/1-3 and by the Intramural Research Program of the National Institutes of Health, National Institute of Allergy and Infectious Diseases.
Author information
Authors and Affiliations
Contributions
A.F., H.W., M.G., H.M.S., and I.V. conceived the experiments. A.F., H.W., R.B., A.H., Y.D., C.P., K.R., S.L. and L.P. conducted the experiments. S.A.P. contributed novel reagents. A.F., H.W., M.G., Y.D., C.P., K.R., S.L. and I.V. analyzed the data. A.F., H.W. and I.V. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fehlinger, A., Wolf, H., Hossinger, A. et al. Prion strains depend on different endocytic routes for productive infection. Sci Rep 7, 6923 (2017). https://doi.org/10.1038/s41598-017-07260-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-07260-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.