Abstract
Bacteria naturally form communities of cells known as biofilms. However the physiological roles of biofilms produced by non-pathogenic microbiota remain largely unknown. To assess the impact of a biofilm on host physiology we explored the effect of several non-pathogenic biofilm-forming bacteria on Caenorhabditis elegans. We show that biofilm formation by Bacillus subtilis, Lactobacillus rhamnosus and Pseudomonas fluorescens induces C. elegans stress resistance. Biofilm also protects against pathogenic infection and prolongs lifespan. Total mRNA analysis identified a set of host genes that are upregulated in response to biofilm formation by B. subtilis. We further demonstrate that mtl-1 is responsible for the biofilm-mediated increase in oxidative stress resistance and lifespan extension. Induction of mtl-1 and hsp-70 promotes biofilm-mediated thermotolerance. ilys-2 activity accounts for biofilm-mediated resistance to Pseudomonas aeruginosa killing. These results reveal the importance of non-pathogenic biofilms for host physiology and provide a framework to study commensal biofilms in higher organisms.
Similar content being viewed by others
Introduction
Bacteria colonize their hosts, as they do other natural surfaces, predominantly as biofilms1, 2. In a biofilm, the encased community of bacterial cells is held in a self-produced extracellular matrix1 that serves as a physical and chemical diffusion barrier, but allows for cell differentiation and specialization3, 4. Biofilms thus offer bacteria strong competitive advantages under various environmental challenges2. For example, biofilms render bacteria less susceptible to host immunity and to antibiotics5. As biofilms contribute to the recalcitrance of chronic infections, they have been studied primarily with respect to their effects on human pathogens, such as Pseudomonas aeruginosa 5, whereas the impact of non-pathogenic biofilms on the host remains largely unknown.
The enormous complexity of the mammalian microbiota presents significant challenges to deciphering the specific mechanisms by which it influences host health and fitness. It has been difficult to establish whether changes that occur in microbiota are the cause or effect of concomitant pathologies and age-related transformations in well-controlled mouse models, and it is even more difficult in human subjects6, 7. Moreover, sessile bacteria are notoriously more difficult to study than their planktonic counterparts.
C. elegans represents a robust model for the study of host-microbiota interaction, as it is easily manipulated in the laboratory, has a short lifespan, is genetically tractable, and its colonization by bacteria can be controlled8. We used a defined model system consisting of C. elegans and B. subtilis to study bacterial biofilm-host interaction. In its natural habitat, such as decomposing plant material, C. elegans proliferates on live, actively metabolizing bacteria, including Bacillus 9,10,11 (see Supplemental Information), which serve not only as a food source but also contribute to the worm’s health and lifespan12,13,14,15. In the same natural environment Bacillus forms biofilms that have been well characterized genetically and biochemically16, 17. Moreover, it was recently reported that C. elegans fed on spores of undomesticated B. subtilis live longer and exhibit higher stress resistance as compared to worms fed on spores of a domesticated strain18. Here we demonstrate that biofilm formation by B. subtilis improves C. elegans resistance to infection and stress and prolongs lifespan. Using biofilm-deficient mutants of B. subtilis, we also identify specific C. elegans genes that confer biofilm-mediated phenotypes. Additionally, we were able to visualize biofilm formed by B. subtilis in C. elegans intestine and show that other non-pathogenic bacterial biofilms, such as those produced by L. rhamnosus and P. fluorescens, also promote stress resistance in worms.
Results
Biofilm formation by B. subtilis augments C. elegans resistance to stress
The ability to withstand stress usually indicates that an organism is in a healthy state. Thus, to investigate the effect of B. subtilis biofilm formation on C. elegans physiology we first examined thermotolerance of worms fed the undomesticated B. subtilis isolate NCBI3610 and its biofilm-deficient derivatives, ΔepsH and ΔtasA, each of which lacks a different extracellular matrix component (exopolysaccharide and amyloid-like protein fibers, respectively) essential for biofilm formation19, 20. The use of two different biofilm-deficient strains eliminates the indirect effect of each individual mutant on heat-shock resistance. Bacterial colonization of C. elegans reaches its maximum by day 5 of adulthood21, therefore we subjected day-5-adults grown on the described B. subtilis strains to lethal heat shock at 33 °C. Biofilm-forming B. subtilis significantly improved the animals’ thermotolerance as compared to biofilm-deficient bacteria (Fig. 1a). Interestingly, when worms were subjected to lethal heat-shock at earlier stages of adulthood, we did not observe any significant biofilm-dependent survival advantages (Supplementary Fig. S1a,b), indicating that the biofilm’s beneficial effect requires time for the biofilm to actually form in the animal’s intestine. Complementation of the ΔtasA mutation with a copy of tasA inserted at a distal locus (amyE) restored biofilm formation and also increased the C. elegans stress resistance to the level observed with wild-type bacteria (Fig. 1b).
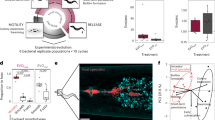
Biofilm enhances C. elegans stress resistance. Each graph represents mean values ± SD from at least free independent biological replicates. Each biological replicate was performed with at least 60 worms per condition. (a) B. subtilis biofilm increases C. elegans thermotolerance. left panel, Day-five-adult worms grown on B. subtilis NCBI3610 (biofilm) or its biofilm-deficient mutants, ΔepsH and ΔtasA, were subjected to heat shock at 33 °C for 3 hours. Surviving animals were scored after 20 hours of recovery at 20 °C. Mean values ± SD are plotted, n = 5. right panel, C. elegans were grown on B. subtilis CYBS-5 (biofilm) or its biofilm-deficient derivative (ΔepsH) until day 5 of adulthood and subjected to heat shock at 33 °C for 3 hours. Surviving animals were scored after 20 hours of recovery at 20 °C. T-test p-value = 0.0270, n = 3. (b) Complementation of B. subtilis tasA deficiency restores biofilm-mediated enhancement of C. elegans thermotolerance. Worms were grown on either biofilm-forming NCBI3610 B. subtilis (biofilm), its biofilm-deficient derivative (ΔtasA), or ∆tasA complimented with a copy of tasA inserted at a distal locus (+tasA (complement)). Five-day old adult worms were heat shocked and scored as described in (a), n = 3. (c) The Lactobacillus rhamnosus biofilm enhances worms resistance to elevated temperatures. C. elegans were grown on Lactobacillus rhamnosus GG (biofilm) or its biofilm-deficient derivative (ΔspaCBA) until day 5 of adulthood and subjected to heat shock at 36 °C for 3 hours. After 20 hours of recovery at 20 °C it was unclear if animals were dead due to residual movement, thus surviving animals were scored after 40 hours of recovery at 20 °C. T-test p-value = 0.0389, n = 3. (d) The Pseudomonas fluorescens biofilm promotes worm heat shock-resistance. C. elegans were grown on Pseudomonas fluorescens Pf0-1(biofilm) or its biofilm-deficient derivative (ΔlapA) until day 5 of adulthood and subjected to heat shock at 35 °C for 4 hours. Surviving animals were scored after 20 hours of recovery at 20 °C. T-test p-value = 0.0144, n = 3. (e) The biofilm renders C. elegans more resistant to oxidative stress. Five-day old C. elegans, grown on NCBI3610 (biofilm) or its biofilm-deficient derivative (ΔepsH) were transferred to plates supplemented with juglone (90 μM) and scored for survival every hour. For statistical data see Supplementary Table S1, n = 4. (f) B. subtilis biofilm protects against pathogenic bacteria. Five-day old worms fed NCBI3610 (biofilm) or its biofilm-deficient derivatives, ΔepsH and ΔtasA, were transferred to P. aeruginosa PA14 seeded plates and scored for survival every 6 hours. LT50, the time at which 50% of animals were scored as dead, was taken from Kaplan-Meier survival curves (median survival data). For the log-rank test p-values see Supplementary Table S2, n = 6.
To substantiate our results we analyzed another natural biofilm-forming isolate of B. subtilis, CYBS-5. Similar to the experiment with NCBI3610, worms were more susceptible to heat stress when grown on biofilm-deficient derivative of CYBS-5 (Fig. 1a, right panel), suggesting that beneficial effects of the B. subtilis biofilm were not specific to any particular bacterial genetic background.
To address if the beneficial effect of biofilm is a unique property of B, subtilis or is also a feature of other non-pathogenic bacteria, we studied thermotolerance of C. elegans fed on another Gram-positive bacterium Lactobacillus rhamnosus GG and its biofilm-deficient counterpart (ΔspaCBA) (Fig. 1c) and the Gram-negative bacterium Pseudomonas fluorescens Pf0-1 and its biofilm-deficient mutant (ΔlapA) (Fig. 1d). Both Lactobacillus and Pseudomonas were found in the C. elegans natural microbiome10, 11, 22. Lactobacillus rhamnosus were shown to be beneficial for worms23, 24 and its biofilm was recently described25. Pseudomonas fluorescens Pf0-1 in standard growth conditions is also non-pathogenic for C. elegans 26 and its biofilm formation has been studied in detail27, 28. We found that L. rhamnosus and P. fluorescens biofilms also significantly promote C. elegans heat-resistance (Fig. 1c and d).
To further investigate the potential of a biofilm to improve C. elegans stress resistance, we assayed the animals’ survival after exposure to the oxidative agent juglone. We found that day-5-adults grown on biofilm-forming B. subtilis exhibited much greater resistance to oxidative stress compared to animals fed on biofilm-deficient bacteria (Fig. 1e and Supplementary Table S1). These results demonstrate that the biofilm produced by non-pathogenic bacterial is beneficial for host stress resistance.
Biofilm formation by B. subtilis renders C. elegans resistant to pathogenic infection
Commensal bacteria are known to protect mammalian hosts from colonization by opportunistic pathogens29. To determine whether biofilm-producing B. subtilis can also reduce susceptibility to pathogenic infection, we examined the survival of C. elegans exposed to P. aeruginosa. We used the slow killing assay in which worms die from intestinal pathogenic colonization26. We grew worms on B. subtilis until day 5 of adulthood and then transferred them to agar plates seeded with P. aeruginosa PA14. Worms that were fed biofilm-forming bacilli displayed a much greater resistance to P. aeruginosa infection than did those that were fed biofilm-deficient strains (Fig. 1f and Supplementary Table S2).
Recently, it was suggested that spores of biofilm-forming B. subtilis can germinate and colonize the C. elegans intestine18. Also another wild isolate of B. subtilis GS67 can accumulate in C. elegans gut and protect animals from Gram-positive infection12. To determine whether the protective effect of biofilm-forming bacteria against lethal infection by pathogenic P. aeruginosa was due to their superior intestinal retention, we compared gut colonization by wild-type and biofilm-deficient B. subtilis, and found, by either colonization or guillotine assays, no significant difference in the number of cells of each B. subtilis strain in the gut lumen (Supplementary Fig. S2a and b). These results suggest that commensal biofilm-producing B. subtilis may not compete directly with pathogenic Pseudomonas in the C. elegans gut, but rather mobilize host defense system(s) against the pathogen.
Biofilm formation by B. subtilis extends the C. elegans lifespan
Higher resistance to stress and pathogens often correlate with lifespan extension14, 30. To examine the potential benefit of the B. subtilis biofilm we measured its effect on C. elegans lifespan. The biofilm-deficient ΔepsH and ΔtasA mutants reduced median lifespan of C. elegans on average by, respectively, 20.5% and 15.1% as compared with the biofilm-producing parental strain (Fig. 2a and Supplementary Fig. S3a and b and Supplementary Table S3; Supplementary Discussion). Other physiological aspects of C. elegans, such as post-embryonic development and rate of egg production were not significantly affected (Supplementary Fig. S3c). Complementation of the ΔtasA mutation with a wild type copy of tasA restored C. elegans longevity to a wild-type biofilm level (Supplementary Fig. S3b and Supplementary Table S3), confirming that the bacterial effect on the worms’ lifespan was biofilm-specific.
B. subtilis biofilm extends C. elegans lifespan independently of caloric restriction. (a) C. elegans N2 were fed either biofilm-forming B. subtilis NCBI3610 (biofilm) or its biofilm-deficient derivatives (ΔepsH or ΔtasA). The graph is representative of three independent biological replicates. Median lifespan, days: biofilm–19; ΔepsH–16; ΔtasA–16. The log-rank test p-value for each experiment is ≤0.05. Also see Supplementary Table S3. (b) Worms fed on biofilm-producing B. subtilis exhibit slower age-associated decline in motility. The graph represents linear regression of the median motility values of day-2 and day-8-adults grown of biofilm-forming (biofilm) or biofilm-deficient (ΔepsH, ΔtasA) B. subtilis. The median values were calculated as described in Supplemental Fig. S3d and mean values ± SD from 3 independent experiments are plotted. (c) The biofilm prolongs lifespan of dietary restricted worms DA1116 (eat-2). Median lifespan, days: biofilm–34; ΔepsH–29; ΔtasA–28. The graph is representative of three independent biological replicates. The log-rank test p-value is ≤0.05, Supplementary Table S3. (d) pha-4 is induced in response to dietary restriction, but not by the B. subtilis biofilm. RT-PCR analysis of pha-4 expression in C. elegans eat-2 (dashed bars) and N2 (empty bars) fed biofilm-forming (biofilm) and biofilm deficient (ΔepsH, ΔtasA) B. subtilis strains. Mean ± SD from three independent experiments are plotted, n = 100 per experiment per condition. One-way ANOVA: p-value = 0.6696 for N2 worms; p-value = 0.1945 for eat-2 worms.
To assess the impact of the biofilm on C. elegans healthspan we also studied its effect on motility. The rate of decline in motor activity and longevity are inversely correlated in worms31,32,33. We therefore measured the rate of decline in motility of animals grown on biofilm-forming and biofilm-deficient B. subtilis between day 2 and 8 of adulthood (Fig. 2b and Supplementary Fig. S3d). To avoid a possible bacterial strain-specific behavioral bias affecting C. elegans locomotion on the plate (either with or without food) we used a thrashing assay to study worm motility (Supplementary Fig. S3d). We found that biofilm formation slows down the age-associated decline in the C. elegans thrashing rate (Fig. 2b). These results, taken together with the beneficial effects of the biofilm on worm stress-resistance and lifespan, suggest that biofilm formation by a non-pathogenic bacteria improves the C. elegans healthspan.
Biofilm-mediated effects do not entail dietary restriction
The nutritional status of bacteria as a food source is an important parameter that affects worm physiology15, 34. If biofilm formation diminishes the dietary value of B. subtilis cells, feeding C. elegans wild-type bacilli may induce a calorie restriction-like response, which is known to enhance lifespan and stress resistance35, 36. Moreover, domesticated B. subtilis 168 sporulates very efficiently on NGM, making it a poor nutritional source for C. elegans 13, 37. To address the nutritional issue, we first determined that the sporulation rates of undomesticated wild-type and its biofilm-deficient derivative are similar: both maintained over 70% vegetative bacteria under conditions used in our experiments (Supplementary Fig. S4a). We next studied the lifespan of an eat-2 genetic mimetic of dietary restricted worms (DA1116). Biofilm-deficient B. subtilis decreased the lifespan of eat-2 worms to about the same extent as that of N2 worms (Fig. 2c, Supplementary Table S3), arguing that caloric restriction is not a potential cause of biofilm-mediated phenotypes. We also showed that the mRNA levels of the pha-4 transcription factor, which increases upon dietary restriction38, do not change in response to the biofilm (Fig. 2d). Moreover, the biofilm did not affect post-embryonic development (Supplementary Fig. S3c), animal size (Supplementary Fig. S4b and Supplementary Table S4) or the rate of pharyngeal pumping (Supplementary Fig. S4c). Together these results argue that the worms do not undergo caloric restriction in response to biofilm-forming B. subtilis.
The biofilm exerts its effects from within the C. elegans intestinal lumen
To confirm that B. subtilis was forming biofilm inside C. elegans we first used a strain that expresses the red fluorescent protein mKate2 under the control of a biofilm-specific promoter (P tasA -mKate2). Worms grown on B. subtilis P tasA -mKate2 exhibited red fluorescence in their intestinal lumen (Fig. 3a), indicating that biofilm matrix genes were being expressed. Moreover, mutation in the upstream regulator of biofilm synthesis sinI (B. subtilis PtasA-mKate2 ΔsinI) completely abrogated fluorescence in the intestine, demonstrating the specificity of the fluorescent signal to the induction of extracellular matrix genes inside C. elegans (Fig. 3a and b). To visualize biofilm formation, we also stained for biofilm exopolysaccharide in the gut of 5-day old worms with FITC-conjugated wheat germ agglutinin (WGA-FITC), a lectin. WGA-FITC was previously shown to bind to exopolysaccharides produced by Yersinia pestis, Yersinia pseudotuberculosis, and Staphylococcus epidermidis biofilms formed in association with C. elegans 39, 40. Also, WGA specifically binds to poly-N-acetylglucosamine, which is the major constituent of the B. subtilis biofilm matrix41. We therefore used it to label B. subtilis biofilm within C. elegans. As shown in Fig. 3c and d, WGA-FITC produced a strong signal in the presence of B. subtilis biofilms. Stained exopolysaccharides were detected only in the intestine of the worms fed with wild type B. subtilis, while in worms grown on ΔepsH we observed only minor intestinal staining due to the binding of WGA-FITC to worms’ glycocalyx42, and possibly due to weak binding to the B. subtilis cell wall.
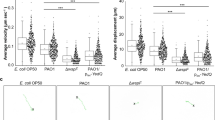
Beneficial effects likely require intestinal biofilm production. (a) Induction of biofilm matrix gene expression in the C. elegans intestine. Representative image of worms grown on the indicated B. subtilis strains were visualized at day 5 of adulthood. The red fluorescent signal indicates tasA expression in B. subtilis cells. (b) Fluorescence quantification of five-day old adults (n = 15) grown on corresponding bacterial strain. (c) Biofilm matrix exopolysaccharides are present in the C. elegans intestinal lumen. Representative image of FITC-conjugated wheat germ agglutinin (WGA-FITC) stained C. elegans grown on the indicated B. subtilis strains at day 5 of adulthood. Green fluorescence indicates biofilm matrix. (d) Fluorescence quantification of adult worms (n = 15) grown on corresponding bacterial strains until day 5 of adulthood and stained with WGA-FITC. Error bars show means ± SD from three independent experiments. (e) The anti-aging effect of biofilm requires live metabolizing bacteria. B. subtilis strains were grown on NGM plates overnight at 20 °C and then treated with a mixture of antibiotics (100 μg/ml kanamycin and 500 μg/ml carbenicillin). The graph is representative of three independent biological replicates. Median lifespan, days: biofilm–21; ΔepsH–21; ΔtasA–21. The log-rank test p-value for each experiment is >0.05. Supplementary Table S3. (f) Biofilm-induced thermotolerance requires live bacteria. B. subtilis lawns were treated with a mixture of antibiotics (100 μg/ml kanamycin and 500 μg/ml carbenicillin) prior to transferring C. elegans to them. Five-day old adult worms were heat shocked as in (a) for the indicated period of time. Each experiment includes at least 60 worms per condition per experiment. Mean values ± SD are plotted, n = 5. One-way ANOVA: p-value = 0.8759 (3 h), p-value = 0.9310 (4 h).
To obtain independent visual support for biofilm formation in C. elegans, we examined worms fed with biofilm-producing B. subtilis (biofilm) or its ΔepsH mutant by electron microscopy and found vegetative cells surrounded by a dense matrix only in animals fed on biofilm-producing B. subtilis (Supplementary Fig. 5). Taken together, our data demonstrate that B. subtilis forms biofilms in the C. elegans intestine.
To determine whether the observed beneficial effects of the biofilm require live, metabolizing bacteria, we treated plated bacteria with a mix of antibiotics to eliminate vegetative cells, leaving previously-synthesized components of the biofilm intact (see Supplementary Discussion). Elimination of live bacteria abrogated all the positive effects of the biofilm on lifespan and thermotolerance (Fig. 3e and f and Supplementary Table S3). These results argue that it is not the extracellular matrix per se, but the metabolic state of biofilm-forming bacteria that benefits the worms.
MTL-1 induction accounts for biofilm-mediated longevity and stress resistance
To understand the mechanism underlying the effects of the biofilm on worm physiology we determined the changes it causes in C. elegans gene transcription. Total mRNA sequencing revealed that a small group of C. elegans genes were differentially expressed in response to biofilm-forming vs. biofilm-deficient B. subtilis (Supplementary Table S5), demonstrating a specific response to the biofilm. To focus on bona fide biofilm-dependent changes, we assessed mRNA levels of identified genes in worms grown on the two biofilm-deficient mutants versus those in worms grown on wild-type B. subtilis (Supplementary Fig. S6). We also focused only on those genes that showed the same reproducible changes in response to both biofilm-deficient mutants used in our study (Supplementary Fig. S6, marked with asterisks). Based on supportive evidence from Real-Time PCR analyses and the relevance to observed C. elegans phenotypes, we selected four genes, mtl-1, ilys-2, F44E5.4 and F44E5.5 for in-depth analysis (Table 1) (see Supplementary Discussion).
mtl-1 encodes metallothionein–a small cysteine-rich metal-binding protein that has been implicated in longevity and proteotoxic stress43. It is regulated by DAF-16, a FOXO transcription factor broadly involved in lifespan regulation44, 45. Indeed, biofilm-producing bacteria failed to extend the lifespan of either mtl-1 or daf-16 mutant worms (Fig. 4a and Supplementary Fig. S7a), indicating that MTL-1 plays a primary role in biofilm-mediated longevity.
Biofilm acts via induction of metallothionein in C. elegans. In each case the average values ± SD from at least three independent experiments are plotted. Each experiment includes at least 60 worms per condition. (a) MTL-1 is required for biofilm-dependent lifespan extension. C. elegans mtl-1 (tm1770) were fed either biofilm-forming B. subtilis (biofilm, blue) or its biofilm-deficient derivatives (ΔepsH and ΔtasA, red and green, respectively). The graph is representative of three independent biological replicates. Median lifespan, days: biofilm–15; ΔepsH–15; ΔtasA–15. The log-rank test p-value for each experiment is >0.05, Supplementary Table S3. (b) MTL-1 is required for biofilm-mediated thermotolerance. Wild-type (N2) and mtl-1 (tm1770) C. elegans were grown in parallel on indicated bacterial strains until day 5 of adulthood and then subjected to heat shock at 33 °C for 3 hours. Surviving animals were scored after 20 hours of recovery at 20 °C. The graph represents mean values ± SD from free independent biological replicates. One-way ANOVA p-value = 0.4800. (c) MTL-1 is required for biofilm-mediated resistance to oxidative stress. Five-day old wild-type (N2) and mtl-1 (tm1770) C. elegans were grown on indicated bacterial strains, transferred to plates supplemented with juglone (90 μM) and scored for survival every hour. The graph represents mean values ± SD from free independent biological replicates. Also see Supplemental Table S1.
MTL-1 is known to be upregulated by elevated temperature46, although its role in the heat shock response remains unclear. We found that the biofilm-mediated increase in thermotolerance was abrogated in mtl-1 knockout worms (Fig. 4b), thus implicating MTL-1 in biofilm-mediated heat resistance. Curiously, this effect on thermotolerance was independent of DAF-16 (Supplementary Fig. S7b), suggesting alternative modalities of mtl-1 regulation in response to the biofilm.
Mammalian metallothioneins function in the oxidative stress response47. We, therefore, examined whether the observed upregulation of MTL-1 by biofilm-forming B. subtilis improves C. elegans oxidative stress resistance. Indeed, mtl-1 knock-out worms were not protected from juglone by biofilm-forming bacteria (Fig. 4c and Supplementary Table S1).
To further support the hypothesis that MTL-1 plays the principle role in biofilm-mediated effects we studied a C. elegans mutant that overexpresses mtl-1 (mtl-1 OE) (Fig. 5a). In the background of higher mtl-1 expression, the B. subtilis biofilm failed to promote worm thermotolerance or oxidative stress resistance (Fig. 5b and c).
mtl-1 overexpression abrogates biofilm-mediated effects. In each case the average values ± SD from at least three independent experiments are plotted. Each experiment includes at least 60 worms per condition. (a) Relative levels of mtl-1 expression. The relative level of mtl-1 mRNA was calculated as the fold change from the expression level in C. elegans N2 fed on wild-type B. subtilis (biofilm) using RT-PCR. (b) The B. subtilis biofilm fails to increase the thermotolerance of C. elegans overexpressing mtl-1. Five-day old mtl-1 OE (WU1394) grown on B. subtilis NCIB3610 (biofilm) or its biofilm-deficient derivative (ΔepsH) were heat shocked at 33 °C for 5 hours. The duration of heat shock was increased from 3 to 5 hours because 100% of mtl-1 OE worms survived a 3 hour heat-shock treatment regardless of the bacterial strain. Surviving animals were scored after 20 hours of recovery at 20 °C. One-way ANOVA p-value = 0.2877. (c) The B. subtilis biofilm fails to promote resistance to oxidative stress in worms overexpressing mtl-1. Five-day old mtl-1 OE (WU1394) grown on B. subtilis NCBI3610 (biofilm) or its biofilm-deficient derivative (ΔepsH) were transferred to plates supplemented with juglone (90 μM) and scored for survival every hour. See Supplementary Table S1.
Taken together, these results demonstrate that upregulated MTL-1 expression accounts for biofilm-associated C. elegans phenotypes, including extended lifespan and increased resistance to physical and chemical stress.
Induction of HSP70 by a commensal biofilm improves C. elegans heat resistance
F44E5.4 and F44E5.5 encode two isoforms of inducible heat shock protein 70 (HSP70), a family of evolutionarily-conserved molecular chaperones that are crucial for survival at elevated temperatures48. The F44E5.4 and F44E5.5 genes have overlapping promoters from which transcription proceeds in divergent directions49. Their transcription is, at least partially, controlled by HSF-1, a master regulator of the heat-shock response50 that has been shown to contribute to C. elegans lifespan extension51. We investigated whether the B. subtilis biofilm improves thermotolerance and increases lifespan via HSF-1-dependent activation of heat shock proteins. Thermotolerance and lifespan assays in an hsf-1 mutant showed that HSF-1 contributes to biofilm-activated resistance to heat-shock, but not to biofilm-induced lifespan extension (Supplementary Fig. S7d and c). We next examined the effect of the biofilm on the thermotolerance of F44E5.4/F44E5.5 double mutant worms and found that the biofilm was unable to improve their thermotolerance (Fig. 6a). Thus, our results demonstrate that the commensal biofilm activates HSF-1, which in turn leads to induction of specific hsp70 genes and, as a result, causes increased resistance to elevated temperatures.
HSP70 and lysozyme contribute to biofilm-mediated resistance to heat and infection. (a) C. elegans hsp-70 (F44E5.4 and F44E5.5) are required for biofilm-induced thermotolerance. Wild-type (N2) and hsp70-deficient C. elegans were grown in parallel on biofilm-forming (biofilm) and biofilm-deficient (ΔepsH, ΔtasA) B. subtilis. Five-day old worms were subjected to heat shock as in Fig. 1a. Mean values ± SD from three independent biological replicates are plotted. Each biological replicate includes at least 60 worms per condition. One-way ANOVA p-value = 0.1210. (b) ILYS-2 is dispensable for the biofilm-mediated increase in C. elegans thermotolerance. N2 and ilys-2 worms were grown in parallel on indicated bacterial strains until day 5 of adulthood and then subjected to heat shock at 33 °C for 3 hours. Surviving animals were scored after 20 hours of recovery at 20 °C. (c) Biofilm-dependent induction of ilys-2 is essential for resistance to P. aeruginosa infection. Five-day old wild-type (N2) and ilys2 worms grown in parallel on indicated B. subtilis strains were transferred to P. aeruginosa PA14 seeded plates and survival was scored every 6 hours. LT50 was taken from Kaplan-Meier survival curves (median survival data). For the log-rank test p-values see Supplementary Table S2. (d) MTL-1 and HSP70 are not required for biofilm-dependent resistance to P. aeruginosa infection. Five-day old wild-type (N2), mtl-1 (tm1770) and hsp70 knockout worms grown in parallel on indicated B. subtilis strains were transferred to P. aeruginosa PA14 seeded plates and survival was scored every 6 hours. LT50 was taken from Kaplan-Meier survival curves (median survival data). For the log-rank test p-values see Supplementary Table S2. (e) Biofilm-induced signaling in C. elegans. Feeding C. elegans biofilm-forming B. subtilis induces expression of MTL-1, Hsp70 (F44E5.4 and F44E5.5) and ILYS-2, partly via HSF-1 and DAF-16 activation (blue and red arrows). Pentagons indicate transcription factors. Upregulation of these biofilm responsive genes in host organism results in enhanced lifespan, thermotolerance, and resistance to oxidative stress and pathogenic infection.
ILYS-2 is required for biofilm-mediated protection against P. aeruginosa infection
As described earlier, the B. subtilis biofilm promotes C. elegans resistance to P. aeruginosa (Fig. 1e). A commensal microbiota is known to stimulate the immune response of higher organisms, thereby promoting their survival during infection52. Among the few genes upregulated in response to the biofilm was ilys-2 (Table 1), which encodes one of the invertebrate lysozymes that function in the C. elegans innate immune response53. We, therefore, examined whether ILYS-2 contributed to biofilm-mediated resistance against P. aeruginosa infection. We constructed C. elegans ilys-2 knockout mutant worms and subjected them to the P. aeruginosa slow killing assay, as described earlier. In the absence of ilys-2 the B. subtilis biofilm was unable to protect against infection (Fig. 6c, Supplementary Table S2 and Supplementary Fig. S8). Notably, neither MTL-1 nor HSP70s (F44E5.4 and F44E5.5) were required for the biofilm-dependent increase in resistance to P. aeruginosa (Fig. 6d, Supplementary Table S2 and Supplementary Fig. S8). Likewise, biofilm was still able to augment thermotolerance of the ilys-2 knockout (Fig. 6b). Thus, biofilm-dependent induction of ilys-2 specifically promotes C. elegans resistance to pathogenic infection.
Discussion
Metazoan hosts and their commensal microbiota have coevolved and established intimate relationships for more than 500 million years. Only recently have we begun to understand the complexity of these interactions7. Germ-free mice display a plethora of physiological and pathological traits, including decreased digestive activity, body fat, muscle wall thickness, and cytokine and serum immunoglobulin production accompanied by increased susceptibility to infection54. Colonization of germ-free mice with a single human symbiotic species, Bacteroides thetaiotaomicron, alters the transcriptional response of various tissues, resulting in improved nutrient uptake, metabolic changes, and restoration of the mucosal barrier function55. Although it is recognized that the majority of bacteria on the gut epithelium and within the mucus layer exist as biofilms56, little is known of the effects of these bacterial populations on the health and disease susceptibility of their hosts.
We utilized a defined model system composed of B. subtilis and C. elegans to investigate the role of non-pathogenic biofilm-host interaction in host physiology and fitness. Both organisms proliferate in the same ecological niche and potentially represent a natural system that was established long before the evolutionary advent of vertebrates9, 16 (Supplementary Discussion). Our data demonstrate that the B. subtilis biofilm endows C. elegans with important survival advantages, such as increased resistance to heat, oxidants, and pathogenic infection. The biofilm also augments C. elegans longevity under undisturbed growth conditions. Notably, these beneficial effects are not due to better colonization of the C. elegans host, as we did not detect any biofilm-dependent difference in the lumenal bacterial load (Supplementary Fig. 2). Rather, we found that specific changes in host gene expression, such as induction of the stress (mtl-1 and hsp-70) and innate immune (ilys-2) responses, underlie the mechanism of beneficial biofilm-host interactions (Fig. 6e). Lysozymes are important in controlling bacterial population in C. elegans intestine53 (Fig. 6c and Supplementary Fig. S8). Therefore, ilys-2 induction may help to curb the accumulation of biofilm-proficient B. subtilis in the gut (Supplementary Fig. S2).
Our work demonstrates that the beneficial effects of biofilm on the worm require live bacteria (Fig. 3). Recently, the positive influence of biofilm-specific bacterial metabolites on C. elegans was also reported18. Notably, in this work C. elegans were fed B. subtilis spores, which is a poor food source that delays animals development and, thus, potentially induces a caloric restriction response37. It was proposed that nitric oxide and a quorum sensing factor are the metabolites responsible for biofilm mediated effects18. However, we have demonstrated that B. subtilis makes NO and extends C. elegans lifespan independently of biofilm formation13. Furthermore, the deletion of a biofilm matrix gene (bslA) and quorum-sensing factor (csf) additively shorten C. elegans lifespan18, indicating that CSF increases the lifespan independently of biofilm. Further studies are required to find the biofilm-specific bacterial metabolites that are responsible for the observed C. elegans phenotypes.
We found that the expression of a small group of C. elegans genes was specifically altered by biofilm formation (Table 1, Supplementary Table S5). Both DAF-16 and SKN-1 are important for C. elegans oxidative stress resistance43, 57. As we did not detect any significant change of SKN-1-dependent genes (Supplementary Table S5) or SKN-1 cellular localization (not shown) in response to biofilm we assumed that skn-1 is dispensable for biofilm-mediated effects. In contrast, DAF-16-dependent mtl-1 was upregulated by biofilm (Table 1 and Supplementary Table S5). MTL-1, is a member of the family of metallothioneins, a group of small cysteine-rich proteins with high affinity for metal ions that are widely distributed among species ranging from bacteria to mammals58. Metallothioneins function to maintain metal and redox homeostasis and to protect cells against oxidative stress and various genotoxic and proteotoxic agents58. In mammals, metallothioneins have also been implicated in neuronal tissue regeneration, modulation of metabolic activity, and suppression of pro-inflammatory and pro-apoptotic responses59, 60. The levels of metallothionein are elevated in various tissues of certain strains of long-lived dwarf mice61, and the targeted overexpression of metallothionein in cardiac tissue extended the lifespan of wild type mice by 14%62. In flies, upregulation of MTL in either motor neurons or the peripheral nervous system was correlated with an approximately 40% increase in lifespan and increased resistance to iron and cadmium toxicity63.
C. elegans harbors two metallothionein genes (mtl-1 and mtl-2). MTL-1 is constitutively and inducibly expressed, respectively, in the terminal bulb and intestine64. MTL-1 is under DAF-16 control43, 44, but its role in C. elegans longevity remains unclear. We identify MTL-1 as a major determinant of increased C. elegans lifespan and stress resistance in response to a commensal biofilm (Figs 4 and 5). DAF-16 is required for biofilm-dependent lifespan extension, but is dispensable for augmented thermotolerance (Supplementary Fig. S7a and b), suggesting that other transcription factors stimulate the biofilm-stimulated expression of mtl-1 to render C. elegans resistant to heat stress (Fig. 6e). Further studies will address the molecular details of biofilm-mediated regulation of DAF-16, and potential tissue-specificity of biofilm effects.
Evolutionarily conserved HSP70 proteins also contribute to the biofilm-mediated increase in C. elegans thermotolerance (Fig. 6a). We show that two HSP70 encoding genes, F44E5.4 and F44E5.5 that are transcribed from overlapping divergent promoters are induced by the B. subtilis biofilm (Table 1). The HSP70 family of chaperones performs a multitude of cytoprotective functions in all organisms48. The C. elegans genome contains six full-length cytosolic HSP70-encoding genes49. It was shown that the expression of F44E5.4 and F44E5.5 increases upon heat shock65, hypoxic stress66, and pathogen infection67. The products of these genes have not been well studied and their specific role in C. elegans remains unknown. Our data implicate these HSP70 paralogs in thermotolerance, which is controlled by the commensal biofilm via HSF1 (Fig. 6a and Supplementary Fig. S7c). Interestingly, Donato et al. reported that HSF-1 is required for biofilm mediated life span extension, while our results show that HSF-1 is only required for biofilm-mediated thermotolerance (Supplementary Fig. S7c and d). It has been shown that HSF-1 is involved in a dietary restriction response68. B. subtilis spores may trigger the dietary restriction response because they are a poor nutritional source37. This may explain the apparent discrepancy between our results and those reported by Donato et al., who fed worms on bacterial spores.
Natural microbiota primes and balances the host immune system52. Activation of the innate response by commensal bacteria leads to accumulation of diverse types of antimicrobial factors, including lactic acid and bacteriocins, in the gastrointestinal tract to restrain the bacterial load54. The invertebrate lysozyme, ILYS-2, functions in the C. elegans innate immune system53. It was shown to be upregulated during infection, suggesting its role in organismal defense against pathogens. However, whether ILYS-2 actually contributes to the animals’ survival upon infection has not been determined. Our data show that the B. subtilis biofilm specifically stimulates ilys-2 expression, increasing the resistance of C. elegans to killing by Pseudomonas (Fig. 6c and e and Table 1).
In this work we focus on the overall response of the host to non-pathogenic biofilm, leaving the identification of inter-species and inter-tissue signaling for future studies. The beneficial effects of bacterial biofilm can be mediated by C. elegans intestinal cells and by non-autonomous cell signaling. Although the expression patterns of mtl-1 and ilys-2 in the intestinal cells have been characterized, little is known about F44E5.4 and F44E5.5 44, 53. Overall, considering the high structural and functional conservation of metallothioneins, HSP70, and lysozymes throughout evolution, and the ancient origin of the regulatory networks with which they are associated (Fig. 6e), it is tempting to hypothesize that biofilms formed by the mammalian microbiota also confer beneficial effects on their host via similar regulatory pathways.
Matherials and Methods
C. elegans strains and growth conditions
Wild-type C. elegans (N2), PS3551 (hsf-1(sy441) I), CF1038 (daf-16(mu86) I), DA1116 (eat-2 (ad1116) II) strains were obtained from the Caenorhabditis Genetics Center. The C. elegans mtl-1 knock-out (tm1770) strain was obtained from National Bioresource Project for the Experimental Animal “Nematode C. elegans” (Tokyo, Japan) and outcrossed to C. elegans N2 6 times before use in experiments. The C. elegans mtl-1 overexpression strain WU1394 (pKD8 amEx183(Pmtl-1(WT)::MTL-1::GFP::mtl-1 3’UTR; myo-3::mCherry) was kindly provided by Kerry Kornfeld, Washington University in St. Louis69. C. elegans ilys-2 and hsp70 knock-out mutants were constructed in this work (Supplementary Table S6). Nematodes were handled according to standard method70. Worms were grown on NGM (Research Product International Corp.) at 20 °C, unless otherwise indicated, and routinely maintained on E. coli OP50. To transfer the nematodes to B. subtilis strains eggs were purified by the alkaline hypochlorite method71. In all assays animals were grown on the indicated B. subtilis strains for at least one, but not more than three, generations prior to use in experiments.
Bacterial strains and growth conditions
Wild-type undomesticated B. subtilis NCIB3610, CYBS-5 and their biofilm-deficient derivatives ΔepsH::tet (DS76), ΔtasA::spec (SSB505), the ΔtasA complemented strain (ΔtasA::spec, amyE::PyqxM-yqxM-sipW-tasA (FC202)), CYBS-5 ΔepsH::tet and B. subtilis sacA::PtasA-mKate2 (TMN503) were isolated or constructed previously17, 72,73,74. B. subtilis PtasA-mKate2 ∆sinI (ASD390) was constructed in this study by transducing TMN1075 with phage from TMN503. TMN1075 is a markerless ∆sinI strain which was constructed using pTMN99474. P. aeruginosa PA14 was kindly provided by Fred Ausubel26. Pseudomonas fluorescens Pf0-1 and its ΔlapA derivative were the generous gift of George A. O’Toole, Geisel School of Medicine at Dartmouth. L. rhamnosus GG (ATCC53103) and its ΔspaCBA (CMPG5357) counterpart were kindly provided by Sarah Leeber, University of Antwerp25. E. coli OP50 was obtained from the Caenorhabditis Genetics Center (Supplementary Table S6).
Overnight bacterial cultures were grown in LB media at 30 °C with agitation and 50 µl were spread atop NGM agar plates. Seeded plates were incubated at 25 °C for ~20 hours and then for 2 hours at 20 °C before worms were transferred onto them.
For the experiments with antibiotic-treated agar plates, overnight B. subtilis lawns on NGM plates were treated with a mixture of antibiotics (500 μg/ml carbenicillin and 100 μg/ml kanamycin in final concentration) and dried for 1.5 hours at room temperature. Plates were shifted to 20 °C for 2 hours before larval stage L4 worms were transferred onto them.
Worms were transferred to freshly treated plates every other day.
For experiments with L. rhamnosus bacterial cultures were grown in MRS medium at 30 °C without agitation and 150 µl were spread atop NGM agar plates. The plates were equilibrated at 20 °C for 2 hours and used immediately.
Lifespan analysis
Lifespans were monitored at 20 °C without FUDR as described previously75 with the following changes. Previously age –synchronized L4 nematodes were transferred to fresh agar plates of the same bacterial strain and the following day was counted as day one in all lifespan measurements. Nematodes were judged as dead when they ceased pharyngeal pumping and did not respond to prodding with a platinum wire. Escaped animals or animals with internal hatching were not included in the lifespan calculations. Kaplan-Meier survival curves were generated using the GraphPad Prism6 statistical analysis software package. Kaplan-Meier curves were compared with the Log-rank (Mantel-Cox) statistical test.
Thermotolerance assay
Thermotolerance assays were performed as described previously for the single-point survival assay76 with the following changes: previously age–synchronized L4 animals were transferred to fresh agar plates of the same bacterial strain and subsequently transferred every other day. Day-5-adults were subjected to heat shock in a water bath at 33 °C (unless otherwise indicated) for the indicated time. After heat shock, worms were shifted to an air incubator at 20 °C and surviving animals were scored after an approximate 20 hour recovery (unless otherwise indicated).
For the experiments with L. rhamnosus GG (ATCC53103) animals were grown on E. coli OP50 until L4 stage, since Lactobacillus do not support C. elegans development. L4-staged worms were treated 3 times with the mix of antibiotics (4 mg/ml carbenicillin and 0.8 mg/ml kanamycin) and then transferred to either L. rhamnosus GG or its biofilm-deficient derivative (ΔspaCBA). Consequently animals were transferred to the fresh plates with corresponding bacterial strains every day until day 5 of adulthood and plates were monitored for potential E. coli OP50 contamination. Worms were subjected to the heat-shock as described earlier.
Oxidative stress resistance assay
Oxidative stress resistance was determined as described77 with the following changes: Fresh 2 mg/ml juglone solution was prepared by dissolving juglone in ethanol (200 proof for molecular biology, Sigma) for 30 min. We used a motorized pestle to facilitate the dilution process. Agar plates containing juglone (90 μM) were freshly prepared before each experiment using fresh juglone solution and were dried in the sterile hood for 1 hour. Worms were age-synchronized as described earlier and transferred to fresh plates with the indicated B. subtilis strain every other day. Day 5 adult animals were transferred to juglone containing plates without bacteria and surviving animals were counted every hour. The percentages of surviving animals was fit to a sigmoidal curve and the LT50 was determined and analyzed for statistical difference using the GraphPad Prism6 statistical analysis software package.
P. aeruginosa slow killing assay
The P. aeruginosa slow killing assay was performed essentially as described in ref. 26. C. elegans were age-synchronized and grown until day 5 of adulthood as described earlier. An overnight culture of P. aeruginosa PA14 in BHI broth was spread on modified NGM (0.35% peptone) agar plates, incubated at 37 °C for 24 hours and then at 25 °C for 18 hours. 30–50 five-day old adult worms were transferred to the P. aeruginosa PA14 plates and examined for survival at 25 °C every 6 hours.
Plasmid construction
An ilys-2 homology repair template, pBADilys2, was designed to delete the first three exons of ilys-2. Two 1-kb homologous regions were PCR-amplified from C. elegans N2 genomic DNA (for primer sequences see Supplementary Table S7).
The Unc-119(+) rescue construct was amplified from punc-119cbr (Addgene #32568). The resulting DNA fragments were gel-purified and assembled into the pBAD cloning vector using the Gibson assembly kit (NEB).
The Hsp-70 homology repair template pBADhsp70 was designed to delete the common promoter region of F44E5.4 and F44E5.5. Using C. elegans N2 genomic DNA as a template, two homologous regions (1.81-kb and 1.877) were PCR-amplified (for primer sequences see Supplementary Table S6).
The Unc-119(+) rescue construct was amplified from punc-119cbr (Addgene #32568). The resulting DNA fragments were gel-purified and assembled into the pBAD cloning vector using the Gibson assembly kit (NEB).
sgRNA encoding vectors: pSgRilys2 and pSgRhsp70. sgRNA sequences were designed using the Benchling Genome Engineering software and cloned into pDD162 (Addgene #47549) with forward primer 5′-N19GTTTTAGAGCTAGAAATAGCAAGT-3′ and reverse primer 5′-CAAGACATCTCGCAATAGG-3, where:
N19 pSgRilys2 = atataagccgtcaaggtag,
N19 pSgRhsp70 = gcatcaaatactgtattctc
ilys-2 and hsp-70 mutants construction
ilys-2 and hsp70 mutants were constructed using CRISPR-Cas9 essentially as described in ref. 78. Briefly, C. elegans HT1593 were injected with a plasmid mixture containing a homology repair template (10 ng/μl), Cas9-a (50 ng/μl), and co-injection markers. Injections were performed by Knudra Transgenics (http://www.knudra.com/). Animals with an unc-119 rescued phenotype that do not carry the co-injection markers were selected at 20 °C and subjected to PCR-genotyping (for primer sequences see Supplementary Table S7). Resulting mutant strains were outcrossed 6 times to wild-type N2.
RNA isolation, next-generation sequencing, and differential expression analysis
Approximately 200 five-day-old adult worms grown on wild-type or ∆epsH B. subtilis were collected and washed in S-buffer, and total RNA was isolated as described in ref. 79. A TrueSeq RNA Sample Preparation Kit v2 (Illumina) was used to prepare cDNA libraries for RNA-Seq from 1 μg of total RNA. Three independent biological replicates were used for each experimental condition. The reference genome and annotation data for C. elegans (Ensembl assembly based on WS220 build) were downloaded from the Illumina websitm (ftp://igenome:G3nom3s4u@ussd-ftp.illumina.com/Caenorhabditis_elegans/Ensembl/WS220/Caenorhabditis_elegans_Ensembl_WS220.tar.gz). To estimate the expression level of transcripts and test for differential expression between different experimental conditions, the Tophat/Cufflinks/Cuffdiff pipeline was used80. Briefly, the RNA-seq reads were trimmed of adaptor sequences and then mapped to the C. elegans transcriptome with the Tophat software package81 using the bowild-typeie2 aligner and default parameters. The transcripts were assembled and their abundances estimated using the Cufflinks package82. A statistical test for differential gene expression was performed using the Cuffdiff tool in the Cufflinks package with a q value (p value adjusted for multiple testing83) threshold of 0.05. Analyses of the differential expression data were performed with the R software package (version 2.15.1) using the cummeRbund library (version 2.0).
Reverse transcription and real-time PCR
Total RNA from 100 worms grown on indicated bacterial strains was extracted as described earlier. Total RNA samples were treated with turbo DNAse (Ambion), and total RNA concentration was measured using NanoDrop1000 (Thermoscientific). Equal amounts of total RNA were reverse transcribed with random primers using SuperScriptII (Invitrogen). 1 μl of the resulting cDNA solution was used for RT-PCR with specific primers (Supplementary Table S6) and Power SYBR Green PCR master mix 2 × (Applied Biosystems) for 40 cycles in a 7300 real-time PCR system (Applied Biosystems) according to the manufacturer’s instructions. The fold change of mRNA levels of each target gene was normalized to the geometric mean of fold change of control genes (pmp-3 and cdc-42).
Self-brood size and rate of egg production
Experiments were performed essentially as described84. Eggs from strain N2 nematodes were isolated, treated with hypochlorite, and incubated for 20 hours at 20 °C in S-buffer without cholesterol85. A synchronized population of L1 arrested worms was then placed on NGM agar plates seeded with wild-type or ΔepsH B. subtilis. Five stage L4 animals were picked manually and transferred to a new plate. Worms were then transferred twice a day to prevent overcrowding until egg-laying ceased. The progeny were counted 3 days after removal of the parents.
Post-embryonic development
Experiments were performed essentially as described in ref. 84. Worms were grown on NGM agar plates seeded with wild-type or ΔepsH B. subtilis. Unstaged eggs were placed at 20 °C and allowed to hatch for 4 hours. Larvae that hatched during that period were placed singly on fresh agar plates and monitored every 5 hours until they began to lay eggs.
Fluorescent microscopy and WGA-FITC staining
To fluorescently label exopolysaccharides we used FITC-conjugated wheat germ agglutinin, a lectin from Triticum vulgaris (Sigma). The labeling was performed as following: C. elegans were grown on B. subtilis strains until day 5 of adulthood and then incubated in 50 μl of WGA-FITC solution in PBS (1 μg/ml) for 5 h with agitation.
For microscopic visualization of mKate2 expression and WGA-FITC labeled exopolysaccharides worms were anesthetized with a drop of 5 mM levamisole. Images of 15 worms for each variant were captured at a fixed exposure time using a Zeiss AxioZoom.V16 equipped for fluorescence illumination. Fluorescence intensity was quantified using ImageJ software.
Motility (thrashing) assay
C. elegans were grown on the corresponding B. subtilis strains as described previously and trashing rate was measured as in (http://www.phage.dk/plugins/download/wrMTrck.pdf) with the following modifications. At day 2 and day 8 of adulthood 10 worms were placed into the 20 μl drop of M9 buffer on the surface of NGM plate without bacteria and left for 1 min to calibrate. 1 minute videos were taken using Zeiss AxioZoom.V16 equipped with AxioCam HRc camera and body bends per second per worm were calculated using ImageJ software and wrMTrck plugin86.
Colonization assays
The colonization assay was adapted from87. 10 worms were picked and placed in a drop of M9 buffer and subsequently washed in 3 drops of M9, the last of which contained 4 mM levamisole for paralysis. Nematodes were then collected and washed in M9 with 1% Triton X-100 and twice with M9 buffer. An aliquot of the final M9 buffer wash was plated as a control for the presence of non-intestinal bacteria. Worms were mechanically disrupted using a motorized pestle. The resulting worm homogenates were serially diluted and plated on LB agar. To calculate the percentage of spores, aliquots of worm homogenate were incubated at 80 °C for 20 min and plated on LB agar. The number of resulting colony forming units (CFU) on LB agar plates was counted after 20 hours at 37 °C. The number of vegetative bacterial cells per worm was calculated by subtracting the CFUs of the non-intestinal control the intestinal spore count from the total number of CFUs and divided by the number of worms per sample.
Alternatively, we used a “guillotine method”, adapted from88 which eliminates bacteria stuck in the pharynges, to determine bacterial colonization. Worms were collected and washed as described for the colonization assay and the pharynx was dissected under the microscope. Worms were collected individually in 50 μl of M9 buffer and mechanically disrupted using a motorized pestle. Aliquots of the homogenates of individual worms were plated on LB agar plates. The aliquots were also plated after having been heated at 80 °C for 20 min to evaluate the percentage of spores. The number of vegetative bacterial cells per worm was calculated as described earlier.
Pumping rate measurements
Pumping rate was measured as the rate of pharyngeal terminal bulb contractions as described (http://www.wormbook.org/chapters/www_measurepharyngeal/measurepharyngeal.html). Briefly, 10 day-one adult worms grown on indicated bacterial strains were examined for 10 seconds, and the number of terminal bulb contractions were counted.
Worm size measurement
The size of C. elegans on B. subtilis plates was measure as described in ref. 89. Briefly, the images of the worms grown on bacterial plates were taken at L4 larval stage and day 1, 3, 4 of adulthood using Zeiss AxioZoom.V16 with the set magnification. Next, surface area of individual worms was determined by means of ImageJ software and WormSizer plugin. The recognition of each individual worm by software was reevaluated by eye.
Transmission Electron Microscopy
5 to 8 living worms were put into the center of 100 µm deep planchette hats that is filled with yeast paste. The hats were coated with hexadecane, sealed in the planchette holder and high pressure freezing commenced with a Wohlwend Compact HPF-01 High Pressure Freezer (Bal-Tec AG, Liechtenstein). The frozen hats were transferred into a mixture of 2% osmium tetroxide, 0.1% uranyl acetate and 2% ddH2O in acetone at liquid nitrogen, and freeze freeze substitution were using Leica EM AFS2 unit. The samples were left in the –90 °C for 96 hours, raised 5 °C per hour to –60 °C and incubated for 12 hours, then to –30 °C for an additional 12 hours, and finally to a temperature of 0 °C for 4 hrs. Three times 1 hour exchanges of pure acetone were used to rinse out the osmium at 0 °C. Infiltration at room temperature began with a mixture of acetone and Embed 812 (Electron Microscopy Sciences, Hatfield, PA) at 1 to 1 for 1 hour, and 1 to 2 overnight. The samples were allowed to sit in pure resin for 4 hours before embedding. The worms were flat embedded with Aclar embedding film and polymerized at 60 °C. Serial semi-thin sections were cut (UC6 microtome; Leica Microsystems) at 1 mm and stained with 1% toluidine blue to evaluate the quality of preservation and find the area of interest. 60 nm ultrathin sections were cut and stained with uranyl acetate and lead citrate by standard methods. Stained grids were examined under Philips CM-12 electron microscope and photographed with a Gatan (4k × 2.7 k) digital camera.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Lopez, D., Vlamakis, H. & Kolter, R. Biofilms. Cold Spring Harbor perspectives in biology 2, a000398, doi:10.1101/cshperspect.a000398 (2010).
Hall-Stoodley, L., Costerton, J. W. & Stoodley, P. Bacterial biofilms: from the natural environment to infectious diseases. Nature reviews. Microbiology 2, 95–108, doi:10.1038/nrmicro821 (2004).
Branda, S. S., Vik, S., Friedman, L. & Kolter, R. Biofilms: the matrix revisited. Trends in microbiology 13, 20–26, doi:10.1016/j.tim.2004.11.006 (2005).
Hobley, L., Harkins, C., MacPhee, C. E. & Stanley-Wall, N. R. Giving structure to the biofilm matrix: an overview of individual strategies and emerging common themes. FEMS microbiology reviews 39, 649–669, doi:10.1093/femsre/fuv015 (2015).
Hall-Stoodley, L. & Stoodley, P. Evolving concepts in biofilm infections. Cellular microbiology 11, 1034–1043, doi:10.1111/j.1462-5822.2009.01323.x (2009).
Langille, M. G. et al. Microbial shifts in the aging mouse gut. Microbiome 2, 50, doi:10.1186/s40168-014-0050-9 (2014).
Cho, I. & Blaser, M. J. The human microbiome: at the interface of health and disease. Nature reviews. Genetics 13, 260–270, doi:10.1038/nrg3182 (2012).
Cabreiro, F. & Gems, D. Worms need microbes too: microbiota, health and aging in Caenorhabditis elegans. EMBO molecular medicine 5, 1300–1310, doi:10.1002/emmm.201100972 (2013).
Felix, M. A. & Braendle, C. The natural history of Caenorhabditis elegans. Current biology: CB 20, R965–969, doi:10.1016/j.cub.2010.09.050 (2010).
Samuel, B. S., Rowedder, H., Braendle, C., Felix, M. A. & Ruvkun, G. Caenorhabditis elegans responses to bacteria from its natural habitats. Proceedings of the National Academy of Sciences of the United States of America 113, E3941–3949, doi:10.1073/pnas.1607183113 (2016).
Dirksen, P. et al. The native microbiome of the nematode Caenorhabditis elegans: gateway to a new host-microbiome model. BMC biology 14, 38, doi:10.1186/s12915-016-0258-1 (2016).
Iatsenko, I., Yim, J. J., Schroeder, F. C. & Sommer, R. J. B. subtilis GS67 protects C. elegans from Gram-positive pathogens via fengycin-mediated microbial antagonism. Current biology: CB 24, 2720–2727, doi:10.1016/j.cub.2014.09.055 (2014).
Gusarov, I. et al. Bacterial nitric oxide extends the lifespan of C. elegans. Cell 152, 818–830, doi:10.1016/j.cell.2012.12.043 (2013).
Garsin, D. A. et al. Long-lived C. elegans daf-2 mutants are resistant to bacterial pathogens. Science 300, 1921, doi:10.1126/science.1080147 (2003).
MacNeil, L. T., Watson, E., Arda, H. E., Zhu, L. J. & Walhout, A. J. Diet-induced developmental acceleration independent of TOR and insulin in C. elegans. Cell 153, 240–252, doi:10.1016/j.cell.2013.02.049 (2013).
Earl, A. M., Losick, R. & Kolter, R. Ecology and genomics of Bacillus subtilis. Trends in microbiology 16, 269–275, doi:10.1016/j.tim.2008.03.004 (2008).
Chen, Y. et al. Biocontrol of tomato wilt disease by Bacillus subtilis isolates from natural environments depends on conserved genes mediating biofilm formation. Environ Microbiol 15, 848–864, doi:10.1111/j.1462-2920.2012.02860.x (2013).
Donato, V. et al. Bacillus subtilis biofilm extends Caenorhabditis elegans longevity through downregulation of the insulin-like signalling pathway. Nature communications 8, 14332, doi:10.1038/ncomms14332 (2017).
Elsholz, A. K., Wacker, S. A. & Losick, R. Self-regulation of exopolysaccharide production in Bacillus subtilis by a tyrosine kinase. Genes & development 28, 1710–1720, doi:10.1101/gad.246397.114 (2014).
Branda, S. S., Chu, F., Kearns, D. B., Losick, R. & Kolter, R. A major protein component of the Bacillus subtilis biofilm matrix. Molecular microbiology 59, 1229–1238, doi:10.1111/j.1365-2958.2005.05020.x (2006).
Portal-Celhay, C., Bradley, E. R. & Blaser, M. J. Control of intestinal bacterial proliferation in regulation of lifespan in Caenorhabditis elegans. BMC microbiology 12, 49, doi:10.1186/1471-2180-12-49 (2012).
Berg, M. et al. Assembly of the Caenorhabditis elegans gut microbiota from diverse soil microbial environments. The ISME journal. doi:10.1038/ismej.2015.253 (2016).
Grompone, G. et al. Anti-inflammatory Lactobacillus rhamnosus CNCM I-3690 strain protects against oxidative stress and increases lifespan in Caenorhabditis elegans. PloS one 7, e52493, doi:10.1371/journal.pone.0052493 (2012).
Ikeda, T., Yasui, C., Hoshino, K., Arikawa, K. & Nishikawa, Y. Influence of lactic acid bacteria on longevity of Caenorhabditis elegans and host defense against salmonella enterica serovar enteritidis. Applied and environmental microbiology 73, 6404–6409, doi:10.1128/AEM.00704-07 (2007).
Lebeer, S. et al. Functional analysis of Lactobacillus rhamnosus GG pili in relation to adhesion and immunomodulatory interactions with intestinal epithelial cells. Applied and environmental microbiology 78, 185–193, doi:10.1128/AEM.06192-11 (2012).
Tan, M. W., Mahajan-Miklos, S. & Ausubel, F. M. Killing of Caenorhabditis elegans by Pseudomonas aeruginosa used to model mammalian bacterial pathogenesis. Proceedings of the National Academy of Sciences of the United States of America 96, 715–720 (1999).
Hinsa, S. M., Espinosa-Urgel, M., Ramos, J. L. & O’Toole, G. A. Transition from reversible to irreversible attachment during biofilm formation by Pseudomonas fluorescens WCS365 requires an ABC transporter and a large secreted protein. Molecular microbiology 49, 905–918 (2003).
Boyd, C. D. et al. Structural features of the Pseudomonas fluorescens biofilm adhesin LapA required for LapG-dependent cleavage, biofilm formation, and cell surface localization. Journal of bacteriology 196, 2775–2788, doi:10.1128/JB.01629-14 (2014).
Silva, M. J. et al. The multifaceted role of commensal microbiota in homeostasis and gastrointestinal diseases. Journal of immunology research 2015, 321241, doi:10.1155/2015/321241 (2015).
Johnson, T. E., Lithgow, G. J. & Murakami, S. Hypothesis: interventions that increase the response to stress offer the potential for effective life prolongation and increased health. The journals of gerontology. Series A, Biological sciences and medical sciences 51, B392–395 (1996).
Herndon, L. A. et al. Stochastic and genetic factors influence tissue-specific decline in ageing C. elegans. Nature 419, 808–814, doi:10.1038/nature01135 (2002).
Hsu, A. L., Feng, Z., Hsieh, M. Y. & Xu, X. Z. Identification by machine vision of the rate of motor activity decline as a lifespan predictor in C. elegans. Neurobiology of aging 30, 1498–1503, doi:10.1016/j.neurobiolaging.2007.12.007 (2009).
Hosono, R., Sato, Y., Aizawa, S. I. & Mitsui, Y. Age-dependent changes in mobility and separation of the nematode Caenorhabditis elegans. Experimental gerontology 15, 285–289 (1980).
Avery, L. & Shtonda, B. B. Food transport in the C. elegans pharynx. The Journal of experimental biology 206, 2441–2457 (2003).
Walker, G., Houthoofd, K., Vanfleteren, J. R. & Gems, D. Dietary restriction in C. elegans: from rate-of-living effects to nutrient sensing pathways. Mechanisms of ageing and development 126, 929–937, doi:10.1016/j.mad.2005.03.014 (2005).
Eric Greer, A. B. In Handbook of the Biology of Aging Handbook of the Biology of Aging (ed Steven N. Austad Edward J. Masoro) Ch. 1, 3–23 (Academic Press, 2010, 2011).
Laaberki, M. H. & Dworkin, J. Role of spore coat proteins in the resistance of Bacillus subtilis spores to Caenorhabditis elegans predation. Journal of bacteriology 190, 6197–6203, doi:10.1128/JB.00623-08 (2008).
Panowski, S. H., Wolff, S., Aguilaniu, H., Durieux, J. & Dillin, A. PHA-4/Foxa mediates diet-restriction-induced longevity of C. elegans. Nature 447, 550–555, doi:10.1038/nature05837 (2007).
Begun, J. et al. Staphylococcal biofilm exopolysaccharide protects against Caenorhabditis elegans immune defenses. Plos Pathog 3, e57, doi:10.1371/journal.ppat.0030057 (2007).
Tan, L. & Darby, C. A movable surface: formation of Yersinia sp. biofilms on motile Caenorhabditis elegans. Journal of bacteriology 186, 5087–5092, doi:10.1128/JB.186.15.5087-5092.2004 (2004).
Roux, D. et al. Identification of Poly-N-acetylglucosamine as a Major Polysaccharide Component of the Bacillus subtilis Biofilm Matrix. The Journal of biological chemistry 290, 19261–19272, doi:10.1074/jbc.M115.648709 (2015).
Borgonie, G. et al. Internal lectin binding patterns in the nematodes Caenorhabditis elegans, Panagrolaimus superbus and Acrobeloides maximus. Fund Appl Nematol 20, 173–186 (1997).
Murphy, C. T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424, 277–283, doi:10.1038/nature01789 (2003).
Zhang, P., Judy, M., Lee, S. J. & Kenyon, C. Direct and indirect gene regulation by a life-extending FOXO protein in C. elegans: roles for GATA factors and lipid gene regulators. Cell metabolism 17, 85–100, doi:10.1016/j.cmet.2012.12.013 (2013).
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R. A. C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464, doi:10.1038/366461a0 (1993).
Freedman, J. H., Slice, L. W., Dixon, D., Fire, A. & Rubin, C. S. The novel metallothionein genes of Caenorhabditis elegans. Structural organization and inducible, cell-specific expression. The Journal of biological chemistry 268, 2554–2564 (1993).
Colangelo, D., Mahboobi, H., Viarengo, A. & Osella, D. Protective effect of metallothioneins against oxidative stress evaluated on wild type and MT-null cell lines by means of flow cytometry. Biometals: an international journal on the role of metal ions in biology, biochemistry, and medicine 17, 365–370 (2004).
Hartl, F. U. & Hayer-Hartl, M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295, 1852–1858, doi:10.1126/science.1068408 (2002).
Nikolaidis, N. & Nei, M. Concerted and nonconcerted evolution of the Hsp70 gene superfamily in two sibling species of nematodes. Molecular biology and evolution 21, 498–505, doi:10.1093/molbev/msh041 (2004).
Chiang, W. C., Ching, T. T., Lee, H. C., Mousigian, C. & Hsu, A. L. HSF-1 regulators DDL-1/2 link insulin-like signaling to heat-shock responses and modulation of longevity. Cell 148, 322–334, doi:10.1016/j.cell.2011.12.019 (2012).
Morley, J. F. & Morimoto, R. I. Regulation of longevity in Caenorhabditis elegans by heat shock factor and molecular chaperones. Mol Biol Cell 15, 657–664, doi:10.1091/Mbc.E03-07-0532 (2004).
O’Hara, A. M. & Shanahan, F. The gut flora as a forgotten organ. EMBO reports 7, 688–693, doi:10.1038/sj.embor.7400731 (2006).
Gravato-Nobre, M. J., Vaz, F., Filipe, S., Chalmers, R. & Hodgkin, J. The Invertebrate Lysozyme Effector ILYS-3 Is Systemically Activated in Response to Danger Signals and Confers Antimicrobial Protection in C. elegans. Plos Pathog 12, e1005826, doi:10.1371/journal.ppat.1005826 (2016).
Shanahan, F. The host-microbe interface within the gut. Best practice & research. Clinical gastroenterology 16, 915–931 (2002).
Xu, J. & Gordon, J. I. Honor thy symbionts. Proceedings of the National Academy of Sciences of the United States of America 100, 10452–10459, doi:10.1073/pnas.1734063100 (2003).
Macfarlane, S. & Dillon, J. F. Microbial biofilms in the human gastrointestinal tract. Journal of applied microbiology 102, 1187–1196, doi:10.1111/j.1365-2672.2007.03287.x (2007).
Tullet, J. M. et al. Direct inhibition of the longevity-promoting factor SKN-1 by insulin-like signaling in C. elegans. Cell 132, 1025–1038, doi:10.1016/j.cell.2008.01.030 (2008).
Miles, A. T., Hawksworth, G. M., Beattie, J. H. & Rodilla, V. Induction, regulation, degradation, and biological significance of mammalian metallothioneins. Critical reviews in biochemistry and molecular biology 35, 35–70, doi:10.1080/10409230091169168 (2000).
Inoue, K., Takano, H., Shimada, A. & Satoh, M. Metallothionein as an anti-inflammatory mediator. Mediators of inflammation 2009, 101659, doi:10.1155/2009/101659 (2009).
Leung, Y. K. et al. Metallothionein induces a regenerative reactive astrocyte phenotype via JAK/STAT and RhoA signalling pathways. Experimental neurology 221, 98–106, doi:10.1016/j.expneurol.2009.10.006 (2010).
Swindell, W. R. Gene expression profiling of long-lived dwarf mice: longevity-associated genes and relationships with diet, gender and aging. BMC genomics 8, 353, doi:10.1186/1471-2164-8-353 (2007).
Yang, X. et al. Metallothionein prolongs survival and antagonizes senescence-associated cardiomyocyte diastolic dysfunction: role of oxidative stress. FASEB journal: official publication of the Federation of American Societies for Experimental Biology 20, 1024–1026, doi:10.1096/fj.05-5288fje (2006).
Bahadorani, S., Mukai, S., Egli, D. & Hilliker, A. J. Overexpression of metal-responsive transcription factor (MTF-1) in Drosophila melanogaster ameliorates life-span reductions associated with oxidative stress and metal toxicity. Neurobiology of aging 31, 1215–1226, doi:10.1016/j.neurobiolaging.2008.08.001 (2010).
Moilanen, L. H., Fukushige, T. & Freedman, J. H. Regulation of metallothionein gene transcription. Identification of upstream regulatory elements and transcription factors responsible for cell-specific expression of the metallothionein genes from Caenorhabditis elegans. The Journal of biological chemistry 274, 29655–29665 (1999).
GuhaThakurta, D. et al. Identification of a novel cis-regulatory element involved in the heat shock response in Caenorhabditis elegans using microarray gene expression and computational methods. Genome research 12, 701–712, doi:10.1101/gr.228902 (2002).
Shen, C., Nettleton, D., Jiang, M., Kim, S. K. & Powell-Coffman, J. A. Roles of the HIF-1 hypoxia-inducible factor during hypoxia response in Caenorhabditis elegans. The Journal of biological chemistry 280, 20580–20588, doi:10.1074/jbc.M501894200 (2005).
Pukkila-Worley, R., Ausubel, F. M. & Mylonakis, E. Candida albicans infection of Caenorhabditis elegans induces antifungal immune defenses. Plos Pathog 7, e1002074, doi:10.1371/journal.ppat.1002074 (2011).
Steinkraus, K. A. et al. Dietary restriction suppresses proteotoxicity and enhances longevity by an hsf-1-dependent mechanism in Caenorhabditis elegans. Aging cell 7, 394–404, doi:10.1111/j.1474-9726.2008.00385.x (2008).
Roh, H. C. et al. A modular system of DNA enhancer elements mediates tissue-specific activation of transcription by high dietary zinc in C. elegans. Nucleic Acids Res 43, 803–816, doi:10.1093/nar/gku1360 (2015).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Stiernagle, T. Maintenance of C. elegans. WormBook: the online review of C. elegans biology, 1-11, doi:10.1895/wormbook.1.101.1 (2006).
Kearns, D. B., Chu, F., Branda, S. S., Kolter, R. & Losick, R. A master regulator for biofilm formation by Bacillus subtilis. Molecular microbiology 55, 739–749, doi:10.1111/j.1365-2958.2004.04440.x (2005).
Chu, F., Kearns, D. B., Branda, S. S., Kolter, R. & Losick, R. Targets of the master regulator of biofilm formation in Bacillus subtilis. Molecular microbiology 59, 1216–1228, doi:10.1111/j.1365-2958.2005.05019.x (2006).
Norman, T. M., Lord, N. D., Paulsson, J. & Losick, R. Memory and modularity in cell-fate decision making. Nature 503, 481–486, doi:10.1038/nature12804 (2013).
Greer, E. L. et al. An AMPK-FOXO pathway mediates longevity induced by a novel method of dietary restriction in C. elegans. Current biology: CB 17, 1646–1656, doi:10.1016/j.cub.2007.08.047 (2007).
Lithgow, G. J., White, T. M., Melov, S. & Johnson, T. E. Thermotolerance and extended life-span conferred by single-gene mutations and induced by thermal stress. Proceedings of the National Academy of Sciences of the United States of America 92, 7540–7544 (1995).
de Castro, E., Hegi de Castro, S. & Johnson, T. E. Isolation of long-lived mutants in Caenorhabditis elegans using selection for resistance to juglone. Free radical biology & medicine 37, 139–145, doi:10.1016/j.freeradbiomed.2004.04.021 (2004).
Dickinson, D. J., Ward, J. D., Reiner, D. J. & Goldstein, B. Engineering the Caenorhabditis elegans genome using Cas9-triggered homologous recombination. Nature methods 10, 1028–1034, doi:10.1038/nmeth.2641 (2013).
Reinke, V. et al. A global profile of germline gene expression in C. elegans. Molecular cell 6, 605–616 (2000).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nature protocols 7, 562–578, doi:10.1038/nprot.2012.016 (2012).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111, doi:10.1093/bioinformatics/btp120 (2009).
Roberts, A., Pimentel, H., Trapnell, C. & Pachter, L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 27, 2325–2329, doi:10.1093/bioinformatics/btr355 (2011).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proceedings of the National Academy of Sciences of the United States of America 100, 9440–9445, doi:10.1073/pnas.1530509100 (2003).
Wong, A., Boutis, P. & Hekimi, S. Mutations in the clk-1 gene of Caenorhabditis elegans affect developmental and behavioral timing. Genetics 139, 1247–1259 (1995).
Fabian, T. J. & Johnson, T. E. Production of age-synchronous mass cultures of Caenorhabditis elegans. J Gerontol 49, B145–156 (1994).
Nussbaum-Krammer, C. I., Neto, M. F., Brielmann, R. M., Pedersen, J. S. & Morimoto, R. I. Investigating the spreading and toxicity of prion-like proteins using the metazoan model organism C. elegans. Journal of visualized experiments: JoVE, 52321, doi:10.3791/52321 (2015).
Portal-Celhay, C. & Blaser, M. J. Competition and resilience between founder and introduced bacteria in the Caenorhabditis elegans gut. Infection and immunity 80, 1288–1299, doi:10.1128/IAI.05522-11 (2012).
Laaberki, M. H. & Dworkin, J. Death and survival of spore-forming bacteria in the Caenorhabditis elegans intestine. Symbiosis 46, 95–100 (2008).
Moore, B. T., Jordan, J. M. & Baugh, L. R. WormSizer: high-throughput analysis of nematode size and shape. PloS one 8, e57142, doi:10.1371/journal.pone.0057142 (2013).
Acknowledgements
We thank Kerry Kornfeld, Fred Ausubel, George A. O’Toole, Sarah Leeber and Caenorhabditis Genetics Center (University of Minnesota) for materials, NYULMC OCS Microscopy Core for the electron microscopy services. We thank Timur Artemev for his contribution. This work was supported by the Howard Hughes Medical Institute and Blavatnik Family Foundation (E.N.).
Author information
Authors and Affiliations
Contributions
E.N. conceptualized the study. O.S., I.G., and E.N. designed the experiments. O.S., L.G., performed the experimental work. O.S., I.G., A.S.D., and R.L. discussed the results and commented on the manuscript. I.S. performed the bioinformatics analysis. O.S and E.N. wrote the paper. R.L. and E.N. supervised the research.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Smolentseva, O., Gusarov, I., Gautier, L. et al. Mechanism of biofilm-mediated stress resistance and lifespan extension in C. elegans . Sci Rep 7, 7137 (2017). https://doi.org/10.1038/s41598-017-07222-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-07222-8
This article is cited by
-
Vitamin B12 produced by gut bacteria modulates cholinergic signalling
Nature Cell Biology (2024)
-
Endogenous DAF-16 spatiotemporal activity quantitatively predicts lifespan extension induced by dietary restriction
Communications Biology (2023)
-
Bacillus Species as Direct-Fed Microbial Antibiotic Alternatives for Monogastric Production
Probiotics and Antimicrobial Proteins (2023)
-
Bacillus subtilis biofilm formation and social interactions
Nature Reviews Microbiology (2021)
-
C. elegans: A biosensor for host–microbe interactions
Lab Animal (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.