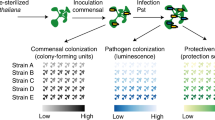
Abstract
Arabidopsis thaliana mlo2 mlo6 mlo12 triple mutant plants exhibit complete immunity against infection by otherwise virulent obligate biotrophic powdery mildew fungi such as Golovinomyces orontii. While this phenotype is well documented, the interaction profile of the triple mutant with other microbes is underexplored and incomplete. Here, we thoroughly assessed and quantified the infection phenotypes of two independent powdery mildew-resistant triple mutant lines with a range of microbes. These microorganisms belong to three kingdoms of life, engage in diverse trophic lifestyles, and deploy different infection strategies. We found that interactions with microbes that do not directly enter leaf epidermal cells were seemingly unaltered or showed even enhanced microbial growth or symptom formation in the mlo2 mlo6 mlo12 triple mutants, as shown for Pseudomonas syringae and Fusarium oxysporum. By contrast, the mlo2 mlo6 mlo12 triple mutants exhibited reduced host cell entry rates by Colletotrichum higginsianum, a fungal pathogen showing direct penetration of leaf epidermal cells comparable to G. orontii. Together with previous findings, the results of this study strengthen the notion that mutations in genes MLO2, MLO6 and MLO12 not only restrict powdery mildew colonization, but also affect interactions with a number of other phytopathogens.
Similar content being viewed by others
Introduction
Powdery mildew is a widespread fungal disease of many angiosperm plants1. It is caused by ascomycetes of the order Erysiphales. The more than 400 known powdery mildew species can infect ~10,000 plant species, including the dicotyledonous reference species Arabidopsis thaliana 2. Currently, four powdery mildew species have been reported to be able to complete their asexual life cycle on Arabidopsis host plants, i.e. Erysiphe cruciferarum, Golovinomyces cichoracearum, G. orontii and Oidium neolycopersici 3, 4.
The major naturally occurring source of resistance effective against powdery mildews in Arabidopsis is RESTANCE TO POWDERY MILDEW8 (RPW8)5,6,7. This complex locus shows extensive intraspecific genetic variation and confers dominantly inherited resistance against multiple powdery mildew species. The respective genes encode non-canonical resistance proteins that lead to arrest of fungal pathogenesis after host cell penetration (“post-penetration resistance”). Effective resistance correlates with the encasement of the fungal feeding structures (haustorial complexes) in a callose-containing cell wall matrix8.
A different type of powdery mildew resistance is conferred by recessively inherited loss-of-function mutations in specific MILDEW RESISTANCE LOCUS O (MLO) genes. These genes, which encode integral membrane proteins of unknown biochemical activity, comprise a family of 15 members in Arabidopsis9. Loss-of-function mutations in MLO2 (At1g11310) result in incomplete resistance against powdery mildew attack that is characterized by a reduction in host cell entry rates by ~50%. This coincides with an arrest of hyphal growth prior to the formation of conidiophores in mlo2 plants, resulting in almost entirely abolished sporulation10, 11. Mutations in MLO6 (At1g61560) and MLO12 (At2g39200) do not affect powdery mildew interactions on their own. However, they co-operatively enhance mlo2-conditioned resistance, and in combination with a mutation in MLO2 cause a complete lack of host cell penetration by fungal sporelings, leading to complete immunity (“pre-penetration resistance”). This type of powdery mildew resistance is best known from barley, where natural and induced mlo mutants have been discovered more than 70 years ago and have been successfully employed in agriculture for over 35 years12, 13. More recently, mlo-based resistance was described in several other monocotyledonous and dicotyledonous plant species such as, amongst others, pea14, tomato15 and wheat16, 17. Hence, mlo-based resistance is a seemingly universal phenomenon within angiosperm plant species that are hosts to powdery mildew fungi18.
At the phenotypical and molecular level, mlo resistance resembles the highly effective defence against non-adapted powdery mildews (“non-host resistance”19). The mlo2 mlo6 mlo12 mutant shows a spectacular level of resistance against different powdery mildews11, 15. Conversely, it revealed slightly enhanced disease symptoms–and in part also pathogen proliferation–in interactions with some hemibiotrophic/necrotrophic pathogens such as Alternaria alternata, A. brassicicola and Phytophthora infestans 11. These phenotypes might be the indirect consequence of deregulated mesophyll cell death in leaves of the mlo2 mlo6 mlo12 mutant. Similar to the barley mlo mutant, leaves of the triple mutant are subject to spontaneous deposition of callose-containing cell wall appositions and ultimately premature senescence11, 20,21,22.
Apart from powdery mildews, the model plant Arabidopsis can serve as a host for a number of additional microbial pathogens and/or endophytes. Well-established and widely studied patho-systems comprise the interaction of Arabidopsis with bacteria (e.g. Pseudomonas syringae 23) and oomycetes (e.g. Hyaloperonospora arabidopsidis 24 and Albugo spp.25, 26), which cause the bacterial speck, downy mildew and white rust disease, respectively. More recently established patho-systems include, amongst others, the interaction of Arabidopsis with the anthracnose fungus Colletotrichum higginsianum 27 and the causal agent of the vascular wilt and root rot disease, Fusarium oxysporum 28, 29. Apart from pathogenic microbes, Arabidopsis harbors a broad range of bacterial and fungal endophytes in the rhizo- and phyllosphere when grown under natural conditions (leaf and root microbiota and mycobiota)30, 31. A well-studied root-colonizing endophyte of many plant species, including Arabidopsis, with a reported growth-promoting activity is Serendipita indica (syn: Piriformospora indica)32,33,34.
While the role of mlo mutants in providing resistance to powdery mildew fungi is well-documented18, the effect of mutations in MLO genes on colonization with other microbes has not yet been explored systematically in a quantitative manner. To fill this existing gap in knowledge, we performed comprehensive infection assays with two independent powdery mildew-resistant mlo2 mlo6 mlo12 T-DNA knockout lines of Arabidopsis and a range of microbial species that exhibit different lifestyles and diverse modes of plant colonization. Quantitative assessment in relation to control genotypes revealed altered infection phenotypes of the mlo2 mlo6 mlo12 mutants in the case of the fungal parasites F. oxysporum and C. higginsianum and the bacterial pathogen P. syringae.
Results
In order to establish a comprehensive interaction profile of the Arabidopsis mlo2 mlo6 mlo12 triple mutant, we challenged individuals of two independent triple mutant lines with a broad panel of microorganisms that are known to be virulent on the Arabidopsis Col-0 accession. These two mutant lines represent two entirely independent allele combinations in the genetic background of Col-0. The resulting infection phenotypes were compared with that of Col-0 wild type, which served as control in all experiments. At the taxonomic level, the panel of tested microorganisms comprised fungal (G. orontii, F. oxysporum, C. higginsianum, S. indica), oomycete (H. arabidopsidis, A. laibachii) and bacterial (P. syringae) species, which engage in obligate biotrophic (G. orontii, H. arabidopsidis, A. laibachii), and hemibiotrophic (P. syringae, C. higginsianum and F. oxysporum) interactions, respectively. We also included in our set of experiments a fungal endophyte (S. indica) that is known to have a growth-promoting effect on colonized Arabidopsis plants.
A novel mlo2 mlo6 mlo12 triple mutant line with near-complete powdery mildew resistance
We previously showed that the triple mutant line mlo2-5 mlo6-2 mlo12-1 is fully resistant to the adapted powdery mildew pathogen, G. orontii 11. We generated a second triple mutant line, mlo2-6 mlo6-4 mlo12-8, which is based on different T-DNA insertions in the three AtMLO genes (Fig. 1A; see Materials and Methods for further details). As revealed by reverse transcriptase-polymerase chain reaction (RT-PCR) analysis, the T-DNA insertions in these lines result in a lack of full-length MLO2, MLO6 and MLO12 transcripts (Fig. 1B). The two triple mutant lines grow similarly as Col-0 wild-type plants, but suffer from early leaf senescence as described before in detail for mlo2-5 mlo6-2 mo12-111, 20. This phenotype is evident by the slightly chlorotic rosette leaves of the two mlo2 mlo6 mlo12 lines at the age of approximately six weeks (Fig. 1C) We analysed the triple mutants upon challenge with G. orontii and found that line mlo2-6 mlo6-4 mlo12-8 fully resembles line mlo2-5 mlo6-2 mlo12-1 with respect to the macroscopic and microscopic infection phenotypes (Fig. 1C–E). Unlike the Col-0 control plants, which showed abundant fungal sporulation at 8 days post inoculation (dpi), individuals of both lines lacked visible powdery mildew symptoms (Fig. 1C). This finding was consistent with analysis at the microscopic level at 48 hours post inoculation (hpi), which revealed an early abortion of fungal pathogenesis at the level of host cell entry in the two triple mutants, while Col-0 control plants showed extensive mycelial growth (Fig. 1D). We noted, however, that line mlo2-5 mlo6-2 mlo12-1 was entirely resistant, lacking any recognizable host cell penetration (0% entry rate as judged by the absence of secondary hyphae and discernible haustoria) in our experiments, while line mlo2-6 mlo6-4 mlo12-8 allowed the occasional formation of fungal micro-colonies (ca. 1% entry rate; Fig. 1E). With our sample size, this minor difference between the two mlo2 mlo6 mlo12 genotypes was not statistically significant. Thus, the two mlo2 mlo6 mlo12 triple mutants are essentially equivalent with regard to the level of resistance against the obligate biotrophic powdery mildew pathogen, G. orontii.
The G. orontii resistance phenotype of the mlo2-6 mlo6-4 mlo12-8 triple mutant is indistinguishable from the mlo2-5 mlo6-2 mlo12-1 triple mutant. Six-week-old Arabidopsis plants were touch-inoculated with G. orontii conidiospores. (A) Scheme depicting the T-DNA insertion sites in MLO2, MLO6 and MLO12. Rectangles represent exons, black lines introns. Triangles symbolize the T-DNA insertion sites of the various mlo alleles. Lines flanked by inverted arrows (primer binding sites) below the gene models indicate the RT-PCR amplicons used to test for MLO transcript accumulation in the mutant lines. (B) RT-PCR analysis of MLO2, MLO6 and MLO12 transcript accumulation. Primer pairs covering the regions indicated in panel A were used to amplify the respective transcript amplicons from cDNA of lines mlo2-5 mlo6-2 mlo12-1 and mlo2-6 mlo6-4 mlo12-8 (two individuals each) as well as Col-0 wild-type plants (positive control). RT-PCR reactions without reverse transcription (control 1) and amplification without template (control 2) served as negative controls. White arrowheads indicate RT-PCR products of the expected size in case of Col-0 wild-type plants. (C) Representative macroscopic infection phenotypes at 8 dpi. (D) Light micrographs visualizing fungal pathogenesis at 48 hpi. Leaf samples were cleared in destaining solution and fungal infection structures subsequently stained with Coomassie Brillant Blue. Bars = 100 µm. (E) Quantitative assessment of host cell entry. Data show the mean ± standard error of the mean (SEM) from three experiments. In each experiment, at least 100 interaction sites from 1-3 leaves of five independent plants per genotype were assessed (total of > 500 interaction sites per genotype and experiment). *** Indicates a statistically significant difference from Col-0 (P < 0.001) according to a GLM test (binomial distribution) on n = 3 independent experimental replicates.
mlo2 mlo6 mlo12 plants show unaltered susceptibility to downy mildew
We next assessed the infection phenotype of the mlo2 mlo6 mlo12 triple mutants with the obligate biotrophic oomycete H. arabidopsidis (isolate Noco2) upon spray inoculation of 16-day-old seedlings. At 7 dpi, we scored the formation of conidiospores in comparison to the susceptible accession Col-0, the resistant accession Landsberg erecta (Ler) and the super-susceptible eds1-2 mutant35, 36 (in Col-0 genetic background) using a hemocytometer-based assay. Owing to considerable experiment-to-experiment variation regarding the absolute numbers of conidiospores per gram seedlings, we normalized the results of the five independent experiments to the values obtained with Col-0, set as 100%. In comparison to Col-0 (100% ± 15%), the two mlo2 mlo6 mlo12 triple mutants showed unaltered levels of conidiospore formation (96% ± 21% and 115% ± 32%, respectively; no statistically significant difference from Col-0), while Ler(0% ± 0%) and the eds1-2 mutant (240% ± 66%) revealed the expected resistant and super-susceptible phenotypes (Fig. 2). These data demonstrate that the infection phenotype with H. arabidopsidis isolate Noco2 is unaltered in the mlo2 mlo6 mlo12 triple mutants.
The mlo2 mlo6 mlo12 triple mutants show an unaltered H. arabidopsidis phenotype. Sixteen-day-old seedlings were spray-inoculated with H. arabidopsidis (isolate Noco2) and spore formation (spores per g FW) was quantitatively assessed at 7 dpi. Sporulation was normalized relative to accession Col-0 set as 100%. Data show the mean ± SEM based on five independent experiments. Accession Ler served as resistant control, the mutant eds1-2 as super-susceptible control. ** Indicates a statistically significant difference from Col-0 (P < 0.01) according to a Wilcoxon-Mann-Whitney rank sum test on n = 5 independent experimental replicates.
mlo2 mlo6 mlo12 plants show unaffected susceptibility to white rust
We then analysed the interaction of the mlo2 mlo6 mlo12 triple mutants with another obligate biotrophic oomycete, A. laibachii (white rust pathogen; isolate Nc14), following spray inoculation with zoospores. The susceptible accession Col-0 and the resistant accession Keswick (Ksk-137) were included as controls. At 10 dpi, infection phenotypes were scored (Fig. 3A) and infection rates were quantitatively assessed as the percentage of uninfected (showing no signs of white rust sporulation) versus infected (showing abundant sporulation) leaves. This experiment revealed infection rates of 82% ± 6% for Col-0 and 0% ± 0% for Ksk-1, compared to 81% ± 12% for mlo2-5 mlo6-2 mlo12-1 and 74% ± 5% for mlo2-6 mlo6-4 mlo12-8 (Fig. 3B). The values for the two mlo2 mlo6 mlo12 triple mutants were not significantly different from the Col-0 control plants, as supported by our statistical analysis. Taken together, the data indicate that the susceptibility to A. laibachii does not differ between Col-0 and the two mlo2 mlo6 mlo12 triple mutants.
The mlo2 mlo6 mlo12 triple mutants show an unaltered A. laibachii infection phenotype. Six-week-old plants were spray-inoculated with A. laibachii (isolate Nc14) and infection phenotypes were assessed at 10 dpi. (A) Representative examples of inoculated rosette leaves of accessions Col-0, Ksk-1 and the two mlo2 mlo6 mlo12 triple mutant lines. Insets show a magnification of an inoculated leaf area. (B) Quantitative evaluation of the infection rate. Data show the mean ± standard deviation (SD) of the proportion of rosette leaves with disease symptoms (macroscopically visible white rust pustule formation) based on 5-6 plants per genotype and 12-30 evaluated leaves per plant. ** Indicates a statistically significant difference from Col-0 (P < 0.01) according to a Wilcoxon-Mann-Whitney rank sum test.
mlo2 mlo6 mlo12 plants show enhanced resistance to C. higginsianum
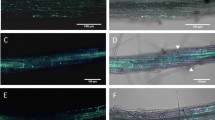
We further extended our study by analysing infection of the mlo2 mlo6 mlo12 triple mutants with the hemibiotrophic ascomycete pathogen C. higginsianum (strain IMI349063). At 3 dpi, the foliar fungal pathogen triggered local host cell death (necrotic lesions) in rosette leaves of mlo2 mlo6 mlo12 plants, the extent of which was seemingly lower in the mutants than in Col-0 control plants (Fig. 4A). Consistent with this observation, quantitative cytological analysis at 3 dpi revealed reduced levels of host cell entry (successful host cell penetration and formation of intracellular biotrophic hyphae) in both mlo2 mlo6 mlo12 triple mutant lines compared to Col-0 control plants (Fig. 4B). While both mutant lines showed a comparable decrease in host cell entry in this experiment, there was a statistically significant difference between the two triple mutant lines in a second independent experiment (Fig. S1). Taken together, both host cell penetration and symptom formation are reduced in both mlo2 mlo6 mlo12 mutant lines.
The mlo2 mlo6 mlo12 triple mutants show decreased host cell entry by C. higginsianum. Infection phenotypes of Col-0, mlo2-5 mlo6-2 mlo12-1 and mlo2-6 mlo6-4 mlo12-8 at 3 dpi with C. higginsianum (isolate IMI349063A). Plants were spray-inoculated with spore suspension (5 × 105 spores ml−1). (A) Representative examples of whole leaves cleared and stained with Trypan blue (left column) and light micrographs showing leaf epidermal cells after clearing in choral hydrate (right column). Biotrophic hyphae (arrows) are visible beneath some melanised appressoria. Bars = 30 μm. (B) Quantitative assessment of host cell entry. Data show the mean ± SD from counts of at least 140 appressoria from each leaf (one leaf from each of 3 different plants), i.e. at least 420 appressoria per plant genotype. ***Indicates a statistically significant difference from Col-0 (P < 0.001) according to a GLM test (Poisson distribution) on n = 3 technical replicates (individual plants). The experiment was repeated once with similar results (Fig. S1).
mlo2 mlo6 mlo12 plants show enhanced disease symptoms to F. oxysporum
We then turned our attention to the hemibiotrophic root-infecting fungal pathogen F. oxysporum. Following dip inoculation of roots of 2-week-old plants in spore suspensions of F. oxysporum isolate Fo5176, disease progression was scored at 5, 7 and 10 dpi by semi-quantitative classification of disease symptoms into six categories and by calculating an “average disease index” based on this classification38. Fifteen to forty plants were assessed per experimental replicate, genotype and time point. This experimental setup revealed more severe disease symptoms in the two mlo2 mlo6 mlo12 mutants, typically recognizable as a tendency already at 5 dpi, becoming more evident at 7 dpi and being pronounced at 10 dpi (Figs 5 and S2). Taking into account all three experimental replicates, there was no apparent difference in hypersusceptibility between the two mlo2 mlo6 mlo12 mutant lines.
The mlo2 mlo6 mlo12 triple mutants show enhanced disease symptoms upon challenge with F. oxysporum. Two-week-old Arabidopsis seedlings were inoculated with a spore suspension of F. oxysporum (isolate Fo5176). (A) Infection phenotypes were scored at 5, 7 and 10 dpi by assigning a disease index on a 0 (no symptoms) to 5 (severe disease symptoms) scale. Data shown are from a representative experiment and based on 15-20 seedlings per genotype. Each symbol in the categorical scatter plot (circle, square or triangle) represents the infection phenotype of one seedling. The crosses indicate the mean values ± SEM. *, ** and *** indicate statistically significant differences from Col-0 (P < 0.05, P < 0.01 and P < 0.001, respectively) according to a GLM test (Poisson distribution) on n = 15-20 technical replicates (individual seedlings). The experiment was repeated twice with similar results (Fig. S2). (B) Representative macroscopic infection phenotypes at 7 dpi.
mlo2 mlo6 mlo12 plants show enhanced susceptibility to P. syringae
Next, we examined the infection phenotype of the two mlo2 mlo6 mlo12 triple mutant lines with the bacterial pathogen P. syringae pv. maculicola (strain ES4326). For these experiments, we deployed a strain that carries a chromosomal integration of the Photorhabdus luminescens luciferase gene (here designated as P. syringae pv. maculicola lux). Luminescence emitted by this strain reliably reports bacterial growth in infected Arabidopsis leaves39. We infiltrated bacterial suspensions into leaves of Col-0 wild type and mlo2 mlo6 mlo12 mutant plants and further included sid2-2/eds16-140 and edr1 41 mutants as additional control plants, which were previously reported to show super-susceptibility and enhanced resistance, respectively, in response to infection with P. syringae. Luminescence was recorded shortly after inoculation (0 dpi) and at 3 dpi. In the case of the two triple mutants, we observed increased luminescence at 3 dpi, corresponding to approximately two-fold higher bacterial titres compared to Col-0, while the sid2-2 and edr1 controls showed the expected phenotypes of intensely increased susceptibility in sid2-2 (approx. 8-fold increase) and a tendency to an increased resistance in edr1 (Fig. 6). A slight but statistically significant increase in bacterial titres in the mlo2 mlo6 mlo12 mutant plants was found in three out of four additional experimental replicates of this pathogen assay (Fig. S3). Taken together, both mlo2 mlo6 mlo12 mutant lines show somewhat increased susceptibility to P. syringae pv. maculicola.
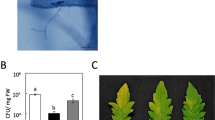
The mlo2 mlo6 mlo12 triple mutants show an elevated bacterial titre upon challenge with P. syringae. Five-week-old Arabidopsis plants were infiltrated with P. syringae pv. maculicola lux (OD600 = 0.0005) and luminescence (RLU cm−2; corresponding to bacterial titre) determined at 0 dpi (to ensure an equal start inoculum) and 3 dpi. (A) The boxplot shows data from one representative experiment based on n = 7 (0 dpi) and n = 11 plants (3 dpi) per genotype, with each plant value representing the mean of three leaves. Centre lines mark the medians, upper and lower box limits indicate the 25th and 75th percentiles, respectively; upper and lower whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles, respectively; and dots represent outliners. Letters indicate statistically different groups (at least P < 0.05) according to a GLM test (quasi-Poisson distribution) on n = 11 technical replicates (individual plants). The experiment was repeated four times with n = 7-13 plants per genotype in each experiment and in two different controlled environments (see Materials and Methods for details) with comparable results (Fig. S3). (B) Representative macroscopic infection phenotypes at 3 dpi. Yellow arrows indicate the inoculated leaves.
mlo2 mlo6 mlo12 plants show an unaltered infection phenotype with S. indica
We finally focused our attention on the fungal root endophyte S. indica, belonging to the Basidiomycota, which exerts a beneficial effect on the growth of Arabidopsis32. Interaction assays with this species (isolate DSM11827) were performed with sterile seedlings in petri dishes, and the extent of S. indica colonization was assessed relative to accession Col-0 via genomic DNA content by quantitative PCR analysis at 3 and 7 dpi34. Based on three independent experiments, we observed no statistically significant difference in fungal DNA content at any time point between Col-0 control plants and the two mlo2 mlo6 mlo12 triple mutant lines (Fig. 7).
The mlo2 mlo6 mlo12 triple mutants show unaltered colonization by S. indica. Roots of 2-week-old plants were inoculated with S. indica (isolate DSM11827) and the rate of root colonization determined by quantifying the relative amount of fungal genomic DNA by qPCR analysis. Data show the mean ± SEM based on three experiments with 240 seedlings per genotype and experiment. There is no statistically significant difference between genotypes according to GLM (quasi-Poisson distribution) on n = 3 independent experimental replicates.
A. laibachii cannot break powdery mildew resistance of mlo2 mlo6 mlo12 plants
Although Arabidopsis is a nonhost plant for the oomycete P. infestans, the causal agent of late blight in tomato and potato, a recent study reports that pre-infection with A. laibachii can render Arabidopsis susceptible to colonization by P. infestans 42. Based on this finding, we wondered whether pre-infection with A. laibachii could also overcome mlo-based resistance in Arabidopsis and turn an otherwise incompatible interaction between mlo2 mlo6 mlo12 triple mutants and the powdery mildew pathogen G. orontii into a compatible one. To test this speculation, we spray-inoculated 5-week-old Col-0 and mlo2 mlo6 mlo12 mutant plants with either water (mock control) or A. laibachii (isolate Nc14) zoospores and at 8-9 dpi challenged the pre-inoculated leaves with G. orontii. We then scored the penetration success of the powdery mildew pathogen at 48 hpi by microscopic analysis. We found (based on two independent experiments) that the pre-inoculation of plants with A. laibachii did not change the G. orontii entry levels. Col-0 plants remained fully susceptible (>92% host cell entry) under these conditions, while the triple mutant plants remained either fully (mlo2-5 mlo6-2 mlo12-1; 0% host cell entry) or nearly fully (mlo2-6 mlo6-4 mlo12-8; 2-6% host cell entry) resistant (Table 1). These values are equivalent to the figures obtained in standard G. orontii inoculation experiments with these genotypes (Fig. 1E). Thus, pre-inoculation with A. laibachii fails to break resistance conditioned by mlo triple mutants in Arabidopsis.
Discussion
We studied the infection phenotype of two mlo2 mlo6 mlo12 triple T-DNA mutants based on independent allele combinations (mlo2-5 mlo6-2 mlo12-1 and mlo2-6 mlo6-4 mlo12-8) with a number of plant-colonizing microbes in a quantitative manner. These species, with the exception of the endophyte S. indica, are mostly pathogenic, comprised representatives from three kingdoms of life (bacteria, oomycetes, fungi) that utilize different trophic lifestyles (obligate biotrophs and hemibiotrophs) and diverse infection strategies. They thus constitute a broad panel of microbial species that can colonize the model plant A. thaliana.
As expected and previously shown for one of the two mutant lines11, both mlo2 mlo6 mlo12 triple mutants were highly resistant against the adapted powdery mildew pathogen G. orontii (Fig. 1). We noted, however, that the newly generated mlo2-6 mlo6-4 mlo12-8 mutant showed a very low level of host cell entry (1–6%), whereas the previously reported mlo2-5 mlo6-2 mlo12-1 mutant was fully immune (0% host cell entry; Fig. 1E and Table 2). This minute yet reproducible difference in host cell penetration could be due to residual levels of truncated MLO mRNA in the mlo2-6 mlo6-4 mlo12-8 mutant. This concerns particularly MLO6, for which the T-DNA insertion site in the mlo6-4 mutant is at the very distal end of the gene (Fig. 1A). Partial functionality of C-terminally truncated protein variants was previously demonstrated in the case of barley Mlo43. Alternatively, anonymous second-site T-DNA insertions in the mlo2-6 mlo6-4 mlo12-8 mutant may modulate the powdery mildew infection phenotype in this line. Potent powdery mildew resistance of the mlo2-5 mlo6-2 mlo12-1 mutant line was previously also found with respect to another adapted powdery mildew fungus, the tomato pathogen O. neolycopersici 15. In addition, the triple mutant is also resistant to two non-adapted powdery mildew fungi, the pea pathogen Erysiphe pisi and Blumeria graminis f.sp. hordei (Bgh), a pathogen of barley11 (Table 2).
In comparison to Col-0, both mlo2 mlo6 mlo12 triple mutant lines exhibited unaltered infection phenotypes upon challenge with the obligate biotrophic oomycetes H. arabidopsidis and A. laibachii and following inoculation with the fungal endophyte S. indica (Figs 2, 3 and 7). Thus, mutations in genes MLO2, MLO6 and MLO12 apparently do not affect colonization of Arabidopsis by these microbes. However, we only tested one particular Arabidopsis accession (Col-0) with single strains/isolates of the respective microorganisms and performed the assays under standard laboratory conditions. In addition, only certain parameters of plant colonization such as successful sporulation (H. arabidopsidis and A. laibachii) or the amount of fungal biomass (S. indica) were assessed. Therefore, we cannot rule out that the respective infection phenotypes of mlo2 mlo6 mlo12 mutant plants might be altered in different settings or upon deployment of other host or microbial genotypes.
In contrast to the examples outlined above, infection phenotypes with the fungal pathogens C. higginsianum and F. oxysporum and the bacterial pathogen P. syringae were overall consistently affected in both mlo2 mlo6 mlo12 mutant lines. In the case of F. oxysporum and P. syringae, the triple mutant plants showed enhanced disease symptoms as exemplified by either extended tissue chlorosis and necrosis (F. oxysporum; Fig. 5) or more pronounced leaf chlorosis (P. syringae; Fig. 6). In the case of F. oxysporum, a pathogen that colonizes plants via roots44, the enhanced disease symptoms could be caused by both root or shoot effects of the mlo triple mutant. Since at the onset of symptom development (5–7 dpi; Fig. 5A) the fungus typically has reached shoot tissue45, we are at present unable to discriminate between these two possibilities. For P. syringae we recorded about two-fold higher bacterial titres in the mutants (Fig. 6). This contrasts with the situation for C. higginsianum, where we found lower levels of host cell penetration with the fungal pathogen in mutant plants (Fig. 4). Thus, infection phenotypes of virulent pathogens can be modulated in either direction, towards enhanced susceptibility (as observed for F. oxysporum and P. syringae) or towards increased resistance (as found for G. orontii and C. higginsianum) in mlo2 mlo6 mlo12 mutant plants (Table 2). The higher titres of P. syringae pv. maculicola strain ES4326 in the triple mutant were somewhat unexpected since a previous study reported enhanced resistance of mlo2-6 and mlo2-7 single mutant plants upon challenge with P. syringae pv. tomato DC300046. The seeming discrepancy might be explained by the usage of different bacterial pathovars and/or differences between mlo2 single and mlo2 mlo6 mlo12 triple mutant plants. Notably, except maybe for C. higginsianum (Fig. S2), the two distinct mlo2 mlo6 mlo12 mutant lines behaved in an indistinguishable manner in response to the various pathogenic and endophytic microbes, suggesting that the observed phenotypes reflect the authentic effect of the mlo2 mlo6 mlo12 mutations and are not caused by any second-site mutations in these lines.
The data of the present study complement former analyses of the mlo2-5 mlo6-2 mlo12-1 mutant line with an extended range of pathogens. Previously, we reported enhanced powdery mildew resistance (G. orontii, O. neolycopersici, E. pisi, Bgh), but also enhanced disease symptoms and/or an increased pathogen biomass upon challenge with the adapted necrotrophic fungi A. alternata and A. brassicicola and the non-adapted hemibiotrophic oomycete P. infestans 11 (Table 2). Likewise, we discovered apart from powdery mildews one further example of enhanced disease resistance for the mlo2-5 mlo6-2 mlo12-1 mutant. This was a camalexin-sensitive isolate of the necrotrophic fungal pathogen Botrytis cinerea, which caused less disease symptoms on the mutant line than on Col-0 wild-type plants20 (Table 2). In most of these cases, the mlo2-5 mlo6-2 mlo12-1 triple mutant showed stronger alterations of the infection phenotype than the respective mlo2-5 single mutant, suggesting a general cooperative effect of the three Arabidopsis MLO genes in modulation of immunity (Table 2). In the present work, we extended these former studies conducted with only one mlo2 mlo6 mlo12 triple mutant, which are less conclusive due to the presence of potential second-site mutations in the genetic background of a single line. In addition, most of the previous pathogen assays lacked quantification; thus the present dataset is more complete and robust than the former analyses.
Apart from the Arabidopsis mlo2 mlo6 mlo12 triple mutants studied here in detail, the barley mlo mutant has been most extensively investigated regarding the infection phenotypes with phytopathogens other than powdery mildew (Bgh). It was for example reported that interactions with obligate biotrophic rust fungi (leaf rust, Puccinia hordei; stripe rust, Puccinia striiformis; stem rust, Puccinia graminis f.sp. tritici), the take-all fungus (Gaeumannomyces graminis) and the scald fungus (Rhynchosporium secalis) were not affected by mutations in barley Mlo 47. In that study it was, however, noted that the premature leaf senescence phenotype of mlo mutants, which is associated with necrotic leaf spotting, may somewhat reduce the extent of colonization by rusts (i.e., necrotic areas not being colonized by these obligate biotrophic fungi). Though that report provides valuable information regarding a number of common barley diseases, it rests in part on “personal communications” and lacks any cytological assessment and/or quantitative data.
In contrast to these pathogens, leading to seemingly unaltered infection phenotypes, a number of phytopathogens have been described that show an aberrant outcome of colonization on barley mlo genotypes. A prominent example is the interaction of barley with the rice blast fungus, Magnaporthe oryzae. While Mlo wild-type plants show limited colonization by the fungal pathogen, mlo mutant plants are hyper-susceptible to M. oryzae 48. In addition, mlo mutant lines display more severe disease symptoms than Mlo genotypes in response to Ramularia collo-cygni, the causal agent of Ramularia leaf spot disease49. Furthermore, it has been reported that the mutant shows accelerated spread of the fungal pathogen Fusarium graminearum in infected ears50. However, a recent study did not find an increased DNA content of the latter two pathogens on barley mlo genotypes upon natural infections in the field51. An interesting example of a seemingly developmentally controlled effect of mlo mutations on infection by the hemibiotrophic pathogen Phytophthora palmivora has been reported lately52. The authors of this study found that the oomycete shows delayed colonization of young tissues in barley mlo plants. In addition to encounters with pathogens, the interaction with the beneficial mycorrhizal fungus Funneliformis mosseae (syn. Glomus mosseae) also seems to be affected in barley mlo mutant plants. Colonization intensity and arbuscule abundance were found to be reduced in a mlo null mutant compared to its isogenic parental Mlo wild type line53.
Based on the recent study, reporting that pre-infection of Arabidopsis with A. laibachii can break non-host resistance against the Irish famine pathogen, P. infestans 42, we explored whether pre-inoculation of our mlo triple mutant lines with A. laibachii can likewise result in a breakdown of mlo resistance. We found that pre-inoculation with A. laibachii did not affect the resistance phenotype of mlo2 mlo6 mlo12 triple mutant plants under our conditions (Table 1). This outcome is consistent with the fact that A. laibachii also failed to overcome non-host resistance of Arabidopsis against the barley powdery mildew pathogen42. Maybe the defence suppression exerted by the white blister rust pathogen is not effective in the leaf epidermis, the target tissue of powdery mildew pathogens. Alternatively, or in addition, the lack of compatibility conferred by mlo triple mutants might be too powerful to be disabled by A. laibachii. This would be in line with the results of recent genetic analysis, demonstrating that single or even multiple defence pathways are dispensable for resistance of the mlo2-5 mlo6-2 mlo12-1 triple mutant54.
Taken together, mutations in genes MLO2, MLO6 and MLO12 in Arabidopsis and Mlo in barley, which condition broad-spectrum powdery mildew resistance in these species, also modify the outcome of infections with some other, but not all, microbial species (Table 2). This situation is consistent with an authentic and broadly acting role of the respective MLO proteins in plant immunity. Data from co-expression networks support such a role for MLO proteins in immunity55. The changed infection phenotypes could thus be the result of perturbed regulation of immunity in the absence of MLO proteins, which may be to the benefit of some microbes and to the disadvantage of others.
It remains, however, puzzling that no consistent pattern is recognizable on first view regarding the altered infection phenotypes. The tested microbes belong to different kingdoms of life (bacteria, oomycetes and fungi) and engage in different modes of plant colonization and trophic lifestyles (obligate biotrophs, hemibiotrophs and necrotrophs). We found a tendency for enhanced susceptibility in mlo2 mlo6 mlo12 mutant plants to correlate with a hemibiotrophic/necrotrophic lifestyle (Table 2). The exception from this trend is the hemibiotrophic fungus C. higginsianum, which shows reduced host cell entry on mlo2 mlo6 mlo12 genotypes (Fig. 4B). An obvious commonality in the mode of infection between powdery mildews and C. higginsianum is that both phytopathogens directly enter epidermal host cells via appressoria/infection pegs that breach the cuticle and underlying cell wall (refs 27, 56 and Table 2). By contrast, leaf-colonizing A. laibachii and P. syringae enter host tissue through stomata26, 57, while H. arabidopsidis penetrates via anticlinal cell walls of neighboring epidermal pavement cells24. The remaining two microbes, F. oxysporum and S. indica, gain access to plant tissues via the roots34, 44, which lack a cuticle. Thus, the mode of host cell entry might be a key factor determining the modulation of the infection phenotype on mlo2 mlo6 mlo12 mutant plants. This notion is largely consistent with results from previous patho-assays with this mutant (Table 2). A. alternata, A. brassicicola and P. infestans may be considered as possible exceptions in this respect. However, these pathogens have been reported to invade by both entry through stomata and direct host cell penetration, depending on environmental factors such as temperature and humidity58, 59.
It nevertheless remains unclear why mlo mutants typically show a spectacular level of resistance against powdery mildews, while infection phenotypes to other pathogens are generally only mildly impacted. One possibility is that powdery mildew fungi actively target MLO proteins for defence suppression, as previously suggested60, 61, or interfere with membrane dynamics or vesicle trafficking for compatibility62. Notably, it also appears that mlo genotypes of some plant species (such as Arabidopsis and barley) show altered interactions with pathogens apart from powdery mildews, whereas in other plant species (e.g. pea and tomato) this seems not to be the case14, 15. It thus remains a future task to disentangle which molecular parameters determine whether a microbial species shows a modified infection phenotype on mlo plants and why some plant species are affected by this phenomenon and others are not.
Materials and Methods
Plant materials
Triple mutants mlo2-5 mlo6-2 mlo12-1 and mlo2-6 mlo6-4 mlo12-8 were generated by crossing and are based on the following T-DNA insertion single mutants (all in the genetic background of accession Col-0): mlo2-5 (SAIL_878_H12), mlo2-6 (SALK_050191), mlo6-2 (SAIL_0523_D09), mlo6-4 (SALK_039680), mlo12-1 (SLAT 24-21) and mlo12-8 (SALK_041042). The mlo2-5 mlo6-2 mlo12-1 mutant has been described before11. In addition to the mlo triple mutant lines, Arabidopsis accessions Columbia (Col-0), Landsberg erecta (Ler) and Keswick (Ksk-1) as well as mutants edr1 41, eds1-236, npr1-163, pmr4-110, sid2-164 and sid2-2/eds16-140 were used in this study. All mutant lines are in the Arabidopsis thaliana Col-0 background.
RT-PCR analysis
Total RNA was extracted from rosette leaves of 6-week-old plants using the Nucleospin RNA plus kit (Macherey-Nagel, Düren, Germany). cDNA was generated from total RNA (1 µg) with the High Capacity RNA-to-cDNA kit (Thermo Fisher Scientific, Waltham, MA, USA) following the manufacturer’s instructions. RT-PCR analysis (35 cycles) was performed with the following primer pairs: AtMLO2 (2 C 5′-ATG GAG ATG GAG ATA AAC CCG GTC-3′ and 2bw1 5′-ACT AGT ATC TAG GAG AAG GAG-3′; amplicon size ca. 1.2 kb), AtMLO6 (6 G 5′-GCT TTC TTT GTC TGG AGT ACG-3′ and 6B4 5′-CAA GAA CTG GTT TCA TTT AGC-3′; amplicon size ca. 0.65 kb) and AtMLO12 (18 K 5′-AAA GTA GCA TTA GTA TCT GCC-3′ and 18 L 5′-ATG TCT TCT GTT TTG TGG TGG-3′; amplicon size ca. 0.8 kb).
G. orontii infection assays
Plants were grown under short-day conditions (8 h light, 22 °C; 16 h dark, 20 °C; 80 μmol m−2 s−1) and inoculated at the age of six weeks. Heavily G. orontii-infected leaves of accession Col-0 were used for leaf-to-leaf contact inoculation. Samples for microscopy were taken at 48 hpi and cleared in destaining solution (1:2 mixture of stock solution and ethanol; stock solution 1:2:1 mixture of lactic acid, glycerol and deionized H2O) at room temperature. Fungal structures were visualized with Coomassie Brilliant Blue staining (0.6% Coomassie Brilliant Blue R-250 (Carl Roth GmbH, Karlsruhe, Germany) in ethanol) by briefly dipping the specimens into the staining solution shortly prior to mounting the samples for microscopy. For quantification of fungal host cell entry, the proportion of germinated fungal sporelings that developed secondary hyphae served as an approximation of penetration success.
H. arabidopsidis infection assays
H. arabidopsidis isolate Noco2 was kindly provided by Jane Parker (Max Planck Institute for Plant Breeding Research, Cologne, Germany) and maintained on Arabidopsis Col-0 by weekly transfer to healthy 14- and 21-day-old seedlings. For inoculation, healthy 16-day-old seedlings were evenly sprayed with a spore suspension (4 × 104 spores ml−1). Plants were allowed to dry and sprayed once more with the spore suspension. After drying, plants were kept under a sealed lid (100% relative humidity) in a growth chamber at 16 °C with a 10 h photoperiod (100 μmol m−2 s−1). Seven days after inoculation, aerial parts of the plants were harvested and the fresh weight (FW) was determined. Evaluation of H. arabidopsidis propagation was performed by quantifying conidiospores with a hemocytometer as described65 and represented as the number of conidiospores per gram FW. For normalization, the corresponding value for Col-0 was set as 100%. Ler and eds1-2 plants were used as resistant and super-susceptible controls, respectively.
A. laibachii infection assays
Plants were grown in short-day conditions (10 h light, 22 °C, 65% humidity/14 h dark, 21 °C, 60% humidity, 40 μmol m−2 s−1) and inoculated at the age of six weeks. A. laibachii (isolate Nc1466) zoospores obtained from propagation on Arabidopsis accession Ws-0 were suspended in water (105 spores ml−1) and incubated on ice for 30 min. The spore suspension was filtered through Miracloth (Calbiochem, San Diego, CA, USA) and sprayed onto the plants using a spray gun (~700 μl/plant), both on the adaxial and abaxial sides of rosette leaves. Plants were incubated at 8 °C in a cold room in the dark overnight. Inoculated plants were kept under 10 h light/14 h dark cycles with a 20 °C day and 16 °C night temperature. Infection rates were determined at 10 dpi for 5–6 individuals per genotype by counting numbers of infected versus uninfected leaves.
C. higginsianum infection assays
For C. higginsianum infection assays, cultures of the fungal isolate IMI349063 and Arabidopsis plants were grown as described previously67. Plants were spray-inoculated with conidial suspension (5 × 105 spores ml−1), covered in sealed propagators to maintain 100% humidity and incubated in a controlled environment (16 h photoperiod, 80 μmol m−2 s−1, 25 °C). Methods for Trypan blue-lactophenol staining of infected leaves and subsequent clearing of the stained tissues in chloral hydrate were adapted from68. To quantify the frequency of appressorial penetration, Arabidopsis leaves were cleared in ethanol:chloroform (3:1) and mounted on slides in lactophenol. Appressorium-based penetration was scored as the presence of a visible hypha in the underlying epidermal cell at 3 dpi.
F. oxysporum infection assays
Arabidopsis plants used for inoculation with F. oxysporum were grown in controlled conditions with 11 h light/13 h darkness (90 μmol m−2 s−1) at 22 °C. Two-week-old seedlings were uprooted from soil, cleaned with tap water and placed in a spore suspension of F. oxysporum (isolate Fo517669) for 5 min. Spores were harvested from a 5-day-old liquid culture38 by filtering through three layers of Miracloth. Spores were resuspended in tap water to 106 spores/ml for Arabidopsis inoculation. Autoclaved tap water was used for mock treatment. Inoculated and mock-treated seedlings were replanted into the original soil, placed under a transparent plastic hood to maintain humidity, and grown under controlled conditions with 11 h light/13 h darkness (28 °C, 70% relative humidity). The disease index was scored at 5, 7 and 10 dpi on a scale ranging from 0 to 5 (0 = no disease symptoms; 1 = 1–2 leaves with yellow veins or plant is smaller; 2 = some fully developed leaves show chlorosis or yellow veins; 3 = most fully developed leaves show chlorosis; 4 = all fully developed leaves show chlorosis; 5 = dead; adapted from38).
S. indica infection assays
Serendipita indica (syn. Piriformospora indica) isolate DSM1182733 was obtained from the German collection of microorganisms and cell cultures in Braunschweig (Germany). The fungus was grown on complete medium70 supplemented with 1.5% agar at 24 °C in the dark. Arabidopsis plants were grown under sterile conditions (ca. 60 seedlings per plate) in a growth cabinet (120 μmol m−2 s−1; 10 h light; 22 °C/18 °C (light/dark); 60% humidity) on vertically placed square petri dishes containing growth medium (5 mM KNO3, 2.5 mM KH2PO4, 3 mM MgSO4, 3 mM, Ca(NO3)2, 50 μM Fe-EDTA 70 μM H3BO3, 14 μM MnCI2, 0.5 μM CuSO4, 1 μM ZnSO4, 0.2 μM NaMoO4, 10 μM NaCl, 0.01 μM CoCl2), modified as described71 and supplemented with 0.45% Gelrite (Duchefa Biochemie, Haarlem, The Netherlands). Roots of 2-week-old plants were inoculated with 5 × 105 S. indica chlamydospores (1 ml per petri dish). The inoculated roots were harvested at 3 and 7 days after inoculation for subsequent DNA extraction. Genomic plant and fungal DNA was obtained as described72. qRT-PCR analysis was performed on a Bio-Rad iCycler iQ5 using a standard protocol. Forty ng of genomic DNA served as template with 20 µL of SYBR Green JumpStart Taq ReadyMix (Sigma-Aldrich, Taufkirchen, Germany) and 350 nM oligonucleotides. Fungal colonization was determined by the comparative C T method73 by subtracting the raw cycle threshold values of S. indica Internal Transcribed Spacer (5′-CAACACATGTGCACGTCGAT-3′/5′-CCAATGTGCATTCAGAACGA-3′) from those of AtUbi5 (At3g62250; 5′-CCAAGCCGAAGAAGATCAAG-3′/5′-ATGACTCGCCATGAAAGTCC-3′).
P. syringae infection assays
Arabidopsis seeds were stratified and then grown on soil (Dachstaudensubstrat SoMi 513; Hawita). Growth conditions for experiments 1-4 were: 10 h day (80 μmol m−2 s−1)/14 h night cycle, 21 °C, and a relative humidity of 68%; experiment 5: 10 h day (80 μmol m−2 s−1) at 21 °C/14 h night cycle at 18 °C, and a relative humidity of 68%. For experiment 5, the soil was sterilized by microwaving for 10 min before usage. Fourteen-day-old healthy seedlings were individually transferred to bigger pots (9 × 9 × 8 cm).
P. syringae pv. maculicola strain ES4326 carrying the luxCDABE operon from Photorhabdus luminescens under the control of a constitutive promoter (Psm lux 39; kindly provided by Prof. Jürgen Zeier, Heinrich Heine University Düsseldorf, Germany) was cultivated at 28 °C in King’s B medium supplemented with 50 μg ml−1 rifampicin and 25 μg ml−1 kanamycin. Overnight log phase liquid cultures were diluted to an OD600 of 0.001 (experiments 1 and 2) or 0.0005 (experiments 3-5) in 10 mM MgCl2. Leaf inoculations were performed with 5-week-old plants of uniform, healthy appearance. Using a needleless 1 ml syringe, three mature leaves per plant were infiltrated from the abaxial side. The bacterial growth in the leaves was determined via the bioluminescence of the Psm lux stain at 0 dpi (following infiltration) and at 3 dpi. The measurement was carried out on leaf discs (r = 3 mm, adaxial side up) from inoculated and control leaves, individually placed in 250 µl MgCl2 in 96-well microtiter plates (flat bottom). The bioluminescence of each leaf disc was monitored as a 10 s averaged measurement per well with a 10 s delay at the start of each plate using the ‘Centro XS³ LB 960 Microplate Luminometer’ and the corresponding ‘MikroWin 2000 software’ (Berthold Technologies, https://www.berthold.com/). The average auto-luminescence (background of untreated leaves) was subtracted from the luminescence values of the treated leaves, and the average of three leaf discs per plant was determined. Increasing luminescence values (indicated by relative light units, RLU) correspond to increasing bacterial titres and reflect bacterial growth determined by the standard plate assay39.
Statistical analysis
Statistical analyses were carried out using R v3.3.2 software for Windows74 (http://www.r-project.org/). Owing to limited sample size, non-normal distribution and unequal variance, we performed non-parametric statistical tests. Datasets for G. orontii, F. oxysporum, C. higginsianum, S. indica and P. syringae were analysed by Generalized Linear Modeling (GLM), assuming Poisson or quasi-Poisson distribution (continuous datasets) or binomial distribution (ratio/percentage datasets), as recommended for this type of data75. The other datasets (H. arabidopsidis and A. laibachii) were additionally analysed by Wilcoxon-Mann-Whitney rank sum test.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files). Respective raw data are available from the corresponding author upon request.
References
Glawe, D. A. The powdery mildews: A review of the world’s most familiar (yet poorly known) plant pathogens. Annu. Rev. Phytopathol. 46, 27–51 (2008).
Takamatsu, S. Phylogeny and evolution of the powdery mildew fungi (Erysiphales, Ascomycota) inferred from nuclear ribosomal DNA sequences. Mycoscience 45, 147–157 (2004).
Kuhn, H. et al. Biotrophy at its best: Novel findings and unsolved mysteries of the Arabidopsis-powdery mildew pathosystem. The Arabidopsis Book 14, e0184 (2016).
Micali, C., Göllner, K., Humphry, M., Consonni, C. & Panstruga, R. The powdery mildew disease of Arabidopsis. a paradigm for the interaction between plants and biotrophic fungi. The Arabidopsis Book 6, e0115 (2008).
Xiao, S. Y. et al. Broad-spectrum mildew resistance in Arabidopsis thaliana mediated by RPW8. Science 291, 118–120 (2001).
Orgil, U., Arakit, H., Tangchaiburana, S., Berkey, R. & Xiao, S. Intraspecific genetic variations, fitness cost and benefit of RPW8, a disease resistance locus in Arabidopsis thaliana. Genetics 176, 2317–2333 (2007).
Göllner, K., Schweizer, P., Bai, Y. & Panstruga, R. Natural genetic resources of Arabidopsis thaliana reveal a high prevalence and unexpected phenotypic plasticity of RPW8-mediated powdery mildew resistance. New Phytol. 177, 725–742 (2008).
Wang, W. M., Wen, Y. Q., Berkey, R. & Xiao, S. Y. Specific targeting of the Arabidopsis resistance protein RPW8.2 to the interfacial membrane encasing the fungal haustorium renders broad-spectrum resistance to powdery mildew. Plant Cell 21, 2898–2913 (2009).
Devoto, A. et al. Molecular phylogeny and evolution of the plant-specific seven-transmembrane MLO family. J. Mol. Evol. 56, 77–88 (2003).
Vogel, J. & Somerville, S. Isolation and characterization of powdery mildew-resistant Arabidopsis mutants. Proc. Natl. Acad. Sci. U.S.A. 97, 1897–1902 (2000).
Consonni, C. et al. Conserved requirement for a plant host cell protein in powdery mildew pathogenesis. Nat. Genet. 38, 716–720 (2006).
Jørgensen, J. H. Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 63, 141–152 (1992).
Lyngkjær, M. F., Newton, A. C., Atzema, J. L. & Baker, S. J. The barley mlo -gene: an important powdery mildew resistance source. Agronomie 20, 745–756 (2000).
Humphry, M., Reinstädler, A., Ivanov, S., Bisseling, T. & Panstruga, R. Durable broad-spectrum powdery mildew resistance in pea er1 plants is conferred by natural loss-of-function mutations in. PsMLO1. Mol. Plant Pathol. 12, 866–878 (2011).
Bai, Y. L. et al. Naturally occurring broad-spectrum powdery mildew resistance in a central American tomato accession is caused by loss of Mlo function. Mol. Plant-Microbe Interact. 21, 30–39 (2008).
Wang, Y. et al. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 32, 947–951 (2014).
Acevedo-Garcia, J. et al. mlo-based powdery mildew resistance in hexaploid bread wheat generated by a non-transgenic TILLING approach. Plant Biotechnol. J. 15, 367–378 (2017).
Kusch, S. & Panstruga, R. mlo-based resistance: An apparently universal “weapon” to defeat powdery mildew disease. Mol. Plant-Microbe Interact. 30, 179–189 (2017).
Humphry, M., Consonni, C. & Panstruga, R. mlo-based powdery mildew immunity. Silver bullet or simply non-host resistance? Mol. Plant Pathol. 7, 605–610 (2006).
Consonni, C. et al. Tryptophan-derived metabolites are required for antifungal defense in the Arabidopsis mlo2 mutant. Plant Physiol. 152, 1544–1561 (2010).
Wolter, M., Hollricher, K., Salamini, F. & Schulze-Lefert, P. The mlo resistance alleles to powdery mildew infection in barley trigger a developmentally controlled defense mimic phenotype. Mol. Gen. Genet. 239, 122–128 (1993).
Peterhänsel, C., Freialdenhoven, A., Kurth, J., Kolsch, R. & Schulze-Lefert, P. Interaction analyses of genes required for resistance responses to powdery mildew in barley reveal distinct pathways leading to leaf cell death. Plant Cell 9, 1397–1409 (1997).
Katagiri, F., Thilmony, R. & He, S. Y. The Arabidopsis thaliana-Pseudomonas syringae interaction. The Arabidopsis Book 1, e0039 (2002).
Slusarenko, A. J. & Schlaich, N. L. Downy mildew of Arabidopsis thaliana caused by Hyaloperonospora parasitica (formerly Peronospora parasitica). Mol. Plant Pathol. 4, 159–170 (2003).
Thines, M. et al. A new species of Albugo parasitic to Arabidopsis thaliana reveals new evolutionary patterns in white blister rusts (Albuginaceae). Persoonia 22, 123–128 (2009).
Holub, E. B. et al. Phenotypic and genotypic variation in the interaction between Arabidopsis thaliana and Albugo candida. Mol. Plant-Microbe Interact. 8, 916–928 (1995).
O’Connell, R. et al. A novel Arabidopsis-Colletotrichum pathosystem for the molecular dissection of plant-fungal interactions. Mol. Plant-Microbe Interact. 17, 272–282 (2004).
Diener, A. C. & Ausubel, F. M. RESISTANCE TO FUSARIUM OXYSPORUM 1, a dominant Arabidopsis disease-resistance gene, is not race specific. Genetics 171, 305–321 (2005).
Berrocal-Lobo, M. & Molina, A. Arabidopsis defense response against Fusarium oxysporum. Trends Plant Sci. 13, 145–150 (2008).
García, E., Alonso, Á., Platas, G. & Sacristán, S. The endophytic mycobiota of Arabidopsis thaliana. Fungal Divers. 60, 71–89 (2013).
Bai, Y. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369 (2015).
Peskan-Berghöfer, T. et al. Association of Piriformospora indica with Arabidopsis thaliana roots represents a novel system to study beneficial plant-microbe interactions and involves early plant protein modifications in the endoplasmic reticulum and at the plasma membrane. Physiol Plant 122, 465–477 (2004).
Verma, S. et al. Piriformospora indica, gen. et sp. nov. a new root-colonizing fungus. Mycologia 90, 896 (1998).
Jacobs, S. et al. Broad-spectrum suppression of innate immunity is required for colonization of Arabidopsis roots by the fungus Piriformospora indica. Plant Physiol. 156, 726–740 (2011).
Parker, J. E. et al. Characterization of eds1, a mutation in Arabidopsis suppressing resistance to Peronospora parasitica specified by several different RPP genes. Plant Cell 8, 2033–2046 (1996).
Bartsch, M. et al. Salicylic acid-independent ENHANCED DISEASE SUSCEPTIBILITY1 signaling in Arabidopsis immunity and cell death is regulated by the monooxygenase FMO1 and the nudix hydrolase NUDT7. Plant Cell 18, 1038–1051 (2006).
Agler, M. T. et al. Microbial hub taxa link host and abiotic factors to plant microbiome variation. PLoS Biol. 14, e1002352 (2016).
Gawehns, F. et al. The Fusarium oxysporum effector Six6 contributes to virulence and suppresses I-2-mediated cell death. Mol. Plant-Microbe Interact. 27, 336–348 (2014).
Fan, J., Crooks, C. & Lamb, C. High-throughput quantitative luminescence assay of the growth in planta of Pseudomonas syringae chromosomally tagged with Photorhabdus luminescens luxCDABE. Plant J. 53, 393–399 (2008).
Dewdney, J. et al. Three unique mutants of Arabidopsis identify eds loci required for limiting growth of a biotrophic fungal pathogen. Plant J. 24, 205–218 (2000).
Frye, C. A. & Innes, R. W. An Arabidopsis mutant with enhanced resistance to powdery mildew. Plant Cell 10, 947–956 (1998).
Belhaj, K. et al. Arabidopsis late blight: infection of a nonhost plant by Albugo laibachii enables full colonization by Phytophthora infestans. Cell Microbiol. (2016).
Elliott, C. et al. Conserved extracellular cysteine residues and cytoplasmic loop-loop interplay are required for functionality of the heptahelical MLO protein. Biochem. J. 385, 243–254 (2005).
Rodríguez-Gálvez, E. & Mendgen, K. The infection process of Fusarium oxysporum in cotton root tips. Protoplasma 189, 61–72 (1995).
Lyons, R. et al. Fusarium oxysporum triggers tissue-specific transcriptional reprogramming in Arabidopsis thaliana. PLoS One 10, e0121902 (2015).
Lewis, J. D. et al. Quantitative Interactor Screening with next-generation Sequencing (QIS-Seq) identifies Arabidopsis thaliana MLO2 as a target of the Pseudomonas syringae type III effector HopZ2. BMC Genomics 13, 8 (2012).
Jørgensen, J. H. Spectrum of resistance conferred by ML-O powdery mildew resistance genes in barley. Euphytica 26, 55–62 (1977).
Jarosch, B., Kogel, K. H. & Schaffrath, U. The ambivalence of the barley Mlo locus: Mutations conferring resistance against powdery mildew (Blumeria graminis f. sp. hordei) enhance susceptibility to the rice blast fungus Magnaporthe grisea. Mol. Plant-Microbe Interact. 12, 508–514 (1999).
McGrann, G. R. D. et al. A trade off between mlo resistance to powdery mildew and increased susceptibility of barley to a newly important disease, Ramularia leaf spot. J. Exp. Bot. 65, 1025–1037 (2014).
Jansen, C. et al. Infection patterns in barley and wheat spikes inoculated with wild-type and trichodiene synthase gene disrupted Fusarium graminearum. Proc. Natl. Acad. Sci. USA 102, 16892–16897 (2005).
Hofer, K., Linkmeyer, A., Textor, K., Hückelhoven, R. & Hess, M. MILDEW LOCUS O mutation does not affect resistance to grain infections with Fusarium spp. and Ramularia collo-cygni. Phytopathology 105, 1214–1219 (2015).
Le Fevre, R., O’Boyle, B., Moscou, M. J. & Schornack, S. Colonization of barley by the broad-host hemibiotrophic pathogen Phytophthora palmivora uncovers a leaf development-dependent involvement of Mlo. Mol. Plant-Microbe Interact. 29, 385–395 (2016).
Ruiz-Lozano, J. M., Gianinazzi, S. & Gianinazzi-Pearson, V. Genes involved in resistance to powdery mildew in barley differentially modulate root colonization by the mycorrhizal fungus Glomus mosseae. Mycorrhiza 9, 237–240 (1999).
Kuhn, H. et al. Key components of different plant defense pathways are dispensable for powdery mildew resistance of the Arabidopsis mlo2 mlo6 mlo12 triple mutant. Front. Plant Sci. 8, 1006 (2017).
Humphry, M. et al. A regulon conserved in monocot and dicot plants defines a functional module in antifungal plant immunity. Proc. Natl. Acad. Sci. U.S.A. 107, 21896–21901 (2010).
Bélanger, R. R., Bushnell, W. R., Dik, A. J., Carver, T. L. (eds). The powdery mildews. A comprehensive treatise. (APS Press, St. Paul, 2002).
Melotto, M., Underwood, W., Koczan, J., Nomura, K. & He, S. Y. Plant stomata function in innate immunity against bacterial invasion. Cell 126, 969–980 (2006).
Hohl, H. R. & Suter, E. Host–parasite interfaces in a resistant and a susceptible cultivar of Solanum tuberosum inoculated with Phytophthora infestans: leaf tissue. Can. J. Bot. 54, 1956–1970 (1976).
Sharma, P., Deep, S., Bhati, D. S., Sharma, M. & Chowdappa, P. Penetration and infection processes of Alternaria brassicicola on cauliflower leaf and Alternaria brassicae on mustard leaf. A Histopathological Study. Plant Pathol. J. 13, 100–111 (2014).
Panstruga, R. & Schulze-Lefert, P. Corruption of host seven-transmembrane proteins by pathogenic microbes: a common theme in animals and plants? Microbes Infect. 5, 429–437 (2003).
Panstruga, R. Serpentine plant MLO proteins as entry portals for powdery mildew fungi. Biochem. Soc. Trans. 33, 389–392 (2005).
van Schie, C. C. N. & Takken, F. L. W. Susceptibility genes 101: how to be a good host. Annu. Rev. Phytopathol. 52, 551–581 (2014).
Cao, H., Bowling, S. A., Gordon, A. S. & Dong, X. N. Characterization of an Arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance. Plant Cell 6, 1583–1592 (1994).
Nawrath, C. & Metraux, J. P. Salicylic acid induction-deficient mutants of Arabidopsis express PR-2 and PR-5 and accumulate high levels of camalexin after pathogen inoculation. Plant Cell 11, 1393–1404 (1999).
Asai, S., Shirasu, K. & Jones, J. Hyaloperonospora arabidopsidis (downy mildew) infection assay in Arabidopsis. Bio Protoc. 5 (2015).
Kemen, E. et al. Gene gain and loss during evolution of obligate parasitism in the white rust pathogen of Arabidopsis thaliana. PLoS Biol. 9 (2011).
Huser, A., Takahara, H., Schmalenbach, W. & O’Connell, R. Discovery of pathogenicity genes in the crucifer anthracnose fungus Colletotrichum higginsianum, using random insertional mutagenesis. Mol. Plant-Microbe Interact. 22, 143–156 (2009).
Keogh, R. C., Deverall, B. J. & McLeod, S. Comparison of histological and physiological responses to Phakopsora pachyrhizi in resistant and susceptible soybean. Trans. Br. Mycol. Soc. 74, 329–333 (1980).
Thatcher, L. F., Gardiner, D. M., Kazan, K. & Manners, J. M. A highly conserved effector in Fusarium oxysporum is required for full virulence on Arabidopsis. Molecular plant-microbe interactions: MPMI 25, 180–190 (2012).
Pham, G. H. et al. In Plant surface microbiology, edited by A. Varma, L. Abbott, D. Werner & Hampp R (Springer, Berlin Heidelberg New York, 2004).
Estelle, M. A. & Somerville, C. Auxin-resistant mutants of Arabidopsis thaliana with an altered morphology. Mol. Gen. Genet. 206, 200–206 (1987).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108 (2008).
R Core Team. R: A language and environment for statistical computing. Available at http://www.R-project.org/.
Crawley, M. J. Statistics. An introduction using R. (John Wiley & Sons, Inc, Chichester, West Sussex, UK, 2014).
Acknowledgements
We thank Jane Parker (Max Planck Institute for Plant Breeding Research, Cologne, Germany) for providing H. arabidopsidis isolate Noco2, Jürgen Zeier (Heinrich-Heine University Düsseldorf, Germany) for sharing the Psm lux strain and Uwe Conrath (RWTH Aachen University) for making available edr1 and sid2-2 mutant seeds.
Author information
Authors and Affiliations
Contributions
J.A.G., K.G., A.R., A.K., L.C., M.U.R., R.O.C. and H.K. performed the pathogen assays, R.P. and J.A.G. planned the project, E.K., F.L.W.T., P.S., R.O.C. and R.P. analysed the data, S.K. performed the statistical analysis and R.P. wrote the manuscript. All authors proofread and approved the document.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Acevedo-Garcia, J., Gruner, K., Reinstädler, A. et al. The powdery mildew-resistant Arabidopsis mlo2 mlo6 mlo12 triple mutant displays altered infection phenotypes with diverse types of phytopathogens. Sci Rep 7, 9319 (2017). https://doi.org/10.1038/s41598-017-07188-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-07188-7
This article is cited by
-
Tomato leaves under stress: a comparison of stress response to mild abiotic stress between a cultivated and a wild tomato species
Plant Molecular Biology (2021)
-
A stable DNA-free screening system for CRISPR/RNPs-mediated gene editing in hot and sweet cultivars of Capsicum annuum
BMC Plant Biology (2020)
-
Arabidopsis mlo3 mutant plants exhibit spontaneous callose deposition and signs of early leaf senescence
Plant Molecular Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.