Abstract
Invasiveness and the impacts of introduced plants are known to be mediated by plant-microbe interactions. Yet, the microbial communities associated with invasive plants are generally poorly understood. Here we report on the first comprehensive investigation of the bacterial and fungal communities inhabiting the rhizosphere and the surrounding bulk soil of a widespread invasive tree, Acacia dealbata. Amplicon sequencing data indicated that rhizospheric microbial communities differed significantly in structure and composition from those of the bulk soil. Two bacterial (Alphaproteobacteria and Gammaproteobacteria) and two fungal (Pezizomycetes and Agaricomycetes) classes were enriched in the rhizosphere compared with bulk soils. Changes in nutritional status, possibly induced by A. dealbata, primarily shaped rhizosphere soil communities. Despite a high degree of geographic variability in the diversity and composition of microbial communities, invasive A. dealbata populations shared a core of bacterial and fungal taxa, some of which are known to be involved in N and P cycling, while others are regarded as plant pathogens. Shotgun metagenomic analysis also showed that several functional genes related to plant growth promotion were overrepresented in the rhizospheres of A. dealbata. Overall, results suggest that rhizosphere microbes may contribute to the widespread success of this invader in novel environments.
Similar content being viewed by others
Introduction
Invasive trees are a global concern because they pose a direct threat to natural ecosystems and cause important economic losses, although they also provide goods and services that sustain human populations1. Australian acacias are among the most invasive trees worldwide2. In their invasive ranges, acacias have had numerous and severe environmental impacts3, 4. For example, Acacia dealbata (silver wattle) is an aggressive invader in southern Europe, the Americas and southern Africa2. In South Africa, this species reduces water availability5, and alters faunal6 and microbial7 community diversity and composition. Acacia dealbata also reduces the abundance and cover of native plant species under its understory8 and modifies soil basal respiration and soil enzymatic activities9, two important proxies for ecosystem functioning.
Plant species traits (e.g., high seed outputs, adaptability), abiotic factors (e.g., similar climate and soil chemistry in native and invaded ranges), native range biogeography, and deliberate and accidental human intervention10, have been commonly used to explain why many acacia species have become invasive in new environments11.
It is also likely that rhizosphere microbes (native or introduced), such as bacteria and fungi, play an important role in the establishment and invasion success of acacias12. Microbial communities influence many important ecosystem processes, including plant nutrient acquisition and nitrogen and carbon cycling13, which regulate plant diversity and productivity. For example, acacias develop symbiotic associations with rhizobia that convert atmospheric nitrogen into plant available ammonium14. Recent work has shown that invasive acacias in South Africa’s Cape Floristic Region (CFR) associate with a diverse assemblage of Bradyrhizobium strains that are not commonly associated with native legumes15. Acacias also interact with arbuscular mycorrhizal (AMF) and/or ectomycorrhizal (EMF) fungi16 which modify the root system and enhance mobilization and the uptake of several essential elements, especially phosphorus17. On the other hand, microbial parasites and pathogens, through their effects on plant health and productivity, may impact negatively on plant invasiveness18.
Apart from the interactions of acacias with rhizobia and mycorrhiza, there is currently little knowledge about the identity of the overall soil microbial communities associated with acacia species. For instance, for A. dealbata only limited data are available from studies employing low resolution methods and/or low sample sizes19, 20 (but see ref. 7). In addition, most of these studies were carried out in soils in the proximity of plants, but not in the narrow zone of soil that surrounds and is most influenced by plant roots (i.e., the rhizosphere). As rhizosphere microbes can directly and/or indirectly affect the composition and biomass of plant communities21, increasing our knowledge on the root microbiome is important to better understand the factors contributing to plant invasiveness.
Here, we use Illumina MiSeq data (targeting 16S rRNA genes and ITS regions) and shotgun metagenomics to investigate microbial (bacterial and fungal) communities and functional processes associated with the roots of this species in various invasive populations of A. dealbata across South African grasslands, and compare them with those from the bulk soil. More specifically, we evaluate (i) how abiotic (i.e., soil chemistry) factors may shape the diversity and structure of these communities, (ii) identify prominent taxa that may contribute to the success of this invader and (iii) investigate whether rhizosphere and bulk soil microbial communities differ in their metabolic capacity.
Results and Discussion
Analysing the microbial communities associated with the rhizospheres of A. dealbata, we obtained 2,444,152 and 694,434 total high-quality reads, which resulted in 29,678 and 14,471 OTUs (97% cut-off) for bacteria and fungi, respectively. The majority of all prokaryotic OTUs in the rhizosphere were also present in the bulk soils (Supplementary Fig. S2a), whereas a larger proportion of the eukaryotic OTUs were unique to the sample type (Supplementary Fig. S2b). For both bacterial and fungal communities the OTUs shared between rhizosphere and bulk soil samples accounted for the majority of the reads (97.2% and 93% respectively; Supplementary Fig. S2a,b). The percentage of OTUs shared between all eight sampling sites was considerably smaller for fungal data sets than for the bacterial data sets (1.6% and 12%, respectively). This is in agreement with the view that fungal communities generally show more geographic structure than bacterial communities22.
OTU accumulation curves indicated reasonable sequence saturation at a regional level, especially for bacteria (Supplementary Fig. S3). The levels of microbial diversity (richness, Shannon, inverse Simpson, Pielou’s evenness) tended to be higher in the bulk soil than in the rhizosphere samples, although they did not differ significantly (Supplementary Fig. S4). This is in contrast to previous studies19, which showed A. dealbata-invaded zones containing higher bacterial richness and lower fungal richness compared to uninvaded zones. However, it is now accepted that the community fingerprinting method used in that study (i.e., DGGE) is not well suited to accurately estimate microbial richness23.
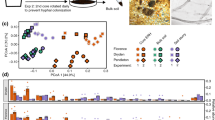
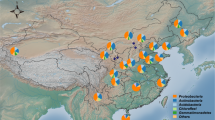
A total of 30 distinct bacterial phyla were detected across all samples. The most abundant sequences were affiliated with the phylum Proteobacteria (27% of total relative abundance), followed by Actinobacteria (23%), Acidobacteria (18%), Planctomycetes (8%) and Bacteroidetes (5%) (Fig. 1). These phyla have been shown to be widely represented in bulk and rhizosphere soil samples (e.g., see review by ref. 21), including those from other invasive species such as Japanese barberry24. At the class level, Actinobacteria (22%), Acidobacteria (17%), Alphaproteobacteria (17%), Planctomycetia (8%) and Betaproteobacteria (6%) were represented by the majority of sequences. Rhizosphere soil samples were enriched for Alphaproteobacteria (Kruskal-Wallis χ2 = 4.4; P < 0.05) and Gammaproteobacteria (Kruskal-Wallis χ2 = 7.4; P < 0.01), and depauperate in Spartobacteria (Verrucomicrobia; Kruskal-Wallis χ2 = 9.1; P < 0.01), compared with the bulk soil. Members of the Alphaproteobacteria and Gammaproteobacteria are usually defined as copiotrophic25; that is, they compete successfully when organic resources are abundant. The high abundance of these OTUs in the rhizosphere of A. dealbata plants could reflect their ability to proliferate in the presence of plant-derived polysaccharides. Spartobacteria are highly abundant in grassland soils26, probably because they associate with nematodes, which are typically abundant in grasslands27. Whether the decrease in Spartobacteria in this study results directly from the effects of wattles or indirectly through trophic cascades needs to be further investigated.
Mean relative abundances of taxa (phylum/class levels) within each location. The abundance of each taxon was calculated as the percentage of sequences per location for a given microbial group. The group ‘Other’ encompasses unclassified sequences together with classes representing ≤0.5% of total sequences. B, bulk soil; R, rhizosphere soil.
Fungal communities were dominated by the phylum Ascomycota (74%), while Basidiomycota represented only 16% of the reads. This corroborates what has been found across grasslands worldwide28. At the class level, the majority of the sequences could be assigned to Sordariomycetes (29%), Dothideomycetes (22%), Agaricomycetes (10%) and Eurotiomycetes (10%). Other groups such as Pezizomycetes (1.5%) and Glomeromycetes (AMF, 1.2%) were also represented, albeit in lower proportions. Rhizosphere soil samples were enriched for Pezizomycetes (Kruskal-Wallis χ2 = 11.0; P < 0.001) and Agaricomycetes (Kruskal-Wallis χ2 = 25.0; P < 0.001). High Pezizomycetes abundance has also been found in the rhizosphere of other trees such as willows29. The majority of Agaricomycetes sequences were identified as Inocybe sp., a widely distributed ectomycorrhizal fungus (EMF)30. This is not surprising, as Inocybe species have been found in the rhizosphere of other acacias31. Glomeromycetes, mainly the genera Gigaspora, Acaulospora, Dentiscutata and Glomus, were more abundant in bulk than in rhizosphere soils (Kruskal-Wallis χ2 = 6.4; P < 0.05). There are at least two possible explanations for this. First, it could be argued that AMF were less abundant in rhizosphere soils for historical reasons; that is, they were historically less dominant in the areas invaded by A. dealbata. However, we find this to be unlikely because historical differences in AMF abundance would be unlikely to be maintained over time at small spatial distances (10–20 m separated rhizosphere and bulk soil samples in a given location) due to the efficient dispersal of these fungi32. Therefore, it is likely that the lower AMF abundance results as a consequence of the invasion process12. The inhibition of some AMF by allelopathic compounds has previously been shown in other invasive plants33, but it is uncertain in A. dealbata where the allelopathic effects seem to be dependent on ecosystem type34. Alternatively, it has also been suggested that AMF in plants with dual associations (i.e., plants in association with both AMF and EMF) are only found in substantial abundance when these plants are growing in disturbed habitats or flooded soils, or are present as young seedlings35.
Soil chemistry differed significantly between rhizosphere and bulk soil samples (PERMANOVA F1,76 = 10.93; P < 0.001). Comparisons between rhizosphere and bulk soils showed that Total C, Total N, NH4 and NO3 values were higher for rhizosphere than for bulk soils (Supplementary Fig. S5). Acacia dealbata is able to alter the spatial distribution of nutrients in a wide variety of habitats19, 36 and our results are consistent with these observations. It has been suggested that the massive detritus production under the A. dealbata canopy results in increased accumulation of these nutrients in invaded soils37. Nitrogen levels are further improved due to the association of A. dealbata with N2-fixing bacteria. In contrast to what was shown in previous studies, we did not find A. dealbata to have lowered the pH and/or increased the P content of the rhizosphere soil. Since soil acidification and P increase seem to reach their maxima 10–25 years after invasion20, it is possible that our data reflect relatively recent invasion events.
The presence of A. dealbata had a significant effect on microbial community structure and composition based on Bray-Curtis distances (PERMANOVABacteria F1,76 = 3.44; PERMANOVAFungi F1,76 = 2.60; both P < 0.001). Similar patterns were observed for the 16S rRNA gene data set using unweighted (PERMANOVABacteria F1,76 = 2.84; P < 0.001) and weighted (PERMANOVABacteria F1,76 = 3.33; P < 0.001) UniFrac distances. We did not perform PERMANOVA for the ITS data set with UniFrac distances because of known issues with the accuracy of phylogenetic trees generated from this hypervariable region38. Redundancy analysis suggested that soil chemistry changes (especially in pH, C, NO3, NH4, P and Mg) around acacia roots played an important role in shaping the structure and composition of microbial communities (Fig. 2). Results showed that the different classes of bacteria and fungi were significantly correlated with these and other soil parameters (Fig. 3). Factors such as pH and nutrient status are the main drivers controlling composition and diversity of soil microbial communities39. Additional factors, for instance, the modification of soil structure by the root system40 and the release of allelopathic compounds40, 41 might also lead to changes in microbial communities.
Distance-based redundancy analysis (db-RDA) biplot of (a) bacterial and (b) fungal communities and soil chemistry parameters. Only the environmental variables that significantly (P < 0.05) explained variability in microbial community structure are shown (arrows). The direction of the arrows indicates the direction of maximum change of that variable, whereas the length of the arrow is proportional to the rate of change. Instead of plotting each sampling point, ellipses are shown that represent the standard error around location centroids.
Correlation between soil properties (aggregate data for rhizosphere and bulk soils) and the different bacterial and fungal classes shown in Fig. 1. Significance level P < 0.05.
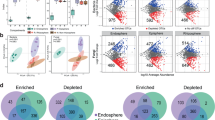
The effects of soil chemistry on variation in microbial community structure were further investigated by taking into account the confounding effects of geography (spatial distance between samples). Variation partitioning models explained between 23–25% (fungi) and 33–36% (bacteria) of the community variation, with soil chemistry and spatial distance significantly explaining 4–17% and 4–9% of the variation, respectively (Fig. 4; P < 0.001, based on 999 Monte Carlo permutation tests). We note that a large proportion of the variation remains unexplained. Although environmental variables that were not recorded may contribute to the high fraction of unexplained variation, we speculate that this could also be caused by ecological drift (i.e., changes in species abundances caused by the stochastic processes). Ecological drift seems to be an important driver for microbial community assembly42. Interestingly, bulk soil communities were more affected by geographic distance than rhizosphere soil communities, contradicting previous findings43. Overall, these results suggest that spatial distance, soil properties and A. dealbata-mediated soil interactions shape the organization of microbial communities on a regional scale.
Partitioning of the variation in bacterial and fungal community structure. The specific effects of soil chemistry and geographic distance and the total co-variation (i.e., the variance jointly explained by the soil chemistry and geographic distance) are represented. Statistical significance is indicated by ***P < 0.001. B, bulk soil; R, rhizosphere soil.
Despite the influence of biogeography on microbial community composition and diversity, we could identify a core microbiome associated with A. dealbata (sensu ref. 44). In total, 1,900 bacterial OTUs (6% of total OTUs, 72% of total reads) and 109 fungal OTUs (0.7% of total OTUs, 43% of total reads) were identified in rhizosphere soils in all eight locations, suggesting that there is a set of A. dealbata-associated microbes across a wide array of environments. Of those core OTUs, a total of 141 bacterial taxa and 14 fungal taxa were identified as biomarkers of the rhizosphere (Fig. 5 and Supplementary Dataset S2). The most important bacterial biomarkers of the rhizosphere were Proteobacteria classified as Bradyrhizobium (Alphaproteobacteria, 2 OTUs) and Burkholderia (Betaproteobacteria, 2 OTUs). Several species of the genus Burkholderia are known for their plant growth-promoting activities45 and ability to form rhizobial symbiotic relationships with several South African legumes46. Bradyrhizobium species are the predominant N2-fixing rhizobia associated with acacias (e.g., refs 47 and 48). N2-fixing symbionts enhance growth and competitive ability of their host plants49, which consequently influences plant invasiveness50. Interestingly, it has been reported that seven of nine selected acacia species grew better in association with Bradyrhizobium phylotypes than with members of the genera Burkholderia, Rhizobium and Ensifer 51.
Two fungal OTUs belonging to the families Pezizaceae (Pezizomycetes, 1 OTU) and Telephoraceae (Agaricomycetes, 1 OTU), together with an OTU classified as Inocybe sp. (Agaricomycetes), were the most abundant in the rhizosphere of A. dealbata. Numerous Pezizaceae and Telephoraceae fungi and members of the genus Inocybe are defined as EMF22. EMF associate with a large number of woody plants, facilitating plant nutrient uptake, especially phosphorus52. In addition, EMF may indirectly contribute to the global N cycle by providing phosphorus and other nutrients to N2-fixing legumes, particularly in natural ecosystems that are often P limited53. Several fungal genera (Clonostachys, Truncatella and Clonostachys) catalogued as probable plant pathogens54 using FUNGuild were also found as biomarkers of the rhizosphere (Supplementary Dataset S2). Pathogens can potentially reduce the performance, and therefore impacts, of invasive species. Alternatively, pathogen accumulation may exacerbate the effects of invasions if pathogens negatively affect co-occurring native species and reduce their performance and competitive ability55.
Furthermore, functional analyses of shotgun sequencing data suggest significant enrichment of approximately 300 genes in rhizosphere compared to bulk soils (Supplementary Dataset S3), among them, genes associated with membrane transport, signal transduction, xenobiotic degradation and metabolisms, and genes encoding for the metabolism of carbohydrates, amino acids, iron and nitrogen. Similar findings have been reported in other studies56, 57, supporting the view that microbial communities in the rhizosphere are selected based on the functions they perform.
In summary, we have shown that A. dealbata associates with different bacterial and fungal communities, depending on biogeography and soil nutritional status. Nonetheless, a large group of taxa were shared across all locations; these taxa represented >50% (for bacteria) and >40% (for fungi) of the rhizosphere community. Some of these microbes are likely involved in nutrient cycling, whereas others may act as plant pathogens. This is relevant because it shows that different mutualists of the root microbiome of A. dealbata have the potential to complement each other in acquiring different limiting nutrients, which concomitant with the possible accumulation of plant pathogens of native plants may contribute to the widespread success of this invader in novel environments.
Materials and Methods
Study sites and sample acquisition
Soil was collected in October 2015 from eight sites (between 3 to 532 km apart) in the grassland biome of South Africa (Table 1 and Supplementary Fig. S1). In South Africa, A. dealbata mainly invades grassland riparian habitats58, 59. At each site two types of samples were collected: rhizosphere soil, which was firmly attached to the roots, and bulk soil, gathered 10–20 m away from any conspecific tree to avoid ongoing plant-soil microbial feedbacks60. Sampling and processing equipment was sterilized with 90% ethanol after each sampling regime. A 2-mm sieve was used to remove leaves and other coarse material prior to sample homogenization. Soil sub-samples for genetic analysis were stored at −80 °C, within three days of sampling, until further analysis. A total of 80 samples were collected (8 populations ×5 individuals ×2 habitat types (rhizosphere or bulk soil)).
Soil chemistry analysis
Soil chemical analysis was performed according to standard methods at BemLab (SANAS Accredited Testing Laboratory, Cape Town, South Africa). pH was determined in saturated soil extracts (SSE). Total N and C were measured by combustion at 1350 °C. Total phosphorus, potassium, calcium and magnesium were extracted using HCl-HNO3 after combustion (3 h, 550 °C), followed by quantification by inductively coupled plasma optical emission spectrometry (ICP-OES). Ammonium and nitrate were extracted with 2 M KCl and diluted prior to determination by a flow injection analyser (FIA). Soil characteristics and all metadata are presented in Supplementary Dataset S1.
DNA extraction and sequencing
Genomic DNA was isolated from the 80 samples using a PowerSoil DNA isolation kit (MO Bio Laboratories, Inc. Carlsbad, CA) following the manufacturer’s instructions. The internal transcribed spacer (ITS) region was amplified using fungal-specific primers61: ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′). Bacterial 16S rRNA gene amplicons were amplified using primers 515F (5′-GTGYCAGCMGCCGCGGRA-3′) and 909R (5′-CCCCGYCAATTCMTTTRAG-3′) as in Oloo, et al.62. DNA regions were amplified using the HotStarTaq Plus Master Kit (Qiagen, Valencia, CA). Amplicons from different samples were mixed in equal concentrations and purified using Agencourt Ampure beads (Agencourt Bioscience Corporation, USA). Paired-end 2 × 250 bp sequencing was performed on an Illumina MiSeq instrument (Illumina Inc., San Diego, CA, USA).
Additionally, 4 bulk soil and 4 rhizosphere samples were randomly selected for shotgun metagenomic sequencing. Shotgun libraries were prepared using a Nextera DNA sample preparation kit (Illumina Inc., San Diego, CA, USA) following the manufacturer’s instructions. Libraries were sequenced using an Illumina Hiseq-2000 using paired-end technology (2 × 150 bases). All sequencing was performed by the Molecular Research LP next generation sequencing service (http://www.mrdnalab.com).
Amplicon sequencing analysis
Sequence data were analysed using QIIME version 1.9.163. Sequences that were <200 bp, contained more than 2 ambiguous characters, had quality scores <25, or contained more than one mismatch to the sample-specific barcode or to the primer sequences were excluded from further downstream analyses. Chimeric sequence detection and OTU selection at 97% sequence similarity were conducted using USEARCH v6.164. Taxonomies were assigned to each OTU using the RDP Naïve Bayesian Classifier65 with the SILVA-ARB (release 123) and UNITE-INSD (release 7) databases for bacteria and fungi, respectively. OTUs whose classifications did not match their expected taxonomic kingdoms (fungi and bacteria, respectively) were removed. Singletons were excluded and each sample was randomly subsampled (rarefied) to the same number of sequences per sample; that is, 29,678 bacterial and 8,093 fungal sequences. Two samples with a low number of sequences were excluded, yielding a total of 78 (39 rhizosphere and 39 bulk soil) samples that were used for downstream analyses.
Shotgun metagenomic analysis
Paired-end sequences were quality filtered with PRINSEQ66 and joined using FLASH67. Combined reads were aligned against the NCBI-NR protein database using DIAMOND BLASTX v0.7.1 (E-value cut-off at 1e-5)68. Functional annotation was performed based on KEGG pathways and SEED subsystems in MEGAN v5.0.369. The matrix of raw counts of functional annotations were normalized to account for the unequal sequence coverage between samples.
Statistical analyses
OTU richness and diversity indices (richness, Shannon, inverse Simpson and Pielou’s evenness), together with accumulation curves were calculated using the vegan R package70, 71. We applied a mixed model ANOVA to determine significant differences in microbial diversity and soil chemistry between rhizosphere and bulk soils using the phia package72. In these analyses, location was specified as a random factor. Abiotic data were standardized and pair-wise distances computed based on Euclidean distances. Community data matrices were Hellinger-transformed and the Bray-Curtis distance measure was used to generate a dissimilarity matrix. Weighted and unweighted UniFrac dissimilarities were also obtained73. The effect of abiotic data in explaining variation in microbial community structures was assessed by distance-based redundancy analysis. First, we checked for collinearity among variables. This led to the removal of Na (correlated with NO3; r2 = 0.64) and Ca (correlated with pH; r2 = 0.80). Then, we performed forward selection to select the best set of variables that could explain the variation in community composition. Variation partitioning analyses were performed to determine the respective effects of the environment (soil chemistry) and geographic distances on the variation in microbial community composition74. A permutational analysis of variance (PERMANOVA)75 was carried out to test for differences in composition between habitats (bulk and rhizosphere soil) using the ‘adonis’ function (strata = location) in vegan. Selection for putative major microbial players in the rhizosphere communities was carried out as follows. First, we identified rhizosphere bacterial and fungal taxa that were present in all eight locations. We then performed linear discriminant analysis (LDA) effect size (LEfSe)76 to discriminate between microbial markers of the rhizosphere and bulk soil. We hypothesised that if microorganisms are selected within a given habitat on the basis of their functional capacities, microbial markers should be among the most influential microbial taxa in their respective habitat. Finally, we plotted the average relative abundance and frequency of occurrence of those taxa across each sample type to infer their putative ecological relevance. FUNGuild77 was used to assign fungal phylotypes to one of three trophic modes (saprotroph, symbiont or pathogen) where possible. To determine statistical differences between the functional potential of the rhizosphere and the bulk soil samples, the Statistical Analysis of Metagenomic Profiles (STAMP78) software package was used. P-values were calculated using a two-sided Fischer’s exact test and corrected using Benjamini-Hochberg false discovery rate.
Data availability
The raw sequencing reads for this project were submitted to the National Center for Biotechnology Information Short Read Archive under accession no. SRP090490 (targeting sequencing) and SRP098951 (shotgun sequencing).
References
Richardson, D. M., Hui, C., Nuñez, M. A. & Pauchard, A. Tree invasions: patterns, processes, challenges and opportunities. Biol Invasions 16, 473–481 (2014).
Rejmánek, M. & Richardson, D. M. Trees and shrubs as invasive alien species–2013 update of the global database. Divers Distrib 19, 1093–1094 (2013).
Le Maitre, D. et al. Impacts of invasive Australian acacias: Implications for management and restoration. Divers Distrib 17, 1015–1029 (2011).
Richardson, D. M. & Rejmánek, M. Trees and shrubs as invasive alien species–a global review. Divers Distrib 17, 788–809 (2011).
de Neergaard, A. et al. Australian wattle species in the Drakensberg region of South Africa–An invasive alien or a natural resource? Agric Syst 85, 216–233 (2005).
Coetzee, B., van Rensburg, B. & Robertson, M. Invasion of grasslands by silver wattle, Acacia dealbata (Mimosaceae), alters beetle (Coleoptera) assemblage structure. Afr Entomol 15, 328–339 (2007).
Slabbert, E., Jacobs, S. M. & Jacobs, K. The soil bacterial communities of South African Fynbos riparian ecosystems invaded by Australian Acacia species. PloS ONE 9, e86560 (2014).
Fuentes-Ramírez, A., Pauchard, A., Cavieres, L. A. & García, R. A. Survival and growth of Acacia dealbata vs. native trees across an invasion front in south-central Chile. For Ecol and Manage 261, 1003–1009 (2011).
Souza-Alonso, P., Novoa, A. & González, L. Soil biochemical alterations and microbial community responses under Acacia dealbata Link invasion. Soil Biol Biochem 79, 0e108 (2014).
Le Roux, J. J. et al. Phylogeographic consequences of different introduction histories of invasive Australian Acacia species and Paraserianthes lophantha (Fabaceae) in South Africa. Divers Distrib 17, 861–871 (2011).
Richardson, D. M. et al. Human‐mediated introductions of Australian acacias–a global experiment in biogeography. Divers Distrib 17, 771–787 (2011).
Traveset, A. & Richardson, D. M. Mutualistic interactions and biological invasions. Annu Rev Ecol Evol Syst 45, 89–113 (2014).
Wagg, C., Bender, S. F., Widmer, F. & van der Heijden, M. G. Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc Natl Acad Sci USA 111, 5266–5270 (2014).
Rodríguez‐Echeverría, S., Le Roux, J. J., Crisóstomo, J. A. & Ndlovu, J. Jack‐of‐all‐trades and master of many? How does associated rhizobial diversity influence the colonization success of Australian Acacia species? Divers Distrib 17, 946–957 (2011).
Le Roux, J. J., Mavengere, N. R. & Ellis, A. G. The structure of legume-rhizobium interaction networks and their response to tree invasions. AoB plants 8, plw038 (2016).
Brundrett, M. C. Mycorrhizal associations and other means of nutrition of vascular plants: understanding the global diversity of host plants by resolving conflicting information and developing reliable means of diagnosis. Plant Soil 320, 37–77 (2009).
van der Heijden, M. Mycorrhizal ecology and evolution: the past, the present, and the future. New Phytol 205, 1406–1423 (2015).
Van der Putten, W. H., Klironomos, J. N. & Wardle, D. A. Microbial ecology of biological invasions. ISME J 1, 28–37 (2007).
Lorenzo, P., Rodríguez-Echeverría, S., González, L. & Freitas, H. Effect of invasive Acacia dealbata Link on soil microorganisms as determined by PCR-DGGE. Appl Soil Ecol 44, 245–251 (2010).
Souza-Alonso, P., Guisande-Collazo, A. & González, L. Gradualism in Acacia dealbata Link invasion: Impact on soil chemistry and microbial community over a chronological sequence. Soil Biol Biochem 80, 315–323 (2015).
Philippot, L., Raaijmakers, J. M., Lemanceau, P. & van der Putten, W. H. Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11, 789–799 (2013).
Bonito, G. et al. Plant host and soil origin influence fungal and bacterial assemblages in the roots of woody plants. Mol Ecol 23, 3356–3370 (2014).
Bent, S. J. & Forney, L. J. The tragedy of the uncommon: understanding limitations in the analysis of microbial diversity. ISME J 2, 689–695 (2008).
Coats, V., Pelletreau, K. & Rumpho, M. Amplicon pyrosequencing reveals the soil microbial diversity associated with invasive Japanese barberry (Berberis thunbergii DC.). Mol Ecol 23, 1318–1332 (2014).
Leff, J. W. et al. Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc Natl Acad Sci USA 112, 10967–10972 (2015).
Bergmann, G. T. et al. The under-recognized dominance of Verrucomicrobia in soil bacterial communities. Soil Biol Biochem 43, 1450–1455 (2011).
Fierer, N., Strickland, M. S., Liptzin, D., Bradford, M. A. & Cleveland, C. C. Global patterns in belowground communities. Ecol Lett 12, 1238–1249 (2009).
Prober, S. M. et al. Plant diversity predicts beta but not alpha diversity of soil microbes across grasslands worldwide. Ecol Lett 18, 85–95 (2015).
Bell, T. H. et al. Linkage between bacterial and fungal rhizosphere communities in hydrocarbon-contaminated soils is related to plant phylogeny. ISME J 8, 331–343 (2014).
Matheny, P. B. et al. Out of the Palaeotropics? Historical biogeography and diversification of the cosmopolitan ectomycorrhizal mushroom family. Inocybaceae. J Biogeogr 36, 577–592 (2009).
Birnbaum, C., Bissett, A., Thrall, P. H. & Leishman, M. R. Invasive legumes encounter similar soil fungal communities in their non-native and native ranges in Australia. Soil Biol Biochem 76, 210–217 (2014).
Davison, J. et al. Global assessment of arbuscular mycorrhizal fungus diversity reveals very low endemism. Science 349, 970–973 (2015).
Callaway, R. M. et al. Novel weapons: invasive plant suppresses fungal mutualists in America but not in its native Europe. Ecology 89, 1043–1055 (2008).
Lorenzo, P., Pereira, C. S. & Rodríguez-Echeverría, S. Differential impact on soil microbes of allelopathic compounds released by the invasive Acacia dealbata Link. Soil Biol Biochem 57, 156–163 (2013).
Brundrett, M. C. Coevolution of roots and mycorrhizas of land plants. New Phytol 154, 275–304 (2002).
González-Muñoz, N., Costa-Tenorio, M. & Espigares, T. Invasion of alien Acacia dealbata on Spanish Quercus robur forests: impact on soils and vegetation. For Ecol Manage 269, 214–221 (2012).
Lazzaro, L. et al. Soil and plant changing after invasion: The case of Acacia dealbata in a Mediterranean ecosystem. Sci Total Environ 497, 491–498 (2014).
Lindahl, B. D. Fungal community analysis by high-throughput sequencing of amplified markers–a user’s guide. New Phytol 199, 288–299 (2013).
Lauber, C. L., Hamady, M., Knight, R. & Fierer, N. Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75, 5111–5120 (2009).
Bardgett, R. D., Mommer, L. & De Vries, F. T. Going underground: root traits as drivers of ecosystem processes. Trends Ecol Evol 29, 692–699 (2014).
Lorenzo, P. & Rodríguez-Echeverría, S. Influence of soil microorganisms, allelopathy and soil origin on the establishment of the invasive Acacia dealbata. Plant Ecol Divers 5, 67–73 (2012).
Bahram, M. et al. Stochastic distribution of small soil eukaryotes resulting from high dispersal and drift in a local environment. ISME J 10, 885–896 (2015).
Hovatter, S. R., Dejelo, C., Case, A. L. & Blackwood, C. B. Metacommunity organization of soil microorganisms depends on habitat defined by presence of Lobelia siphilitica plants. Ecology 92, 57–65 (2011).
Shade, A. & Handelsman, J. Beyond the Venn diagram: The hunt for a core microbiome. Environ Microbiol 14, 4–12 (2012).
Compant, S., Nowak, J., Coenye, T., Clément, C. & Barka, E. A. Diversity and occurrence of Burkholderia spp. in the natural environment. FEMS Microbiol Rev 32, 607–626 (2008).
Lemaire, B. et al. Symbiotic diversity, specificity and distribution of rhizobia in native legumes of the Core Cape Subregion (South Africa). FEMS Microbiol Ecol 91, 1–17 (2015).
Hoque, M. S., Broadhurst, L. M. & Thrall, P. H. Genetic characterization of root-nodule bacteria associated with Acacia salicina and A. stenophylla (Mimosaceae) across south-eastern Australia. Int J Syst Evol Microbiol 61, 299–309 (2011).
Ndlovu, J., Richardson, D. M., Wilson, J. R. & Le Roux, J. J. Co‐invasion of South African ecosystems by an Australian legume and its rhizobial symbionts. J Biogeogr 40, 1240–1251 (2013).
Klock, M. M., Barrett, L. G., Thrall, P. H. & Harms, K. E. Host promiscuity in symbiont associations can influence exotic legume establishment and colonization of novel ranges. Divers Distrib 21, 1193–1203 (2015).
Parker, M. A., Malek, W. & Parker, I. M. Growth of an invasive legume is symbiont limited in newly occupied habitats. Divers Distrib 12, 563–571 (2006).
Bever, J., Broadhurst, L. & Thrall, P. Microbial phylotype composition and diversity predicts plant productivity and plant-soil feedbacks. Ecol Lett 16, 167–174 (2013).
Wallander, H. Uptake of P from apatite by Pinus sylvestris seedlings colonised by different ectomycorrhizal fungi. Plant Soil 218, 249–256 (2000).
Elser, J. J. et al. Global analysis of nitrogen and phosphorus limitation of primary producers in freshwater, marine and terrestrial ecosystems. Ecol Lett 10, 1135–1142 (2007).
Tedersoo, L. et al. Global diversity and geography of soil fungi. Science 346, 1256688 (2014).
Luke, F. S. & Clay, K. Pathogen accumulation and long‐term dynamics of plant invasions. J Ecol 101, 607–613 (2013).
Zarraonaindia, I. et al. The soil microbiome influences grapevine-associated microbiota. MBio 6, e02527–02514 (2015).
Mendes, L. W., Kuramae, E. E., Navarrete, A. A., Van Veen, J. A. & Tsai, S. M. Taxonomical and functional microbial community selection in soybean rhizosphere. ISME J 8, 1577–1587 (2014).
Henderson, L. Invasive, naturalized and casual alien plants in southern Africa: a summary based on the Southern African plant invaders atlas (SAPIA). Bothalia 37, 215–248 (2007).
Holmes, P., Richardsonb, D., Eslerb, K., Witkowski, E. & Fouriee, S. A decision-making framework for restoring riparian zones degraded by invasive alien plants in South Africa. S Afr J Sci 101, 553 (2005).
Callaway, R. M., Bedmar, E. J., Reinhart, K. O., Silvan, C. G. & Klironomos, J. Effects of soil biota from different ranges on Robinia invasion: acquiring mutualists and escaping pathogens. Ecology 92, 1027–1035 (2011).
White, T. J., Bruns, T., Lee, S. & Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics (eds. Innis, M., Gelfand, D., Sninsky, J., White, T.) 315–322 (Academic Press, 1990).
Oloo, F. et al. Habitat heterogeneity and connectivity shape microbial communities in South American peatlands. Sci Rep 6 (2016).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naïve bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73, 5261–5267 (2007).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat Methods 12, 59–60 (2015).
Huson, D. H. & Mitra, S. Introduction to the analysis of environmental sequences: Metagenomics with MEGAN. Methods Mol Bio 856, 415–429 (2012).
Oksanen, J. et al. Vegan: Community Ecology Package, vR package version 2.0-2, http://cran.r-project.org/package=vegan (2007).
R Development Core Team. R: A language and environment for statistical computing. R foundation for statistical computing, http://www.r-project.org/ (2013).
De Rosario-Martinez, H. phia: Post-Hoc Interaction Analysis. R package version 0.2-0, https://cran.r-project.org/package=phia (2015).
Lozupone, C. & Knight, R. UniFrac: a New Phylogenetic Method for Comparing Microbial Communities. Appl Environ Microbiol 71, 8228–8235 (2005).
Stomeo, F. et al. Hypolithic and soil microbial community assembly along an aridity gradient in the Namib Desert. Extremophiles 17, 329–337 (2013).
Anderson, M. J. Distance‐based tests for homogeneity of multivariate dispersions. Biometrics 62, 245–253 (2006).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biolo 12, 1 (2011).
Nguyen, N. H. et al. FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20, 241–248 (2016).
Parks, D. H. & Beiko, R. G. Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26, 715–721 (2010).
Mucina, L. & Rutherford, M. The vegetation of South Africa, Lesoto and Swaziland. South African National Biodiversity Institute, Pretoria (2006).
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25, 1965–1978 (2005).
Acknowledgements
Funding for this research was provided by the National Research Foundation, South Africa (Grant No. CPRR14071676470 to AV). SV is the recipient of a postdoctoral research fellowship from the Claude Leon Foundation, South Africa. DMR and JJLR received support from the DST-NRF Centre of Excellence for Invasion Biology. DMR and MG acknowledge funding from the National Research Foundation of South Africa (grants 85417 and 98889, respectively). We thank Lesley Henderson, Fanus Venter and Emma Steenkamp for the information on A. dealbata populations.
Author information
Authors and Affiliations
Contributions
A.V., T.P.M., J.J.LR., D.R., D.C. designed research. C.N.K., G.K.N performed research. C.N.K., S.V., A.V. analysed the data. A.V. and C.N.K wrote the manuscript. All authors commented on the manuscript at all stages.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kamutando, C.N., Vikram, S., Kamgan-Nkuekam, G. et al. Soil nutritional status and biogeography influence rhizosphere microbial communities associated with the invasive tree Acacia dealbata . Sci Rep 7, 6472 (2017). https://doi.org/10.1038/s41598-017-07018-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-07018-w
This article is cited by
-
Does Host Plant Drive Variation in Microbial Gut Communities in a Recently Shifted Pest?
Microbial Ecology (2023)
-
Plant Growth Promoting Bacterial Consortium Induces Shifts in Indigenous Soil Bacterial Communities and Controls Listeria monocytogenes in Rhizospheres of Cajanus cajan and Festuca arundinacea
Microbial Ecology (2022)
-
Driving Factors, Co-occurrence Networks, and Metabolic Profiles of Soil Bacterial Communities Within the Root Proximity of Amaranthus palmeri
Journal of Soil Science and Plant Nutrition (2022)
-
Impact of crop cultivation, nitrogen and fulvic acid on soil fungal community structure in salt-affected alluvial fluvo-aquic soil
Plant and Soil (2021)
-
A core of rhizosphere bacterial taxa associates with two of the world’s most isolated plant congeners
Plant and Soil (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.