Abstract
Divergent selection plays a critical role not only as a speciation driver but also in maintaining post-speciation divergence. In the absence of direct evidence, ancestral interspecific gene flow between incipient species can reflect ancient selective pressure for ecological speciation. In the present study, two late-Pleistocene diverged species endemic to Taiwan, Scutellaria playfairii and S. tashiroi, were spatially and ecologically partitioned with partial overlap. Multilocus genome-scan analyses and in silico evaluation revealed ancestral interspecific gene flow but distinct genetic compositions, implying that adaptive divergence contributed to their speciation. Ecological niche modeling and principal component analysis suggested incomplete divergent niches between the two species; the species distribution is therefore consistent with Hutchinson’s metaphor of multidimensional hypervolume niches rather than attributable to a single factor. Constraint ordination analysis supported this inference of a combination of variables explaining the genetic structure. The rare occurrence of hybrids in the sympatric population suggested hybrid breakdown, providing further evidence of divergent selection blocking gene flow. The correlation of environmental variables with integrated genetic components demonstrated that environmental heterogeneity maintains the species and population differentiation. This study highlights the importance of environmental heterogeneity and divergent selection for the rapid speciation and recent diversification of island plants.
Similar content being viewed by others
Introduction
In addition to accelerating speciation, divergent selection is also responsible for maintaining differences between species. Studies have increasingly shown that environmental variables, rather than geographic distance, are responsible for genetic divergence1,2,3, indicating the importance of surrounding environments for species evolution. Via4 proposed two stages of selective pressure on ecological speciation and divergence. Stage 1 involves selection against effective migration, and stage 2 involves independent responses to selection within the new lineages. Stage 1 was suggested to clearly define the difference between allopatric and sympatric speciation4, 5, whereas directional selection could further independently act on the diverged lineages to reduce the potential for interspecific mating upon physical contact after speciation4, 6, 7. However, because past environmental conditions are difficult to record, present environmental heterogeneity can usually explain post-speciation divergence but not the actual driving force for ecological speciation, i.e., only stage 2 of Via can be verified4.
Since ecological speciation is a time-dependent process, i.e., genomic barriers for gene flow gradually increase between adaptively divergent species5, 8, detecting the degrees of interspecific gene flow through time can support or reject ecological speciation. A selective process leads to species (or population) divergence more rapidly than a stochastic drift process9, 10. Hence, gene flow will rapidly be blocked if the selection continues to maintain or enhance the divergence. The development of modeling-based simulation analyses such as the approximate Bayesian computation (ABC) has enabled the evaluation of the best-fit evolutionary scenario of isolation vs. gene flow to resolve the speciation hypothesis. In combination with correlation analysis between genetic variation and environmental variables, the effect of environmental heterogeneity on species differentiation and local adaption can be clarified.
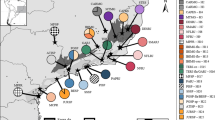
To investigate the degree to which the genetic diversity may be influenced by the primary speciation process and subsequent environmental pressures, we selected as experimental materials two herb species, Scutellaria playfairii Kudo and S. tashiroi Hayata, which both have a relatively limited spatial distribution. Scutellaria playfairii and S. tashiroi are endemic to Taiwan and are sparsely distributed among low-elevation hills. The S. playfairii are mainly distributed in the southern Taiwan, while S. tashiroi are common in the stream valleys of eastern Taiwan. Despite most populations of these two species were allopatrically distributed, there are still some sympatric populations being found in the southeastern Taiwan (Fig. 1). The inflorescence, which can be used to distinguish Scutellaria species11, is mostly axillary, with rare terminal racemes and dark-purple corollas in S. tashiroi and terminal, loose racemes and whitish-purple corollas in S. playfairii (Supplementary Table S1). Transcriptomic-based analysis has shown that the growth form and flower color-related transcription factors (MYBs) are positively selected and are possibly related to the differentiation of flower color and speciation12. Divergent selection and functional subdivision of anthocyanin-biosynthetic genes have also been suggested to be responsible for the rapid evolution and diversification of Scutellaria species in the island of Taiwan13, 14. In addition, biogeographic analyses have indicated a very recent divergence time, within 0.2 Mya, of these two species, and their divergence was suggested to be a consequence of a series of dispersal and vicariance events corresponding to climate shifts through the late Pleistocene15. Natural selection and biogeographic expansion may together explain the rapid diversification of these two island species.
Distribution range of S. playfairii (light blue) and S. tashiroi (yellow) in Taiwan. Exact distribution of specimen records is shown in Fig. 5. Despite overlapping distribution ranges in the eastern Taiwan, sympatric populations were found in the southeastern Taiwan only. The map was constructed using the packages maptools69 implemented in R.
Because geo-environmental heterogeneity can act as a selective agent for species and population differentiation, ecologically divergent selection and local adaptation may be valid explanations of species differentiation and population structure. Quantifying the effects of environmental factors on geographic and genetic distributions can aid the clarification of the mechanisms driving and maintaining species/population differentiation. For example, multivariate analyses that using linear dependent similarity matrix can assess how does the constraint factors (e.g. environmental or/and geographic distances) affect the response (e.g. genetic distance)16, 17. Furthermore, modeling species distribution using climatic variables (ecological niche modeling, ENM) can be used for exploring the ecological association and discriminating environmental requirements of each species, which are widely used in testing hypotheses of ecological divergence and speciation18, 19. The hypothetico-deductive evolutionary analyses, such as the ABC approach, provide powerful statistical verification method to evaluate the best speciation model by using summary statistics and simulation without exact likelihood calculations20, 21. The ABC is widely used in unravelling complex evolutionary history such as the relict cycad species22, phylogenetic closely related skullcap flowers23, Dysosma versipellis-pleiantha complex24, etc. Through integration of these frameworks, we can test the ecological divergence of S. tashiroi and S. playfairii, and evaluate the relative importance of each environmental factor on their genetic divergence. These integrated approaches has successfully applied in another two Taiwanese Scutellaria species to confirm their taxonomic relationship23.
In this paper, we assessed the ecological speciation and post-speciation adaptive divergence of these two recently diverged Scutellaria species through in silico speciation-processing modeling and correlations between climatic variables and genetics. Here, we extract environmental factors that cause or sustain genetic differences and highlight the important role of integrated environmental elements in plant genetic make-up.
Results
Genetic variation and population structure
According to the genetic assessment of 19 microsatellite loci, S. playfairii and S. tashiroi showed similar genetic diversity with respect to expected heterozygosity (ranges: 0.231 ± 0.061~0.340 ± 0.067 in S. playfairii and 0.253 ± 0.069~0.371 ± 0.069 in S. tashiroi) and the inbreeding coefficient (fixation index: 0.431 ± 0.085~0.727 ± 0.053 in S. playfairii and 0.387 ± 0.061~0.663 ± 0.043 in S. tashiroi) (Table 1). The moderate heterozygosity levels and inbreeding coefficients suggest self-compatibility in both species, consistent with the inference of a selfing or sibling-mating tendency in small and isolated populations of S. montana 25 and S. indica 26. The BayeScan approach found three positive outlier loci and one negative outlier locus (Supplementary Fig. S1), which were suggested to be candidate loci under positive selection and balancing selection, respectively27. Analysis of molecular variance (AMOVA) based on the remaining 15 neutral loci revealed a non-significant difference in genetic variation between species but a significant divergence between populations (Table 2). Less than half of the total genetic variation was due to between-species differences in total loci and neutral loci (47.30% and 39.88%, respectively), whereas 90.51% of the genetic variation was due to between-species differences in adaptive loci, with a smaller contribution from population effects (Table 2). However, non-significant genetic differences were detected between species in all three datasets (P = 0.107, 0.093, and 0.096, respectively, Table 2). By contrast, significant genetic differences were detected between populations within species, implying adaptive divergence among populations (local adaptation) for both plant species.
We then performed Bayesian clustering analysis (BCA) and discriminant analysis of principal components (DAPC) to illustrate the grouping patterns of these five populations. BCA suggested that the best clustering number was two (K = 2), according to the highest ΔK (lnP(K) = −2465.31, ln’(K) = 1393.59, |ln”(K)| = 1005.47, ΔK = 3633.97, Fig. 2A), corresponding to groups of species in both the adaptive- and neutral-locus datasets (Fig. 2B). In the neutral-locus dataset, K = 3 was also optimal, separating the Wulu population of S. tashiroi from the other two S. tashiroi populations (Fig. 2B), a pattern similar to that of the DAPC for neutral loci. DAPC showed clear differentiation between the species when only the first two PCs were considered (Fig. 3A,B). At the population level, genetic admixture was observed between the two populations of S. playfairii, whereas the Wulu population of S. tashiroi was distinguished from the other two populations (Fig. 2C,D). The DAPC scatterplot revealed similar genetic compositions of the Lanyu and Taroko populations of S. tashiroi (Fig. 3C) due to similar admixture patterns of genetic components (Fig. 3D). It is worth to notice that the Wulu population of S. tashiroi has distinguished but non-admixed genetic component from other populations of both species under K = 3 in BCA and DAPC (Figs 2B and 3D). Genotypes of S. tashiroi in Wulu neither exhibit intermediate form (i.e. heterozygotes or chimera) between any populations of two species, means that the distinguished genetic cluster of the Wulu population of S. tashiroi is not attributed to hybridization between two species.
Results of Bayesian clustering analysis conducted by STRUCTURE. (A) The changes in ΔK and lnP(K) in different clustering (K) situations in neutral loci. The ΔK plot shows that the highest ΔK value occurs at K = 2. This plot also shows lnP(K), which demonstrates the increase in the posterior probability of K. (B) The clustering patterns of genetic components by two groups (K = 2) in adaptive loci (positive outliers) and the patterns by two (K = 2) and three groups (K = 3) in neutral loci.
Genetic structure of the sampled populations as estimated by discriminant analysis of principal components (DAPC). The results show discriminable groups at the species level in the scatter plot (A), with small fractions of genetic admixture in the compoplot (B), and clusters of populations with partial overlap (C) and obvious admixture patterns between populations of each species, except the Wulu population of S. playfairii (D).
All three positive outlier loci were homozygous (Supplementary Table S2); however, loci aus9-2 and aus9-5 are species-private alleles. The other locus, aus9-3, has three fixed genotypes corresponding to the Wulu and Wutai populations of S. playfairii and the species S. tashiroi. These results suggest an adaptive divergence of the two species and local adaptation among populations of S. playfairii. DAPC analysis could not be performed using positive outlier loci due to the homozygosity of all samples. Incongruent grouping by neutral (S. playfairii; Wulu of S. tashiroi; Taroko and Lanyu of S. tashiroi) and adaptive loci (Wulu of S. playfairii; Wutai of S. playfairii; S. tashiroi) suggested that drift (neutral) processes and selective pressures affected population genetic structure and species divergence simultaneously but not synchronously.
Speciation model
An ABC framework permitted an evaluation of speciation scenarios for S. playfairii and S. tashiroi. Among four speciation scenarios (see Methods), the ancestral migration (AM) model (marginal density: 2.66 × 10−8, relative density: 92.20%) describing early interspecific gene flow after divergence best explained the current genetic divergence of S. tashiroi and S. playfairii relative to the complete isolation (CI), secondary contact (SC), and continuous migration (CM) models (Fig. 4). We detected equal rates of bidirectional gene flow (Mode of M tas→pla = M pla→tas = 0.100, i.e., 10% of individuals are migrants per generation, Supplementary Table S3) at the beginning of speciation under the AM model. The estimated time of gene flow (t 1 mode = 134474 generations, t 1 median = 150300 generations) was similar to the divergence time (t 2 mode = 111448 generations, t 2 median = 280534 generations). Scutellaria species are annual herbs. Consequently, the divergence and early interspecific gene flow can be traced back to the early Pleistocene. This estimate of the divergence time is thus consistent with the estimate of <0.2 Mya based on nuclear and chloroplast genes15. The detailed parameter estimates of the AM model are listed in Supplementary Table S3.
Evolutionary scenarios of speciation between the phylogenetically close species S. tashiroi (tas) and S. playfairii (pla) and the estimated consequences of approximate Bayesian computation: (A) the CI model; (B) the AM model; (C) the SC model; and (D) the CM model. The AM scenario had the highest posterior probability. Distributions of the estimated parameters for the best scenario (AM) are presented: (E) migration rate, (F) effective population sizes, (G) divergent time and the time of gene flow, and (H) average mutation rate of 15 neutral microsatellite loci. MD, marginal density; RD, relative density.
Niche differentiation between S. playfairii and S. tashiroi
After removing variables of multicollinearity, the remaining five environmental variables with the smallest variance inflation factor [VIF, including altitude (alt), mean actual annual evapotranspiration (AET), isothermality (bio3) and precipitation in the wettest (bio13) and driest months (bio14)] revealed partial overlap of niches between the two species in principal component analysis (PCA) space (Supplementary Fig. S2). The first three PCs explained 44.20%, 33.68%, and 10.87% of the environmental variance. The Kruskal-Wallis rank sum (KW) test was further performed to test significant differences between the species. Only PC1 revealed significant species differentiation (KW test: χ 2 = 7.839, df = 1, P = 0.0051, Supplementary Fig. S3), whereas the other two PCs did not indicate significant differences between the species (PC2: KW test: χ 2 = 0.833, df = 1, P = 0.3614; PC3: KW test: χ 2 = 0.364, df = 1, P = 0.5463, Supplementary Fig. S3). The general linear model (GLM) further revealed that all five environmental variables were significantly correlated with the species-divergent PC1 (Supplementary Table S4). These correlations indicate that these five variables are all responsible for the explanation of niche divergence on PC1. We next examined whether these environmental parameters explain the niche differentiation between species together or independently by performing multivariate logistic regression to assess the relationships between the dependent variables (i.e., species) and each independent variable (i.e., environmental variables or the PCs), controlling for the other independent variables. None of the environmental variables directly predicted species differentiation alone, but the reduction axis PC1 of these environmental variables had a significant effect on predicting species divergence (Z = 2.626, P = 0.009, Table 3). This result suggests that the niche differentiation between species cannot not be explained by any single environmental factor but rather by the contributions of a variety of environmental factors.
Ecological niche modeling
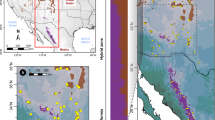
To model the niches of S. playfairii and S. tashiroi and compare ecological associations between these two phylogenetically close species, the potential species distribution was modeled on a map of Taiwan according to total 71 environmental variables (ENM1, Supplementary Data) or five remaining variables without multicollinearity (ENM2). The average test of the cross-validated area under the curve (AUC) for predicting the distribution of S. playfairii was 0.830 (±0.171) and 0.841 (±0.093) for replicate runs of ENM1 and ENM2, respectively; the AUC for predicting the distribution of S. tashiroi was 0.921 (±0.025) and 0.888 (±0.047) in ENM1 and ENM2, respectively. The high AUC value indicates good performance in modeling the distributions of both Scutellaria species. The range of species distribution predicted by ENM1 is closer to the actual recorded sites of S. playfairii, with a likely distribution in low-altitude mountains of eastern Taiwan and lowlands of southwestern Taiwan (Fig. 5A), whereas the likely distribution of S. tashiroi is in the lowlands of eastern Taiwan and islets off southeastern Taiwan (Fig. 5B). When the predicting variables were reduced to five (ENM2), the potential distribution of S. playfairii expanded to the south and included more lowland areas of southeastern and southwestern Taiwan (Fig. 5C). By contrast, the potential distributions of S. tashiroi predicted by ENM1 and ENM2 were similar (Fig. 5D). ENM1, which used all available predictors, provided a better fit to the real distributions than ENM2, indicating that the present distributions of the extant populations of these two species might be sensitively impacted by local- or micro-environmental variation.
Ecological niche modeling for predicting the potential distributions of S. playfairii and S. tashiroi. (A,B) are the potential distributions inferred by all 71 collected environmental variables; (C,D) were predicted by the remaining five variables after removing variables of multicollinearity. The map was constructed using the packages maptools69 implemented in R.
Explanation of the role of environmental variables in plant genetic diversity
According to dbRDA, environmental variables explained 90.83% of the genetic variation of the two species (Supplementary Table S5), and four constraint variables (alt, AET, bio3, and bio14) significantly influenced the genetic structure of the plant populations (type II analysis of variance (ANOVA), P < 0.001, Supplementary Table S6). By contrast, bio13 did not significantly explain the genetic variation in all repeated analyses in which any other variable was removed (not shown). The distribution of these four significantly explanatory variables along the ordination axes was further examined by GLM, which revealed a significant correlation (P < 0.05) of all variables along axis 1 and axis 2, except a marginal significance in bio3 along axis 1 (P = 0.0717, Table 4). Four environmental variables had significant F statistics (adjusted R 2 = 0.704, 0.475, 0.362 and 0.749 for alt, AET, bio3, and bio14, respectively, Table 4). The high adjusted R 2 indicates that these estimates sufficiently explain these two axes of dbRDA. Consistent estimates of influence were obtained even when the order of predictors was altered; thus, these four variables are highly relevant for explaining the genetic structure of the plant populations.
The distributions of the sampled populations in the space of constraint ordination analysis by dbRDA were similar to the DAPC clustering patterns (Fig. 3C,D). The scatter and ordisurf plots for each environmental variable congruently showed broader ranges of environmental contours in populations of S. tashiroi compared with populations of S. playfairii (Supplementary Fig. S4). The Wulu population of S. tashiroi exhibited a different distribution of the space of dbRDA along axis 1 but similar contour ranges of environmental variables compared with the populations of S. playfairii, reflecting its sympatric distribution with S. playfairii in Wulu. By contrast, the distributions of the allopatric Lanyu and Taroko populations of S. tashiroi were too close to be distinguished in the dbRDA space and were positively associated with the explanatory elements AET and bio14 (Supplementary Fig. S4). Ordisurf plots clearly illustrated the niche differentiation between S. playfairii and S. tashiroi. However, it must be noted that this conclusion is only based on the results of estimation from a small number of populations (two populations of S. playfairii and three populations of S. tashiroi).
Discussion
Genetic evidence of non-allopatric speciation
This study provides a comprehensive assessment of the role of evolutionary background and environmental effects on the current genetic distribution of plant populations. The state-of-the-art ABC method was used to investigate the process of reproductive isolation (RI) (i.e., barriers to gene flow) between S. playfairii and S. tashiroi. Interspecific gene flow was detected at the beginning of speciation (Fig. 4), implying that divergent pressure occurred gradually rather than instantaneously28. In this ancestral gene-flow mode of speciation, divergent ecological adaptation may play a critical role in driving and maintaining RI29. However, the divergence of S. playfairii and S. tashiroi, which have very small estimated bidirectional migration rates and overlapping time estimates of divergence and gene flow (Fig. 4 and Supplementary Table S3), could be driven by integrated mechanisms, including adaptation and demographic change, e.g., vicariance15. In other words, adaptive and stochastic (drift) processes are not mutually exclusive for explaining the speciation of these two Scutellaria species. Although S. playfairii and S. tashiroi recently diverged (roughly 105 generations ago, Fig. 4 and Supplementary Table S3), the differentiated habitat preference likely increased the successful RI between these species. Such niche differentiation is attributable not to a single environmental factor but to the overall environmental heterogeneity (Table 3), consistent with Hutchinson’s metaphor of a multidimensional hypervolume niche30. However, the environmental variables used for niche analysis were present-day data rather than from ancient time; thus, the factors that we detected as responsible for species divergence are not the driving force for speciation but are likely mechanisms for maintaining or even for secondary accumulation of genetic divergence after speciation31.
The distribution of plants has been suggested to be synchronously affected by postglacial colonization and environmental heterogeneity32, 33. Drastic climatic change in the late Pleistocene may have resulted in distributional divergence between S. playfairii and S. tashiroi, which diverged roughly 0.2 Mya (as estimated by nuclear and chloroplast DNA sequences15) or <0.15 Mya (as estimated by microsatellite markers in this study). Larger-scale studies have demonstrated that both climate and geo-historical factors can explain the plant diversity of eastern Asian archipelagos, reflecting as well the interaction of recent ecological and evolutionary processes34,35,36. The inference of late Pleistocene divergence with initial gene flow (Fig. 4) is congruent with the inference of climate-dependent dispersal/vicariance events by Chiang et al.15. Furthermore, Scutellaria exhibit passive seed dispersal by water37 as well as high selfing or inbreeding rates due to the occasional cleistogamous flower26, 38. Such dispersal and mating mechanisms limit gene flow and favor divergence between populations (Table 2). The characteristic of self-compatibility also reduces the dilemma of fitness reduction of incipient species due to loss of pollinators cf. 39.
Ecological divergence maintains post-speciation reproductive isolation
After speciation, cyclic glacial/interglacial epochs influenced the population genetic structure of Scutellaria species in Taiwan15. The late Pleistocene climate cannot be obtained directly, but our results indicate a significant role of current environmental heterogeneity in shaping the genetic structure of Scutellaria populations (Table 4 and Supplementary Table S6). Several studies have also inferred that the spatial heterogeneity of local climates is responsible for genetic structure and demography in a variety of organisms40,41,42,43. Recent studies have also suggested that several environmental variables are together involved in RI between species44, 45 or in the genetic structure within species in Taiwan46.
Micro-environmental differences reduce the chance of population inter-diffusion and prevent genetic secondary contact if physical contact occurs47. An example is the sympatric populations of both species in Wulu, where both species bear similar environmental pressure (e.g., temperature and precipitation), with slight differences in micro-environments (e.g., loam substrate and more humid in the microhabitat of S. playfairii; sand substrate and more arid in the microhabitat of S. tashiroi, Supplementary Table S1 and Supplementary Data). In our field observations, a few hybrids were found but were not widely spread in the distribution range of Wulu, suggesting that the fitness reduction of hybrids reduced the intensity of hybrid expansion (i.e., extrinsic postzygotic isolation or hybrid breakdown). Although we cannot completely rule out the probability that the unique genetic cluster of Wulu population in S. tashiroi in BCA (K = 3, Fig. 2B) is a consequence of asymmetric gene flow from the sympatric population of S. playfairii, the low observed heterozygosity (0.098), no chimera individuals, and lower marginal density of recent contact model (SC and CM, Fig. 4) in ABC together reduce the possibility of unequal rates of introgression leading unique genetic cluster in Wulu population of S. tashiroi. The better explanation for the unique genotype of Wulu should be caused by a lack of effective gene flow with other S. tashiroi populations over a long term, rather than the acquisition of genes from other species. These observations support our speculation that the ecological heterogeneity between these two species plays a critical role in maintaining species divergence.
Notably, the integrated variable (i.e., PC1, Table 3) but no single environmental variable differentiated the species, implying that the observed niche differentiation can be explained as a consequence of adaptive divergence instead of as a driving force for speciation. Adaptive divergence for speciation is usually initiated by single or simple environmental differences, followed by the expansion of the accumulating genetic divergence to the whole genome and subsequent niche divergence5, 48. This integration of environmental factors in the niche-allocation explanation is also reflected in the results of the more accurate spatial distribution prediction using multiple environmental predictors (Fig. 5). In this study, we used neutral markers to represent the genetic background, and hence the estimated population/species divergence is a surrogate for the degree of RI cf. 47. The significant correlation between the environmental variables and neutral genetic diversity suggests that the demographic dynamics and population structure are governed by environmental variation (Table 4 and Supplementary Tables S5 and S6). The lack of correlation among genetic, geographic, and environmental distances (Supplementary Note online) also implies that the genetic structure of the current population cannot be explained only by the current geographic distribution and environments but should also be jointly explained by the evolutionary history (paleodistribution, ancient selective pressures, etc.).
In our sampled population, the nonsignificant genetic differentiation between species estimated by AMOVA (Table 2) is contrary to the population structure estimation by BCA (Fig. 2) and DAPC (Fig. 3). This discrepancy is likely attributable to masking of the genetic delimitation of species in AMOVA by the high proportion of common ancestral polymorphisms due to recent divergence. However, the significant population differentiation estimated by AMOVA of either neutral loci or positive-outlier loci (Table 2) supports the inference that local adaptive divergence has already facilitated population differentiation. Traditionally, the genetic diversity of neutral genes is expected to be correlated with geographic distance and demographic events, whereas local environmental pressures act on certain “adaptive” genes under natural selection49. However, the lower fitness of immigrants could increase the genetic divergence between nearby populations and further result in correlations of neutral genetic variation with local environmental heterogeneity by facilitating genetic drift cf. 47, 50, 51. In other words, our data suggest that local environmental heterogeneity extends the ecological divergence among populations, i.e., post-speciation adaptive divergence.
Conclusions
Adaptation divergence is usually involved in speciation on islands with rugged topography and varied environments. It is difficult to detect the source of natural selection that leads to species divergence at the initial stage of speciation, but adaptive divergence can be genetically detected via heterogeneous genomic differentiation52 and ancestral gene flow between species5. In addition, environmental pressures for maintaining and enhancing species divergence, even local adaption between populations, can be tested by correlation analysis of genetic variation and environmental variables47. Here, we provided a case of ecological speciation and local adaptive divergence between two island herb species with limited distributions and clarified the mechanisms influencing the population genetics of these plants. We inferred that the evolutionary history, i.e., late Pleistocene divergence marked by early interspecies gene flow, and the incorporation of climate pressures shaped the current genetic distribution of Scutellaria species in eastern Taiwan. Integrated environmental factors jointly maintained the species divergence and shaped the current genetic spatial distribution. Our study provides a comprehensive assessment of the historical and environmental effects impacting the local adaptation of insular plants.
Materials and Methods
Study sites for genetic analyses
Two populations of S. playfairii (Wutai and Wulu) and three populations of S. tashiroi (Taroko, Wulu, and Lanyu) were sampled for genetic accession. These populations all differed in terms of habitat characteristics (e.g., soil types, shade, and precipitation; Supplementary Table S1). Wulu is the only location in which both species are sympatrically distributed. Lanyu is a small island off southeastern Taiwan. Hence, our study system includes two different plant species with both allopatric and sympatric distributions, enabling an understanding of how different environmental conditions affect the genetic divergence of plants.
Genetic analyses
The evaluation of the genetic diversity of plants was based on 19 selected microsatellite loci, adapted from Chiang et al.53 (Supplementary Data). Neutral loci and candidate adaptive loci (positive outliers) were determined based on a multinomial-Dirichlet distribution under simulations of reversible-jumped Markov chain Monte Carlo (rjMCMC) using BayeScan 2.154. Outlier loci were determined after running ×105 iterations with a thinning interval of 1000, posterior odds (PO) >10 (strong selection P = 0.90), and a q-value < 0.05 for the false-discovery rate as the threshold settings. We conducted DAPC and BCA using the function dapc in the R55 package adegenet56, and we used STRUCTURE 2.3.457 to discriminate genetic clusters. DAPC is a “without a priori” method using partial synthetic variables to minimize variation within groups58, 59, whereas BCA is a population-model-based approach based on the Hardy-Weinberg and linkage equilibria60. For DAPC, groups of “species” and “populations” were set to conduct the DAPC separately, and the first two principal components (PCs) were selected to discriminate groups. For BCA, the individual assignment was conducted under admixture model without incorporating locality information (i.e. without LOC prior). The correlated allele frequency model was assumed for BCA. Total 10 independent runs were conducted for BCA. Each run contains 1 million MCMC simulation followed by 100 thousand steps burn-in. These 10 runs were aligned and summarized using CLUMPP61. Analysis of molecular variance (AMOVA) by hierarchical groupings of species and populations permitted an assessment of population structure.
Evaluating speciation models by approximate Bayesian computation
We tested four scenarios by ABC: (1) CI model: gene flow was not permitted during divergence (Fig. 4A); (2) AM model: gene flow persisted only during initial divergence (Fig. 4B); (3) SC model: gene flow occurred in late periods of divergence (Fig. 4C); (4) CM model: gene flow persisted continuously during divergence (Fig. 4D). We set the migration rate as a free parameter in both directions in AM, SC, and CM models, which let both situations of symmetric and asymmetric interspecific gene flow (introgression) to be considered in each migration model. For performing the ABC, we used 15 neutral microsatellite loci and discarded four outlier loci for analysis, because these outlier loci could be adaptive under positive or balancing selection62 and could exhibit different evolutionary scenarios to neutral loci52, 63, 64. One million simulations were generated using the package simcoal2 implemented in ABCtoolbox65. The summary statistics were reduced to five partial least-squares components following the instructions in ABCtoolbox. Using the R package ABCestimator, we performed a general linear model post-sampling regression adjustment on the best 5,000 simulations to obtain the marginal density of each model. We selected the best model scenario using the Bayes factor and ratio of marginal densities for each model.
Niche differentiation between species estimated by reduction of environmental factors
To infer the niche distributions of S. playfairii and S. tashiroi, a total of 71 climatic variables were retrieved from WorldClim [elevation (alt), monthly precipitation (prec), temperature (tmin and tmax), water vapor pressure (vapr) and bioclim, http://www.worldclim.org/bioclim] and Consultative Group for International Agricultural Research - Consortium for Spatial Information (CGIAR-CSI) [global aridity, potential evapotranspiration (PET), and actual evapotranspiration (AET), http://www.cgiar-csi.org/data] (Supplementary Data). The occurrence data of both species were obtained from our sampling sites, herbarium records, and Global Biodiversity Information Facility (GBIF, http://www.gbif.org). Total 75 (S. playfairii) and 78 (S. tashiroi) records have been obtained. Species occurrence data were checked and cleaned. We followed instruction in dismo packages implemented in R66. Repeated records (multiple records of the same species in the same grid) of occurrence sites were removed to leave one record only in order to avoid unnecessary weight. Furthermore, each specimen was checked if available, and occurrence records of misidentified species were discarded. Occurrence data that did not project onto land were removed. Finally, twenty-three and sixteen records were preserved for S. playfairii and S. tashiroi, respectively. Variables with a high VIF were removed to reduce multicollinearity. We repeated this procedure until the VIF values of all remaining variables were <10, and finally, five environmental variables, alt, AET, bio3, bio13, and bio14, were retained for further analyses.
First, PCA was conducted to reduce the dimensionality of these five variables defining the niche space, thus enabling comparison of the integrity of Grinnellian niches between S. playfairii and S. tashiroi. Next, we performed the Kruskal-Wallis rank-sum test to detect differences in environmental variables between species. Then, we analyzed the distribution of environmental variables along the PCs of significant species differences by GLM to confirm the significance of the correlation of variables on the PCs. Multivariate logistic regression was further conducted to test whether the environmental variables significantly predicted species differentiation.
Ecological niche modeling to predict the species distribution pattern
To demonstrate the spatial distributions of the environmental niches of the two species, ecological niche modeling (ENM, also known as species distribution modeling) was performed under the maximum entropy model implemented in MaxEnt67 and the R package raster68. For ENM, 71 environmental variables (ENM1) and five retaining variables (ENM2) were used to predict the potential species distributions. All variables were re-scaled to a resolution of 30 arc-sec (approximately 1 km × 1 km). A maximum of 2000 iterations was conducted for each species, and the species occurrence data were randomly divided (10%) to train the model. One regularization multiplier and 10,000 background points were set to create models for each species set. The logistic output consisting of a grid map with a probability value ranging from 0 to 1 was generated and visualized with the R package maptools69. The predicted models were evaluated by the AUC value, which is a ranked approach for evaluating model fit and determines the probability that a presence location is ranked higher than a random simulation.
Redundancy analyses of environmental variables for assessing influence on genetic variation
To test environmental effects on the genetic variation of the sampled populations of S. tashiroi and S. playfairii, dbRDA was conducted with the first two principal components (PC1 and PC2) of the microsatellite data as the genetic response and the remaining five environmental variables (alt, AET, bio3, bio13, and bio14) after VIF analysis as the environmental predictors. However, since only five populations were sampled for genotyping, which limits the number of variables applying to the genetic response, we repeated the dbRDA several times by discarding one of the environmental predictors. Finally, we found that bio13 could not obtain a significant explanation in any analysis. Thus, bio13 was excluded, leaving only the four other environmental variables as predictors for the dbRDA. These four constrained environmental variables were then transformed to the Euclidean distance to perform the RDA (i.e., dbRDA). Type II ANOVA was then used to test the significance of each environmental variable for explaining the genetic composition under 9999 permutations. We further analyzed the distribution of environmental variables along the ordination axes using the GLM. GLM analysis facilitates the elucidation of the importance of variables in the explanation of genetic composition under the dbRDA. We fit counters of current climate variables onto the ordination space of dbRDA plots to show the association of environmental variables with genetic divergence.
References
Shafer, A. B. A. & Wolf, J. B. W. Widespread evidence for incipient ecological speciation: a meta-analysis of isolation-by-ecology. Ecol. Lett. 16, 940–950, doi:10.1111/ele.12120 (2013).
Sexton, J. P., Hangartner, S. B. & Hoffmann, A. A. Genetic isolation by environment or distance: which pattern of gene flow is most common? Evolution 68, 1–15, doi:10.1111/evo.12258 (2014).
Coyne, J. A. Speciation in a small space. Proc. Natl. Acad. Sci. USA 108, 12975–12976, doi:10.1073/pnas.1110061108 (2011).
Via, S. Natural selection in action during speciation. Proc. Natl. Acad. Sci. USA 106, 9939–9946, doi:10.1073/pnas.0901397106 (2009).
Wu, C. I. The genic view of the process of speciation. J. Evol. Biol 14, 851–865, doi:10.1046/j.1420-9101.2001.00335.x (2001).
Zitari, A., Scopece, G., Helal, A. N., Widmer, A. & Cozzolino, S. Is floral divergence sufficient to maintain species boundaries upon secondary contact in Mediterranean food-deceptive orchids? Heredity 108, 219–228, doi:10.1038/hdy.2011.61 (2012).
Palumbi, S. R. Speciation and the evolution of gamete recognition genes: pattern and process. Heredity 102, 66–76, doi:10.1038/hdy.2008.104 (2009).
Scopece, G., Musacchio, A., Widmer, A. & Cozzolino, S. Patterns of reproductive isolation in Mediterranean deceptive orchids. Evolution 61, 2623–2642, doi:10.1111/j.1558-5646.2007.00231.x (2007).
Ellstrand, N. C. & Elam, D. R. Population genetic consequences of small population size: Implications for plant conservation. Annu. Rev. Ecol. Syst. 24, 217–242, doi:10.1146/annurev.es.24.110193.001245 (1993).
Slatkin, M. Gene flow and the geographic structure of natural populations. Science 236, 787–792, doi:10.1126/science.3576198 (1987).
Hsieh, T. H. & Huang, T. C. Notes on the Flora of Taiwan (20) - Scutellaria (Lamiaceae) in Taiwan. Taiwania 40, 35–56 (1995).
Huang, B. H. et al. Positive selection and functional divergence of R2R3-MYB paralogous genes expressed in inflorescence buds of Scutellaria species (Labiatae). Int J Mol Sci 16, 5900–5921, doi:10.3390/Ijms16035900 (2015).
Huang, B. H., Chen, Y. W., Huang, C. L., Gao, J. & Liao, P. C. Imbalanced positive selection maintains the functional divergence of duplicated DIHYDROKAEMPFEROL 4-REDUCTASE genes. Sci Rep 6, 39031, doi:10.1038/srep39031 (2016).
Huang, B. H., Chen, Y. W., Huang, C. L., Gao, J. & Liao, P. C. Diversifying selection of the anthocyanin biosynthetic downstream gene UFGT accelerates floral diversity of island Scutellaria species. BMC Evol. Biol. 16, 191, doi:10.1186/s12862-016-0759-0 (2016).
Chiang, Y. C., Huang, B. H. & Liao, P. C. Diversification, biogeographic pattern, and demographic history of Taiwanese Scutellaria species inferred from nuclear and chloroplast DNA. PLOS ONE 7, e50844, doi:10.1371/journal.pone.0050844 (2012).
He, Q., Edwards, D. L. & Knowles, L. L. Integrative testing of how environments from the past to the present shape genetic structure across landscapes. Evolution 67, 3386–3402, doi:10.1111/evo.12159 (2013).
Balkenhol, N., Waits, L. P. & Dezzani, R. J. Statistical approaches in landscape genetics: an evaluation of methods for linking landscape and genetic data. Ecography 32, 818–830, doi:10.1111/j.1600-0587.2009.05807.x (2009).
Blair, M. E., Sterling, E. J., Dusch, M., Raxworthy, C. J. & Pearson, R. G. Ecological divergence and speciation between lemur (Eulemur) sister species in Madagascar. J. Evol. Biol 26, 1790–1801, doi:10.1111/jeb.12179 (2013).
Glor, R. E. & Warren, D. Testing ecological explanations for biogeographic boundaries. Evolution 65, 673–683, doi:10.1111/j.1558-5646.2010.01177.x (2011).
Csillery, K., Blum, M. G., Gaggiotti, O. E. & Francois, O. Approximate Bayesian Computation (ABC) in practice. Trends Ecol Evol 25, 410–418, doi:10.1016/j.tree.2010.04.001 (2010).
Bertorelle, G., Benazzo, A. & Mona, S. ABC as a flexible framework to estimate demography over space and time: some cons, many pros. Mol. Ecol. 19, 2609–2625, doi:10.1111/j.1365-294X.2010.04690.x (2010).
Yang, Y.-Q., Huang, B.-H., Yu, Z.-X. & Liao, P.-C. Inferences of demographic history and fine-scale landscape genetics in Cycas panzhihuaensis and implications for its conservation. Tree Genet. Genom. 11, 1–15, doi:10.1007/s11295-015-0894-6 (2015).
Hsiung, H.-Y. et al. Local climate heterogeneity shapes population genetic structure of two undifferentiated insular Scutellaria species. Front Plant Sci 8, 159, doi:10.3389/fpls.2017.00159 (2017).
Wang, Y. H. et al. Quaternary climate change drives allo-peripatric speciation and refugial divergence in the Dysosma versipellis-pleiantha complex from different forest types in China. Scientific Reports 7, 40261, doi:10.1038/srep40261 (2017).
Cruzan, M. B. Population size and fragmentation thresholds for the maintenance of genetic diversity in the herbaceous endemic Scutellaria montana (Lamiaceae). Evolution 55, 1569–1580 (2001).
Sun, M. Cleistogamy in Scutellaria indica (Labiatae): effective mating system and population genetic structure. Mol. Ecol. 8, 1285–1295 (1999).
Beaumont, M. A. Adaptation and speciation: what can F st tell us? Trends Ecol. Evol. 20, 435–440, doi:10.1016/j.tree.2005.05.017 (2005).
Roux, C. et al. Shedding light on the grey zone of speciation along a continuum of genomic divergence. PLoS Biol. 14, e2000234, doi:10.1371/journal.pbio.2000234 (2016).
Schuler, H., Hood, G. R., Egan, S. P. & Feder, J. L. Modes and mechanisms of speciation. Reviews in Cell Biology and Molecular Medicine 2, 60–93, doi:10.1002/3527600906.mcb.201600015 (2016).
Hutchinson, G. E. Homage to Santa Rosalia or why are there so many kinds of animals? Am Nat 93, 145–159 (1959).
Wolf, J. B. & Ellegren, H. Making sense of genomic islands of differentiation in light of speciation. Nat Rev Genet. doi:10.1038/nrg.2016.133 (2016).
Svenning, J. C. & Skov, F. Mesoscale distribution of understorey plants in temperate forest (Kalo, Denmark): the importance of environment and dispersal. Plant Ecol. 160, 169–185, doi:10.1023/A:1015885321263 (2002).
Pearson, R. G. & Dawson, T. P. Predicting the impacts of climate change on the distribution of species: are bioclimate envelope models useful? Global Ecol. Biogeogr. 12, 361–371, doi:10.1046/j.1466-822X.2003.00042.x (2003).
Kubota, Y., Hirao, T., Fujii, S. J. & Murakami, M. Phylogenetic beta diversity reveals historical effects in the assemblage of the tree floras of the Ryukyu Archipelago. J. Biogeogr. 38, 1006–1008, doi:10.1111/j.1365-2699.2011.02491.x (2011).
Kubota, Y., Hirao, T., Fujii, S., Shiono, T. & Kusumoto, B. Beta diversity of woody plants in the Japanese archipelago: the roles of geohistorical and ecological processes. J. Biogeogr. 41, 1267–1276, doi:10.1111/jbi.12290 (2014).
Kubota, Y., Shiono, T. & Kusumoto, B. Role of climate and geohistorical factors in driving plant richness patterns and endemicity on the east Asian continental islands. Ecography 38, 639–648, doi:10.1111/ecog.00981 (2015).
Williams, P. A. Ecology of the endangered herb Scutellaria novae-zelandiae. N. Z. J. Ecol. 16, 127–135 (1992).
Denda, T. Cleistogamy in Scutellaria rubropunctata Hayata (Labiatae) from the Ryukyu Archipelago. Biol Mag Okinawa 40, 7–14 (2002).
Roels, S. A. B. & Kelly, J. K. Rapid evolution caused by pollinator loss in Mimulus guttatus. Evolution 65, 2541–2552, doi:10.1111/j.1558-5646.2011.01326.x (2011).
Liao, P. C. et al. Historical spatial range expansion and a very recent bottleneck of Cinnamomum kanehirae Hay. (Lauraceae) in Taiwan inferred from nuclear genes. BMC Evol. Biol. 10, 124, doi:10.1186/1471-2148-10-124 (2010).
Chen, S. F., Rossiter, S. J., Faulkes, C. G. & Jones, G. Population genetic structure and demographic history of the endemic Formosan lesser horseshoe bat (Rhinolophus monoceros). Mol. Ecol. 15, 1643–1656, doi:10.1111/j.1365-294X.2006.02879.x (2006).
Huang, J. P. & Lin, C. P. Diversification in subtropical mountains: Phylogeography, Pleistocene demographic expansion, and evolution of polyphenic mandibles in Taiwanese stag beetle, Lucanus formosanus. Mol. Phylogen. Evol. 57, 1149–1161, doi:10.1016/j.ympev.2010.10.012 (2010).
Huang, J. P. & Lin, C. P. Lineage-specific late pleistocene expansion of an endemic subtropical gossamer-wing damselfly, Euphaea formosa, in Taiwan. BMC Evol. Biol. 11, 94, doi:10.1186/1471-2148-11-94 (2011).
Huang, C. L. et al. Disentangling the effects of isolation-by-distance and isolation-by-environment on genetic differentiation among Rhododendron lineages in the subgenus Tsutsusi. Tree Genet. Genom. 12, 53, doi:10.1007/s11295-016-1010-2 (2016).
Chen, C. Y. et al. Demography of the upward-shifting temperate woody species of the Rhododendron pseudochrysanthum complex and ecologically relevant adaptive divergence in its trailing edge populations. Tree Genet. Genom 10, 111–126, doi:10.1007/s11295-013-0669-x (2014).
Huang, C. L. et al. Influences of environmental and spatial factors on genetic and epigenetic variations in Rhododendron oldhamii (Ericaceae). Tree Genet. Genom. 11, 823, doi:10.1007/S11295-014-0823-0 (2015).
Lee, C. R. & Mitchell-Olds, T. Quantifying effects of environmental and geographical factors on patterns of genetic differentiation. Mol. Ecol. 20, 4631–4642, doi:10.1111/j.1365-294X.2011.05310.x (2011).
Nosil, P., Harmon, L. J. & Seehausen, O. Ecological explanations for (incomplete) speciation. Trends Ecol. Evol. 24, 145–156, doi:10.1016/j.tree.2008.10.011 (2009).
Holderegger, R., Kamm, U. & Gugerli, F. Adaptive vs. neutral genetic diversity: implications for landscape genetics. Landscape Ecol. 21, 797–807, doi:10.1007/s10980-005-5245-9 (2006).
Thibert-Plante, X. & Hendry, A. P. When can ecological speciation be detected with neutral loci? Mol. Ecol. 19, 2301–2314, doi:10.1111/j.1365-294X.2010.04641.x (2010).
Nosil, P., Egan, S. P. & Funk, D. J. Heterogeneous genomic differentiation between walking-stick ecotypes: “Isolation by adaptation” and multiple roles for divergent selection. Evolution 62, 316–336, doi:10.1111/j.1558-5646.2007.00299.x (2008).
Nosil, P., Funk, D. J. & Ortiz-Barrientos, D. Divergent selection and heterogeneous genomic divergence. Mol. Ecol. 18, 375–402, doi:10.1111/j.1365-294X.2008.03946.x (2009).
Chiang, Y. C. et al. Characterization of 24 transferable microsatellite loci in four skullcaps (Scutellaria, Labiatae). Am. J. Bot. 99, E24–E27 (2012).
Foll, M. & Gaggiotti, O. A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180, 977–993, doi:10.1534/genetics.108.092221 (2008).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, 2015).
Jombart, T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405, doi:10.1093/bioinformatics/btn129 (2008).
Hubisz, M. J., Falush, D., Stephens, M. & Pritchard, J. K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 9, 1322–1332, doi:10.1111/j.1755-0998.2009.02591.x (2009).
Manel, S., Gaggiotti, O. E. & Waples, R. S. Assignment methods: matching biological questions techniques with appropriate. Trends Ecol. Evol. 20, 136–142, doi:10.1016/j.tree.2004.12.004 (2005).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet. 11, 94, doi:10.1186/1471-2156-11-94 (2010).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Jakobsson, M. & Rosenberg, N. A. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806, doi:10.1093/bioinformatics/btm233 (2007).
Stephan, W. Signatures of positive selection: from selective sweeps at individual loci to subtle allele frequency changes in polygenic adaptation. Mol. Ecol. 25, 79–88, doi:10.1111/mec.13288 (2016).
Minder, A. M. & Widmer, A. A population genomic analysis of species boundaries: neutral processes, adaptive divergence and introgression between two hybridizing plant species. Mol. Ecol. 17, 1552–1563, doi:10.1111/j.1365-294X.2008.03709.x (2008).
de Lafontaine, G., Prunier, J., Gérardi, S. & Bousquet, J. Tracking the progression of speciation: variable patterns of introgression across the genome provide insights on the species delimitation between progenitor–derivative spruces (Picea mariana × P. rubens). Mol. Ecol. 24, 5229–5247, doi:10.1111/mec.13377 (2015).
Wegmann, D., Leuenberger, C., Neuenschwander, S. & Excoffier, L. ABCtoolbox: a versatile toolkit for approximate Bayesian computations. BMC Bioinformatics 11, doi:10.1186/1471-2105-11-116 (2010).
Hijmans, R. J., Phillips, S., Leathwick, J. & Elith, J. Dismo: Species distribution modeling. R package version 0.8–17 (2013).
Phillips, S. J. & Dudík, M. Modeling of species distributions with Maxent: new extensions and a comprehensive evaluation. Ecography 31, 161–175, doi:10.1111/j.0906-7590.2008.5203.x (2008).
Hijmans, R. J. & van Etten, J. raster: Geographic data analysis and modeling. R package version 2, 15 (2014).
Lewin-Koh, N. J. et al. Maptools: Tools for reading and handling spatial objects. R package version 0.8-10, URL http://CRAN.R-project.org/package=maptools (2011).
Acknowledgements
We thank the National Center for Genome Medicine of the National Core Facility Program for Biotechnology, the Ministry of Science and Technology, Taiwan, for technical and bioinformatics support. This research was financially supported by the Ministry of Science and Technology in Taiwan (MOST 105-2628-B-003 -002 -MY3 & MOST 105-2628-B-003 -001 -MY3) and was also subsidized by the National Taiwan Normal University (NTNU), Taiwan.
Author information
Authors and Affiliations
Contributions
P.C.L. conceived and designed the experiments. B.H.H., C.L.H., and P.C.L. contributed plant materials. B.H.H. performed the laboratory experiments. B.H.H. and P.C.L. analyzed the data. P.C.L. wrote the paper. B.H.H. and C.W.H. critically reviewed the manuscript. All authors participated in the discussion and read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, BH., Huang, CW., Huang, CL. et al. Continuation of the genetic divergence of ecological speciation by spatial environmental heterogeneity in island endemic plants. Sci Rep 7, 5465 (2017). https://doi.org/10.1038/s41598-017-05900-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-05900-1
This article is cited by
-
Localized environmental heterogeneity drives the population differentiation of two endangered and endemic Opisthopappus Shih species
BMC Ecology and Evolution (2021)
-
Changing Only Slowly: The Role of Phylogenetic Niche Conservatism in Caviidae (Rodentia) Speciation
Journal of Mammalian Evolution (2020)
-
Propagule pressure and hunting pressure jointly determine genetic evolution in insular populations of a global frog invader
Scientific Reports (2019)
-
Differential genetic responses to the stress revealed the mutation-order adaptive divergence between two sympatric ginger species
BMC Genomics (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.