Abstract
Previous studies suggested that Hd1 promoted heading under short-day conditions (SD) and delayed heading under long-day conditions (LD). However in this study, Hd1 was demonstrated to consistently promote heading date in Zhenshan 97 (ZS97) background by upregulating Ehd1, Hd3a and RFT1 expression under both SD and LD. While the high photoperiod sensitivity of Hd1 was observed in Minghui 63 (MH63) background, with heading being suppressed in LD but promoted in SD. Comparative analysis of two sets of near isogenic lines of Hd1 in MH63 and ZS97 backgrounds indicated that the alternative functions of Hd1 in promoting or suppressing heading under LD are dependent on the previously cloned flowering repressor gene Ghd7. The interaction between proteins Ghd7 and Hd1 occurred through binding of the CCT domain of Ghd7 to the transcription-activating domain of Hd1, resulting in suppression of Ehd1 and florigen gene expression. The involvement of the transcription-activating domain of Hd1 in this protein-protein interaction probably blocked or weakened its transcriptional activity. These findings suggest that Hd1 alone essentially acts as a promoter of heading date, and the protein interaction between Ghd7 and Hd1 determines photoperiod sensitivity and integrated Hd1-mediated and Ehd1-mediated flowering pathways in rice.
Similar content being viewed by others
Introduction
Rice (Oryza sativa L.) is a facultative short-day plant. It perceives the day length and flowers faster in short-day (SD) than in long-day (LD) conditions. Rice is cultivated widely across the globe, ranging from 45° north latitude to 35° south latitude1. Thus, rice shows high natural variations in heading date, which influences the responses of this crop to different photoperiods and temperatures. Heading date (HD) is a crucial determinant of regional and seasonal adaptation in rice2. Understanding the genetic basis of heading date will aid in the design of breeding schemes for specific regions and cropping seasons and contribute to safe rice production.
Since 2000, many heading date-related QTLs have been mapped using different genetic populations and methods3,4,5,6,7,8,9,10,11. Several major heading date QTLs have been cloned to date using a map-based cloning strategy12,13,14,15,16,17,18,19. Hd1, the ortholog of Arabidopsis CONSTANS (CO), is a major gene controlling heading date in rice. Hd1 inhibits Hd3a expression and delays heading date under LD and promotes Hd3a expression and accelerates heading date under SD19. Hd3a, the ortholog of Arabidopsis FT, is a rice florigen gene that is regulated by Hd1 13. Ehd1, which encodes a B-type response regulator, is a rice-specific heading date gene functioning upstream of Hd3a for which no ortholog has been identified in Arabidopsis 20. Several genes with pleiotropic effects on heading date, plant height, grain number and grain yield have been identified, such as Ghd7, Ghd8/DTH8 and Ghd7.1 15,16,17,18. These genes delay heading date by suppressing Ehd1 and increasing grain yield under LD. Two genes, Hd6 and Hd16, encoding casein kinases14, 21, regulate heading at protein level, in which Hd6 and Hd16 phosphorylate Ghd7 and Ghd7.1 in vitro, respectively, and delay heading date in LD22. In addition, several heading date genes, such as OsGI, RID1/OsId1/Ehd2, OsMADS51, Ehd3 and HAF1, have been isolated through reverse genetics23,24,25,26,27. This research progress contributes to our understanding of the genetic control of heading date in rice.
The molecular mechanism underlying photoperiod flowering has been well characterized in Arabidopsis and rice28,29,30,31,32. Arabidopsis is a typical long-day plant, and its photoperiodic flowering pathway is mediated by the GI-CO-FT pathway28. In contrast, two independent pathways have been identified in rice: the OsGI-Hd1-Hd3a pathway is conserved, sharing high similarity with the GI-CO-FT pathway in Arabidopsis 30, while the other pathway is mediated by OsGI-Ehd1-Hd3a, a unique flowering pathway in rice. The genes of Ghd7, Ghd8/DTH8, Ghd7.1 and OsCOL4 have been identified in the latter pathway. These genes repress the transcription of Hd3a and RFT1 (Rice Flowering Locus T 1) by downregulating Ehd1 expression independent of Hd1 in LD30. The OsGI-Hd1-Hd3a and OsGI-Ehd1-Hd3a pathways have been demonstrated to function as independent pathways in controlling heading date at the early stage20. However, recent studies have shown that Hd1 represses the expression of Ehd1 in LD33, 34, suggesting that Hd1- and Ehd1-mediated pathways are not always independent. In a recent report, Heading Date Repressor 1 (HDR1) and OsK4 were found to suppress heading date by upregulating Hd1 and downregulating Ehd1, and the HDR1-OsK4 interaction complex can phosphorylate Hd135.
Zhang et al.36 showed that in a mixed Zhenshan 97 (ZS97) and Miyang 46 genetic background, the Hd1 allele of ZS97 expresses photoperiod sensitivity and promotes heading in SD but delays heading in LD36. In contrast, in the ZS97 background, the Hd1 allele of Miyang46 exhibits photoperiod insensitivity, delayed flowering and increased plant height and grain productivity under either LD or SD conditions36. Therefore, the dual functions of Hd1 are dependent on the genetic background. However, it is not clear which gene in the genome background coordinates the photoperiod sensitivity of Hd1. In this study, 2 sets of Hd1 near isogenic lines in ZS97 and Minghui 63 (MH63) backgrounds exhibiting distinct photoperiod patterns are used as basic materials. Our objectives are (1) to accurately characterize the photoperiod of Hd1; (2) to identify which gene is interacted with Hd1; and (3) to elucidate how the function of Hd1 reverses between LD and SD.
Results
Validation of Ghd6
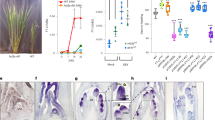
A BC4F1 plant with ZS97 as the recurrent parent and Teqing (TQ) as the donor parent headed later than ZS97. Genotyping of 150 SSR markers evenly distributed throughout the whole genome showed that the plant carried a heterozygous region between RM19746 and RM19795 on chromosome 6, and no other heterozygous regions were checked. A BC4F2 population of 219 plants were developed by selfing the BC4F1 plant. The progeny test of the BC4F2 population confirmed the co-segregation for spikelets per panicle, heading date and plant height under natural long-day conditions (NLD) (Supplementary Figure 1). The segregation ratio fitted a 3:1 for a single dominance gene (χ2 = 0.95 < χ2 0.05 = 3.84), indicating a pleiotropic gene between RM19746 and RM19795 controlling these trait variations. Subsequently, the gene was designated grains per panicle, plant height and heading date 6 (Ghd6). QTL analysis showed that Ghd6 explained approximately 78.5% of the observed heading date variance, with an additive effect of 6.0 d; 81.2% of plant height variance, with an additive effect of 7.7 cm; and 50.9% of the variance in spikelets per panicle, with an additive effect of 22.1 in the BC4F2 population (Supplementary Table 1). Ghd6 exhibited partial dominance (Supplementary Table 1). Homozygous plants containing the ZS97 region of Ghd6 (NIL-ZS) headed earlier, showing a shorter culm and smaller panicles than the TQ homozygotes (NIL-TQ) (Fig. 1A–C).
Performance of near isogenic lines of Ghd6/Hd1 and map-based cloning of Ghd6/Hd1. (A) Phenotype of the whole plants for NIL-ZS (left) and NIL-TQ (right), (A) the picture was taken when NIL-TQ was flowering; (B) the picture was taken when NIL-ZS was ripening; (C) main panicles for NIL-ZS (left) and NIL-TQ (right); (D) Map-based Cloning of Hd1; (E) comparison sequences of Hd1 of Teqing/Minghui 63 allele and Zhenshan 97 allele. The red rectangle represents the coding region of Hd1. The black region means the CCT domain of Hd1 from +1012 bp to +1146 bp. The black line means the intron. Diamond indicates the position of a 4-bp (AAAG) deletion in CCT domain on +1089 site. (F) The frame shift mutation in CCT domain in Teqing and Minghui 63 allele.
Fine mapping and cloning of Ghd6
A total of 3300 BC4F2 individuals were assessed for heading date, plant height and spikelets per panicle. A total of 576 late-heading plants with a tall culm were selected for Ghd6 fine mapping. First we screen the recombinants between Ghd6 and the flanking makers RM19746 and RM19795. In total, 10 and 14 recombinants were selected using the markers RM19746 and RM19795, respectively (Fig. 1D). Chromosome walking showed that Ghd6 co-segregated with markers of Ghd6-12 and S56, indicating that Ghd6 might localize to the region flanked by these two markers. The gene Hd1/Loc_Os06g16370 encoding a zinc finger transcription factor located in the target region has been reported to control heading date. Probably it was the candidate gene of Ghd6. Hence, we sequenced the Hd1 alleles of ZS97, TQ, NIL-ZS and NIL-TQ. ZS97 and NIL-ZS shared the same Hd1 sequence, which differed from that shared by TQ and NIL-TQ. Compared with the ZS97 allele, the TQ allele harbored a 4-bp (AAAG) deletion in the CCT domain at the +1089 bp site, leading to a frame-shift mutation resulting in loss of Hd1 function (Fig. 1E,F). All these 576 late heading plants exhibited the same genotype as TQ with the 4-bp deletion. Therefore, Hd1 was the most likely candidate gene for Ghd6.
Complementation test of Hd1
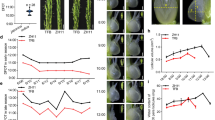
To determine whether Hd1 was Ghd6, the 1450-bp fragment upstream of the translation start codon (ATG) and the complete coding region of Hd1 from ZS97 were introduced into the PFA1300-GFP vector and transformed into NIL-TQ. In the T1 family of T0 plant 85, the positive transformed plants exhibited a significantly earlier heading date than the negative plants in Hainan (natural short-day conditions, NSD) (Fig. 2A). All but unhealthy one of the 7 negative plants displayed a taller plant height (Fig. 2B). One positive homozygous line and one negative line from T2 generation of T0 plant 85 were chosen to recheck the phenotype variation, the positive homozygous lines still exhibited a significantly earlier heading date, shorter plant height and fewer spikelets per panicle than the negative lines under NLD (Fig. 2C–F). Generations T1 and T2 from T0 plant 86 displayed a similar phenotype in NSD and NLD (Supplementary Table 2). In addition, several Hd1 knockout mutants in NIL-ZS were created using the CRISPR-Cas9 method. As expected, knockout mutants 24 and 25, harboring different mutations, headed later than NIL-ZS by 18.7 and 16.9 days, respectively. Their heading dates were similar to NIL-TQ, which flowered later than NIL-ZS by 18.2 days (Fig. 2G). Taken together, the results indicated that Hd1 was the pleiotropic gene underlying Ghd6.
Phenotypes of Hd1 transformation plants. (A) The heading date of T1 individuals from T0 plant 85 by transforming ProHd1:Hd1:GFP into NIL-TQ; (B) The plant height of T1 individuals from T0 plant 85; (C) a picture for homozygous positive and negatives T2 plants from T0 plant 85; (D) the heading date of T2 plants by selfing 85; (E) the plant height of T2 plants by selfing 85; (F) the spikelet per panicle of T2 plants by selfing 85; (G) the heading date of Hd1 knock out plants 24 and 25 in T1 generation. “+” and “−” means the positive and negative transformed plants, respectively.
Response of Ghd6 to photoperiod
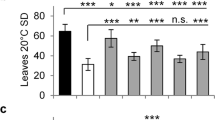
When NIL-ZS and NIL-TQ plants were treated under LD and SD (8 or 10 hours of natural light, followed by 16 or 14 hours of dark), they generally did not react to the photoperiod, as neither genotype exhibited a significant difference in heading between SD and LD. Moreover, NIL-ZS consistently headed approximately 13–15 days earlier than NIL-TQ in 2013, regardless the day-length conditions under which the plants were grown, and 15–17 days earlier in 2014 (Table 1). In addition, the differences in the single plant yield and spikelets per panicle remained stable (40% approximately increasing in NIL-TQ) across LD and SD (Table 1). The results indicated that Hd1/Ghd6 was insensitive to the photoperiod, which was not consistent with a previous report on Hd1 19.
Hd1 upregulates Ehd1 expression in ZS97 background, regardless of day length
In ZS97 background, Hd1 promoted heading date in both LD and SD. To address this phenomenon, the expression of Hd1, Ehd1, Hd3a and RFT1 in NIL-ZS and NIL-TQ was investigated under LD and SD. The expression levels of Ehd1, Hd3a and RFT1 in NIL-ZS were significantly higher than that in NIL-TQ under both LD and SD, indicating that Hd1 promoted the expression of Hd3a and RFT1 through upregulation of Ehd1 (Fig. 3A,B), resulting in early flowering, regardless of day length.
Photoperiod sensitivity of Hd1 in MH63 background
A previous study showed that Hd1 delayed heading date in LD but promoted heading date in SD. These inconsistent results indicated that genes hidden in the genome control alternative photoperiod sensitivity. To identify the gene(s) involved, two reciprocal Hd1 introgression lines, ZS-hd1 and MH-Hd1, from a cross between ZS97 and MH63 were screened. ZS-hd1 carried a non-functional MH63 hd1 allele in ZS97 background, which shared the same coding region as the hd1 allele, similar to TQ (Fig. 1E,F). MH-Hd1 carried a functional Hd1 from ZS97 in MH63 background. To understand the genetic background of ZS-hd1 and MH-Hd1, the RICE6K SNP array was employed for the analysis of both genotypes (Supplementary Figure 2). Few regions segregated in both genotypes. However, no QTLs other than Hd1 were located in the introgression segments, indicating that Hd1 was the unique factor responsible for the altered heading dates. Again, ZS97 harboring functional Hd1 headed earlier than ZS-hd1 in both NLD and NSD, by 11.3 and 17.4 days, respectively (Supplementary Figure 3A), consistent with the situation observed between NIL-ZS and NIL-TQ in LD and SD. However, compared with MH63, MH-Hd1 showed clear photoperiod sensitivity, with heading being delayed under NLD by 13.3 days and promoted under NSD by 23.7 days (Supplementary Figure 3A). Expression analysis showed that in LD, Hd1 upregulated Ghd7 and downregulated Ehd1, Hd3a and RFT1 in the morning (8:30 am), when these genes should be highly expressed (Supplementary Figure 3B). This expression pattern is consistent with the delayed heading performance observed in LD. In SD, Hd1 upregulated Ehd1, Hd3a and RFT1 in the morning (8:30 am) and at midnight (0:30 am) (Supplementary Figure 3C) and led to MH-Hd1 heading earlier than MH63. Hence, the photoperiod sensitivity of Hd1 was dependent on the genetic background.
Genetic interaction between Hd1 and Ghd7
The genetic interaction between Hd1, Ghd8 and Ghd7 has been reported to greatly delay flowering34. ZS97 and MH63 do not carry a functional Ghd8 gene, while ZS97 harbors a functional Hd1 and a deletion of Ghd7, and MH63 possesses Ghd7 and a non-functional hd1 16, 18, 34. We hypothesized that Ghd7 is the background determinant affecting the photoperiod sensitivity of Hd1. To confirm this hypothesis, we developed two F2 populations by crossing ZS-hd1 with ZS-Ghd7 carrying a functional Ghd7 from MH63 and crossing MH-Hd1 with MH-ghd7 possessing a non-functional ghd7 from ZS97. In NLD, Ghd7 significantly interacted with Hd1 in determining the heading date in ZS97 background (Table 2). Under LD, Hd1 promoted heading date by 14.7 and 5.6 days without Ghd7 in ZS97 and MH63 backgrounds, respectively, and Hd1 correspondingly repressed heading date by 8.3 and 12.3 days in the presence of Ghd7 (Fig. 4A,B). Accordingly, the effect of Ghd7 on delaying heading date was significantly altered between the Hd1 and hd1 backgrounds. Ghd7 in the ZS97 background with Hd1 delayed heading by 29.3 days, but the delay was decreased to 6.3 days without Hd1, while Ghd7 in MH63 background with Hd1 delayed heading by 14.8 days, but promoted heading by 2–3 days without Hd1 (Fig. 4A,B). In SD, regardless of the presence or absence of Ghd7, Hd1 consistently promoted the heading date by 14.1 and 19 days in the ZS97 background and by 11.6 and 7.4 days in the MH63 background (Fig. 4A,B). In the ZS97 background, Ghd7 repressed heading date by 16 days with Hd1 or 11 days without Hd1 (Fig. 4A,B). In MH63 background, the effect of Ghd7 in repressing heading date sharply decreased, regardless of Hd1 (Fig. 4A,B).
Genetic interaction on heading date between Ghd7 and Hd1 under long day and short day conditions. The heading date of 4 homozygous combinations between Ghd7 and Hd1 in Zhenshan 97 background (A) and Minghui 63 background (B). “NLD”, nature long days in Wuhan, Hubei; “NSD”, nature short days in Lingshui, Hainan. “LD”, controlled long day-length of 14 hours light; “SD”, controlled short day-length of 8 hours light.
Expression analysis of genes involved in the photoperiod pathway
After detecting the genetic interaction between Hd1 and Ghd7, we examined whether the expression patterns of key flowering genes also exhibited subtle changes in the presence of different gene combinations. Hence, gene expression was assessed in four homozygotes for Hd1 and Ghd7 in ZS97 background. In SD, no transcriptional regulation was observed between Ghd7 and Hd1 (Supplementary Figure 4). However, Hd1 upregulated Ghd7 in LD (Supplementary Figures 3B and 4). In LD, Ghd7Hd1 showed strongly suppressed expression of Ehd1 and Hd3a than Ghd7hd1 and Ghd7Hd1, resulting in the latest heading. In SD, Ghd7Hd1 exhibited moderately suppressed expression of Ehd1 and Hd3a, resulting in the second earliest heading among these 4 genotypes. The expression patterns of florigen genes were obviously dependent on the combination of Ghd7 and Hd1 genes. RFT1 is also an important florigen gene in the heading date pathway; however, a previous study showed that RFT1 is a nonfunctional allele in ZS97 background37. Therefore, we did not examine the expression of the RFT1 in ZS97 background, and a yeast one-hybrid assay showed that both Ghd7 and Hd1 could not bind to the promoter of Ehd1 from −1341 bp to +47 bp (Supplementary Figure 5).
Physical interaction between Ghd7 and Hd1
Ghd7 has been reported to repress heading via transcriptional regulation of Ehd1, but not of Hd1 16. It was considered most likely for Ghd7 to interact with Hd1 at the protein level. To confirm this potential interaction, the yeast two-hybrid system was used. Analysis of the transcriptional activation of Ghd7 revealed that Ghd7 presented the ability to undergo self-activation, conferred by the segment from amino acids 1 to 186 (Fig. 5A). Subsequently, the segment of Ghd7 (aa187–257) containing the CCT domain was used to test hybridization with Hd1 in a yeast two-hybrid assay. Significant interactions were observed between the Ghd7 (aa187–257) and Hd1 (aa32–111) segments harboring the zinc finger domain and the Hd1 segment (aa112–337) between the zinc finger domain and the CCT domain (Fig. 5B).
Ghd7 inhibits the transcriptional activation activity of Hd1
In CONSTANS (CO), the ortholog of Hd1 in Arabidopsis, transcriptional activation is conferred by the region between the zinc finger and CCT domains38. Thus, we suggested that Hd1 acts as a transcription activator and that the region (aa112–337) between the zinc finger and CCT domain confers this activity. Subsequently, several variants of Hd1 were cloned in-frame into the effector vector containing the GAL4 DNA-binding domain of yeast and co-transformed with a reporter vector into rice protoplasts for transcriptional activation activity assays (Supplementary Figure 6A). The relative LUC activity activated by GAL4-Hd1ZS, GAL4-Hd1NIP, GAL4-Hd1TQ, GAL4-Hd1ZS337 and GAL4-Hd1 ZS111 was 117.2-, 50.4-, 2.1-, 828.5- and 1.8-fold higher than in the GAL4-YFP control, respectively (Supplementary Figure 6B). These results suggested that Hd1 possessed transcriptional activation conferred by the region (aa112–337) between the zinc finger and CCT domains. The transcriptional activation activity of the TQ allele of Hd1 was 4.2 percent that of the ZS97 allele(Supplementary Figure 6B), indicating that the mutation in the CCT domain of Hd1 in the TQ allele might destroy its transcriptional activation activity, leading NIL-TQ to undergo late flowering compared with NIL-ZS flowering.
Accordingly, to survey whether Ghd7 affected the transcriptional activation of Hd1, the 35 S:Ghd7-CFP and GAL4-Hd1ZS vectors were co-transformed at different dosages into rice protoplasts, and analysis was performed via transcriptional activation assays. The relative LUC activity activated by GAL4-Hd1ZS gradually decreased from 7.0 to 0.8 with an increasing dosage of Ghd7-CFP (Supplementary Figure 6B), indicating that Ghd7 repressed the transcriptional activation activity of Hd1.
Discussion
Hd1 essentially acts as a promoter of the heading date in rice
In the present study, the heading date gene Ghd6 in ZS97 was found to be insensitive to photoperiod, promoting heading regardless of day length. Gene cloning showed that Hd1 was the gene underlying Ghd6. Thus Hd1 is photoperiod insensitivity in ZS97, which was not consistent with a previous report that Hd1 promotes heading in SD but delays heading in LD19. Photoperiod insensitivity of Hd1 was also previously observed in Hd1 mutant that delayed heading regardless of day length as compared to its wild type39. Moreover, the Hd1 allele from ZS97 promotes heading in both LD and SD36. Here, we further confirmed that Hd1 upregulated Ehd1, Hd3a and RFT1 in LD and SD and acted as a transcriptional activator; these findings are identical to the most recent study in which Hd1 was shown to activate the expression of Hd3a40. Thus, Hd1 essentially acts as a promoter of heading date in rice without Ghd7 (Supplementary Figure 6B).
The alternative functions of Hd1 are dependent on Ghd7 in LD
It is clear that Hd1 possesses opposite functions in the regulation of heading date under different day-length conditions19. Additional surveys have consistently confirmed the bifunctional character of Hd1 20, 36, 39, 41. In this study, we found that Hd1 essentially acts as a promoter of heading date in rice, but it delays heading in some genetic backgrounds such as the MH63 background in LD. These indicate function of Hd1 is genetic background dependent. The heading date performance of two sets of Hd1 NILs in the present study indicated that Ghd7 is the determinant of Hd1 bi-functionality. Without Ghd7, Hd1 promotes heading date in LD and SD by upregulating its downstream genes, including Ehd1, Hd3a and RFT1. Under LD, proteins of Ghd7 and Hd1 assemble into a complex through binding of the Ghd7 CCT domain to the transcription activation domain of Hd1. It has been suggested that the interaction between Ghd7 and Hd1 might block or weaken the transcriptional activation activity of Hd1 and release the transcriptional repression activity of Ghd7, thereby repressing the expression of downstream genes of Ehd1, Hd3a and RFT1, consequently delaying heading date in LD. It has been reported that Ghd7 expression is significantly lower in SD than in LD16. Thus, we hypothesized that this situation likely impairs or weakens the interaction between Ghd7 and Hd1 in SD. Therefore Hd1 still promotes the heading date in a Ghd7 background under SD. More evidence from biochemistry and structural biology analyses is needed to confirm this hypothesis.
In the presence of a functional allele of Hd1, Ghd7 exerts a large effect on heading date. Without Hd1, Ghd7 in ZS97 background has a much smaller effect on delaying heading, and the effect of Ghd7 is almost absent in MH63 background (Fig. 4A,B). It is likely for this reason that a few varieties with strong Ghd7 alleles have been observed in northeast China and Japan34, 42. Additionally, when Ghd7 segregating population in ZS97 background was used for gene cloning, its effects were enhanced that made Ghd7 cloning work feasible and successful16. Taken together, these findings indicate that the photoperiod sensitivities of Ghd7 and Hd1 are dependent on each other.
Integration of Hd1-mediated and Ehd1-mediated photoperiod pathways
Rice exhibits two photoperiod pathways that regulate heading, mediated by Hd1 and Ehd1. Previous studies suggested that these two pathways are independent20. Recent studies have indicated the Ehd1-mediated pathway is dominant, while Hd1-mediated pathway likely functions via the Ehd1 pathway40. The Ghd7Hd1 genotype displays significantly delayed heading under LD. At the transcriptional level, Ghd7Hd1 significantly downregulates Ehd1 in LD. Moreover, interactions were detected between Ghd7 and Hd1. However, these proteins do not directly bind to the promoter of Ehd1. ChIP analysis showed that Ghd7 was enriched at the promoter of Ehd1, but there were no cis-elements in the promoter of Ehd1 40. It is likely that Ghd7 and Hd1 are involved in a repression complex containing components that directly bind to the promoter of Ehd1. Ghd7 acts as a repressor of Ehd1, independent of Hd1 40, whereas Hd1 is an activator of Ehd1. When these proteins interacted under LD, the repression function of Ghd7 was observed, while under SD, the promotion function of Hd1 was primarily recorded. Considering these findings together, we suggest the existence of a regulatory network involving Ghd7 and Hd1 (Fig. 6). In SD, the interaction between Ghd7 and Hd1 was not observed. Hd1 promotes Hd3a expression either directly or via the upregulation of Ehd1, resulting in early heading. Under LD, without Ghd7, Hd1 alone upregulated Ehd1 and further upregulated Hd3a, ultimately leading to early heading. Without Hd1, Ghd7 alone suppressed Ehd1 and Hd3a, resulting in late heading. In the presence of Ghd7, Hd1 upregulated Ghd7, consistent with a previous report33, and a repressor protein complex including Ghd7, Hd1 and components of unknown identity was formed and suppressed Ehd1 and Hd3a/RFT1, resulting in markedly late heading.
In summary, Hd1 promotes heading without Ghd7 regardless of day length but represses heading through interactions with Ghd7 under LD. The interaction between Ghd7 and Hd1 determines their corresponding photoperiod sensitivities. The involvement of the transcription-activating domain of Hd1 in the protein-protein interaction likely abolishes its transcription-activating activity. The interaction between Ghd7 and Hd1 integrates the Hd1-mediated and Ehd1-mediated photoperiod flowering pathways. These findings provide new insight into the function of Hd1 and Ghd7 in the photoperiodic flowering pathway in rice.
Materials and Methods
Plant materials
A BC4F2 population showing varied heading dates was developed using ZS97 as the recurrent parent and Teqing (TQ) as the donor parent. The BC4F2 plants exhibiting homozygous TQ and ZS97 introgression fragments containing Ghd6/Hd1 were designated NIL-TQ and NIL-ZS, respectively. A total of 3300 BC4F2 plants were used to identify the gene underlying the variation of heading date. Introgression lines of Hd1 in the ZS97 background and MH63 background were scanned from reciprocal advanced chromosome segment substitution lines (CSSLs) between ZS97 and MH636] and will hereafter be referred to as ZS-hd1 (MH63 homozygous Hd1 alleles in ZS97 background) and MH-Hd1 (ZS97 homozygous Hd1 alleles in MH63 background), respectively. ZS-Ghd7, a NIL carrying MH63 Ghd7 alleles in ZS97 background16, was crossed with ZS-hd1, and a near isogenic F2 population was obtained to analyze the genetic interaction between Hd1 and Ghd7. A NIL without Ghd7 in MH63 background was also selected from the CSSLs, which was designated MH-ghd7.
Field experiments and photoperiod treatment
The plant materials were sown in the middle of May, and 25-day-old seedlings were transplanted to the field at the same planting density, with a distance of 16.5 cm between plants within a row and 26.5 cm between rows. The plants were subsequently grown under natural long-day field conditions (NLD), in which the day length was more than 13.5 hours, from the middle of May to the beginning of August at the experimental station of Huazhong Agriculture University, Wuhan, China (31°N latitude). For the experiments involving natural short-day field conditions, the experimental materials were sown in Lingshui, Hainan (18°N latitude) at the beginning of December and were transplanted to the field after 1 month, at the same planting density as in Wuhan, and grown under an average day-length of less than 12 hours from December to the middle of March. For the photoperiod treatment, the seedlings were initially grown in the natural field for 20 days under natural long days. Subsequently, 5 plants of each genotype were transplanted in parallel to the fields under NLD (LD) and short-day conditions (SD), with a day length of 8 or 10 hours and darkness of 16 or 14 hours in the field, and were covered with a black cloth until the growth phase transition of late heading plants was initiated.
Trait measurement and data analysis
HD was individually measured as the days from sowing to the emergence of the first panicle in the plant. The number of spikelets per panicle (SPP) was recorded as the total number of spikelets divided by the number of panicles. The grain yield (YD) was calculated as the grain weight per plant. Plant height (PH) was measured from the surface to the top of the main panicle. For the fine mapping of Ghd6 in the BC4F2 population, interval mapping was employed to identify the QTL underlying heading date and plant height using MAPMAKER/EXP and MAPMER/QTL43. The statistical significance of the two-way genetic interaction was assessed via an orthogonal contrast test using the program STATISTICA 8.0. Clustal Omega 1.2.2 was employed for comparisons of the Hd1 protein sequence (http://www.clustal.org/omega/).
Transformation of Hd1
To construct the ProHd1:Hd1:GFP vector, a 1450-bp fragment upstream of the coding start site and the CDS without the terminator of Hd1 from ZS97 were cloned into the PFA1300-GFP vector using the primers ProHd1-1450 and Hd1-GFP (Supplementary Table 3). PFA1300-GFP is a remolded PCAMBIA1300 vector fused with the GFP sequence as a tag. The ProHd1:Hd1:GFP plasmid was subsequently transformed into NIL-TQ by means of Agrobacterium-mediated transformation44.
To obtain the hd1 mutant, we used the vector pCXUN-Cas9 to knock out functional Hd1 in NIL-ZS lines. The target sequence was obtained from the website http://cbi.hzau.edu.cn/cgi-bin/CRISPR 45. The target sequence started with an “A” base, since we used the OsU3 promoter (Supplementary Table 3). Subsequently, we employed the overlapping PCR method to obtain gRNA expression cassettes46. The pOsU3-gRNA plasmid was used as a template for two rounds of PCR. The first round of PCR was performed using the primers Hd1-CRP-F and OsU3-R, and the second was performed used the primers Hd1-CRP-R and OsU3-F (Supplementary Table 3). We mixed the two obtained products for subsequent extraction and purification. Finally, the mixed product was employed for a third round of PCR, which was conducted using the primers OsU3-R and OsU3-F. Daniel Gibson’s enzymatic assembly method was employed to clone the gRNA fragment into the pCXUN-Cas9 vector, which was linearized using FastDigest KpnI (Thermo Scientific, USA)47. Finally, the plasmids were introduced into Agrobacterium strain EHA105 by electroporation and subsequently transformed into NIL-ZS callus according to a previous study44. After the Hd1 mutants were obtained, the primers Hd1-CRP-SeqF and Hd1-CRP-SeqR were used to examine the Hd1 mutations (Supplementary Table 3).
Analysis of diurnal gene expression
Seedlings were grown in pots under natural LD for 20 days and were subsequently transferred to an S10H growth chamber (Conviron, Canada), with half of the plants being grown under LD and the other half under SD. The growth conditions were set as follows: 14 h light and 10 h dark for LD, 10 h light and 14 h dark for SD; light intensity was set at 10,000 1x; and the temperature was 30 °C in the light period and 26 °C in the dark period. After treatment for 7 days, leaf samples for RNA extraction were collected from the LD and SD treatments at 3-h intervals over a 24-h period starting at 08:30. For each time point, leaves from three different plants were harvested as biological replicates. Total RNA was extracted using the TRIzol reagent (Invitrogen, San Diego, CA, USA) and treated with DNase I (Invitrogen). cDNA was synthesized from 3 μg of RNA using SuperScript III Reverse Transcriptase (Invitrogen). The quantitative analysis of gene expression was performed with SYBR Premix Ex Taq (TakaRa, Otsu, Japan) on an Applied Biosystems 7500 Real-time PCR System (Applied Biosystems, Foster City, CA, USA). The data were analyzed using the relative quantification method. Three biological replicates for every sample were prepared at every time point, and every sample was examined with three technical repeats. The mean data across the three technical replicates were regarded as the original data for one biological replicate, and the mean data across the three biological replicates were regarded as the final data for the comparative analysis. In the present study, four major heading date genes were analyzed during diurnal expression in LD and SD. The primers used for real-time PCR are listed in Table Supplement 3.
Yeast one-hybrid assay
We followed the method described by Wang et al.48 to conduct the yeast one-hybrid assay48. The −1786 to +47 region of Ehd1 from Nippobare was truncated into 4 fragments via PCR and inserted into the Placzi2μ vector (P1:LacZ, P2:LacZ, P3:LacZ, and P4:LacZ) (Supplement Table 3). The CDSs of Ghd7 (MH63 allele) and Hd1 (ZS97 allele) were cloned into the PJG4-5 vector (Ghd7-AD and Hd1-AD) using the primers Ghd7-Y1H and Hd1-Y1H (Supplement Table 3). Subsequently, Ghd7-AD, Hd1-AD and empty PJG4-5 were separately co-transfected with the five reporter vectors (the empty Placzi2μvector, P1:LacZ, P2:LacZ, P3:LacZ, and P4:LacZ) into the yeast strain EGY48. Empty PJG4-5 was also used as a control.
Yeast two-hybrid assay for Ghd7 and Hd1
For the yeast two-hybrid assay, the variants of Ghd7 and Hd1 were cloned in-frame into the pGBKT7 (Ghd7-BDs) and pGADT7 (Hd1-ADs) vectors, respectively (Fig. 5). For the Ghd7 self-activation experiment, the Ghd7-BD plasmids were individually co-transformed with pGADT7 into yeast strain AH109. Subsequently, the cells were grown on -2 SD medium lacking Leu and Trp (-LT) and -4 SD medium lacking Leu, Trp, His and Ade (-LTHA). For the yeast two-hybrid analysis of Ghd7 and Hd1, Ghd7-BD (187–257) was co-transformed with each of the Hd1-AD variants to AH109 to perform the assay of Ghd7 self-activation. All primers used are shown in Supplementary Table 3.
Transcriptional activation activity analysis of Hd1
We followed a previously reported analysis method for assessing the transcriptional activation activity of Hd148. The CDSs of Hd1 were amplified from varieties ZS97 and Nipponbare with primers GAL4-Hd1-ZS. The CDS of TQ were amplified with primers GAL4-Hd1-TQ. Two truncated Hd1 of ZS97 were amplified with GAL4-Hd1-337 and GAL4-Hd1-111. Then all Hd1 fragments cloned into effect vectors fused with the GAL4-binding domain using primers (Supplementary Table 3; Supplementary Figure 6A). YFP was also cloned into effect vectors as a control check (CK). Subsequently, the effect vectors, the reporter vector carrying Luciferase and the internal control vector carrying GUS were co-transformed into protoplasts of NIL-TQ at a ratio 2:2:1 (μg). Simultaneously, the 35 S:Ghd7:CFP vector18, was co-transformed with the GAL4-Hd1effect vector (including the full-length CDS of Hd1 from ZS97), the reporter vector and control vector into the protoplasts of NIL-TQ at dosages of 2:2:2:1 (μg), 4:2:2:1 (μg) and 10:2:2:1 (μg), to investigate the influence of Ghd7 on the transcriptional activation activity of Hd1.
References
Monfreda, C., Ramankutty, N. & Foley, J. A. Farming the planet: 2. Geographic distribution of crop areas, yields, physiological types, and net primary production in the year 2000. Global Biogeochem Cy 22 (2008).
Zhong, Z. Z. et al. Fine mapping of a minor-effect QTL, DTH12, controlling heading date in rice by up-regulation of florigen genes under long-day conditions. Mol Breeding 34, 311–322 (2014).
Huang, X. H. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42, 961–U976 (2010).
Jin, L. et al. Genetic diversity and population structure of a diverse set of rice germplasm for association mapping. Theor Appl Genet 121, 475–487 (2010).
Li, Z. K. et al. Genome-wide introgression lines and their use in genetic and molecular dissection of complex phenotypes in rice (Oryza sativa L.). Plant Mol Biol 59, 33–52 (2005).
Shen, G. J. & Xing, Y. Z. Two Novel QTLs for Heading Date Are Identified Using a Set of Chromosome Segment Substitution Lines in Rice (Oryza sativa L.). J Genet Genomics 41, 659–662 (2014).
Xing, Y. Z., Xu, C. G., Hua, J. P., Tan, Y. F. & Sun, X. L. Mapping and isolation of quantitative trait loci controlling plant height and heading date in rice. Acta Bot Sin 43, 721–726 (2001).
Yano, M. et al. Identification of quantitative trait loci controlling heading date in rice using a high-density linkage map. Theor Appl Genet 95, 1025–1032 (1997).
Yano, M., Kojima, S., Takahashi, Y., Lin, H. X. & Sasaki, T. Genetic control of flowering time in rice, a short-day plant. Plant Physiol 127, 1425–1429 (2001).
Zhan, X. et al. Genetic mapping of a QTL controlling source-sink size and heading date in rice. Gene 571, 263–270 (2015).
Zhang, Y. S. et al. Heading date QTL in rice derived from an analysis of chromosome segment substitution lines. Plant Breeding 130, 185–191 (2011).
Doi, K. et al. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev 18 (2004).
Kojima, S. et al. Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short-day conditions. Plant Cell Physiol 43, 1096–1105 (2002).
Takahashi, Y., Shomura, A., Sasaki, T. & Yano, M. Hd6, a rice quantitative trait locus involved in photoperiod sensitivity, encodes the alpha subunit of protein kinase CK2. P Natl Acad Sci USA 98, 7922–7927 (2001).
Wei, X. J. et al. DTH8 Suppresses Flowering in Rice, Influencing Plant Height and Yield Potential Simultaneously. Plant Physiol 153, 1747–1758 (2010).
Xue, W. et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40 (2008).
Yan, W. H. et al. Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res 23, 969–971 (2013).
Yan, W. H. et al. A Major QTL, Ghd8, Plays Pleiotropic Roles in Regulating Grain Productivity, Plant Height, and Heading Date in Rice. Mol Plant 4, 319–330 (2011).
Yano, M. et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. The Plant cell 12, 2473–2484 (2000).
Doi, K. et al. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-Iike gene expression independently of Hd1l. Gene Dev 18, 926–936 (2004).
Hori, K. et al. Hd16, a gene for casein kinase I, is involved in the control of rice flowering time by modulating the day-length response. Plant J 76, 36–46 (2013).
Kwon, C. T., Koo, B. H., Kim, D., Yoo, S. C. & Paek, N. C. Casein kinases I and 2alpha phosphorylate oryza sativa pseudo-response regulator 37 (OsPRR37) in photoperiodic flowering in rice. Molecules and cells 38, 81–88 (2015).
Hayama, R., Izawa, T. & Shimamoto, K. Isolation of rice genes possibly involved in the photoperiodic control of flowering by a fluorescent differential display method. Plant Cell Physiol 43, 494–504 (2002).
Kim, S. L., Lee, S. Y., Kim, H. J., Nam, H. G. & An, G. H. OsMADS51 is a short-day flowering promoter that functions upstream of Ehd1, OsMADS14, and Hd3a(1[W][OA]). Plant Physiol 145, 1484–1494 (2007).
Matsubara, K. et al. Ehd3, encoding a plant homeodomain finger-containing protein, is a critical promoter of rice flowering. Plant J 66, 603–612 (2011).
Wu, C. Y. et al. RID1, encoding a Cys2/His2-type zinc finger transcription factor, acts as a master switch from vegetative to floral development in rice. P Natl Acad Sci USA 105, 12915–12920 (2008).
Yang, Y. et al. The RING-Finger Ubiquitin Ligase HAF1 Mediates Heading date 1 Degradation during Photoperiodic Flowering in Rice. The Plant cell 27, 2455–2468 (2015).
Andres, F. & Coupland, G. The genetic basis of flowering responses to seasonal cues. Nature reviews. Genetics 13 (2012).
Hayama, R. & Coupland, G. The molecular basis of diversity in the photoperiodic flowering responses of Arabidopsis and rice. Plant Physiol 135, 677–684 (2004).
Shrestha, R., Gomez-Ariza, J., Brambilla, V. & Fornara, F. Molecular control of seasonal flowering in rice, arabidopsis and temperate cereals. Annals of botany 114, 1445–1458 (2014).
Simpson, G. G. & Dean, C. Flowering - Arabidopsis, the rosetta stone of flowering time? Science 296, 285–289 (2002).
Tsuji, H., Taoka, K. & Shimamoto, K. Regulation of flowering in rice: two florigen genes, a complex gene network, and natural variation. Curr Opin Plant Biol 14, 45–52 (2011).
Song, Y., Gao, Z. & Luan, W. Interaction between temperature and photoperiod in regulation of flowering time in rice. Science China. Life sciences 55, 241–249 (2012).
Zhang, J. et al. Combinations of the Ghd7, Ghd8 and Hd1 genes largely define the ecogeographical adaptation and yield potential of cultivated rice. The New phytologist 208, 1056–1066 (2015).
Sun, X. et al. The Oryza sativa Regulator HDR1 Associates with the Kinase OsK4 to Control Photoperiodic Flowering. PLoS genetics 12, e1005927 (2016).
Zhang, Z. H. et al. Pleiotropism of the photoperiod-insensitive allele of Hd1 on heading date, plant height and yield traits in rice. PloS one 7, e52538 (2012).
Zhao, J. et al. Genetic interactions between diverged alleles of Early heading date 1 (Ehd1) and Heading date 3a (Hd3a)/RICE FLOWERING LOCUS T1 (RFT1) control differential heading and contribute to regional adaptation in rice (Oryza sativa). New Phytologist 208, 936–948 (2015).
Tiwari, S. B. et al. The flowering time regulator CONSTANS is recruited to the FLOWERING LOCUS T promoter via a unique cis-element. The New phytologist 187, 57–66 (2010).
Izawa, T. et al. Phytochrome mediates the external light signal to repress FT orthologs in photoperiodic flowering of rice. Genes Dev 16, 2006–2020 (2002).
Nemoto, Y., Nonoue, Y., Yano, M. & Izawa, T. Hd1, a CONSTANS ortholog in rice, functions as an Ehd1 repressor through interaction with monocot-specific CCT-domain protein Ghd7. Plant J 86, 221–233 (2016).
Su, L., Shan, J. X., Gao, J. P. & Lin, H. X. OsHAL3, a Blue Light-Responsive Protein, Interacts with the Floral Regulator Hd1 to Activate Flowering in Rice. Mol Plant 9, 233–244 (2016).
Yonemaru, J. et al. Genomic regions involved in yield potential detected by genome-wide association analysis in Japanese high-yielding rice cultivars. BMC genomics 15, 346 (2014).
Lander, E. S. & Botstein, D. Mapping mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121, 185–199 (1989).
Hiei, Y., Ohta, S., Komari, T. & Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6, 271–282 (1994).
Lei, Y. et al. CRISPR-P: A Web Tool for Synthetic Single-Guide RNA Design of CRISPR-System in Plants. Mol Plant 7, 1494–1496 (2014).
Sun, Y. et al. Engineering Herbicide-Resistant Rice Plants through CRISPR/Cas9-Mediated Homologous Recombination of Acetolactate Synthase. Mol Plant 9, 628–631 (2016).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nature Methods 6, 343–345 (2009).
Wang, L. et al. Coordinated regulation of vegetative and reproductive branching in rice. Proc Natl Acad Sci U S A 112, 15504–15509 (2015).
Acknowledgements
This work is supported by the National Natural Science Foundation of China (31271315 and 91335201) and the National Special Program for Research of Transgenic Plant of China.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: Y.X. Performed the experiments: Z.Z., W.H., G.S., H.L., Y.H., X.Z. and T.L. Analyzed the data: Z.Z., W.H., G.S. and Y.X. Wrote the paper: Y.X., Z.Z. and W.H. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Z., Hu, W., Shen, G. et al. Alternative functions of Hd1 in repressing or promoting heading are determined by Ghd7 status under long-day conditions. Sci Rep 7, 5388 (2017). https://doi.org/10.1038/s41598-017-05873-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-05873-1
This article is cited by
-
Mapping QTLs for grain iron, zinc, and yield traits in advanced backcross inbred lines of Samba mahsuri (BPT5204)/Oryza rufipogon
Journal of Plant Biochemistry and Biotechnology (2024)
-
Genetic and functional mechanisms of yield-related genes in rice
Acta Physiologiae Plantarum (2024)
-
Genetic and signaling pathways of flowering regulation in rice (Oryza sativa L.)
Brazilian Journal of Botany (2023)
-
Small Auxin Up RNA 56 (SAUR56) regulates heading date in rice
Molecular Breeding (2023)
-
The transcription factor HBF1 directly activates expression of multiple flowering time repressors to delay rice flowering
aBIOTECH (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.