Abstract
Polysialic acid (polySia) is a homopolymeric saccharide that is associated with some neuroinvasive pathogens and is found on selective cell types in their eukaryotic host. The presence of a polySia capsule on these bacterial pathogens helps with resistance to phagocytosis, cationic microbial peptides and bactericidal antibody production. The biosynthesis of bacterial polySia is catalysed by a single polysialyltransferase (PST) transferring sialic acid from a nucleotide-activated donor to a lipid-linked acceptor oligosaccharide. Here we present the X-ray structure of the bacterial PST from Mannheimia haemolytica serotype A2, thereby defining the architecture of this class of enzymes representing the GT38 family. The structure reveals a prominent electropositive groove between the two Rossmann-like domains forming the GT-B fold that is suitable for binding of polySia chain products. Complex structures of PST with a sugar donor analogue and an acceptor mimetic combined with kinetic studies of PST active site mutants provide insight into the principles of substrate binding and catalysis. Our results are the basis for a molecular understanding of polySia biosynthesis in bacteria and might assist the production of polysialylated therapeutic reagents and the development of novel antibiotics.
Similar content being viewed by others
Introduction
Sialic acid (Sia) is a negatively charged monosaccharide that is widely found on eukaryotic cell surface glycoconjugates. These nine carbon sugar residues (N-acetylneuraminic acid being the predominant form) play essential roles in a multitude of biological processes such as cell-cell interaction, bacterial infection, and when inappropriately expressed, several forms of cancer1. Sia can be found on a handful of eukaryotic proteins as α-2,8-linked homo-polymeric structures of up to 400 residues called polysialic acid (polySia)2. Due to its exceptional physicochemical properties, polysialylation strongly promotes cell migration, neuronal plasticity, as well as tumour metastasis. In addition, altered levels of polySia have implications in development, schizophrenia and nerve repair2,3,4,5,6,7. In vertebrates, polysialylation is catalysed by two Golgi-residing polysialyltransferases (PSTs) of the ST8Sia family transferring Sia from the CMP-Neu5Ac donor substrate to the non-reducing termini of sialylated N- and O-linked glycans on glycoproteins8,9,10. The major acceptors of this highly regulated and protein-specific posttranslational modification are neural cell adhesion molecule (NCAM) and synaptic cell adhesion molecule (SynCAM 1) in the nervous system, as well as neuropilin 2 on dendritic cells11.
Apart from their specialized presence in vertebrates, polySia modifications also exist in prokaryotes. So far four genera of neuroinvasive Gram-negative bacteria including Neisseria meningitidis have been identified to synthesize capsular polysaccharides consisting of polySia that resemble the structures found on eukaryotic glycoproteins. The molecular mimicry of these bacterial polySia capsules represents an elegant strategy to evade the host’s immune recognition since they are not considered as foreign12, 13. In addition, they confer a physical barrier protecting the pathogen from killing by the complement system14. Bacterial polySia capsules exist in three different flavours: Escherichia coli K1, N. meningitidis serotype B, Moraxella nonliquefaciens, and Mannheimia haemolytica A2 synthesize α-2,8-linked polySia15,16,17,18, whereas N. meningitidis serotype C produces a α-2,9-linked polymer and E. coli K92 produces polymers with alternating α-2,8 and α-2,9 linkages19,20,21. Unlike those in vertebrates, bacterial polySia structures are covalently linked to the lipid carrier lyso-phosphatidyl glycerol22 and their biosynthesis follows a general concept conserved for all type 2 capsular polysaccharides. The assembly is initiated at the cytoplasmic side of the plasma membrane by the formation of a β-Kdo linker composed of two to nine Kdo monomers, catalysed by the enzymes KpsS and KpsC (E. coli nomenclature)23, 24. Sialic acid priming of the glycan has been proposed to involve the putative sialyltransferase NeuE, but several in vitro studies have suggested the existence of an additional enzyme to synthesize a di-sialylated structure required for initiation of polysialylation25,26,27. In the central step of the biosynthesis, the formation of the linear α-2,8-linked homo-polymer is catalysed by PST (NeuS) utilizing the nucleotide activated donor substrate CMP-Neu5Ac28, 29. The completed, lipid-linked polysaccharide is translocated to the cell surface by the trans-envelope complex KpsDEMT containing the ABC-transporter KpsMT as the driving force30,31,32 (Fig. 1a).
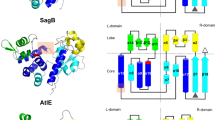
PST catalysed biosynthesis of polysialic acid and structure of MhPST. (a) The biosynthesis of capsular polysialic acid (polySia) is initiated at the cytoplasmic side of the plasma membrane by the transfer of a Kdo linker onto the lipid carrier lyso-phosphatidyl glycerol catalysed by KpsS and KpsC. Extension to polySia requires sialic acid priming involving the action of NeuE, before PST can catalyse the assembly of homopolymeric, α-2,8-linked polySia. The completed, lipid-bound structure is translocated to the cell surface in an ATP-dependent manner by an export complex composed of KpsDEMT. (b) In vitro polysialylation activity of MhPST using a soluble BDP-Sia2Lac acceptor substrate was analysed by TLC. Polysialylation results in suppressed migration of the acceptor (A complete scan of the TLC plate is shown in Supplementary Figure S7). (c) Ribbon diagrams of the Δ20MhPST apo structure showing the GT-B fold. In the N-terminal Rossmann fold, helices are coloured in marine and β-sheets are coloured in purple, whereas in the C-terminal Rossmann fold, helices are coloured in light blue and β-sheets are coloured in pink. The presumed position of the N-terminal membrane anchor is indicated.
Despite the biochemical characterization of bacterial PSTs26, 33, 34, the reaction mechanism of this fundamental enzyme is currently insufficiently understood, mainly due to the lack of structural information. Recent structures of the human ST8SiaIII enzyme provided important insights into polysialylation in mammals35, but bacterial PSTs belong to a completely distinct family of glycosyltransferases (Carbohydrate Active Enzyme (CAZy) designated family GT38 distinct from GT29 for ST8SiaIII)36, which is not specific for particular acceptor proteins, but assembles polySia on a lipid-linked oligosaccharide10, 37.
To understand the unique structural features and the reaction mechanism of bacterial PSTs at a molecular level, we have performed a crystallographic and biochemical characterization of the PST enzyme from M. haemolytica serotype A2. We have determined the X-ray structure of the apo enzyme, as well as the structures of complexes with CDP and the pentasaccharide heparin-mimetic fondaparinux, respectively. In combination with detailed kinetic analyses of active site mutants, this work provides essential insights into the structural architecture, as well as into the molecular principles of substrate binding and catalysis of polysialylation in bacteria.
Results
Structure of M. haemolytica PST
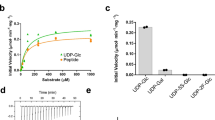
Bacterial PSTs are membrane-associated enzymes acting at the cytoplasmic side of the inner membrane. Since it has been shown that N-terminal truncation of M. haemolytica PST (MhPST) lacking the putative membrane anchor segment results in soluble enzyme34, we expressed the N-terminal truncation Δ20MhPST within the cytoplasm of E. coli. The purified enzyme was able to synthesize a polysialylated product from CMP–Neu5Ac donor and BODIPY-di-sialyllactose (BDP-Sia2Lac) acceptor (Fig. 1b), and our kinetic analysis revealed a Km of 0.6 mM for CMP–Neu5Ac (Table 1) consistent with previous reports33, 34, 38.
To obtain suitably ordered crystals, we introduced two surface entropy reduction mutations (K68A, K69A). We crystallized the resulting Δ20MhPST construct using the microbatch method and solved the X-ray structure of the apo-enzyme to 2.8 Å resolution (Table 2). Co-crystallization of Δ20MhPST with the acceptor substrate analogue di-sialyl-N-acetyllactosamine-6-sulfate (Sia2LacNAc6S) resulted in better-ordered crystals diffracting to 2.2 Å (Table 2), but no clear density for the ligand was observed. This might be a direct consequence of the weak binding of di-sialylated acceptor substrates, as we determined a Km of 2.26 mM for the Sia2Lac acceptor (Table 3). A comparison of the apo-structure with the structure determined in the presence of Sia2LacNAc6S revealed no significant difference in the overall conformation or in the orientation of individual side chains (r.m.s.d. = 0.27 Å over 5695 atoms). However, the Sia2LacNAc6S structure showed unambiguous electron density for two regions that were poorly resolved in the apo-structure (residues M20 to K32 and E231 to K251, neither involved in substrate binding or catalysis, see below). Therefore, we have used this latter more complete structure in our current analysis.
We observed a non-crystallographic dimer of MhPST in the asymmetric unit of all determined crystal structures, where the N-terminal loops intertwine with the opposite monomer providing substantial crystal contacts (Supplementary Fig. S1a). In solution MhPST is monomeric as suggested by size exclusion chromatography (data not shown). Superimposition of the two monomers reveals a slight difference in relative orientation between the N- and C-terminal domains caused by structural flexibility in a hinge region connecting the two domains (r.m.s.d. = 1.91 Å over 3052 atoms). However, the individual N- and C-terminal domains superimpose well with r.m.s.d. values of 0.20 Å and 0.17 Å over 1378 atoms and 1085 atoms, respectively (Supplementary Fig. S1b). The MhPST monomer is composed of two non-identical Rossmann-like α/β/α domains structurally separated by the described hinge region (F227 to N236) (Fig. 1c, Supplementary Fig. S1b). The core of the N-terminal domain is formed by seven parallel β-sheets that are flanked by four α-helices on one side and five α-helices on the other side. The slightly shorter C-terminal Rossmann domain is made up of a six-stranded parallel β-sheet surrounded by three and five α-helices, respectively and contains the nucleotide-binding site (see below) (Supplementary Fig. S2). The observed architecture of MhPST reflects a GT-B fold commonly found for metal independent glycosyltransferases39. The N-terminal Rossmann domain of MhPST is preceded by a 12 amino acid long tail of extended conformation (Supplementary Fig. S1b), which connects the enzyme to the putative membrane anchor (absent in our truncated Δ20MhPST construct), thereby providing sufficient distance to the plasma membrane (Fig. 1c).
A DALI search with the monomeric MhPST structure finds only proteins with low structural similarity (r.m.s.d. values greater than 3.7 Å), including various GTs and non-GT enzymes (e.g. UDP-GlcNAc 2-epimerases)40. Unlike most of these structures, MhPST lacks the GT-B typical C-terminal extension that interacts with the N-terminal domain. Searching the PDB with the individual Rossmann-like domains, resulted in slightly closer matches (r.m.s.d. values around 3.0 Å) and identified, amongst others, bacterial mono-sialyltransferases of the GT80 family, particularly for the search with the C-terminal domain.
Notably MhPST shows no structural similarity to mammalian PSTs of the GT29 family, as the structure of the human ST8SiaIII enzyme exhibits a GT-A fold consisting of a single Rossmann-like domain35.
Nucleotide activated sugar donor binding site
Attempts to obtain a structure with CMP-3FNeu5Ac, a non-hydrolyzable nucleotide activated sugar donor substrate derivative, were not successful. However, we were able to determine the structure of a binary complex with CDP at 3.0 Å resolution (Table 2). Clear electron density for the nucleotide diphosphate was observed, which allowed us to unambiguously model the CDP molecule in a cavity accessible from the cleft between the two Rossmann domains (Fig. 2a).
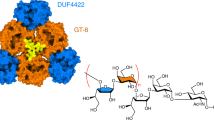
Structures of substrate-bound complexes of MhPST. (a) Experimental Fo – Fc omit electron density map for CDP (yellow sticks) bound to MhPST (ribbon diagram of monomer A) is shown as a blue mesh contoured at 3.0 σ. Structure factor calculation was performed prior to CDP docking. (b) Nucleotide donor binding site of MhPST (monomer A) with residue side chains interacting with bound CDP (yellow sticks) shown in sticks and labelled. Potential hydrogen bonds are indicated as black dashes. (c) Superimposition of donor binding sites of the apo-structure (coloured in cyan) and of the CDP-bound structure (coloured in wheat, pink, and light blue). The shift of the side chain of H291 by 1.3 Å is shown. (a–c) The C-terminal domain is omitted for clarity. (d) Experimental Fo – Fc omit electron density map for fondaparinux (green sticks) bound to MhPST (ribbon diagram of monomer A) is shown as a blue mesh contoured at 3.0 σ. Structure factor calculation was performed prior to ligand docking. (e) Acceptor substrate binding site of MhPST (monomer A) with residue side chains interacting with bound fondaparinux (green sticks) shown in sticks and labelled. The five monosaccharide units of fondaparinux are labelled and potential hydrogen bonds are indicated as black dashes. (a,d) Electron density improved for all ligands upon refinement.
Although CDP binds at the interface of the two domains, it only makes extensive interactions with the C-terminal Rossmann domain (Supplementary Fig. S3), where the pyrimidine ring is inserted into a hydrophobic pocket formed by C256 and P292. The amine group (N4) of the cytosine base forms hydrogen bonds with the backbone carbonyl oxygen of K289 and A257, respectively. The side-chain amine of K289 also provides a hydrogen bond to the non-protonated N3 of the pyrimidine ring (Fig. 2b). This hydrogen-bonding network defines the specificity filter for CMP-activated sugar donor substrates, as none of these interactions would be possible for uracil or thymine bases. Furthermore, the binding pocket does not provide enough space to accommodate a purine base and does not facilitate any unspecific aromatic stacking interactions. However, the hydrogen bonding of the keto group (O2) of the cytosine base to the backbone amide of A322 and to the side chain amine of K289 could also occur with other nucleotides (Fig. 2b). All described interactions between MhPST and cytosine highly resemble the situation observed for Pasteurella multocida mono-sialyltransferase PmST1 and related enzymes of the GT80 family41,42,43.
Binding of the ribose-phosphate moiety of CDP by MhPST also exhibits a common interaction profile conserved amongst GT80 enzymes and lipooligosaccharide sialyltransferase from N. meningitidis representing the GT52 family (Fig. 2b)41, 44. In MhPST, the O2′ and O3′ hydroxyl groups of the ribose are in close contact with the carboxyl group of E323 and form a strong bidentate hydrogen bond pair. The α-phosphate of CDP also makes extensive interactions with the C-terminal Rossmann domain. Phosphate oxygen O1 forms a hydrogen bond to the imidazole ring of H291, whereas O2 is hydrogen bonded by the hydroxyl group of S339. Additionally, the oxygen of the phosphate-phosphate bond connecting the α- and the β-phosphate forms a hydrogen bond to the hydroxyl group of T340. As the naturally occurring leaving group after the glycosyl transfer reaction is CMP and not CDP, T340 probably binds to the O3 oxygen of the α-phosphate in the native reaction. For the β-phosphate, the electron density is less well resolved (Fig. 2a) and no interactions with the protein are observed. This is also reflected by its altered conformation in the two MhPST monomers of the asymmetric unit. While the two CMP moieties of CDP superimpose very well (r.m.s.d. = 0.29 Å), the β-phosphate in monomer B is flipped by 140° as compared to the orientation in monomer A (which is presented in Fig. 2a–c) and points towards H291.
All residues forming specific side-chain interactions with CDP are conserved among bacterial PSTs (Supplementary Fig. S4), suggesting that the donor nucleotide-binding site exhibits identical features in other GT38 enzymes. H291 is not only invariant in bacterial PSTs, but is part of the HP-motif generally conserved in sialyltransferases, as well as in β-Kdo transferases24, 33, 45. Structure-function studies on PmST1 proposed that H311 (H291 in MhPST) is involved in stabilizing the negatively charged CMP leaving group46. Indeed, mutation H291A in MhPST resulted in a seven-fold increased Km for the CMP-Neu5Ac donor substrate and a 38-fold reduced kcat (280-fold reduced catalytic efficiency, kcat/Km), suggesting a significant role in catalysis (Table 1). As expected, the Km for the acceptor substrate Sia2Lac is only marginally affected by the H291A mutation (Table 3).
To our surprise, CDP binding did not result in major conformational changes in the MhPST structure (r.m.s.d. = 0.41 Å over 2802 atoms), and the most significant difference is the movement of the H291 imidazole side chain towards the bound CDP ligand by 1.2 Å (distance between the Nε2 atoms of H291 in the two different conformations; Fig. 2b). This is in strong contrast to other GT-B enzymes, where nucleotide binding usually causes large domain movements in creation of the donor sugar-binding site. In PmST1 for example, CMP binding results in a 23° rotation of the N-terminal Rossmann domain towards the C-terminal domain, thereby closing the catalytic cleft (Supplementary Fig. S6a)41. However, in a recent study on β-Kdo transferase of GT-family 99 that is related to sialyltransferases, CMP binding also did not induce any substantial domain movements24, suggesting that nucleotide binding might not always be sufficient to trigger these rearrangements.
Complex structure with fondaparinux reveals acceptor-binding site
As discussed above, our attempts to crystallize MhPST in the presence of the di-sialylated acceptor substrate Sia2LacNAc6S did not resolve the ligand in the crystal structure. However, we could obtain a complex with fondaparinux, a synthetic polyanionic (heparin) pentasaccharide clinically used as an anticoagulant (Table 2, Supplementary Fig. S5b). Clear electron density was observed only in monomer A, which allowed us to unambiguously model the ligand into electron density lining the deep catalytic cleft between the two Rossmann domains (Fig. 2d). Based on the repeating polyanionic functional groups, we propose fondaparinux maps on to MhPST in a complementary electropositive path similar to that required of the native polySia substrate. We note, however, that fondaparinux carries an additional five anionic functional groups compared to a polySia pentamer and would also be expected to span a comparatively shorter distance than a corresponding nine-carbon sugar sialic acid pentamer would do (Supplementary Fig. S5).
Fondaparinux is bound to MhPST along an electropositive groove (Fig. 3b) by a series of interactions, whereby both carboxyl groups and seven out of the eight sulfate groups are at least partly liganded. In contrast to CDP binding which only involves the C-terminal domain, both Rossmann domains contribute to the fondaparinux binding site (Fig. 2e). Notably, the 6′ carboxyl group of the second saccharide (IdoA2S) is saturated by interactions with the guanidinium group of R259 and the terminal amine of K293. R259 further forms a salt bridge to the 2′ sulfoamino group of the reducing end saccharide (GlcNS6S-OMe). The amine group of K75 on the other side binds to the 2′ sulfoamino group of the non-reducing end saccharide (GlcNS6S), whereas the side-chain of R80 interacts with the 6′ carboxyl group of the fourth saccharide (GlcA), as well as with the 2′ sulfoamino group and the 3′ sulfo group of the third sugar (GlcNS3,6 S) (Fig. 2e). Residues K75, R259, and K293 are strictly conserved in bacterial PST enzymes (Supplementary Fig. S4), suggesting that they also play an important role in binding the natural polySia acceptor substrate.
The MhPST acceptor substrate binding groove at the interface of the N-terminal and the C-terminal domain. (a) Surface representation of MhPST (monomer A in wheat) with bound acceptor ligand analogue fondaparinux shown in green sticks. The surface of residue K293 is coloured in blue and is labelled. (b) Electrostatic surface potential of fondaparinux bound MhPST structure. (c) Electrostatic surface potential of CMP bound structure of mono-sialyltransferase PmST1 (PDB: 3s44). (b,c) Surface potentials were calculated with the Adaptive Poisson Boltzmann Server69, 70 with a PARSE force field, with linear interpolation of colours at intermediate potentials (blue, 4 kT/e; red, −4 kT/e; white, 0 kT/e; probe radius, 1.4 Å).
A surface representation of MhPST further illustrates the central role of K293 in acceptor substrate binding. Its side chain reaches deeply into the catalytic cleft, thereby pinning the bound fondaparinux against the N-terminal domain and locking it in the resulting cavity (Fig. 3a). K293 contacts the pentasaccharide between the second and the third sugar residue, which might explain the preference for tri-sialylated over di-sialylated acceptor substrates (Table 3)34. These observations were corroborated by mutagenesis studies, where mutant K293A had a drastically reduced catalytic efficiency, while the Km value for the CMP-Neu5Ac donor substrate was not negatively affected (Tables 1, 3).
Superimposition of the ligand-free and the fondaparinux-bound structures showed that fondaparinux binding does not cause a movement between the N- and the C-terminal domains (r.m.s.d. = 0.66 Å over 2889 atoms). The observed small conformational rearrangements (<2.0 Å) are primarily located in the catalytic cleft and are required to accommodate the fondaparinux ligand.
Catalytic mechanism of bacterial PST
To generate a modelled composite of a ternary complex for mechanistic analysis, we superimposed the CDP-bound and the fondaparinux-bound structures (Fig. 4a). We propose that the resulting complex resembles a pseudo product complex, in which the reducing end sugar of fondaparinux is in proximity to the α-phosphate of the CDP. We note, however, that the native polySia acceptor binds in the opposite direction with the non-reducing end sugar oriented close to the sugar nucleotide donor (Supplementary Fig. S5). The functional analysis of residue R259, which interacts with the 2′ sulfoamino group of the reducing end saccharide of fondaparinux, supports this hypothesis. Mutant R259A showed a 3-fold increased Km for the CMP-Neu5Ac donor, while the Km for the Sia2Lac acceptor was not affected (Tables 1, 3). Therefore, R259 could, under natural reaction conditions, potentially interact with the sialic acid moiety of the CMP-Neu5Ac donor, which gets transferred to the non-reducing end of the growing polySia chain.
MhPST active site and identification of the catalytic base. (a) Modelled ternary complex by superimposition of the CDP-bound structure and the fondaparinux-bound structure. MhPST (monomer A) is shown as a ribbon diagram with the same color-coding as in Fig. 1c, CDP is shown as yellow sticks, fondaparinux is shown as green sticks and the five monosaccharide units are indicated. Residues with a supposed role in catalysis are shown in sticks and are labelled. Potential hydrogen bonds between E153 and Q41 and Q44, respectively are indicated as black dashes. The location of the mutation K69A is indicated. (b) Superimposition of active site of MhPST (wheat ribbon diagram) and mono-sialyltransferase PmST1 in open conformation (PDB: 3s44, cyan ribbon diagram). The catalytic acid and base are shown in sticks and are labelled for both enzymes, and the distance between E153 and H291 is indicated for MhPST. (c) SN2-like reaction mechanism of MhPST. The catalytic base E153 (red box) abstracts a proton from the C8′ hydroxyl group of the sialic acid acceptor concerted with the nucleophilic attack on the anomeric C2′ carbon (orange circle) of the CMP sialic acid donor substrate, thereby generating an α-2,8 glycosidic linkage. The resulting negatively charged CMP leaving group is stabilized by H291 (blue box) assisted by S339 and T340 (grey boxes). R = α-2,8-linked oligosialyl.
Sialic acid transfer occurs with inversion of configuration (from the β-linked CMP-Neu5Ac donor to the α-2,8-linked polySia), and PST has been proposed to follow a SN2-like direct displacement mechanism45. While H291 could act as a catalytic acid to stabilize the nucleotide phosphate-leaving group, the catalytic base remained unknown. The analysis of the MhPST ternary complex structure did not reveal any residue in proximity to the active site that could serve this purpose. We therefore performed a comparison of our MhPST structure with structures of the PmST1 enzyme, where the catalytic base had been identified46. As described before, PmST1 undergoes large domain movements upon nucleotide donor substrate binding resulting in a closure of the catalytic site (Supplementary Fig. S6a). As a consequence, the distance between the catalytic base (D141) and residue H311 changes from 14.0 Å in the open conformation to 8.4 Å in the closed state (Supplementary Fig. S6b). Structural alignments between MhPST and PmST1 not only indicated that our MhPST structures resemble an open conformation, but also identified residue E153 as a potential catalytic base. E153 is located in an analogous loop to D141 in the PmST1 structure and is 13.1 Å away from residue H291 (Fig. 4b). Furthermore, E153 is part of a previously described D/E-D/E-G motif and is conserved in bacterial PSTs (Supplementary Fig. S4)33, 45. Kinetic analyses of MhPST mutants confirmed that E153 is indeed the catalytic base with an activity too low to assign a Km value to it. The neighbouring mutant E152A was a 300-fold worse catalyst, suggesting that this adjacent residue plays an important role in stabilizing the catalytic loop between strand β5 and helix α5a (Tables 1,3, Fig. 4a). We therefore propose a reaction mechanism for MhPST, in which the carboxyl group of E153 abstracts a proton from the C8′ hydroxyl group of the non-reducing end sialic acid of the acceptor substrate (Fig. 4c) in concert with attack on the anomeric C2′ carbon of the CMP-Neu5Ac donor substrate, forming an α-2,8 glycosidic linkage between the two sialic acid residues. H291 acts as a catalytic acid to stabilize the negative charge at the terminal phosphate of the CMP leaving group. Residues S339 and T340 might further assist in the coordination of the phosphate. Residues Q41 and Q44 which are in proximity to E153 might play a role in activating the catalytic base as reflected by the reduced catalytic performance of mutants Q41A and Q44A (Tables 1, 3, Fig. 4a,c).
Discussion
The structural characterization of MhPST in the presence of the donor substrate analogue CDP and the acceptor substrate mimetic fondaparinux revealed the molecular concepts of substrate binding and catalysis. Residues involved in these interactions are generally conserved among PSTs from different species, despite an overall sequence identity between 28% and 31% (Supplementary Fig. S4) making MhPST a highly valuable model to study the catalytic mechanism of bacterial PSTs. Notably, substrate binding did not cause any significant rotation of the N- and C-terminal Rossmann domains that is frequently observed in other GT-B enzymes, and which leads to a closure of the catalytic cleft. A structural comparison with the mono-sialyltransferase PmST1 revealed that MhPST adopts an open conformation (Fig. 4b). We suggest this may be a reflection of the lack of complete donor substrate CMP-Neu5Ac bound (see below), or the need to accommodate the larger, polymeric PST acceptor substrate. Even though binding of the substrate mimetic fondaparinux did not provoke domain closure in MhPST (perhaps because it is too short or because the precise alignment of sugar units is incompatible with triggering closure), a rotation of the N-terminal domain appears to be necessary to move the proposed catalytic base E153 up to the active site and to position its carboxyl group appropriately to activate the C8′ hydroxyl group of the sialic acid acceptor for nucleophilic attack (Fig. 3c). Therefore, we postulate the existence of MhPST in a closed state during the catalytic cycle. What could be the trigger for such a conformational change? In the case of the mono-sialyltransferase PmST1, CMP binding induced an N-terminal domain movement, in which S143 located in helix α5a (and in proximity to the catalytic base D141) moves 5.2 Å towards the C-terminal domain to form a hydrogen bonding network with Y388 and the terminal phosphate of CMP. Furthermore, the side chain of Y388 is flipped by 180° and the corresponding helix α12b is shifted by 5.5 Å upon CMP binding41. MhPST contains the conserved residues T155 and H382 at corresponding positions that could emulate the function of S143 and Y388 in PmST1, respectively (Fig. 4a, Supplementary Fig. S4). As CDP binding did not facilitate these interactions in MhPST, it is important to note that bound CMP in the complex structure of PmST1 resulted from hydrolysis of the complete donor substrate CMP-Neu5Ac41, and it cannot be excluded that the domain shift occurred before donor hydrolysis. Therefore, binding of the complete sugar donor substrate (or its non-hydrolysable derivative CMP-3FNeu5Ac) might be required to induce the domain shift.
The open conformation of MhPST exhibits a deep cleft between the two Rossmann domains spanning across the entire front of the enzyme. This electropositive groove (~35 Å in diameter) is much more pronounced than in other glycosyltransferases bearing a GT-B fold (Fig. 3b,c) and is concordant with accommodating a polymeric and polyanionic sialic acid acceptor substrate. Interestingly, saturation transfer difference NMR spectroscopy studies on PST from N. meningitidis serotype B (NmBPST) postulated the existence of an extended acceptor-binding site that can accommodate at least six sia residues47. Binding of the acceptor mimetic fondaparinux to MhPST illustrates that the ligand is well aligned between the two domains already in the observed open conformation (Fig. 3a). The formation of the postulated closed enzyme state during catalysis would cause an even more snug fit of the acceptor substrate in the catalytic cleft. These acceptor-binding properties may suggest a processive mechanism of polymerization, in which the growing polySia chain is retained at the active site for addition of multiple sia monomers before product release. However, several in vitro studies using purified PST enzyme proposed a distributive mechanism, where polySia is released from the enzyme after each transfer reaction33, 38. In vitro studies on PST are generally performed on soluble enzyme variants and utilize soluble synthetic acceptor substrates resulting in a reduced local concentration of acceptor substrate, because PST as well as the lipid-linked polySia acceptor are naturally anchored in the inner membrane (Fig. 1a). Therefore, it is not surprising to observe a discrepancy between polySia polymer length in vivo and in vitro 26, 48, 49. A recent study on the polySia product profile of NmBPST proposed that chain elongation in vitro occurs in an abortive processive manner with frequent dissociation of the enzyme-acceptor complex50. Increasing acceptor length resulted in increased enzyme affinity suggesting a continuous binding site able to interact with a 20-mer polySia acceptor. Intriguingly, the authors identified residue K69 as a molecular switch controlling the mechanism of chain elongation and polySia size distribution. Mutations K69Q and K69D changed the chain elongation to a distributive mechanism, yielding reduced product dispersity even for short oligoSia acceptors and a direct interaction of residue 69 with the substrate was proposed50. K69 is also conserved in MhPST (Supplementary Fig. S4), but in order to obtain well-diffracting crystals, it was mutated to alanine. The distance between the methyl group of A69 and the second or third fondaparinux saccharide (IdoA2S or D-GlcNS3,6 S) is more than 10 Å suggesting that even a lysine residue at position 69 would require a domain closure to directly interact with the acceptor substrate (Fig. 4a). However, we cannot exclude that the presence of K69 in MhPST would result in increased acceptor binding. Additional sites of mutation in NmBPST that were found to influence polySia size distribution are not conserved among bacterial PSTs. Even though the effect of these other mutations seems to be specific for NmBPST, the surface potential of MhPST shows two highly electropositive areas at the front of the N-terminal Rossmann domain, which could provide additional interaction surfaces for an extended polySia chain (Fig. 3b). For comparison, the surface of the mono-sialyltransferase PmST1 is lacking a pronounced acceptor-binding groove and mainly shows positive values for the donor-binding site, concordantly with the preference for short uncharged acceptor substrates (Fig. 3c). Therefore, the two positively charged patches on the surface of MhPST could represent an extension of the acceptor-binding groove providing a large interaction interface with low site-specific binding but high avidity for the growing polySia chain. Such a model for acceptor binding would allow substrate translocation from one site to the next and would be in excellent agreement with the higher affinity for long acceptor oligomers observed for NmBPST50.
Strikingly, an analogous mechanism for substrate interaction has been brought forward for mammalian PSTs. The structure of the human ST8SiaIII enzyme also exhibits an extensive positively charged surface groove able to accommodate extended polySia acceptor substrates35. Apart from this conceptual similarity in polySia binding, mammalian and bacterial PSTs share no common features. The two enzymes exhibit completely different folds and none of the conserved motifs defining the active site of mammalian PSTs and of other eukaryotic mono-sialyltransferases of the GT29 family are found in bacterial MhPST belonging to the GT38 family35, 51,52,53. Instead, the molecular principles of substrate binding and catalysis of bacterial PSTs resemble enzymes of CAZy families GT52 and GT8033, 44, 46. Therefore, polySia biosynthesis is a prototype of convergent evolution where bacterial and mammalian enzymes follow different molecular routes to synthesize the identical α-2,8-linked polySia homopolymer. Since polySia biosynthesis is an essential virulence factor for the corresponding pathogens, bacterial PST might be an interesting target for the development of novel antibiotics. Due to the lack of structural similarity between the bacterial and mammalian PSTs shown for the first time here, our insights into MhPST provide encouragement regarding the ability to create bacterial PST-specific therapeutics.
The biosynthesis of polySia has also great potential for various medical applications, and both bacterial and mammalian PSTs represent potential candidates to produce polySia and polysialylated bioconjugates. The broad application of mammalian PSTs is currently limited by their high acceptor protein specificity2, 11, 54, whereas bacterial PSTs exhibit a more relaxed substrate specificity26. Different bacterial enzymes including MhPST have been successfully used to polysialylate a primed version of fetuin as well as different cell surface proteins including NCAM, the most prominent acceptor protein for mammalian PSTs34, 55. Furthermore, an elegant two-step enzymatic polysialylation strategy was applied to site-specifically modify alpha-1-antitrypsin resulting in improved pharmacokinetic properties56. These data illustrate the tremendous value of using bacterial PST enzymes for different therapeutic applications. Our structural characterization of the MhPST enzyme might therefore contribute to the development of specific and tailored polySia-conjugates.
Methods
Cloning and expression of Δ20MhPST
The polysialyltransferase gene from M. haemolytica A2 was cloned as a Δ20 N-terminal truncation into the pCW expression vector as previously described34. To obtain well-ordered crystals, two surface entropy reduction mutations (K68A, K69A) were introduced. This double mutant was referred to as wild-type enzyme and all further mutations in the active site that were used for kinetic studies were based on it. All mutations were introduced by site directed mutagenesis using the Quick Change method.
Δ20MhPST constructs were transformed into E. coli AD202 cells and a single clone was used to inoculate a preculture in LB media supplemented with 100 µg/mL ampicillin. The main culture of 2xYT media supplemented with 100 µg/mL ampicillin was inoculated to an OD600 of 0.05 and cells were grown at 37 °C and 200 rpm until an OD600 of 0.4 to 0.6 was reached. The culture was shifted to 20 °C and expression was induced by addition of 0.5 mM IPTG at an OD600 of 0.8. After incubation for 16 h at 20 °C and 200 rpm, cells were harvested by centrifugation and cell pellets were stored at −80 °C until use.
Purification and crystallization of Δ20MhPST
5 g of frozen cells were resuspended in 25 mL of buffer A consisting of 50 mM HEPES, pH 7.4; 150 mM NaCl; 5 mM β-mercaptoethanol and Complete Mini protease inhibitor cocktail (Roche). Cells were lysed by French Press (2 passes at 1,500 psi) and cell debris were removed by centrifugation at 48,000 × g for 30 min. The supernatant was passed through a filter with 0.45 μm pore size, before the sample was loaded onto a 5 mL Heparin HP column (GE Healthcare) equilibrated with buffer A. The column was washed with 5 column volumes of buffer A, followed by a second wash with 5 column volumes of 15% buffer B consisting of 50 mM HEPES, pH 7.4; 1.5 M NaCl and 5 mM β-mercaptoethanol. The protein was eluted in a linear gradient of 0–100% buffer B over 6 column volumes and 1 mL elution fractions were collected. Fractions containing MhPST were pooled and the buffer was immediately exchanged to buffer C consisting of 50 mM HEPES, pH 7.2 and 100 mM NaCl using a HiPrep desalting column (GE Healthcare). Protein purity was evaluated by SDS-PAGE, and the protein concentration was determined by absorption at 280 nm using an extinction coefficient of 37250 M−1cm−1. Protein monodispersity was analysed by analytical size exclusion chromatography using a Superdex 200 column (GE Healthcare, eluent buffer C). All steps were performed at 4 °C.
For crystallization, the protein was concentrated to 4–5 mg/mL using an Amicon centricon with a molecular weight cut-off of 30 kDa. Initial Δ20MhPST crystals were observed by vapour diffusion in sitting drops under conditions containing PEG 3350. Crystallization conditions were optimized to 17–24% PEG3350 (v/v); 140–250 mM Mg2SO4 and 100 mM MES, pH 7.2 in a 1:1 drop ratio using the microbatch method. Crystals appeared after 1–2 h at 23 °C and grew to final size within 2 days. Δ20MhPST was co-crystallized with 5 mM CDP donor analogue, or 2 mM Sia2LacNAc6S acceptor analogue in the same conditions. Apo crystals were soaked with 2 mM fondaparinux in 200 mM MgSO4; 100 mM MES, pH7.2 and 20% PEG3350 (v/v) for 2 h by transferring crystals to the soaking solution.
Data collection, phasing and refinement
Δ20MhPST crystals were cryoprotected in 200 mM MgSO4; 100 mM MES, pH7.2; 20% PEG3350 (v/v) and 30% glycerol (v/v) and flash frozen in liquid nitrogen. For phasing, 500 mM sodium bromide was added to the cryoprotectant. X-ray diffraction data were collected at both the Advanced Light Source (beamline 5.0.2) and the Canadian Light Source (CMCF beamlines 08ID-1 and 08B1–1). Data were integrated with XDS57 and scaled and merged with Aimless58. Phases for the bromide derivative crystals were solved by SAD using autoSHARP59 and further density modification was carried out using PHENIX60. The initial model was further built manually in Coot61 and refined with REFMAC62, 63 and PHENIX60. Co-crystal structures of Δ20MhPST in complex with Sia2LacNAc6S, CDP, or fondaparinux were solved by molecular replacement with the apo structure of Δ20MhPST using PHASER64. Processing and refinement statistics are summarized in Table 2. (The structures of Δ20MhPST apo, Δ20MhPST + Sia2LacNAc6S and Δ20MhPST + fondaparinux contain R132 as Ramachandran outlier.) The topology diagram was created with PDBsum65 and all structure images were created with PyMOL66.
In vitro polysialylation activity assay
The polysialylation activity of purified MhPST batches was tested in an in vitro reaction containing 0.5 mM BODIPY-diSiaLac acceptor; 10 mM CMP-Neu5Ac donor; 50 mM HEPES, pH 7.4; 10 mM MgCl2 and 0.2 mg/mL enzyme in a total volume of 10 μl. The reaction was incubated at 37 °C for 16 h, before 1 μl of the reaction was applied to a silica gel 60 TLC plate and the sample was separated with a developing phase containing ethylacetate:methanol:H2O:acetic acid in a ratio of 4:2:1:0.1. TLC plates were illuminated under UV light to visualize acceptor substrate conversion.
Chemo-enzymatic synthesis of disialyllactose (Sia2Lac) and trisialyllactose (Sia3Lac)
Both oligosaccharides (Sia2Lac/Sia3Lac) were chemo-enzymatically prepared using bifunctional Cst-II (from Campylobacter jejuni) as previously described67. In brief, 118 mg of lactose (Sigma Aldrich) and 210 mg of CMP-Neu5Ac (Roche) were dissolved in 9 mL of 100 mM Tris, pH 7.9 containing 20 mM MgCl2 at room temperature. The reaction was initiated by addition of 220 μL of Cst-II (stock: 12.5 mg/mL) and 10 µL alkaline phosphatase (Sigma Aldrich) in order to degrade liberated CMP. The reaction was incubated at 25 °C and another 210 mg of CMP-Neu5Ac were added after 2 h and 18 h, respectively. The pH was carefully monitored and adjusted with NaOH as needed. Reaction progress was monitored by TLC (EtOH:n-BuOH:pyridine:H2O:AcOH = 100:10:10:30:3) and the reaction was stopped with 3 mL ice-cold EtOH, centrifuged (10 min at 4,000 × g) and the supernatant was applied to an Amicon centricon with a molecular weight cut-off of 3 kDa to remove remaining protein. Sia2Lac and Sia3Lac were group separated using P-2 size exclusion chromatography (BioRad, eluent: 20% EtOH) and EtOH was removed in vacuo. After MacroQ anion exchange chromatography (GE Healthcare, flow rate: 2 mL/min, gradient: 0% to 100% of 0.4 M ammonium formate in 80 min), Sia2Lac and Sia3Lac fractions were pooled separately, the volume was reduced and both products were ran again on a P-2 column to exchange the buffer to 20% EtOH. Only highly pure fractions were pooled, lyophilized and products were confirmed by ESI mass spectrometry. Final yields were determined with 23% Sia2Lac and 28% Sia3Lac.
Kinetic studies
Kinetic parameters of MhPST mutants were determined with a coupled enzyme assay as described previously68. Briefly, assays were carried out in 96-well plates (half-area wells, Corning) with varying substrate concentrations of Sia2Lac/Sia3Lac (with constant concentration of CMP-Neu5Ac), or CMP-Neu5Ac (with constant concentration of colominic acid as acceptor). Each well contained 100 μL 50 mM HEPES, pH 7.0; 50 mM KCl; 20 mM MgCl2; 1 mg/mL BSA; 2 mM ATP; 2 mM phosphoenolpyruvate; 1 mM NADH; 16 U/mL pyruvate kinase; 29 U/mL lactate dehydrogenase; 0.07 U/mL nucleoside monophosphate kinase (Roche) and appropriate concentration of enzyme mutants. Prior to the addition of MhPST, the assay mixture was incubated at 37 °C until a stable baseline in absorbance at 340 nm (NADH) was achieved. Once MhPST enzyme was added, the rate of NADH consumption was determined by measuring the continuous decrease in absorbance at 340 nm. Rates were converted by using a NADH standard curve in order to calculate kcat (s−1). An extinction coefficient for NADH of 6,300 M−1cm−1 was used for calculations. Since two equivalents of NADH were released per equivalent of CMP-Neu5Ac consumed, the rate of transfer was determined as half the rate of NADH consumption.
Accession codes
Atomic coordinates and structure factors for Δ20MhPST apo, Δ20MhPST + CDP, Δ20MhPST + Sia2LacNAc6S and Δ20MhPST + fondaparinux have been deposited in the Protein Data Bank under PDB codes 5WC8, 5WCN, 5WC6 and 5WD7, respectively.
References
Angata, T. & Varki, A. Chemical diversity in the sialic acids and related alpha-keto acids: an evolutionary perspective. Chem Rev 102, 439–469 (2002).
Sato, C. & Kitajima, K. Disialic, oligosialic and polysialic acids: distribution, functions and related disease. J Biochem 154, 115–136, doi:10.1093/jb/mvt057 (2013).
Schnaar, R. L., Gerardy-Schahn, R. & Hildebrandt, H. Sialic acids in the brain: gangliosides and polysialic acid in nervous system development, stability, disease, and regeneration. Physiol Rev 94, 461–518, doi:10.1152/physrev.00033.2013 (2014).
Doherty, P., Cohen, J. & Walsh, F. S. Neurite outgrowth in response to transfected N-CAM changes during development and is modulated by polysialic acid. Neuron 5, 209–219 (1990).
Rutishauser, U. Polysialic acid in the plasticity of the developing and adult vertebrate nervous system. Nat Rev Neurosci 9, 26–35, doi:10.1038/nrn2285 (2008).
Falconer, R. A., Errington, R. J., Shnyder, S. D., Smith, P. J. & Patterson, L. H. Polysialyltransferase: a new target in metastatic cancer. Curr Cancer Drug Targets 12, 925–939 (2012).
Isomura, R., Kitajima, K. & Sato, C. Structural and functional impairments of polysialic acid by a mutated polysialyltransferase found in schizophrenia. J Biol Chem 286, 21535–21545, doi:10.1074/jbc.M111.221143 (2011).
Eckhardt, M. et al. Molecular characterization of eukaryotic polysialyltransferase-1. Nature 373, 715–718, doi:10.1038/373715a0 (1995).
Angata, K. & Fukuda, M. Polysialyltransferases: major players in polysialic acid synthesis on the neural cell adhesion molecule. Biochimie 85, 195–206 (2003).
Angata, K. et al. Differential biosynthesis of polysialic acid on neural cell adhesion molecule (NCAM) and oligosaccharide acceptors by three distinct alpha 2,8-sialyltransferases, ST8Sia IV (PST), ST8Sia II (STX), and ST8Sia III. J Biol Chem 275, 18594–18601, doi:10.1074/jbc.M910204199 (2000).
Mühlenhoff, M., Rollenhagen, M., Werneburg, S., Gerardy-Schahn, R. & Hildebrandt, H. Polysialic acid: versatile modification of NCAM, SynCAM 1 and neuropilin-2. Neurochem Res 38, 1134–1143, doi:10.1007/s11064-013-0979-2 (2013).
Mazmanian, S. K. & Kasper, D. L. The love-hate relationship between bacterial polysaccharides and the host immune system. Nat Rev Immunol 6, 849–858, doi:10.1038/nri1956 (2006).
Troy, F. A. Polysialylation: from bacteria to brains. Glycobiology 2, 5–23 (1992).
Vogel, U., Hammerschmidt, S. & Frosch, M. Sialic acids of both the capsule and the sialylated lipooligosaccharide of Neisseria meningitis serogroup B are prerequisites for virulence of meningococci in the infant rat. Med Microbiol Immunol 185, 81–87 (1996).
Adlam, C., Knights, J. M., Mugridge, A., Williams, J. M. & Lindon, J. C. Production of colominic acid by Pasteurella haemolytica serotype A2 organisms. FEMS Microbiology Letters 42, 23–25 (1987).
Devi, S. J. et al. Identity between polysaccharide antigens of Moraxella nonliquefaciens, group B Neisseria meningitidis, and Escherichia coli K1 (non-O acetylated). Infect Immun 59, 732–736 (1991).
Puente-Polledo, L., Reglero, A., González-Clemente, C., Rodríguez-Aparicio, L. B. & Ferrero, M. A. Biochemical conditions for the production of polysialic acid by Pasteurella haemolytica A2. Glycoconj J 15, 855–861 (1998).
Silver, R. P. et al. Molecular cloning of the K1 capsular polysaccharide genes of E. coli. Nature 289, 696–698 (1981).
Glode, M. P. et al. Cross-antigenicity and immunogenicity between capsular polysaccharides of group C Neisseria meningitidis and of Escherichia coli K92. J Infect Dis 135, 94–104 (1977).
Lifely, M. R., Nowicka, U. T. & Moreno, C. Analysis of the chain length of oligomers and polymers of sialic acid isolated from Neisseria meningitidis group B and C and Escherichia coli K1 and K92. Carbohydr Res 156, 123–135 (1986).
Devi, S. J., Robbins, J. B. & Schneerson, R. Antibodies to poly[(2,8)-alpha-N-acetylneuraminic acid] and poly[(2,9)-alpha-N-acetylneuraminic acid] are elicited by immunization of mice with Escherichia coli K92 conjugates: potential vaccines for groups B and C meningococci and E. coli K1. Proc Natl Acad Sci USA 88, 7175–7179 (1991).
Willis, L. M. et al. Conserved glycolipid termini in capsular polysaccharides synthesized by ATP-binding cassette transporter-dependent pathways in Gram-negative pathogens. Proc Natl Acad Sci USA 110, 7868–7873, doi:10.1073/pnas.1222317110 (2013).
Willis, L. M. & Whitfield, C. KpsC and KpsS are retaining 3-deoxy-D-manno-oct-2-ulosonic acid (Kdo) transferases involved in synthesis of bacterial capsules. Proc Natl Acad Sci USA 110, 20753–20758, doi:10.1073/pnas.1312637110 (2013).
Ovchinnikova, O. G. et al. Bacterial β-Kdo glycosyltransferases represent a new glycosyltransferase family (GT99). Proc Natl Acad Sci USA 113, E3120–3129, doi:10.1073/pnas.1603146113 (2016).
Andreishcheva, E. N. & Vann, W. F. Gene products required for de novo synthesis of polysialic acid in Escherichia coli K1. J Bacteriol 188, 1786–1797, doi:10.1128/JB.188.5.1786-1797.2006 (2006).
Willis, L. M., Gilbert, M., Karwaski, M. F., Blanchard, M. C. & Wakarchuk, W. W. Characterization of the alpha-2,8-polysialyltransferase from Neisseria meningitidis with synthetic acceptors, and the development of a self-priming polysialyltransferase fusion enzyme. Glycobiology 18, 177–186, doi:10.1093/glycob/cwm126 (2008).
Hobb, R. I., Tzeng, Y. L., Choudhury, B. P., Carlson, R. W. & Stephens, D. S. Requirement of NMB0065 for connecting assembly and export of sialic acid capsular polysaccharides in Neisseria meningitidis. Microbes Infect 12, 476–487, doi:10.1016/j.micinf.2010.02.009 (2010).
Steenbergen, S. M. & Vimr, E. R. Mechanism of polysialic acid chain elongation in Escherichia coli K1. Mol Microbiol 4, 603–611 (1990).
Steenbergen, S. M., Wrona, T. J. & Vimr, E. R. Functional analysis of the sialyltransferase complexes in Escherichia coli K1 and K92. J Bacteriol 174, 1099–1108 (1992).
Cuthbertson, L., Kimber, M. S. & Whitfield, C. Substrate binding by a bacterial ABC transporter involved in polysaccharide export. Proc Natl Acad Sci USA 104, 19529–19534, doi:10.1073/pnas.0705709104 (2007).
Cuthbertson, L., Mainprize, I. L., Naismith, J. H. & Whitfield, C. Pivotal roles of the outer membrane polysaccharide export and polysaccharide copolymerase protein families in export of extracellular polysaccharides in gram-negative bacteria. Microbiol Mol Biol Rev 73, 155–177, doi:10.1128/MMBR.00024-08 (2009).
Cuthbertson, L., Kos, V. & Whitfield, C. ABC transporters involved in export of cell surface glycoconjugates. Microbiol Mol Biol Rev 74, 341–362, doi:10.1128/MMBR.00009-10 (2010).
Freiberger, F. et al. Biochemical characterization of a Neisseria meningitidis polysialyltransferase reveals novel functional motifs in bacterial sialyltransferases. Mol Microbiol 65, 1258–1275, doi:10.1111/j.1365-2958.2007.05862.x (2007).
Lindhout, T., Bainbridge, C. R., Costain, W. J., Gilbert, M. & Wakarchuk, W. W. Biochemical characterization of a polysialyltransferase from Mannheimia haemolytica A2 and comparison to other bacterial polysialyltransferases. PLoS One 8, e69888, doi:10.1371/journal.pone.0069888 (2013).
Volkers, G. et al. Structure of human ST8SiaIII sialyltransferase provides insight into cell-surface polysialylation. Nat Struct Mol Biol 22, 627–635, doi:10.1038/nsmb.3060 (2015).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42, D490–495, doi:10.1093/nar/gkt1178 (2014).
Foley, D. A., Swartzentruber, K. G. & Colley, K. J. Identification of sequences in the polysialyltransferases ST8Sia II and ST8Sia IV that are required for the protein-specific polysialylation of the neural cell adhesion molecule, NCAM. J Biol Chem 284, 15505–15516, doi:10.1074/jbc.M809696200 (2009).
Peterson, D. C., Arakere, G., Vionnet, J., McCarthy, P. C. & Vann, W. F. Characterization and acceptor preference of a soluble meningococcal group C polysialyltransferase. J Bacteriol 193, 1576–1582, doi:10.1128/JB.00924-10 (2011).
Lairson, L. L., Henrissat, B., Davies, G. J. & Withers, S. G. Glycosyltransferases: structures, functions, and mechanisms. Annu Rev Biochem 77, 521–555, doi:10.1146/annurev.biochem.76.061005.092322 (2008).
Holm, L. & Rosenström, P. Dali server: conservation mapping in 3D. Nucleic Acids Res 38, W545–549, doi:10.1093/nar/gkq366 (2010).
Ni, L. et al. Cytidine 5′-monophosphate (CMP)-induced structural changes in a multifunctional sialyltransferase from Pasteurella multocida. Biochemistry 45, 2139–2148, doi:10.1021/bi0524013 (2006).
Huynh, N. et al. Crystal structures of sialyltransferase from Photobacterium damselae. FEBS Lett 588, 4720–4729, doi:10.1016/j.febslet.2014.11.003 (2014).
Iwatani, T. et al. Crystal structure of alpha/beta-galactoside alpha2,3-sialyltransferase from a luminous marine bacterium, Photobacterium phosphoreum. FEBS Lett 583, 2083–2087, doi:10.1016/j.febslet.2009.05.032 (2009).
Lin, L. Y. et al. Structure and mechanism of the lipooligosaccharide sialyltransferase from Neisseria meningitidis. J Biol Chem 286, 37237–37248, doi:10.1074/jbc.M111.249920 (2011).
Audry, M. et al. Current trends in the structure-activity relationships of sialyltransferases. Glycobiology 21, 716–726, doi:10.1093/glycob/cwq189 (2011).
Ni, L. et al. Crystal structures of Pasteurella multocida sialyltransferase complexes with acceptor and donor analogues reveal substrate binding sites and catalytic mechanism. Biochemistry 46, 6288–6298, doi:10.1021/bi700346w (2007).
Freiberger, F. et al. Defining a substrate-binding model of a polysialyltransferase. Chembiochem 14, 1949–1953, doi:10.1002/cbic.201300367 (2013).
Steenbergen, S. M. & Vimr, E. R. Functional relationships of the sialyltransferases involved in expression of the polysialic acid capsules of Escherichia coli K1 and K92 and Neisseria meningitidis groups B or C. J Biol Chem 278, 15349–15359, doi:10.1074/jbc.M208837200 (2003).
Vionnet, J. & Vann, W. F. Successive glycosyltransfer of sialic acid by Escherichia coli K92 polysialyltransferase in elongation of oligosialic acceptors. Glycobiology 17, 735–743, doi:10.1093/glycob/cwm032 (2007).
Keys, T. G. et al. Engineering the product profile of a polysialyltransferase. Nat Chem Biol 10, 437–442, doi:10.1038/nchembio.1501 (2014).
Rao, F. V. et al. Structural insight into mammalian sialyltransferases. Nat Struct Mol Biol 16, 1186–1188, doi:10.1038/nsmb.1685 (2009).
Datta, A. K. & Paulson, J. C. The sialyltransferase “sialylmotif” participates in binding the donor substrate CMP-NeuAc. J Biol Chem 270, 1497–1500 (1995).
Wen, D. X. et al. Primary structure of Gal beta 1,3(4)GlcNAc alpha 2,3-sialyltransferase determined by mass spectrometry sequence analysis and molecular cloning. Evidence for a protein motif in the sialyltransferase gene family. J Biol Chem 267, 21011–21019 (1992).
Hildebrandt, H., Mühlenhoff, M. & Gerardy-Schahn, R. Polysialylation of NCAM. Adv Exp Med Biol 663, 95–109, doi:10.1007/978-1-4419-1170-4_6 (2010).
El Maarouf, A. et al. Enzymatic engineering of polysialic acid on cells in vitro and in vivo using a purified bacterial polysialyltransferase. J Biol Chem 287, 32770–32779, doi:10.1074/jbc.M112.377614 (2012).
Lindhout, T. et al. Site-specific enzymatic polysialylation of therapeutic proteins using bacterial enzymes. Proc Natl Acad Sci USA 108, 7397–7402, doi:10.1073/pnas.1019266108 (2011).
Kabsch, W. XDS. Acta Crystallogr D Biol Crystallogr 66, 125–132, doi:10.1107/S0907444909047337 (2010).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr D Biol Crystallogr 67, 235–242, doi:10.1107/S0907444910045749 (2011).
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol Biol 364, 215–230, doi:10.1385/1-59745-266-1:215 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr 66, 213–221, doi:10.1107/S0907444909052925 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr D Biol Crystallogr 66, 486–501, doi:10.1107/S0907444910007493 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr 53, 240–255, doi:10.1107/S0907444996012255 (1997).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr D Biol Crystallogr 67, 355–367, doi:10.1107/S0907444911001314 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J Appl Crystallogr 40, 658–674, doi:10.1107/S0021889807021206 (2007).
Laskowski, R. A. PDBsum new things. Nucleic Acids Res 37, D355–359, doi:10.1093/nar/gkn860 (2009).
Schrodinger, L. (2015).
Gilbert, M. et al. The genetic bases for the variation in the lipo-oligosaccharide of the mucosal pathogen, Campylobacter jejuni. Biosynthesis of sialylated ganglioside mimics in the core oligosaccharide. J Biol Chem 277, 327–337, doi:10.1074/jbc.M108452200 (2002).
Gosselin, S., Alhussaini, M., Streiff, M. B., Takabayashi, K. & Palcic, M. M. A continuous spectrophotometric assay for glycosyltransferases. Anal Biochem 220, 92–97, doi:10.1006/abio.1994.1303 (1994).
Dolinsky, T. J., Nielsen, J. E., McCammon, J. A. & Baker, N. A. PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res 32, W665–667, doi:10.1093/nar/gkh381 (2004).
Dolinsky, T. J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res 35, W522–525, doi:10.1093/nar/gkm276 (2007).
Acknowledgements
We wish to acknowledge the Canadian Light Source and Advanced Light Source for access to synchrotron data collection facilities. We would like to thank M. Vuckovic for assistance in generating surface entropy reduction mutants. We also acknowledge operating funds from the Canadian Institutes of Health Research (N.C.J.S., S.G.W.), the Howard Hughes Medical Institute International Senior Scholar program (N.C.J.S.) and the Canadian Foundation of Innovation and British Columbia Knowledge Development Fund (N.C.J.S., S.G.W.). N.C.J.S. and S.G.W. are supported as Tier 1 Canada Research Chairs. W.W. wishes to acknowledge operating funds from Ryerson University. C.L. was supported by an advanced Postdoc Mobility fellowship of the Swiss National Science Foundation. L.B. was supported by the Deutsche Forschungsgemeinschaft. G.V. was supported by fellowships from the Michael Smith Foundation for Health Research and the Canadian Department for Foreign Affairs and International Trade.
Author information
Authors and Affiliations
Contributions
C.L., L.J.W. and N.C.J.S. designed the experiments. C.L. performed expression, purification, crystallization, data collection, processing and refinement of crystallographic structures. L.J.W. phased the initial native structure, performed docking of the fondaparinux ligand and assisted with data collection, processing and refinement of various data sets. L.B. synthesized acceptor compounds and performed kinetic experiments. M.P. performed initial expression and purification studies and initial crystallization experiments. G.V. performed initial expression and purification studies and supervised M.P. T.S. assisted in purification of enzyme mutants and helped with final structure refinements. L.S. and W.W. cloned initial MhPST construct and W.W. identified MhPST as a potential candidate for structural studies. S.G.W. supervised kinetic experiments. C.L., L.J.W. and N.C.J.S. analysed the data. C.L. and N.C.J.S. wrote the manuscript. All authors contributed to editing the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lizak, C., Worrall, L.J., Baumann, L. et al. X-ray crystallographic structure of a bacterial polysialyltransferase provides insight into the biosynthesis of capsular polysialic acid. Sci Rep 7, 5842 (2017). https://doi.org/10.1038/s41598-017-05627-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-05627-z
This article is cited by
-
Transition transferases prime bacterial capsule polymerization
Nature Chemical Biology (2024)
-
Bacterial capsules: Occurrence, mechanism, and function
npj Biofilms and Microbiomes (2024)
-
A multi-enzyme machine polymerizes the Haemophilus influenzae type b capsule
Nature Chemical Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.