Abstract
Mitochondria increase in number by the fission of existing mitochondria. Mitochondrial fission is needed to provide mitochondria to daughter cells during cell division. In Arabidopsis thaliana, four kinds of genes have been reported to be involved in mitochondrial fission. Two of them, DRP3 (dynamin-related protein3) and FIS1 (FISSION1), are well conserved in eukaryotes. The other two are plant-specific ELM1 (elongated mitochondria1) and PMD (peroxisomal and mitochondrial division). To better understand the commonality and diversity of mitochondrial fission factors in land plants, we examined mitochondrial fission-related genes in a liverwort, Marchantia polymorpha. As a bryophyte, M. polymorpha has features distinct from those of the other land plant lineages. We found that M. polymorpha has single copies of homologues for DRP3, FIS1 and ELM1, but does not appear to have a homologue of PMD. Citrine-fusion proteins with MpDRP3, MpFIS1 and MpELM1 were localized to mitochondria in M. polymorpha. MpDRP3- and MpELM1-defective mutants grew slowly and had networked mitochondria, indicating that mitochondrial fission was blocked in the mutants, as expected. However, knockout of MpFIS1 did not affect growth or mitochondrial morphology. These results suggest that MpDRP3 and MpELM1 but neither MpFIS1 nor PMD are needed for mitochondrial fission in M. polymorpha.
Similar content being viewed by others
Introduction
Almost all eukaryotes have mitochondria, organelles that are essential for energy production and supply, for the control of diverse metabolic pathways and for regulation of cell death. Mitochondria are dynamic organelles that move around and change their shape1,2,3,4. The morphology of mitochondria varies among species. In yeast and mammalian cells, mitochondria form networks that consists of tubules with multiple branch points5, 6. On the other hand, in plant cells, mitochondria are dispersed with many small spherical or peanut-shaped particles1, 7.
Mitochondria change their morphology depending on the life stage and growth conditions5, 8, 9. Both mitochondrial fission and fusion are needed for maintenance of mitochondrial morphology10, 11. Because mitochondria are not synthesized de novo but are created by the fission of existing mitochondria, mitochondrial fission is fundamental to the existence of eukaryotes12, 13. In yeast, a dynamin-related protein (Dnm1p) is recruited to mitochondrial fission sites where it forms an oligomer that constricts mitochondria to facilitate their fission10, 14,15,16. Dnm1p and its orthologues are well conserved in diverse eukaryotes including yeasts, algae, plants and mammals13. The Arabidopsis thaliana genome has two closely similar dynamin-related proteins, DRP3A and DRP3B, that are the functional orthologs of Dnm1p and that function in a redundant manner7, 17,18,19,20. In the drp3a drp3b double mutant, the mitochondria do not divide and form a massive, elongated network20.
In budding yeast, relocalization of Dnm1p depends on the mitochondrial outer membrane protein Fis1p and the cytosolic adaptor Mdv1p (and its paralog Caf4p), which can bind to both Dnm1p and Fis1p11, 21. In the fis1 mutant, mitochondria were elongated and networked22. Although Fis1p is required in yeast, the roles of Fis1 homologues in mammalian cells are somewhat controversial. Overexpression of hFis1 of Homo sapiens leads to mitochondrial fragmentation, indicating that it has a role in regulating mitochondrial morphology23, 24. However, hFis1 doesn’t appear to be essential for localizing Drp1 to mitochondrial fission sites, as its role can be handled by two other outer-membrane proteins, Mff and MiD49/5125,26,27,28.
The Arabidopsis genome has two closely related homologues of Fis1 (FIS1A/BIGYIN and FIS1B). FIS1A and FIS1B are reported to target to mitochondria, peroxisomes and chloroplasts, and were shown to facilitate mitochondrial fission29,30,31. BLAST searches of the Arabidopsis genome (https://www.arabidopsis.org/Blast/index.jsp) did not reveal any homologues of other mitochondrial fission factors (Mdv1p/Caf4p, Mff or MiD49/51). The closest matches had e-values > 1e-4, indicating very weak matches. On the other hand, it has two plant-specific fission factors, elongated mitochondria1 (ELM1) and peroxisomal and mitochondrial division factors (PMDs). ELM1 was identified as the gene responsible for the elongated mitochondria mutant phenotype in Arabidopsis. In wild-type Arabidopsis, ELM1 surrounds mitochondria and interacts with both DRP3A and DRP3B32. These results suggest that ELM1 is required for relocalization of DRP3A from the cytosol to mitochondrial fission sites. Elongated mitochondria are also observed in the pmd1 mutant, but PMD1 and its paralog PMD2 do not physically interact with DRP3 or FIS1, suggesting that PMD1 facilitates mitochondrial proliferation in a DRP3/FIS1-independent manner33.
Recent molecular phylogenetic analyses of land plants agreed the basal position bryophytes, encompassing liverworts, mosses and hornworts, to vascular plants, and liverworts are considered as one of the earliest diverging distant land plant lineages34,35,36. Therefore, liverworts are could be a key clade to our understanding of the commonalities and differences of mitochondrial fission factors among land plants. Here, we used reverse genetics to study candidate genes for mitochondrial fission in the liverwort Marchantia polymorpha. Marchantia is emerging as an experimental model organism, because it has little genetic redundancy in many cases and is well suited for various strategies for molecular genetics, such as introduction of reporter constructs, gene silencing and targeted gene modification37,38,39,40. Our results demonstrate that mitochondrial fission in a liverwort relies on some factors used in vascular plants (DRP3 and ELM1) but does not seem to rely on other factors (FIS1 and PMDs).
Results
The Marchantia genome has single copies of homologues of DRP3, FIS1 and ELM1
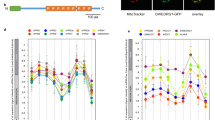
For searches of the M. polymorpha genome, we used the JGI M. polymorpha EST and genome databases ver. 3.1 (https://phytozome.jgi.doe.gov/pz/portal.html). BLASTx searches of the JGI M. polymorpha EST and genome databases for homologues of DRP3, FIS1, ELM1 and PMD revealed no PMD homologue (e-value of closest match > 1e-4), but did reveal single copies of homologues of DRP3 (0.0 or 6e-71), FIS1 (5e-35) and ELM1 (1e-138). We call these homologues MpDRP3 (Mapoly0069s0084), MpFIS1 (Mapoly0147s0019) and MpELM1 (Mapoly0038s0050), respectively. Two different mRNAs of MpDRP3 were found and appear to be transcribed from a single gene (Figs 1a–c and S1a). The predicted splicing variants, MpDRP3s (short) and MpDRP3l (long), differed by 27 bp in exon 13. RT-PCR analysis revealed that the mRNA expression levels of MpDRP3s and MpDRP3l in the thallus were similar (Fig. 1b). The amino acid sequences of MpDRP3s and MpDRP3l are highly similar to their Arabidopsis and Physcomitrella homologues (Fig. S1a). MpDRP3s had 62.4% identity to AtDRP3A and 79.7% identity to PpDRP3. The alternative splice site is not conserved in Arabidopsis and Physcomitrella (Fig. S1a, magenta). In addition, all five genes have predicted GTPase, dynamin middle and GTPase effector domains (Fig. 1c). Similarly, MpFIS1 (Fig. 1d) and MpELM1 (Fig. 1f) had high similarities in length, amino-acid sequences (Fig. S1b and c) and domain structures (Fig. 1e and g) to Arabidopsis and Physcomitrella counterparts, respectively.
Genes and predicted proteins information of MpDRP3, MpFIS1 and MpELM1. (a) Genomic structure of MpDRP3. The splice site variant was identified at exon 13 (denoted by the asterisk). Closed arrows indicate the positions of primers used in (b). (b) RT-PCR analysis of RNA extracted from the detached thallus. A full-length gel image is presented in Figure S6a. (c) Domain structures of AtDRP3A, AtDRP3B, MpDRP3s and MpDRP3l. Each domain is identified and depicted by using the pfam program (http://pfam.sanger.ac.uk/). The magenta colored arrow indicates the splice site. (d) Genomic structure of MpFIS1. (e) Domain structures of AtFIS1A, AtFIS1B and MpFIS1. Each domain is identified and depicted by using the pfam program. (f) Genomic structure of MpELM1. (g) Domain structures of AtELM1 and MpELM1. Each domain is identified and depicted by using the pfam program.
MpDRP3, MpFIS1 and MpELM1 were localized to mitochondria
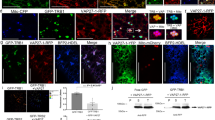
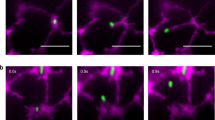
To examine the subcellular localization of MpDRP3s and MpDRP3l, we observed a fluorescent marker Citrine-fused proteins (Citrine-MpDRP3s and Citrine-MpDRP3l) with a confocal laser scanning microscope (CLSM)(Fig. 2a). The fusion genes were given an N-terminal Citrine tag and placed under the control of the CaMV 35 S promoter. In transgenic plants expressing these proteins and stained with MitoTracker, a part of the Citrine-MpDRP3s and Citrine-MpDRP3l signals localized to the constriction sites of mitochondria (Fig. 2a, arrowheads). Other signals were mainly localized to the tips of mitochondria, while some remained in the cytosol. To better see the localizations of MpDRP3s or MpDRP3l, we prepared another reporter protein: RFP fused to the N-terminus of MpDRP3s or MpDRP3l, and observed cells expressing both Citrine-MpDRP3s and RFP-MpDRP3l (or Citrine-MpDRP3l and RFP-MpDRP3s) (Fig. S2a). MpDRP3s and MpDRP3l signals overlapped in each combination of plasmids, suggesting that they took part in the same fission machinery (Fig. S2b). Next, the DNA fragment 35Spro:Citrine:MpFIS1 was introduced into Marchantia to identify the localization of MpFIS1 within the cells. In transgenic plants expressing Citrine-MpFIS1 and stained with MitoTracker, almost all Citrine fluorescence was in the form of green oval-rings. Many of the rings surrounded the MitoTracker fluorescence of mitochondria (Fig. 2b, arrow) while some did not contain the red signals (Fig. 2b, arrowhead). Transient expression of Citrine-MpFIS1 and RFP fused to the peroxisomal targeting signal 2 (PTS2-RFP) showed that a part of Citrine-MpFIS1 colocalized with PTS2-RFP (Fig. S3). In addition, Marchantia thalli were transformed transiently with MpELM1-Citrine and mitochondrial-targeted RFP (MtRFP) by particle bombardment. Although the mitochondria appeared to be enlarged and aggregated, at least some of the MpELM1-Citrine surrounded the RFP fluorescence of mitochondria (Fig. 2c).
Intracellular localization of MpDRP3, MpFIS1 and MpELM1 in Marchantia thallus. (a) CLSM images of the epidermal cell expressing Citrine-MpDRP3s (upper panel) or Citrine-MpDRP3l (lower panel) and stained with MitoTracker (Bars = 10 µm). Magnified images are shown in insets (Bars = 5 µm). The arrowheads indicate Citrine signals localized to the constriction sites of mitochondria. (b) CLSM image of the epidermal cell expressing Citrine-MpFIS1 and stained with MitoTracker (Bars = 10 µm). Magnified images are shown in insets (Bars = 5 µm). The arrow indicates a mitochondrion and the arrowhead indicates a body not mitochondria. (c) Fluorescence image of the epidermal cell transiently transformed with Citrine-MpELM1 and MtRFP by particle bombardment (Bars = 10 µm). Magnified images are shown in insets (Bars = 1 µm). Blue signals were chlorophyll autofluorescence.
Mpelm1 and Mpdrp3 (but not Mpfis1) showed significant growth defects
To clarify the role of each gene in mitochondrial fission and plant growth, we made MpFIS1, MpELM1 and MpDRP3 knockout mutants (named Mpfis1, Mpelm1 and Mpdrp3, respectively) by homologous recombination (HR)-mediated gene targeting (Fig. 3a)38. The knockouts were confirmed by RT-PCR (Fig. 3b). The wild-type plants grew to an area of about 200 mm2 in 2-weeks from a gemma (Fig. 3c). Compared with the wild type, Mpfis1 did not have any noticeable developmental or growth defects (Fig. 3c). In contrast, Mpelm1 and Mpdrp3 showed similar growth retardation, i.e., their areas grew to less than 10 mm2 in 2 weeks from a gemma. In addition, Mpelm1 and Mpdrp3 showed distorted thalli phenotypes (Fig. 3d), and they began to wither before reaching the wild-type size. However, these mutants were able to complete their asexual life cycle, i.e., they generated asexual propagules called gemmae, which develop into adult gametophytes.
Generation of Mpfis1, Mpelm1 and Mpdrp3 plants by homologous recombination and plant phenotypes of the mutants. (a) Schematic representation of the genomic structures of the MpFIS1, MpELM1 and MpDRP3, the disrupting-constructs and their targeting loci. Each primer pair for RT-PCR is indicated by arrows. (b) RT-PCR for MpFIS1, MpELM1 and MpDRP3 mRNA in wild-type (WT), Mpfis1, Mpelm1 and Mpdrp3 plants. MpEF1 (Elongation Factor 1 alpha) was used as an internal control. Full-length gel images are presented in Figure S6b. (c) Photographs of 2-weeks-old plants of wild-type (WT), Mpfis1, Mpelm1 and Mpdrp3. All plants shown in this image were grown on a single plate. Bar = 5 mm. (d) Magnified images of distorted thalli of Mpelm1 and Mpdrp3 by using a stereoscopic microscope. Bars = 1 mm.
Mpelm1 and Mpdrp3 (but not Mpfis1) showed severe morphological phenotypes of mitochondria
We then examined whether knockout of those genes affects mitochondrial morphology. In wild-type and Mpfis1 cells, most mitochondria were observed as particles or narrow (0.5 to 1 µm) rod-like structures (Fig. 4a). Citrine-MpDRP3s and MpELM1-Citrine in Mpfis1 localized to mitochondria (Fig. S4a and S4b respectively), as they did in the wild type (Fig. 2a and c, respectively). By contrast, massively elongated and interconnected mitochondria were observed in Mpelm1 and Mpdrp3 with some lump-like structures (Fig. 4a, arrowhead). To compare the mutants quantitatively by their mitochondrial morphology, we used a morphology scoring assay in which each cell was categorized as having one of four types of mitochondria: 1) circular, 2) tubular, 3) elongated and 4) elongated + enlarged (i.e., mitochondria interconnected with some lump-like structures). Following previous studies10, 15, 26, we compared the phenotypes of mitochondria in similar volumes and the same cell numbers using a cell-based assay, rather than a mitochondrion-based assay. Although there were no “Elongated” cells in the wild-type or Mpfis1, most of the cells in Mpelm1 and Mpdrp3 were categorized as “Elongated” (Fig. 4b), indicating that mitochondrial fission was blocked in Mpelm1 and Mpdrp3, but not in Mpfis1.
Mitochondrial phenotypes of the mutants. (a) CLSM image of the epidermal cell in wild-type and each mutants stained with MitoTracker. The arrowhead indicates an enlarged structure of mitochondria. Blue signals are chlorophyll autofluorescence. Bar = 10 µm. (b) Scoring of mitochondrial morphologies. Each cell was scored into one of four morphological categories. Representative images of each categories are shown in right panels. Data are means ± s.e.m. from three independent experiments. *P < 0.05; n.s., not significant; one-way ANOVA with Tukey’s multiple-comparison test. (c) Transmission electron micrographs of mitochondria in wild-type and each mutants. Lower panels are magnified images of enlarged structures in Mpelm1 and Mpdrp3. The black arrow indicates a vacuolated structure, the black arrowhead indicates a vesicle-like structure and the asterisk indicates the cytosol surrounded by mitochondria (Bars = 500 nm). Fluorescent images of enlarged structures stained with MitoTracker are shown in insets. The yellow arrows indicate the mitochondrial region which has little MitoTracker signals (Bars = 5 µm).
As with the fluorescence microscopy, transmission electron microscopy showed that most of mitochondria in Mpelm1 and Mpdrp3 were much longer than those in the wild-type and Mpfis1 (Fig. 4c). In addition, enlarged structures of mitochondria included vesicle-like structures (black arrowhead) and vacuolated structures (black arrow). These structures might be similar to the dark areas inside mitochondrial red bodies (shown in the insets of Fig. 4c (yellow arrows)), which were not stained with MitoTracker. Although Mpelm1 and Mpdrp3 had enlarged mitochondria, their cristae were similar to those in the wild-type, and they maintained their electrochemical potential because the MitoTracker that we used (Orange CMTMRos) cannot accumulate in mitochondria that do not maintain a membrane potential.
Transient expression of MpDRP3 and MpELM1 in the mutants complemented the morphological phenotypes of mitochondria
Because MpDRP3s and MpDRP3l had similar subcellular localizations, we examined whether both of them were functional. To test this, we examined the ability of the MpDRP3s and MpDRP3l to complement the defective mitochondrial networking phenotype of Marchantia lacking MpDRP3. When Mpdrp3 was transformed transiently with either Citrine-MpDRP3s or Citrine-MpDRP3l by particle bombardment, the elongated mitochondrial morphology reverted to a wild-type-like morphology. Similarly, Mpelm1 cells with MpELM1-Citrine have discrete complemented mitochondria (Fig. 5b).
Complementation analysis of mutants lacking MpDRP3 or MpELM1. (a) CLSM observations of either Citrine-MpDRP3s or Citrine-MpDRP3l and MtRFP in Mpdrp3 by using the particle bombardment (Bars = 5 µm). Those were transiently expressed under the control of the CaMV 35 S promoter. Magnified images are shown in insets (Bars = 1 µm). (b) CLSM Images of MpELM1-Citrine and MtRFP in Mpelm1 (Bars = 5 µm). Blue signals were chlorophyll autofluorescence.
Discussion
In this work, three of four candidate genes for mitochondrial fission (MpDRP3, MpFIS1 and MpELM1) were found in the genomic database of Marchantia polymorpha (Fig. 1). The finding that these three genes (but not PMD) are conserved in Arabidopsis and Marchantia, indicate that MpDRP3, MpFIS1 and MpELM1 have significant roles in land plants. The Arabidopsis genome has two copies of DRP3-, FIS1- and ELM1-type genes13. In contrast, Marchantia has single copies of these genes, which makes it a good model for revealing the mechanism of mitochondrial fission in land plants.
MpDRP3 exists in two alternative splicing isoforms (MpDRP3s and MpDRP3l) that differ by 27 bp in exon 13 (Fig. 1). The mRNA expression levels of MpDRP3s and MpDRP3l in the thallus were similar (Fig. 1b). Furthermore, the mitochondrial morphology in Mpdrp3 was rescued by the expression of either Citrine-MpDRP3s or Citrine-MpDRP3l transiently (Fig. 5a). Together, these findings suggest that MpDRP3s and MpDRP3l are functionally redundant, at least in the observed Marchantia thallus cells.
Because little or no difference was observed in plant growth (Fig. 3c) or mitochondrial morphology (Fig. 4b) between Mpfis1 and the wild-type, MpFIS1 does not appear to have a major role in mitochondrial fission. The mitochondrial localizations of Citrine-MpDRP3s and MpELM1-Citrine in Mpfis1 mutants in Fig. S4a and S4b suggest that MpFIS1, unlike yeast FIS1, is not required for the localization of these mitochondrial fission proteins from the cytosol. Citrine-MpFIS1, which was detected on the mitochondrial surface (Fig. 2b), probably binds by the C-terminal transmembrane domain of MpFIS1, as this domain is crucial for mitochondrial targeting in other eukaryotes11, 23, 24. Furthermore, Citrine-MpFIS1 was localized to ring-shaped bodies (Fig. 2b, left panel, arrowhead), which seems to be peroxisomes (Fig. S3). Although FIS1 also localizes to peroxisomes and is involved in peroxisomal fission in Arabidopsis30, 31, 41, 42, the role of MpFIS1 in peroxisomal fission remains unclear.
Moreover, phenotypic analyses of plant growth (Fig. 3c) and mitochondrial morphology (Fig. 4b) suggested that not only MpDRP3 but also MpELM1 play crucial roles in mitochondrial fission. In Arabidopsis, drp3a drp3b double mutants show a severe growth-defect with network-shaped mitochondria20, but elm1 mutants do not show growth defects32. This is possibly because the latter mutants have residual mitochondrial fission activity, attributed to another ELM1 paralogue (At5g06180) in the Arabidopsis genome32. By contrast, the Marchantia genome has a single copy of ELM1, so that there would be no residual mitochondrial fission activity in Mpelm1 mutants.
The phenotypes in Mpelm1 strongly resemble those in Mpdrp3 (Figs 3c and 4a), which raises the possibility that MpELM1 is involved in an MpDRP3-dependent pathway. In other words, MpDRP3 may require MpELM1, in the same way that Dnm1p requires the cytosolic adaptor Mdv1p/Caf4p, which localizes Dnm1p to the mitochondrial fission sites in budding yeast11, 21 (Fig. 6, left). It is possible that MpDRP3 interacts with MpELM1, since DRP3A and DRP3B interact with ELM1 for their localization to mitochondria in Arabidopsis (two-way arrows in Fig. 6)32. Further biochemical studies in Marchantia are needed to test this model.
Dynamin-related proteins (green shapes in Fig. 6) are universally conserved in eukaryotes, including yeasts, humans and vascular plants7, 14, 43, 44. The present results show that this is also the case in Marchantia. Indeed, MpDRP3 had high identities with DRP3 orthologues from A. thaliana and P. patens (Fig. 1c), suggesting that dynamin-related proteins are also crucial in bryophytes. Although the amino acid sequences of Fis1 homologues (blue bars in Fig. 6) are conserved across diverse eukaryotes and Fis1 plays an integral part for mitochondrial fission in yeast, it remains unclear whether Fis1 has a role in mitochondrial fission in mammals26. Considering that mammalian membrane-anchored proteins (Mff and MiD49/51) act as mitochondrial adaptors for Drp1 in place of (or with) Fis125,26,27,28, it is possible that M. polymorpha has other mitochondrial fission factor(s) on the surface of mitochondria. Because MpFIS1 does not appear to play a role in mitochondrial fission (Fig. 4b), it probably has some other function. This raises the possibility that Fis1s in other eukaryotes might also have some other common function.
The Marchantia genome does not seem to have a PMD homologue. A search of the databases shows that homologues of PMDs are found only in Brassicaceae, a family of vascular plants, and related plants, suggesting that PMD is a mitochondrial fission factor specific to these plants. On the other hand, ELM1-like proteins are evolutionarily conserved across land plants32, but they are not present in the genomes of chlorophytes (Chlamydomonas reinhardtii, Volvox carteri and Chlorella vulgaris) or a rhodophyte (Cyanidioschyzon merolae). The present results show that the amino acid sequence and the function of ELM1 in the vascular plant Arabidopsis are conserved in a bryophyte, indicating that the ancestor of land plants acquired ELM1 as a mitochondrial fission factor.
Experimental procedures
Plant materials and growth conditions
Male, accession Takaragaike-1 (called Tak-1), M. polymorpha were asexually maintained and propagated through gemmae growth at 22 °C by using half-strength Gamborg’s B5 medium containing 1% agar under 16-h of diurnal light or continuous white light (50–100 µmol m−2sec−1)37, 45.
Plasmid constructions
Gateway cloning technology (Life Technologies) was used to construct plasmids. cDNA was amplified by PCR with the following primers (Table S1). The amplified DNA fragments were mixed with an entry vector, pENTR/D-TOPO, and the TOPO reaction were performed according to the manufacturer’s instructions (Life Technologies). In addition, the stop codon of pENTR-MpELM1 was deleted by using Site-Directed Mutagenesis to add the fluorescent protein to their C-terminus. The resulting plasmids were mixed and reacted with an LR clonase with destination vectors, p2RGW746, pMpGWB10547 or pMpGWB10647, to construct binary vectors containing the CaMV 35 S promoter, fluorescent genes and a NOS terminator.
RT-PCR analysis
Total RNA was extracted from sections of thallus. RNA isolation was carried out using the RNeasy Plant Mini Kit (QIAGEN), according to manufacturer’s instructions. First strand cDNA synthesis was carried out starting from 1 µg of total RNA by using SuperScript III Reverse Transcriptase (Invitrogen) and the oligo(dT) primers for reverse transcription. To detect the splicing variants and transcriptional products, RT-PCR analyses were performed using appropriate primers (Table S1). The expression levels of each gene were normalized to the expression level of the MpEF1.
Putative amino-acid sequence and domain structures
Sequence comparisons were performed by using the CLUSTAL W multiple alignment program and GeneDoc Program between sequences of Marchantia, Arabidopsis and Physcomitrella in NCBI database48. To analyse domain structures, we used Conserved Domain Search Service by NCBI (http://www.ncbi.nlm.nih.gov/). SOSUI and TMHMM transmembrane domain searches predicted transmembrane domain in the MpELM1 sequence49, 50.
Agrobacterium-mediated transformation
Transgenic plants were generated with AgarTrap (Agar-utilized Transformation with Pouring Solutions) as described previously40. Transformants were selected on plates containing hygromycin B (10 µg ml−1) and claforan (100 µg ml−1) and cultivated for approximately 1 month to obtain gemmae. Then, a gemma was picked to establish an isogenic line from the respective transgenic lines, as a gemma is developed from a single cell39.
Particle bombardment
Plasmids for visualizing mitochondrial fission factors were introduced via particle bombardment, by using a helium-driven particle accelerator (Bio-Rad, PDS/1000), with all basic adjustments set according to the manufacturer’s recommendations. The bombardment parameters applied were as follows: bombardment pressure, 7.6 × 104 hPa; gold particles of 1.6 µm in diameter; 12 cm between the macrocarrier and plants; decompression vacuum, 915 hPa. After bombardment, plants were incubated for 2-days and observed by using CLSM.
MitoTracker Orange staining
Small sections (2–10 mm2) were cut out of the Marchantia thallus with a sharp razor blade, stained with 0.5 µM MitoTracker Orange CMTMRos (Life Technologies) for about 20 minutes, and washed three times with distilled water.
Microscopic observations and image analysis
Microscopic observations were performed by using a transmission electron microscope (JEOL, JEM-1010), a stereoscopic microscope (Leica, M125), a fluorescence microscope (Nikon, ECLIPSE Ti) and a confocal laser scanning microscope system (Nikon, C1Si). For the electron microscopic analysis, the pre-fixation was performed with phosphate buffer (50 mM) containing 4% paraformaldehyde and 2% glutaraldehyde. The pre-fixed samples were then post-fixed with 2% osmium tetroxide. The specimens were dehydrated in a graded ethanol series and embedded in Spurr resin (Spurr Low Viscosity Embedding Kit, Polysciences). Ultrathin sections (70 nm) were prepared and stained with 2% uranyl acetate and lead solution.
In fluorescence imaging, a 488-nm Ar/Kr laser was used for the excitation of Citrine. A 561-nm diode laser was also used for RFP, MitoTracker Orange and autofluorescence from chloroplasts. Emission signals were detected using a 515/30-nm filter for Citrine, a 590/70-nm filter for RFP and MitoTracker Orange and a 650-nm filter for autofluorescence. All of the data were processed using Adobe Photoshop CS4 Extended 11.0 (Adobe Systems).
Targeted gene knockout
To generate the targeting vectors, pJHY-TMp1 was used38. The 5′ and 3′ homology arms were amplified with the appropriate primer pairs, and the PCR products were cloned into the AscI and PacI sites of pJHY-TMp1, respectively (Tables S1 and S2). Introduction of the targeting construct into M. polymorpha and selection of knockout lines were performed as previously described37, 38.
Quantification of mitochondrial morphology
One experimenter stained the mutant and wild-type plants with MitoTracker Orange, and obtained images of 50 epidermal cells with CLSM in each experiment for three replicates. A second experimenter, blinded to the preparation of the images, categorized each cell into four groups (circular, tubular, elongated and elongated + enlarged structures) based on mitochondrial morphologies. All statistical testing was performed with one-way analysis of variance (ANOVA) followed by Tukey’s multiple-comparison test. P < 0.05 was considered to be statistically significant; n.s., not significant.
Accession numbers
The sequences of the MpEF1 (Elongation Factor 1 alpha), MpDRP3, MpELM1 and MpFIS1 genes are available in DDBJ under the following accession numbers: MpEF1 (KJ146970); MpDRP3 (LC209090); MpELM1 (LC209091); MpFIS1 (LC209092).
References
Logan, D. C. & Leaver, C. J. Mitochondria-targeted GFP highlights the heterogeneity of mitochondrial shape, size and movement within living plant cells. J. Exp. Bot. 51, 865–871 (2000).
Van Gestel, K., Köhler, R. H. & Verbelen, J. P. Plant mitochondria move on F-actin, but their positioning in the cortical cytoplasm depends on both F-actin and microtubules. J. Exp. Bot. 53, 659–667 (2002).
Logan, D. C. Mitochondrial dynamics. New Phytol. 160, 463–478 (2003).
Doniwa, Y., Arimura, S. & Tsutsumi, N. Mitochondria use actin filaments as rails for fast translocation in Arabidopsis and tobacco cells. Plant Biotech. 24, 441–447 (2007).
Bereiter-Hahn, J. & Voth, M. Dynamics of mitochondria in living cells: shape changes, dislocations, fusion, and fission of mitochondria. Micro. Res. and Tech. 27, 198–219 (1994).
Nunnari, J. et al. Mitochondrial transmission during mating in Saccharomyces cerevisiae Is determined by mitochondrial fusion and fission and the intramitochondrial segregation of mitochondrial DNA. Mol. Biol. Cell 8, 1233–1242 (1997).
Arimura, S. & Tsutsumi, N. A dynamin-like protein (ADL2b), rather than FtsZ, is involved in Arabidopsis mitochondrial division. Proc. Natl. Acad. Sci. USA 99, 5727–5731 (2002).
Seguí-Simarro, J. M., Coronado, M. J. & Staehelin, L. A. The mitochondrial cycle of Arabidopsis shoot apical meristem and leaf primordium meristematic cells is defined by a perinuclear tentaculate/cage-like mitochondrion. Plant Physiol. 148, 1380–1393 (2008).
Ruberti, C. et al. Mitochondria change dynamics and morphology during grapevine leaf senescence. PLoS One 9(7), e102012 (2014).
Sesaki, H. & Jensen, R. E. Division versus fusion: Dnm1p and Fzo1p antagonistically regulate mitochondrial shape. J. Cell Biol. 147, 699–706 (1999).
Shaw, J. M. & Nunnari, J. Mitochondrial dynamics and division in budding yeast. Trends Cell Biol 12, 178–184 (2002).
Kuroiwa, T. et al. The division apparatus of plastids and mitochondria. Int. Rev. Cytol 181, 1–41 (1998).
Arimura, S. & Tsutsumi, N. Mitochondrial and peroxisomal division in Molecular Cell Biology of the Growth and Differentiation of Plant Cells (ed. Rose, R.J.) 51–65 (CRC press, 2016).
Otsuga, D. et al. The dynamin-related GTPase, Dnm1p, controls mitochondrial morphology in yeast. J. Cell Biol. 143, 333–349 (1998).
Bleazard, W. et al. The dynamin-related GTPase Dnm1 regulates mitochondrial fission in yeast. Nat. Cell Biol. 1, 298–304 (1999).
Okamoto, K. & Shaw, J. M. Mitochondrial morphology and dynamics in yeast and multicellular eukaryotes. Annu. Rev. Genet. 39, 503–536 (2005).
Arimura, S., Aida, G. P., Fujimoto, M., Nakazono, M. & Tsutsumi, N. Arabidopsis dynamin-like protein 2a (ADL2a), like ADL2b, is involved in plant mitochondrial division. Plant Cell Physiol. 45, 236–242 (2004).
Logan, D. C., Scott, I. & Tobin, A. K. ADL2a, like ADL2b, is involved in the control of higher plant mitochondrial morphology. J. Exp. Bot 55, 783–785 (2004).
Mano, S., Nakamori, C., Kondo, M., Hayashi, M. & Nishimura, M. An Arabidopsis dynamin-related protein, DRP3A, controls both peroxisomal and mitochondrial division. Plant J. 38, 487–498 (2004).
Fujimoto, M. et al. Arabidopsis dynamin-related proteins DRP3A and DRP3B are functionally redundant in mitochondrial fission, but have distinct roles in peroxisomal fission. Plant J. 58, 388–400 (2009).
Westermann, B. Mitochondrial fusion and fission in cell life and death. Mol. Cell Biol. 11, 872–884 (2010).
Mozdy, A. D., Mccaffery, J. M. & Shaw, J. M. Dnm1p GTPase-mediated mitochondrial fission is a multi-step process requiring the novel integral membrane component Fis1p. J. Cell Biol. 151, 367–379 (2000).
Yoon, Y., Krueger, E. W., Oswald, B. J., Mcniven, A. & Mcniven, M. A. The mitochondrial protein hFis1 regulates mitochondrial fission in mammalian cells through an interaction with the dynamin-like protein DLP1. Mol. Cell Biol. 23, 5409–5420 (2003).
Stojanovski, D., Koutsopoulos, O. S., Okamoto, K. & Ryan, M. T. Levels of human Fis1 at the mitochondrial outer membrane regulate mitochondrial morphology. J. Cell Sci. 117, 1201–1210 (2004).
Zhao, J., Liu, T., Jin, S., Lendahl, U. & Niste, M. Human MIEF1 recruits Drp1 to mitochondrial outer membranes and promotes mitochondrial fusion rather than fission. EMBO J. 30, 2762–2778 (2011).
Otera, H. et al. Mff is an essential factor for mitochondrial recruitment of Drp1 during mitochondrial fission in mammalian cells. J. Cell Biol. 191, 1141–1158 (2010).
Palmer, C. S. et al. Adaptor proteins MiD49 and MiD51 can act independently of Mff and Fis1 in Drp1 recruitment and are specific for mitochondrial fission. J. Biol. Chem. 288, 27584–27593 (2013).
Koirala, S. et al. Interchangeable adaptors regulate mitochondrial dynamin assembly for membrane scission. Proc. Natl. Acad. Sci. USA 110, 1342–1351 (2013).
Scott, I., Tobin, A. K. & Logan, D. C. BIGYIN, an orthologue of human and yeast FIS1 genes functions in the control of mitochondrial size and number in Arabidopsis thaliana. J. Exp. Bot. 57, 1275–1280 (2006).
Zhang, X. & Hu, J. Two small protein families, DYNAMIN-RELATED PROTEIN3 and FISSION1, are required for peroxisome fission in Arabidopsis. Plant J. 57, 146–159 (2009).
Ruberti, C., Costa, A., Pedrazzini, E., Schiavo, F. L. & Zottini, M. FISSION1A, an Arabidopsis tail-anchored protein, is localized to three subcellular compartments. Mol. Plant 7, 1393–1396 (2014).
Arimura, S. et al. Arabidopsis ELONGATED MITOCHONDRIA1 is required for localization of DYNAMIN-RELATED PROTEIN3A to mitochondrial fission sites. Plant Cell 20, 1555–1566 (2008).
Aung, K. & Hu, J. The Arabidopsis tail-anchored protein PEROXISOMAL AND MITOCHONDRIAL DIVISION FACTOR1 is involved in the morphogenesis and proliferation of peroxisomes and mitochondria. Plant Cell 23, 4446–4461 (2011).
Qiu, Y. L. et al. The deepest divergences in land plants inferred from phylogenomic evidence. Proc. Natl. Acad. Sci. USA 103, 15511–15516 (2006).
Chang, Y. & Graham, S. W. Inferring the higher-order phylogeny of mosses (Bryophyta) and relatives using a large, multigene plastid data set. Am. J. Bot. 98, 839–849 (2011).
Wickett, N. J. et al. Phylotranscriptomic analysis of the origin and early diversification of land plants. Proc. Natl. Acad. Sci. USA 111, E4859–E4868 (2014).
Ishizaki, K., Chiyoda, S., Yamato, K. T. & Kohchi, T. Agrobacterium-mediated transformation of the haploid liverwort Marchantia polymorpha L., an emerging model for plant biology. Plant Cell Physiol. 49, 1084–1091 (2008).
Ishizaki, K., Johzuka-hisatomi, Y., Ishida, S., Iida, S. & Kohchi, T. Homologous recombination-mediated gene targeting in the liverwort Marchantia polymorpha L. Sci. Rep. 3, 1532, doi:10.1038/srep01532 (2013).
Ishizaki, K., Nishihama, K., Yamato, K. T. & Kohchi, T. Molecular genetic tools and techniques for Marchantia polymorpha research. Plant Cell Physiol. 57, 262–270 (2016).
Tsuboyama, S. & Kodama, Y. AgarTrap: a simplified agrobacterium-mediated transformation method for sporelings of the liverwort Marchantia polymorpha L. Plant Cell Physiol. 55, 229–236 (2014).
Lingard, M. J. et al. Arabidopsis PEROXIN11c-e, FISSION1b, and DYNAMIN-RELATED PROTEIN3A cooperate in cell cycle-associated replication of peroxisomes. Plant Cell 20, 1567–1585 (2008).
Zhang, X. & Hu, J. FISSION1A and FISSION1B Proteins Mediate the Fission of Peroxisomes and Mitochondria in Arabidopsis. Mol. Plant 1, 1036–1047 (2008).
Smirnova, E., Griparic, L., Shurland, D. & Bliek, A. M. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol. Biol. Cell 12, 2245–2256 (2001).
Fujimoto, M., Arimura, S., Nakazono, M. & Tsutsumi, N. A rice dynamin-like protein, OsDRP3A, is involved in mitochondrial fission. Breed. Sci. 54, 367–372 (2004).
Gamborg, O. L., Miller, R. A. & Ojima, K. Nutrient requirements of suspension cultures of soybean root cells. Exp. Cell Res. 50, 151–158 (1968).
Karimi, M., Inzé, D. & Depicker, A. GATEWAYTM vectors for agrobacterium-mediated plant transformation. Trends Plant Sci. 7, 193–195 (2002).
Ishizaki, K. et al. Development of gateway binary vector series with four different selection markers for the liverwort Marchantia polymorpha. PLoS One 10(9), e0138876 (2015).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Hirokawa, T., Boon-Chieng, S. & Mitaku, S. SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14, 378–379 (1998).
Krogh, A., Larsson, B., Heijne, G. V. & Sonnhammer, E. L. L. Predicting transmembrane protein topology with a hidden markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Acknowledgements
We thank Dr. M. Karimi (Ghent University, Belgium) for Gateway destination vectors. We thank Dr. K. Katayama and Dr. M. Fujimoto for technical assistance. We thank Dr. S. Mano (NIBB, Japan) for the PTS2-RFP expression vector. We appreciate Dr. Y. Kodama and Ms. S. Tsuboyama-Tanaka (Utsunomiya University, Japan) for technical advice of AgarTrap. We thank Ms. S. Ishida (Kyoto University, Japan) for the generation of knockout mutants by homologous recombination.
Author information
Authors and Affiliations
Contributions
N.N., A.Y. and S.A. designed the research, interpreted the data and wrote the paper. Molecular cloning and vector constructions were carried out by N.N. and R.K. Gene targeting and other transformations of Marchantia were performed by N.N., A.Y., R.K., K.I. and T.K. Microscopic observations and image analysis were performed by N.N., F.I. and N.T. A.Y. and Y.W. cultivated all plants.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nagaoka, N., Yamashita, A., Kurisu, R. et al. DRP3 and ELM1 are required for mitochondrial fission in the liverwort Marchantia polymorpha . Sci Rep 7, 4600 (2017). https://doi.org/10.1038/s41598-017-04886-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04886-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.