Abstract
Long-read sequencing technologies such as Pacific Biosciences and Oxford Nanopore MinION are capable of producing long sequencing reads with average fragment lengths of over 10,000 base-pairs and maximum lengths reaching 100,000 base- pairs. Compared with short reads, the assemblies obtained from long-read sequencing platforms have much higher contig continuity and genome completeness as long fragments are able to extend paths into problematic or repetitive regions. Many successful assembly applications of the Pacific Biosciences technology have been reported ranging from small bacterial genomes to large plant and animal genomes. Recently, genome assemblies using Oxford Nanopore MinION data have attracted much attention due to the portability and low cost of this novel sequencing instrument. In this paper, we re-sequenced a well characterized genome, the Saccharomyces cerevisiae S288C strain using three different platforms: MinION, PacBio and MiSeq. We present a comprehensive metric comparison of assemblies generated by various pipelines and discuss how the platform associated data characteristics affect the assembly quality. With a given read depth of 31X, the assemblies from both Pacific Biosciences and Oxford Nanopore MinION show excellent continuity and completeness for the 16 nuclear chromosomes, but not for the mitochondrial genome, whose reconstruction still represents a significant challenge.
Similar content being viewed by others
Introduction
The advent of next generation sequencing technologies (NGS) has marked the start of a new era in genomics research. Compared to the previous Sanger technology1, NGS has significantly lowered the cost of sequencing using massively parallel sequencing methods2, 3. In a typical NGS run, DNA molecules are sheared into small fragments and then clonally amplified before being sequenced. After DNA amplification, multiple fragments of the sequences obtained may cover the same genome region, so that computational algorithms can be used to concatenate and assemble such reads like a jigsaw puzzle and generate a consensus to correct for the occasional sequencing errors. The typical length of the DNA fragments sequenced is between 50 and 400 bases long2, and as a result, the assembly obtained from such short reads is fragmented in contigs much smaller than the actual chromosome sizes. In particular, short reads are not able to solve complex genome features like repeated regions (repeats) longer than the fragment length or copy number variations, with the typical outcome that (almost-) identical repeats are collapsed into a single element in the assembly. To overcome the high fragmentation of NGS-based assemblies and to help resolve long repeats, long-read sequencing technologies have been developed and recently adopted by the genomics community. The main characteristic of these new platforms is to work with long DNA molecules and provide reads with lengths up to hundreds of kilobases (kb). Reads of such length can be exploited in various ways. Particularly in the genome assembly field they can be used for de novo assembly with long-read data only, or for scaffolding of NGS-based assemblies by bridging gaps between contigs or spanning long repeats thus resolving them. A major drawback of long-read technologies is the higher rate of sequencing errors (5–20%) compared to NGS data (<1%)2. Such an error profile could negatively affect the assembly accuracy, but because the errors are mostly randomly distributed the majority of long-read assemblers adopt the strategy of correcting base errors algorithmically before attempting to assemble the reads.
The most established long-read technology is from Pacific Biosciences (PacBio), which uses a sequencing by synthesis approach where phospholinked nucleotides are used to synthesize the complement strand of a single stranded DNA template. For the PacBio RS II machine, commercialized since 2011, the reads are characterized by lengths in the 5–60 kb range, with an average length around 12 kb4. The error rate for raw reads is about 13%4 with errors, mostly indels, randomly distributed. After a base error correction step the reads can reach an accuracy of 99.9% and higher across the read length. Throughput can reach up to about 1 Gigabases (Gb) per run4. Presently, the cost of sequencing a large genome with the PacBio technology is still quite high, as it involves a high initial cost for the platform (about $700k) and about $300 per Gigabases4. But PacBio has recently released a new platform, Sequel, that promises to increase the throughput to up to 10 Gb per run, maintaining a long average read length and decreasing the cost per Gb sequenced.
More recently, since 2014, the Oxford Nanopore Technology company (ONT) started distributing a new innovative sequencer, the MinION. The MinION is the first handheld sequencer and measures only 2 cm × 4 cm × 9 cm and weighs about 100 g. Within a MinION flowcell, single-stranded DNA molecules are guided across a membrane through protein-based nanopores. The membrane is immersed in a saline solution with a fixed voltage across it, so that an ionic current constantly passes through the pores. The DNA-strand motion through a pore causes a variation of the current, which is constantly monitored. A basecaller then determines the sequence of bases through the pore according to the observed variation in ionic current. To date, various basecallers are available: from an older Hidden-Markov-Model based (HMM) one to more recent ones based on Recurrent Neural Networks (f.i. Nanonet https://github.com/nanoporetech/nanonet). For the data analysed in this study, we used the HMM-based basecaller, which was the first made available by Oxford Nanopore. This basecalling package infers the sequence of successive k-mer words (5-mer or 6-mer) that passed through a particular pore by analyzing the current variations in time. According to their estimated qualities, the reads are collected into a ‘Fail’ or a ‘Pass’ directory, the latter only including the highest quality bi-directional (2D) reads, for which both strands (template and complement) of a DNA molecule have been sequenced and then merged in a single read. Like PacBio, the MinION sequencing errors are mostly randomly distributed. They can be corrected given enough read depth, with the exception of long sequences of the same base (homomers), that are often collapsed into shorter sequences by the basecaller. Nanopore sequencing is still in its rapid development stage, and since the data produced in this study many progresses have been made regarding error rate and throughput stability. Because of its portable size, low price ($1000 initial investment for the MinION, and <$300/Gb for flowcells bought in bundles), low computing requirements and relatively low wet-lab work load, the MinION has the potential to revolutionize the genomic discipline, in particular with regard to field and clinical sequencing. A number of preliminary attempts have already been reported, ranging from in situ outbreak analysis and control strategy5,6,7, field DNA collection and sequencing8, exploration of metagenomics for clinical use9,10,11, and many more (see https://nanoporetech.com/publications). Moreover, higher-throughput machines that incorporate 5 (GridION, https://nanoporetech.com/products/gridion) or 48 (PromethION, https://nanoporetech.com/products/promethion) flowcells are in early access stage, and they are foreseen to deliver throughput similar to that of Illumina HiSeq per run.
In this paper, we report a comparative study on the yeast genome assemblies from three different sequencing platforms: MiSeq from Illumina (NGS), and the long-read platforms PacBio RS II and ONT MinION. Apart from the reference yeast strain, Saccharomyces cerevisiae S288C, we also sequenced and assembled three other yeast strains, SK1, and the two Saccharomyces paradoxus N44 and CBS432. We explored the results in terms of accuracy, time and memory consumption of a variety of existing pipelines for long-reads-only de novo assembly, and for scaffolding of MiSeq-based assemblies using long reads. For the reference strain S288C, a comparison between the results from an ONT and a PacBio dataset with same read depth and similar read length distribution and accuracy is presented to assess the performance of the various pipelines in correcting the error types typical of the two technologies.
Results
Data Sequencing and Production
We have sequenced the genome of the four yeast strains, N44, SK1, CBS432 and S288C, with three different sequencing platforms: Illumina MiSeq, PacBio RS II, and MinION mk1. For each strain, we obtained about 80X depth of 2 × 150 bp MiSeq paired reads and between 120X and 250X depth from various PacBio runs. For the ONT runs, we obtained a total (‘Fail’ + ‘Pass’) 2D throughput (2D-All) between 12X and 61X depth depending on the strain, but only a throughput between 4X and 31X was achieved for the 2D-Pass data. Most Nanopore runs were performed using the R7.3 flowcells, available at the time of sequencing. For the reference strain S288C, the sequencing reads included two testing runs using the newly released R9 flowcell, in addition to the data of four runs from R7.3. The R9 flowcells generated data with higher accuracy than that of the R7.3 ones (91% against a R7.3 average of 88%). The R9 total 2D-All and 2D-Pass throughputs, 700 Mb and 60 Mb respectively, were too low for an independent study, so we merged the R7.3 and R9 reads to get a dataset of 61X depth 2D-All and a dataset of 31X depth 2D-Pass, upon which our assembly comparisons are based.
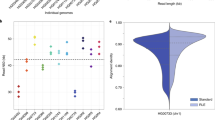
To compare the capabilities of the assembly pipelines we used the S288C strain data, so that assembly quality and accuracy could be easily and directly determined by mapping against its well known reference. Our S288C ONT and PacBio datasets have very different read depths: ONT only 31X (2D-Pass), while PacBio about 120X. By assembling the whole PacBio dataset we obtained contiguous assemblies with accuracy up to 99.95%, as shown in the Supplementary Note. For our pipeline comparison though, we decided to subset the PacBio dataset to the same depth of the ONT dataset, so that differences between platform results are driven by platform differences and not different sample sizes. If we selected the PacBio reads randomly until reaching the desired depth, the subset would follow a similar read length distribution of the original dataset, which has a smaller average read length than ONT. This would make it more difficult to interpret the assembly continuity as due to platform read intrinsic features like error rate and distribution, or different read lengths. Because of this, we decided to reduce the number of variables for assembly assessment by selecting a PacBio subsample of the S288C reads with the same depth (31X) but also similar read length distribution as the S288C 2D-Pass ONT reads: the PacBio ONT-Emu sample (Figure 1-(c)). More information on how we extracted the subset can be found in the Supplementary Note. To study the dependence of the assembly pipelines on read depth, we also selected two subsets with 10X and 20X depth for the S288C ONT and PacBio data, again with similar read length distributions. For the total PacBio S288C dataset, which consists of 120X read-depth, we performed a more detailed depth study shown in the Supplementary Note for subsets of 10X, 20X, 31X, 61X, 80X and the whole sample at 120X depth. Details about the sequenced datasets presented in this paper are summarized in Table 1 for ONT, and in Table 2 for PacBio samples which ‘emulate’ the ONT read length profiles (ONT-Emu). The read length distributions for the S288C, N44, CBS432 and SK1 strains of the 2D-Pass ONT datasets are shown in Fig. 1-(a) and those of the PacBio datasets in Fig. 1-(b).
Read length distributions for the ONT and PacBio datasets. Read length distributions for the four yeast strains, S288C, N44, CBS432 and SK1 of the 2D-Pass ONT datasets in (a) and the PacBio datasets in (b). Comparison of read length distributions for the S288C strain of the 31X datasets ONT 2D-Pass and PacBio ONT-emulating 31X-subset in (c).
De novo Assembly and Scaffolding Pipelines
It has been shown already12 that contiguous and accurate yeast assemblies can be generated de novo solely using ONT data. Here, we focus on assessing the various existing pipelines and on comparing the results obtained from ONT and PacBio data with particular attention to the similarities and differences between the two platforms. The long reads provided by PacBio and ONT can be used to generate a de novo assembly either by themselves or in conjunction with Illumina (NGS) data. In this paper, we show examples of assemblies from long reads only, and from’hybrid’ pipelines that use the Illumina reads either to correct the long reads or to generate a short-read assembly that is later scaffolded with long reads.
We selected eight assembly pipelines for long reads. PBcR13 (with Self- or MiSeq-based error correction: PBcR-Self, PBcR-MiSeq), Canu14 and Falcon15 are based on an Overlap-Layout-Consensus (OLC) algorithm; ABruijn16 is based on a generalized De-Bruijn graph algorithm. All of them include a base-error correction step on the reads before assembling them. SMARTdenovo (available from https://github.com/ruanjue/smartdenovo) is also based on a OLC algorithm, but does not include a base-error correction step. Miniasm17 chooses the best path from a string graph created from the overlap of uncorrected reads: it is the only pipeline here that includes neither a base-error correction nor a consensus step. Racon18 aligns the raw long reads to a Miniasm assembly and generates a consensus, significantly increasing the initial accuracy. We selected and assessed also three scaffolding pipelines: npScarf19, HybridSPAdes20 and SMIS (available from https://github.com/fg6/smis.git), to scaffold an NGS-based assembly from SPAdes21. Details about these pipelines and the parameters used for running them can be found in the Supplementary Note.
Assemblies from different pipelines and platforms
The de novo assemblies of the whole PacBio datasets for S288C and the other strains are shown and discussed in the Supplementary Note, while here we compare the performances of long-read assemblers when run on a 31X depth ONT or PacBio datasets with similar read length distributions. For this purpose we selected a subsample from the PacBio S288C dataset with the same depth as the ONT 2D-Pass sample (31X) and with similar read length distribution, as described above and shown in Fig. 1-(c). The statistic information for the two S288C samples can be compared if looking at the sample S288C ‘2D-Pass: 31X’ in Table 1 for the ONT case, and at the S288C ‘ONT-Emu 31X’ in Table 2 for the PacBio case emulating the ONT sample. As shown in these tables, the two samples also have similar average error rates (92% average identity for PacBio and 93% for ONT datasets). Using these two similar samples, except for normal fluctuations, differences in assembler’s performances can be attributed to the peculiar features of the reads from the two platforms, for instance to error types and distributions.
The assembly information for the ONT dataset ‘2D-Pass 31X’ and for the PacBio ‘ONT-Emu 31X’ can be found in Tables 3 and 4, respectively. With 31X depth the pipelines generated assemblies with similar features when running on PacBio or ONT data.
The fastest pipeline is Miniasm, which does not include a base error correction nor a consensus step. For both PacBio and ONT datasets it only took 4–5 minutes to run but only achieved 89% accuracy. Because of the high number of indels in the ONT and PacBio data, Miniasm reconstructed only about 95–96% of the genome, significantly less than the other pipelines. Globally, the assembly structure from the Miniasm pipeline is correct, indicating that the missing regions are due to local base errors. Part of these base errors are recovered by the other pipelines either by building a consensus from the raw reads (Racon, SMARTdenovo) or during their base-error correction steps (PBcR-MiSeq, PBcR-Self, Canu, Falcon and ABruijn). The assemblies with the highest accuracy, 99.94–99.97% are the ones from the only hybrid assembler, PBcR-MiSeq, that uses MiSeq reads to correct the ONT or PacBio reads. But PBcR-MiSeq also provided the most fragmented assemblies, with Na50s only 270 kb long, where Na50s are the N50s after breaking the contigs at the misassembly points found by Quast22. None of the other assemblers uses Illumina reads, and we refer to them as the non-hybrid pipelines. The non-hybrid pipelines reconstructed 98–99% of the genome, with Na50s in the 400–500 kb range long, and accuracy up to 98.76% for ONT reads and up to 99.93% for the PacBio reads. The highest accuracy between the non-hybrid assemblies for the ONT data is from Racon, with 98.76%, immediately followed by SMARTdenovo and ABruijn with about 98.50%. For the PacBio datasets the Celera-based assemblers (Canu, PBcR-Self) provided the highest accuracy, 99.92–99.93%, followed by ABruijn at 99.87% and Falcon at 99.78%.
To assess the completeness of the newly generated assemblies, we checked for the reconstruction of a list of known S288C genes. From the list of ORF coding sequences reported in the Saccharomyces Genome Database (http://www.yeastgenome.org), we selected 6,615 coding sequences that mapped to the S288C reference for at least 90% of their length with at least 90% mapping identity. We used BWA23 to align each gene against the new assembly under study, and declared a gene ‘reconstructed’ if at least 90% of it was assembled with at least 90% accuracy. Tables 3 and 4 for ONT and PacBio data respectively, show a correlation between the assembly continuity and the number of mapped genes, i.e. an assembly from a particular pipeline with longer contigs is likely to have reconstructed more genes. For example, the SMARTdenovo assemblies consist of 20 contigs when using 31X PacBio reads and 38 contigs when using 20X PacBio data, and the numbers of reconstructed genes are 6,596 and 6,534 respectively. For the ONT data at 31X, the contig number is 28, and it is associated with 6,556 genes; at read coverage of 20X, the contig number is increased to 29, while mapped genes dropped to 6,528.
Even though there is no single pipeline that outperforms at every statistic gathered, SMARTdenovo and Canu assemblies have the longest reference coverage, best or near-best average identity, highest number of genes found and long Na50s for both PacBio and ONT data. Apart from Miniasm, SMARTdenovo is also the pipeline using the least resources, with CPU running time ≤2 h and ≤5 GB of memory. Racon, Falcon and ABruijn are the next fastest pipelines. PBcR is the pipeline typically requiring the longest running times especially when using ONT data.
Missing homomers
Ignoring Miniasm which has no error correction nor a consensus step, and the hybrid pipeline PBcR-MiSeq, which corrects the reads using Illumina data, it is clear that the PacBio assemblies have higher accuracies, from 99.50% to 99.93%, while the ONT assemblies reach a maximum accuracy of 98.76%. Because the average read accuracy, the depth and the read length distribution of the ONT and PacBio datasets used here are very similar, the final higher accuracies for the PacBio assemblies are likely due to platform-intrinsic features of the data. When no base-error correction nor consensus step is performed, like in the Miniasm pipeline, the resulting assemblies are about 89% accurate for both the ONT and PacBio datasets, and present lots of mismatches and indels, with prevalence of indels. Adding a correction or a consensus step like the other non-hybrid pipelines significantly reduces the number of mismatches and indel errors for both ONT and PacBio data. While the number of mismatches is reduced to a similar level for ONT and PacBio, the number of indels in ONT assemblies remains very high. The ONT assembly with fewer indels (Racon, indels = 11 per kb) has about 11 times more indels than the best PacBio case (Canu, indels = 1 per kb), and about 3 times more indels than the worst PacBio case (Racon, indels = 4 per kb). The high accuracy reached with the PacBio data shows that most PacBio errors can be corrected by generating a consensus between the reads that cover the same genomic region, and is a clear indication that PacBio read errors are mostly randomly distributed. For the ONT data the situation is more challenging: while it is true that some of the errors have a random distribution and are corrected by the various pipelines, a good portion of them, mainly indels, seem to escape the correction attempts, pointing to a possible systematic source of errors. Indeed most of the remaining errors are due to missing homomers in the ONT reads. For instance, the assembly from Canu contains far fewer 5-homomers (“AAAAA”, “TTTTT”, “CCCCC” and “GGGGG”) than the reference, as shown in Fig. 2, where the blue bars represent the Reference’s counts and the orange bars Canu’s counts. The figure also shows that if the Canu assembly is polished with Nanopolish24 (green bars) many of the missing homomers are recovered, and the final assembly accuracy reaches 99.57%, as shown in Table 3 for the ‘2D-Pass 31X’ sample. This table also shows the downside of Nanopolish: it needs 1,835 CPU hours to polish the Canu assembly, which makes this pipeline impractical for larger genomes.
Mitochondrial genome reconstruction
While all of the 16 yeast nuclear chromosomes are well reconstructed by every tested pipeline with respect to continuity, accuracy and completeness, many pipelines failed to reconstruct the mitochondrial genome (Supplementary Tables S5 and S6). This chromosome is the smallest one in the reference assembly, only containing 85,779 bases, but appears to be very challenging to assemble, especially with ONT datasets. Using ONT data, only PBcR-MiSeq was able to reconstruct it almost completely (96%), and only Falcon and Canu were able to reconstruct at least half of it (67%, 64%, respectively). Using PacBio data there are three full reconstructions from Racon, SMARTdenovo and Canu, while Miniasm manages to reconstruct 77% of it. There are a number of possible reasons for the mitochondrial genome to be more challenging to reconstruct than the other chromosomes. The GC content in the mitochondrial genome is only 17%, much lower than the average value of 38% for the nuclear chromosomes. There are also a lot of highly repetitive small AT k-mers, which contribute to the difficulty of assembly.
We found that while for the ONT dataset the mitochondrial read depth was consistent with that of the other chromosomes, the PacBio mitochondrial depth was about 6 times higher which could help explain why the mitochondrial genome reconstruction is more complete with the PacBio data. To exclude the possibility that this is due to a bias in our subsetting, we looked at the whole PacBio dataset and found that, similarly to the subset, the mitochondrial genome depth is about 7 times higher than the nuclear chromosomes. This is likely due to the high copy number of mitochondria in a cell. Possibly, the PacBio library preparation preserves this higher copy number, while the ONT library preparation does not. For instance, it is possible that DNA molecules have been less vigorously sheared in the ONT case, so that many of the very short mitochondrial genome remained circular preventing the ligation of the adapters needed for sequencing.
In addition to a lower depth with respect to PacBio, the mitochondrial genome assembly from ONT data is also hindered by an higher than average homomer content: using the S288C reference, we estimated that the mitochondrial genome has at least 30% more homomers than the other chromosomes, for k-mer lengths 3, 4, 5, 6 and 7. Figure 3 shows the 5-homomer counts (#5A + #5T + #5G + #5C) ratios between each chromosome and the mitochondrial genome, each count normalized by the chromosome length in bases.
De novo Assembly by Varying the Read Depth
In order to assess the scalability of the assembly pipelines and to determine how the performances of the assemblers vary with read depth, we selected and analyzed subsets of the ONT and the PacBio full datasets. A dedicated study focusing only on the higher depth PacBio dataset is discussed in the Supplementary Note. Here, we compared the assemblies at 31X mentioned in the previous section, with 20X and 10X subsets from the ONT 2D-Pass and similar subsets from PacBio reads emulating the read length distribution of the ONT subsets, as we did for the 31X case. These datasets are presented in Tables 1 and 2 for ONT and PacBio respectively, where the PacBio subsets are labeled as ONT-Emu. The results of the pipelines are summarized for the 31X, 20X and 10X in Table 3 for ONT and Table 4 for PacBio data. At very low read depth, 10X, the hybrid pipeline, PBcR-MiSeq, was the best performing pipeline for both ONT and PacBio data providing assemblies which cover more than 92% of the genome with accuracies larger than 99.9%, and the longest Na50s, in the 100s of kb. At 20X depth the Na50 is about doubled, and 98–99% of the genes are reconstructed.
For the non-hybrid pipelines, the Celera-based ones, Canu and PBcR-Self, were the best performers at low read depths. At 10X, Canu and PBcR-Self performed better on ONT than PacBio data. These assemblies cover about 90% (75–79%) of the reference with an accuracy of 97% (99%) and a Na50 of 100 (60) kb with ONT (PacBio) data. All the other assemblies cover significantly less proportions of the reference genome, and in particular Falcon seems to have the most difficulties at such low read depth. At 20X, all the Celera-based pipelines still performed very well, with Canu and PBcR-Self providing Na50s in the range of 400s kb, as well as SMARTdenovo. The assembly from Racon distinguishes itself on ONT data for the highest accuracy, immediately followed by ABruijn and SMARTdenovo, while for PacBio data Canu, PBcR-Self and ABruijn assemblies have the highest accuracies. Already at 20X, Canu, PBcR-Self, PBcR-MiSeq and SMARTdenovo reconstructed about 99% of the genes. At 31X depth the non-hybrid pipelines increased the Na50s up to 500s kb and slightly improved their final accuracies.
Genome scaffolding using long reads
We explored the scaffolding performance of three pipelines that use long ONT/PacBio reads to bridge and merge contigs from a NGS assembly. The NGS assembly used here has been generated from a dataset of 80X of 2 × 150 bp MiSeq paired-reads by SPAdes, as one of the scaffolding pipelines is embedded with it (HybridSPAdes). As expected, the assembly from SPAdes is fragmented but has very high accuracy: it has 206 contigs and its N50 is only 125 kb, as shown at the top of Table 5. The scaffolding pipelines were able to bridge a number of contigs significantly increasing the assemblies’ N50s while maintaining a high accuracy for both the PacBio and ONT samples. The npScarf pipeline achieved N50s much longer than HybridSPAdes or SMIS, 771 kb for ONT and 715 kb for PacBio, but the Quast analysis of these assemblies revealed that they are affected by a number of misassemblies, and their final Na50 is 514 kb for the ONT data and 413 kb for PacBio. Even after correcting for the misassemblies npScarf provided the more contiguous assemblies, but the HybridSPAdes and SMIS ones were able to maintain a higher accuracy than npScarf, 99.97–99.98% against 99.91%. The assemblies from HybridSPAdes have a higher coverage over the reference and reconstruct more genes than SMIS or npScarf. The npScarf tool has the advantage of requiring the least resources as it is faster than the other two pipelines and it uses less memory.
We ran the same scaffolding pipelines on the three other yeast strains N44, CBS432 and SK1 with 2D-Pass ONT data at the lower depths of 11X, 9X and 4X, respectively. The scaffolding results are shown in Table 6. As a reference is not available for these strains, we could not evaluate the accuracy and misassemblies of the scaffolds, but, as for the S288C case, we can expect high accuracy, >99.95%, as these assemblies are based on Illumina data. The longer N50s from npScarf are likely significantly affected by misassemblies, as observed in the S288C case, while the N50s estimated by HybridSPAdes and SMIS are expected to be more accurate. A comprehensive structure variation analysis of these strains can be found in ref. 25.
Discussion
We have shown that the yeast genome can be de novo assembled with Na50s up to 550 kb with 31X read depth from PacBio or ONT platforms, reaching an accuracy of up to 99% for PacBio and 98% for ONT, when not using Illumina data. More fragmented assemblies but with an higher accuracy (up to 99.98%) can be achieved when using long reads in conjunction with Illumina reads in hybrid or scaffolding assemblies. Miniasm was the fastest pipeline, requiring only few minutes to run, but because of the lack of a correction or consensus step it generated assemblies with very poor accuracy (~89%). After Miniasm, SMARTdenovo is the pipeline requiring the least resources and in particular the least time for running, <2 h, but providing accuracies close to the highest one achieved without help from Illumina (98.5% for ONT and 99.7% for PacBio). The non-hybrid Celera-based assemblers, PBcR-Self and Canu, generated continuous and accurate assemblies, and appeared to be the most performing pipelines as the read coverage decreased.
In our experience, PacBio and ONT platforms provided reads with similar error rates and lengths in the thousands of bases. The average PacBio lengths were generally smaller than the ONT ones, but the ONT throughput per run was significantly lower than the one from PacBio’s runs. This was mainly because we used early MinION flowcells and chemistries: the throughput improved significantly both in size and stability with more recent MinION kits and software. In addition, a fairer throughput comparison should be done between two benchtop devices, e.g. the PacBio RSII and the Oxford Nanopore GridION, or PromethION, both not available at the time of this study. Both platforms provided error-prone reads, but missing homomers in the ONT data represent a major difference with respect to PacBio. Because of this difference, while PacBio reads could be corrected with enough depth to reach assembly accuracies >99.9%, for ONT data increasing the read depth only helped to reach an accuracy up to 98–99%. To improve further a very time consuming polishing step is needed, at least until the missing homomers issue is reduced or solved, possibly already with a new Neural Network-based basecaller, Scrappie, presently in development at Oxford Nanopore.
Methods
Library Preparation
Library preparations for Oxford Nanopore sequencing
Two ug of genomic DNA was sheared to approximately 18,000 bp by centrifugation at 4000 rpm in a gTUBE. Sequencing libraries were prepared according to the SQK-MAP006 or SQK-NSK007 Sequencing Kit protocol, including the NEBNext FFPE DNA repair step.
MinIONTM flow cell preparation and sample loading
The sequencing mix was prepared with 6 uL of the DNA library, water, the Fuel Mix and the running buffer according to the SQK-MAP006 or the SQK-MAP007 protocols. The sequencing mix was added to the R7 or R9 flowcell for a 24–48 hour run. Typically the flowcells were reloaded after 24 hours as data yield had plateaued.
Pacific Biosciences sequencing library preparation
PacBio sequencing libraries were prepared as follows. Five ug of genomic DNA was sheared to approximately 15,000 bp by centrifugation at 5200 rpm in a gTUBE. DNA was repaired with damage repair reagent and end-repaired using end repair mix before ligation to PacBio blunt end adapter. Unligated material was digested with Exo III and Exo VII then library fragments purified via two consecutive Ampure clean-ups and size selection on Blue Pippin (SageScience, Beverley, MA, USA) with a 0.75% agarose cassette to purify fragments from 12–25 kb.
Illumina PCR-free library preparation and sequencing
DNA (1 ug) was sonicated to a 400 to 600 bp size range using a Covaris LE220 acoustic shearing device (Covaris, Woburn, MA, USA). Fragments were end-repaired using the NEBNext EndRepair Module (New England Biolabs, Ipswich, MA, USA) and A-tailed with the NEBNext dA-Tailing Module. Illumina adapters were added using the NEBNext Quick Ligation Module. Ligation products were purified with AMPure XP beads (Beckman Coulter Genomics, Danvers, MA, USA). Libraries were quantified by qPCR using the KAPA Library Quantification Kit for Illumina Libraries (Kapa Biosystems, Wilmington, MA, USA) and library profiles were assessed using a DNA High Sensitivity LabChip kit on an AgilentBioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Libraries were sequenced on an Illumina MiSeq instrument (San Diego, CA, USA) using paired 150 base read chemistry.
Assembly Assessment and Other Tools
The ONT fastq sequencing reads were extracted from the fast5 files using Poretools26. For the assembly accuracy and other summary statistics we used dnadiff from MUMmer27, while the number of misassemblies and the Na50s, i.e. the N50s after breaking the contigs at the misassembly points, were calculated using Quast22. We ran Quast with default parameters except for the Miniasm assemblies. For Miniasm we required the additional parameter: –min-identity 85, to enable Quast to align the low accuracy assemblies to the reference. Because of such low accuracies, Quast seemed to overestimate the presence of misassemblies for Miniasm, and reported Na50s smaller than the expected ones if comparing the Na50s with those of the higher accuracy Racon assemblies, based on Miniasm. We used the R Biostrings package (available from https://bioconductor.org/packages/release/bioc/html/Biostrings.html) to count the number of homomers in some of the assemblies, using the function oligonucleotideFrequency. Statistics and assessment for all the assemblies have been estimated after eliminating contigs shorter than 1 kb.
References
Sanger, F., Nicklen, S. & Coulson, A. Dna sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. 74(12), 5463–5467 (1977).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Liu, L. et al. Comparison of next-generation sequencing systems. Journal of Biomedicine and Biotechnology 2012, Article ID 251364 (2012).
Glenn, T. 2016 NGS field guide: Overview. http://www.molecularecologist.com/next-gen-fieldguide-2016 (2016).
Quick, J. et al. Real-time, portable genome sequencing for Ebola surveillance. Nature 530, 228–232 (2016).
Hoenen, T. et al. Nanopore sequencing as a rapidly deployable ebola outbreak response tool. Emerg Infect Dis 22 (2015).
Faria, N. R. Zibra project: real-time sequencing of zika virus in brazil. https://nanoporetech.com/publications/zibra-project-real-time-sequencing-zika-virus-brazil (2016).
Parker, J., Helmstetter, A. J., Devey, D. S. & Papadopulos, A. S. T. Field-based species identification in eukaryotes using single molecule, real-time sequencing. bioRxiv (Cold Spring Harbor Labs Journals), doi:10.1101/107656 (2017).
Judge, K., Harris, S. R., Reuter, S., Parkhill, J. & Peacock, S. J. Early insights into the potential of the Oxford Nanopore MinION for the detection of antimicrobial resistance genes. J. Antimicrob. Chemother. 70, 2775–2778 (2015).
Greninger, A. L. et al. Rapid metagenomic identification of viral pathogens in clinical samples by real-time nanopore sequencing analysis. Genome Medicine 7, 99 (2015).
Schmidt, K. et al. Identification of bacterial pathogens and antimicrobial resistance directly from clinical urines by nanopore-based metagenomic sequencing. Journal of Antimicrobial Chemotherapy, doi:10.1093/jac/dkw397 (2016).
Istace, B. et al. de novo assembly and population genomic survey of natural yeast isolates with the oxford nanopore minion sequencer. bioRxiv (Cold Spring Harbor Labs Journals), doi:10.1101/066613 (2016).
Koren, S. et al. Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nature Biotechnology 30, 693–700 (2012).
Koren, S., Walenz, B., Berlin, K., Miller, J. & AM, P. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Research, doi:10.1101/gr.215087.116 (2017).
Chin, C. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nature Methods, doi:10.1038/nmeth.4035 (2016).
Lin, Y., Yuan, J., Kolmogorov, M., Shen, M. & Pevzner, P. Assembly of long error-prone reads using de Bruijn graphs. PNAS 113, 52, doi:10.1101/048413 (2016).
Li, H. Minimap and miniasm: fast mapping and de novo assembly for noisy long sequences. Bioinformatics 32(14), 2103–2110 (2016).
Vaser, R., Sović, I., Nagarajan, N. & Sikic, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Research, doi:10.1101/gr.214270.116 (2016).
Cao, M. et al. Scaffolding and completing genome assemblies in real-time with nanopore sequencing. Nature Communications 8, Article number: 14515 (2017).
Antipov, D., Korobeynikov, A., McLean, J. & Pevzner, P. hybridSPades: an algorithm for hybrid assembly of short and long reads. Bioinformatics 32(7), 1009–15 (2016).
Nurk, S. et al. Assembling genomes and mini-metagenomes from highly chimeric reads. Research in Computational Molecular Biology: 17th Annual International Conference, RECOMB 2013, Beijing, China, April 7-10, 2013. Proceedings 158–170 (2013).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. Quast: quality assessment tool for genome assemblies. Bioinformatics 29(8), 1072–1075 (2013).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with bwa-mem. arXiv: 1303.3997v2 [q–bio.GN] (2013).
Loman, N., Quick, J. & Simpson, J. A complete bacterial genome assembled de novo using only nanopore sequencing data. Nature Methods 12, 733–735 (2015).
Yue, J.-X. et al. Contrasting genome dynamics between domesticated and wild yeasts. Nature Genetics, doi:10.1038/ng.3847 (2017).
Loman, N. & Quinlan, A. Poretools: a toolkit for analyzing nanopore sequence data. Bioinformatics 30(23), 3399–3401 (2014).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biology 5:R12 (2004).
Acknowledgements
This work was supported by the Wellcome Trust (Grant WT098051). The authors would like to thank Oxford Nanopore Technologies for the opportunity to join the MinION early access program.
Author information
Authors and Affiliations
Contributions
R.D. and T.M.K. conceived the project; F.G. carried out the data analysis; J.X.Y., G.L. and J.L. provided DNA samples for sequencing; R.D., T.M.K., D.K.J. and Z.N. managed the project; L.A., M.A.Q. and P.C. conducted the experiments, including DNA extraction, library preparation and running the devices; R.M.D., J.K.B. and G.T. performed data quality assessment; F.G., Z.N. and M.A.Q. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The Wellcome Trust Sanger Institute is a member of the MinION Access Program (MAP) and received flowcells and sequencing kits free of charge from Oxford Nanopore Technologies. The authors declare no other potential conflict of interest.
Additional information
Accession codes: The ONT and MiSeQ data are available from the EBI database with Study accession code PRJEB19900. The PacBio data are available from the EBI database with Study accession code PRJEB7245 (See Supplementary Notes for more details). All the generated assemblies are available from the EBI Bio-Studies database with Study accession code S-BSST17. Scripts to download the final assemblies, the datasets and sample scripts to run the pipelines are available from GitHub: https://github.com/fg6/YeastStrainsStudy.git.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Giordano, F., Aigrain, L., Quail, M.A. et al. De novo yeast genome assemblies from MinION, PacBio and MiSeq platforms. Sci Rep 7, 3935 (2017). https://doi.org/10.1038/s41598-017-03996-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03996-z
This article is cited by
-
Revisiting chloroplast genomic landscape and annotation towards comparative chloroplast genomes of Rhamnaceae
BMC Plant Biology (2023)
-
Testing assembly strategies of Francisella tularensis genomes to infer an evolutionary conservation analysis of genomic structures
BMC Genomics (2021)
-
Consistent ultra-long DNA sequencing with automated slow pipetting
BMC Genomics (2021)
-
LazyB: fast and cheap genome assembly
Algorithms for Molecular Biology (2021)
-
Engineered yeast genomes accurately assembled from pure and mixed samples
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.