Abstract
To help health policy makers gain response time to mitigate infectious disease threats, it is essential to have an efficient epidemic surveillance. One common method of disease surveillance is to carefully select nodes (sentinels, or sensors) in the network to report outbreaks. One would like to choose sentinels so that they discover the outbreak as early as possible. The optimal choice of sentinels depends on the network structure. Studies have addressed this problem for static networks, but this is a first step study to explore designing surveillance systems for early detection on temporal networks. This paper is based on the idea that vaccination strategies can serve as a method to identify sentinels. The vaccination problem is a related question that is much more well studied for temporal networks. To assess the ability to detect epidemic outbreaks early, we calculate the time difference (lead time) between the surveillance set and whole population in reaching 1% prevalence. We find that the optimal selection of sentinels depends on both the network’s temporal structures and the infection probability of the disease. We find that, for a mild infectious disease (low infection probability) on a temporal network in relation to potential disease spreading (the Prostitution network), the strategy of selecting latest contacts of random individuals provide the most amount of lead time. And for a more uniform, synthetic network with community structure the strategy of selecting frequent contacts of random individuals provide the most amount of lead time.
Similar content being viewed by others
Introduction
Despite the medical advances, the outbreak of infectious disease is a persistent major burden to human health. For example, in 2009 the H1N1 influenza epidemic spread across 214 countries, causing an estimated economic loss of USD 3 trillion (nearly 5% of the global GDP) and leaving a death toll of 18,5001,2,3,4. More recently, the Zika virus outbreak—that started in the South and Central America and the Caribbean in 2015—is expected to cost an estimated USD 3.5 billion in these regions and lead to more than USD 1.5 million infections around the world in 20165,6,7,8. To reduce human suffering and economic loss, it is important to discover epidemic outbreaks early. For this purpose, health agencies have designed surveillance systems. These systems not only help to detect outbreak, but also help to forecast the future extent and duration of the outbreak9,10,11,12,13. The most important property of such systems is to identify an emerging outbreak as early as possible. It is a similar problem to network vaccination where the task is to find optimal individuals to vaccinate (or otherwise incapacitate with respect to the disease spreading). This problem is somewhat more well-studied in contact of temporal networks14,15,16. Optimizing the sentinel selection, however, puts an emphasis on early detection and is not concerned with the expected number of others that can be infected in an outbreak starting from a certain individual.
Surveillance strategies, which have been used in surveillance systems for early detection, fall into two general categories—those based on monitoring care-seeking behavior and those mapping out contact behavior. Surveillance systems based on the first category strategies try to monitor individuals who seek outpatient care in sentinel hospitals or online search engines and calculate the number of cases among those individuals. For example, some traditional surveillance systems, such as the Hong Kong Centers for Health Protection (HP) and the United Kingdom Health Protection Agency (HPA), are based on hospital systems. They monitor individuals who see a doctor in sentinel hospitals and report their incidence of infectious disease for early detection17. Some other surveillance systems, such as (currently defunct) Google Flu Trends, are based search engines queries. Such systems monitor users who search for health information online and provide real-time information on disease outbreaks17,18,19,20,21,22.
In contrast to care-seeking surveillance systems, the second contact-behavior category tries to monitor a set of individuals who can catch and transmit the disease much earlier than the average individual. Previous research has designed surveillance strategies for early detection of infectious disease based on topological structures of static contact networks. For example, ref. 23 presented a simple surveillance strategy monitoring friends of randomly selected individuals. They also use this method to detect the spread of flu at Harvard College. The epidemic situation in this strategy occurred about two weeks earlier than the strategy which selects individuals randomly to monitor. Moreover, to find the suitable scenarios for these network-oriented strategies, ref. 11 compared three surveillance strategies (selecting individuals with the highest degree, individuals randomly and friends of random individuals) on three classes of static contact networks (networks with power-law degree distribution, Poisson degree distribution and community structure). They found the optimal strategy depends on the network structures and the basic reproduction number of the disease. If a disease has a low basic reproduction number, the strategy of selecting individuals with the highest degree would provide the earliest information about both the onset and peak of an outbreak. However, if there is no prior knowledge of the network, a practical alternative is selecting friends of random individuals. Surveillance systems based on care seeking can identify an outbreak one or two weeks after the onset or peak of an outbreak or contemporaneously at best. Hypothetically, surveillance systems based on contact-behavior strategies could improve on this. So far such strategies have neglected the fact that contacts are not static24, 25. Contact networks not only have topological structures, but also temporal structures (e.g., cyclic ones—an individual meeting a friend on a daily basis)14.
The temporal structures of networks not only affect the speed of spread and the final outbreak size but also strategies to mitigate the outbreak16, 25,26,27,28. Indeed, the earlier studies have shown that vaccination strategies that exploit temporal structures outperform the neighborhood vaccination strategy to effectively control infectious disease14,15,16. To our knowledge, surveillance strategies that exploit temporal structures to select sentinels for early detection of infectious disease have not been investigated before. That is the goal of this paper.
In this study, we investigate whether vaccination strategies designed for static or temporal networks can serve as suitable strategies for selecting sentinel nodes. To address this problem, we compare two temporal-network strategies for selecting sentinels (sampling the last contact and most frequent contact with random individuals) with two static-network strategies (sampling random contact of random individuals and random individuals) on four temporal networks under a well-studied epidemic scenario (discussed below). Specifically, we first divide the time span of a temporal network into two time windows. In addition, we select sentinels based on different surveillance strategies in the first time window. Finally, we simulate the susceptible-infected (SI) epidemic model to evaluate different surveillance strategies based on lead time in the second time window. We test the efficacy of the surveillance strategies by measuring how long sampling the sentinels increases the lead time in epidemic simulations. The SI model models diseases with relatively long infection times compared to the dynamics of the outbreak.
Materials and Methods
Temporal network data set
In this section, we will present the temporal networks we use as models for the underlying network structure. We will discuss the nodes in this network as individual people. Most of the empirical networks we use are indeed networks of people. In practice, sentinels would rather be healthcare units (wards, clinics, hospitals), or farms (in the case of disease among livestock).
Our first network, Prostitution, was collected from a Brazilian online forum. This dataset includes N = 16,730 individuals and E = 50,632 sexual contacts spanning T = 2,232 days. The contact created at time t between a pair of individuals in the dataset represents the reported sexual encounter between a male sex-buyer and a female escort14, 29. The Dating network comes from an Internet dating community30. This dataset includes N = 28,972 individuals and E = 430,827 sexual contacts and spans T = 512 days. Contacts represent the initial interaction between two people (with the intention to pursue a future off-line relationship)14, 30. The E-mail data was collected from one of the main mail servers of a university31. This dataset includes N = 2,997 email accounts and E = 202,694 contacts and spans T = 82 days. The contact created at time t between a pair of email accounts in the dataset represents one email address sending or receiving a message at a particular time14, 31. In addition, we use a Synthetic model to generate the large-scale temporal networks with community structure to compare with the Prostitution and Dating networks. To generate this data, we use the following approach: First, we use the Lancichinetti-Fortunato-Radicchi model to generate a scale-free static network with community structure32. Then, we add time stamps for every edge in the network generated by the above step based on the varying activity model in refs 14, 33. Since this network model algorithm is stochastic, we average all values over five realizations of the algorithm. This dataset includes N = 20,000 nodes and average E = 1,186,200 edges and spans T = 2,000 time steps. The specific parameters we use in this construction are shown in Table 1.
We summarize some properties of our temporal networks in Table 2 and Fig. 1. The Synthetic entries in Table 2 and Fig. 1 show the average properties of the Synthetic model networks. Regarding the static network structure, the degree distributions of networks of accumulated contacts are qualitatively similar for all four datasets (see Fig. 1(a–d)). Regarding temporal structures, the four datasets all show a high burstiness (the phenomenon that human activities often happen in intense periods separated by periods of quiescence14, see SI text). However, they have different persistence (measured by the fraction of edges that is present both in the first and last 5% of the contacts by the Jaccard similarity coefficient14). The Prostitution and Dating networks are the most dynamic, in the sense that individuals are entering and leaving the system throughout the sampling period. The Email network is less dynamic than the first two networks. The Synthetic is by construction the most uniform in terms of activity. At the same time, this network exhibits not only a broad degree distribution structure, but also obvious community structure (similar as the other three networks).
The statistical characteristics of the temporal networks. In panels (a–d), we plot the probability density p as a function of degree k ((a) Prostitution network, (b) Dating network, (c) Email network and (d) Synthetic network). We plot results for the accumulated network of all contacts. In panel (e), we show the temporal statistics of the datasets—persistence and burstiness.
Surveillance strategies
A surveillance strategy is a method to select a surveillance set υ of sentinels. Here, we focus on two temporal-network surveillance strategies which refer to the immunization strategies on temporal networks14 and two static-network strategies. As mentioned, we divide the time duration of individual contacts T into two parts [0, 3T/4] and [3T/4, T] (Our results are roughly speaking stable in the interval T/2 < t* < 4T/5 when the fraction of individuals in the surveillance set ranges from 10% to 12%, here we settle for t* = 3T/4). The time window [0, 3T/4] is the experience period. We can use the information in this period as a guidance to the sentinel assignment. Then we use the later time window, [3T/4, T], for disease simulation and evaluation of the surveillance strategies. This ex ante methodology circumvents the unreasonable assumption of static network modeling that the past accurately predicts the future (i.e., that the network remains unchanged).
Another important issue is that it is infeasible to assume a complete knowledge of the network even during the experience period. Mapping out a social network is costly, time-consuming and inaccurate34, 35. At best, one can hope to get local information of the network in the neighborhood of some specific interviewees. For this reason, we consider only such local strategies and omit strategies based on the knowledge of the entire (or a large part of) the network. In the spirit of ref. 36, three of our strategies proceed by first inquiring a randomly selected individual about his or her contacts, and base the sentinel selection on that information. Our fourth strategy is a completely random reference strategy.
Our four strategies work as follows (also see Fig. 2):
An illustration of our surveillance strategies. The top diagram illustrates a temporal network of contacts over a time window. There are two time windows, separated at three quarters(round down) of the sampling time, one for training and one for disease spreading simulation. In Panels (a–d), each horizontal line represents an individual and the circles and vertical lines indicate the interaction. The colors of the lines are the same as the vertex colors. The first infected node is marked “seed” and the interviewee selected randomly for information gathering is marked “random”. The sentinel represents selected neighbor of the interviewee selected according to different strategies. In the Acquaintance strategy, the orange red and green nodes can also be selected as sentinels. In the time window for disease spreading, red circle marks infected individual. The epidemic model in this example is the SI model with 100% infection probability.
Recent
We select an individual randomly and add the most recent contact in [0, 3T/4] to υ. We repeat this process until we reach a fraction f of individuals in the surveillance set.
Frequent
We select an individual randomly and monitor his or her most frequently contacted neighbor in [0, 3T/4]. This process is then repeated until we have added a fraction f of individuals to υ.
Acquaintance
We select an individual randomly and a random neighbor (throughout the experience period [0, 3T/4]). We repeat this process until a fraction f of individuals belong to υ.
Random
We let υ be a random fraction f of individuals present in [0, 3T/4].
Disease-spreading model and lead time
We simulate the dynamics of epidemic spreading on temporal networks using the simplest compartmental model—the susceptible-infected (SI) model. Every individual is susceptible to the disease to start with. Upon meeting an infected individual a susceptible node can become infected. Even though this is an extreme situation (in particular with a 100% infection probability), it can provide interesting insights into the role of temporal and topological structure for disease spreading15. In this work, we study the SI model on temporal networks in time window [3T/4, T]. The infection is assumed to originate from one source randomly at time 3T/4. At each time step, each susceptible individual in contact with an infected will become infected with the disease’s per-contact infection probability β. The propagation process will stop when there are no susceptible individuals or the contact time series is exhausted.
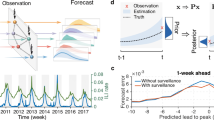
The insight from vaccination studies is that during an epidemic, some individuals could be more efficient spreaders than others. It would be no surprise if this property is correlated with the aptitude of being a sentinel. Fig. 3 gives an illustration of a disease simulation run. To estimate this impact, we ran SI model several times and calculated the average lead time ω υ
where T ρ is the time point at which the outbreak have reached 1% prevalence in the whole population and T υ is the time at which the prevalence within the surveillance set is 1%. This is about the percentage needed to infer an outbreak from sentinel surveillance11, 37, 38. A larger ω υ means that we have more time to respond and fight infection. Note that, first, if the individuals in the surveillance set υ are not infected during the time window [3T/4, T] in which the SI is simulated, then we set T ρ = T υ . Second, if the prevalence in the surveillance set υ does not reach 1% prevalence, then we set T ρ = T υ . Third, when T ρ < T υ , we set T ρ = T υ .
Results
Now we will turn to analyzing our numerical results.
Surveillance strategies evaluation
To evaluate our four surveillance strategies (two temporal-network surveillance strategies and two static-network strategies) on the four temporal networks, we start by analyzing the lead time obtained from simulating SI epidemic model. The main steps are summarized as follows. First, we divide the duration of a temporal network into two consecutive time windows [0, 3T/4] and [3T/4, T]. Second, at time 3T/4, we create a surveillance set υ which is composed of sentinels (because of the small node size of the Email network, the fraction of sentinels, f, ranged from 10% to 20%) based on different surveillance strategies in the time window [0, 3T/4]. Third, at time 3T/4, we simulate the SI epidemic model 10 times to calculate the average lead time ω υ , where the infection probability in the epidemic model is 100%. In Fig. 4, we plot the performance of the average lead time ω υ as a function of the fraction f of the individuals monitored for each surveillance strategy and for each temporal network. We find the Recent, Frequent and Acquaintance surveillance strategies yield similar results for the average lead time ω υ on the Prostitution network, Dating network and the synthetic network (we calculate the average lead time for five temporal networks generated by the Synthetic model). In the SI text, we plot the distribution of the lead time. All these three strategies provide significant average lead time to react to infectious diseases. For example, when f = 10%, the average lead time is about 20 days, 1.5 × 105 seconds and 13 time steps respectively. Though the decrease of ω υ is more gradual, the trend is that it saturates when f > 10%. For example, the average lead time is about 14 days, 1.35 × 105 seconds and 8 time steps separately when f = 20%.
The size of surveillance set impacts performance. How the lead time improves as the surveillance set expands in the (a) Prostitution network, (b) Dating network, (c) Email network and (d) Synthetic network. The surveillance set was chosen by the Random (black), Acquaintance (red), Frequent (green) and Recent (blue) strategies.
The Recent, Frequent and Acquaintance surveillance strategies can provide a substantial lead time on the email network when the surveillance set size is small. For example, the Recent strategy can provide approximately 706 seconds of lead time. The Acquaintance and Frequent strategies can provide approximately 1157 and 2119 seconds of lead time separately when f = 10%.
The Random strategy, which does not exploit any structure, shows the lowest efficacy on the four temporal networks.
To quantify the relative benefits of the strategies (Recent, Frequent), we plot the increase of average lead time with respect to the Acquaintance strategy, ΔΦ (Fig. 5). For example, ΔΦ = 10 if the strategy in question increases the average lead time by 10 time steps relative to Acquaintance strategy. For the Prostitution and Dating networks, the relative advantage of the Recent strategy is strongest. Frequent is similar to or better than Acquaintance strategy. For the Email network, the relative advantage of Frequent strategy is strongest when the surveillance set size is small. For the Synthetic network, the relative advantage of the Frequent strategy is strongest, followed by the Recent strategy. We use two artificial temporal networks to further investigate the effects of the temporal structure on surveillance strategies (see SI text).
The performance of the Recent and Frequent strategies relative to the Acquaintance strategy. The performance measure ΔΦ is the relative average lead time. The shadow regions indicate an improvement on Acquaintance (the more positive values, the better). The four panels correspond to the four datasets (a) Prostitution network, (b) Dating network, (c) Email network, (d) Synthetic network.
We conclude that the Recent and Frequent surveillance strategies both provide a beneficial lead time for early detection of infectious diseases on a network with a strong temporal structure. Moreover, the small f can provide the good effect. With f increasing, more inactive individuals or individuals with activity in the distant future may be selected as sentinels. Thus it decreases the lead time.
Effects of temporal correlations
In a temporal network, the structure in time can impact the dynamical processes in the same way as network structure can33. In order to explore the effect of temporal correlations, we consider using a temporal null model—Randomly permuted times (RP)—to remove correlations between consecutive contacts in original temporal networks. In the RP model, we fix the network’s static topological structures and the numbers of contacts between all pairs of individuals and only reshuffle the contact time stamp randomly15, 33. The main steps are summarized as follows: First, for each contact between a pair of individuals, we reallocate a random time stamp τ with uniform probability. In this method, we get the new list of contact events (i, j, t) describing the contact between i and j at time t. Second, if a contact (i, j, t) appears one or more times, we reallocate the random time τ based on the non-overlapping criterion in the randomization. Third, we apply the same surveillance strategies to this uncorrelated temporal network. Here we mainly focus on two classes of empirical datasets where the Prostitution and Dating networks are similar (we let Prostitution network represent this class of datasets) and the E-mail data shows a unique behavior.
Figure 6 shows the performance of the average lead time ω υ as a function of the fraction f of the population monitored for each surveillance strategy and for each type of randomized network (averaged over five generated networks). On the randomized Prostitution network, the curves in Fig. 6(a) are similar to Fig. 4(a,b). Only the lead time based on the randomized Prostitution network is longer than that based on the original network. For example, when f = 20%, the average lead time (the Recent, Frequent and the Acquaintance surveillance strategies) is about 52 days, whereas the average lead time is about 14 days on the Prostitution network.
The size of surveillance set impacts performance on two randomized temporal networks. The lead time as function of the fraction of sentinels for the (a) randomized Prostitution network and (b) randomized Email network. The colors for the surveillance strategies are the same as in Fig. 4.
On the other hand, for the randomized E-mail network, the curves in Fig. 6(b) are different from those in Fig. 4(c). The strategies (the Recent, Frequent and Acquaintance surveillance strategies) provide amount of lead time. For example, the average lead time is at best about 2.20 × 104 seconds.
We find the changes of the persistence and the burstiness of these two randomized temporal networks are small. The size of the epidemic increases when the temporal correlations are removed from the original networks. In this situation, the chance of selecting a highly active individual is higher with the Recent and Frequent strategies. From the above analyses, we conclude that the Recent, Frequent and Acquaintance surveillance strategies all also provide a significant lead time on a network where the temporal correlations are removed. Furthermore, we see that the temporal correlations presented in the original datasets limit the dynamics of epidemic transmission39.
Effects of the infection probability of disease
To investigate the robustness of our results, we observe the response of the average lead time ω υ to the fraction of surveillance individuals f and the disease’s per-contact infection probability β. We continue to focus on Prostitution network and E-mail network. The main steps of selecting surveillance sentinels and simulating epidemic transmission are almost the same to those discussed above. The only difference is that we run the SI model 10 times with different β-values different from one. In Figs 7 and 8, we plot the variation of the average lead time ω υ with the fraction f of the population monitored and the disease’s infection probability β on Prostitution and Email networks. We find that the amount of lead time depends on the fraction f and the disease’s infection probability β. For example, when f and β are small, the lead time is large. This demonstrates the small infection probability can slow down the speed of epidemic transmission and prolong the duration of the outbreak.
We find that when f and β are small, the relative advantage of Recent strategy is strongest on Prostitution network and the relative advantage of Frequent strategy is strongest on Email network.
We conclude that when the infectious disease is mild with respect to infection probability, we can sample a small fraction of individuals as sentinels exploiting temporal network structures to early detect the onset of infectious diseases.
Conclusion
To design surveillance systems for the early detection of infectious disease outbreak it is important to optimize the set of surveillance sentinels. The understanding of the network structure of individual contacts can help health policy makers to design the optimal surveillance strategy. The contact networks not only have topological structure but also temporal structure. We found that the two temporal-network strategies Recent and Frequent and the Acquaintance strategy provide significant lead time, especially when the network has a very dynamic nature. Further, the Recent is the best strategy for the Prostitution and Dating empirical temporal networks, whereas the Frequent is the better strategy for the Email and the Synthetic networks. In addition, the Recent, Frequent and the Acquaintance strategy provide a greater amount of lead time when the infectious disease is mild. Obtaining this result will help health policy makers design more efficient surveillance systems for the early detection of infectious disease in reality.
To further understand the effect of temporal structure and parameters in an epidemic model, we investigate the network’s temporal correlations and the disease’s infection probability. Specifically, we evaluate two temporal-network strategies and two static-network strategies on two representative networks (Prostitution network and E-mail network). The results show the amount of lead time depends on the temporal structure of networks and the disease’s infection probability. Specifically, the two temporal-network strategies and the Acquaintance strategy provide a greater amount of lead time on a network where the temporal correlations are removed. On the other hand, these three strategies provide a greater amount of lead time when the disease’s infection probability is lower.
This study is a first step effort to analyze the efficiency of the surveillance strategies based on temporal structures of contact networks for the early detection of infectious disease. This study leaves room for future investigations. For example, the factors need to be investigated to well explain the early detection phenomenon on the network where temporal correlations are removed. How the burstiness of the temporal networks affect the performance of the early detection. We mainly focus on whether the two vaccination strategies on temporal networks can serve as surveillance strategies. We will study more surveillance strategies and the relationship to temporal structures.
References
Organization, W. H. et al. Who recommendations for the post-pandemic period. Retrieved October (2010).
Sullivan, S. J., Jacobson, R. M., Dowdle, W. R. & Poland, G. A. 2009 h1n1 influenza. In Mayo Clinic Proceedings, vol. 85, 64–76 (Elsevier, 2010).
Liu, J. & Xia, S. Toward effective vaccine deployment: a systematic study. J Med Syst. 35, 1153–1164 (2011).
Chan, M. World now at the start of 2009 influenza pandemic (2009).
Mlakar, J. et al. Zika virus associated with microcephaly. N. Engl. J. Med. 2016, 951–958 (2016).
Fauci, A. S. & Morens, D. M. Zika virus in the americas—yet another arbovirus threat. N. Engl. J. Med. 374, 601–604 (2016).
Szabo, L. World bank offers $150 million to combat zika. http://www.usatoday.com/story/experience/2016/02/18/world-bank-offers-150-million-combat-zika/80567848/ (Accessed February 18, 2016).
Organization, W. H. et al. Zika situation report: Zika and potential complications (2016).
Khuwaja, S., Mgbere, O., Awosika-Olumo, A., Momin, F. & Ngo, K. Using sentinel surveillance system to monitor seasonal and novel h1n1 influenza infection in houston, texas: Outcome analysis of 2008–2009 flu season. J Community Health. 36, 857–863 (2011).
Yan, W.-R. et al. Establishing a web-based integrated surveillance system for early detection of infectious disease epidemic in rural china: a field experimental study. BMC Med Inform Decis Mak. 12, 4 (2012).
Herrera, J. L., Srinivasan, R., Brownstein, J. S., Galvani, A. P. & Meyers, L. A. Disease surveillance on complex social networks. PLoS Comput. Biol. 12, e1004928 (2016).
Fourquet, F. & Drucker, J. Communicable disease surveillance: the sentinel network. Lancet 349, 794 (1997).
Bai, Y., Du, Z., Yang, B. & Meyers, L. A. Location based surveillance for early detection of contagious outbreaks. In Adjunct Proceedings of the 2015 ACM International Joint Conference on Pervasive and Ubiquitous Computing and Proceedings of the 2015 ACM International Symposium on Wearable Computers, 77–80 (ACM, 2015).
Lee, S., Rocha, L. E., Liljeros, F. & Holme, P. Exploiting temporal network structures of human interaction to effectively immunize populations. PLoS One 7, e36439 (2012).
Starnini, M., Machens, A., Cattuto, C., Barrat, A. & Pastor-Satorras, R. Immunization strategies for epidemic processes in time-varying contact networks. J. Theor. Biol. 337, 89–100 (2013).
Liu, S., Perra, N., Karsai, M. & Vespignani, A. Controlling contagion processes in activity driven networks. Phys. Rev. Lett. 112, 118702 (2014).
Ginsberg, J. et al. Detecting influenza epidemics using search engine query data. Nature 457, 1012–1014 (2009).
Carneiro, H. A. & Mylonakis, E. Google trends: a web-based tool for real-time surveillance of disease outbreaks. Clin. Infect. Dis. 49, 1557–1564 (2009).
Boulos, M. N. K., Sanfilippo, A. P., Corley, C. D. & Wheeler, S. Social web mining and exploitation for serious applications: Technosocial predictive analytics and related technologies for public health, environmental and national security surveillance. Comput Methods Programs Biomed. 100, 16–23 (2010).
Lee, B. K. Epidemiologic research and web 2.0—the user-driven web. Epidemiology 21, 760–763 (2010).
Chew, C. & Eysenbach, G. Pandemics in the age of twitter: content analysis of tweets during the 2009 h1n1 outbreak. PLoS One 5, e14118 (2010).
Broniatowski, D. A., Paul, M. J. & Dredze, M. National and local influenza surveillance through twitter: an analysis of the 2012–2013 influenza epidemic. PLoS One 8, e83672 (2013).
Christakis, N. A. & Fowler, J. H. Social network sensors for early detection of contagious outbreaks. PLoS One 5, e12948 (2010).
Holme, P. Modern temporal network theory: a colloquium. Eur Phys J B. 88, 1–30 (2015).
Masuda, N., Klemm, K. & Eguluz, V. M. Temporal networks: slowing down diffusion by long lasting interactions. Phys. Rev. Lett. 111, 188701 (2013).
Rocha, L. E., Liljeros, F. & Holme, P. Simulated epidemics in an empirical spatiotemporal network of 50,185 sexual contacts. PLoS Comput. Biol. 7, e1001109 (2011).
Lloyd-Smith, J. O., Schreiber, S. J., Kopp, P. E. & Getz, W. M. Superspreading and the effect of individual variation on disease emergence. Nature 438, 355–359 (2005).
Shaw, L. B. & Schwartz, I. B. Enhanced vaccine control of epidemics in adaptive networks. Phys Rev E. 81, 046120 (2010).
Rocha, L. E., Liljeros, F. & Holme, P. Information dynamics shape the sexual networks of internet-mediated prostitution. Proc. Natl. Acad. Sci. USA 107, 5706–5711 (2010).
Holme, P., Edling, C. R. & Liljeros, F. Structure and time evolution of an internet dating community. Social Networks 26, 155–174 (2004).
Eckmann, J.-P., Moses, E. & Sergi, D. Entropy of dialogues creates coherent structures in e-mail traffic. Proc. Natl. Acad. Sci. USA 101, 14333–14337 (2004).
Lancichinetti, A., Fortunato, S. & Radicchi, F. Benchmark graphs for testing community detection algorithms. Phys Rev E. 78, 046110 (2008).
Holme, P. Temporal networks (Springer, 2014).
Bernard, H. R., Killworth, P., Kronenfeld, D. & Sailer, L. The problem of informant accuracy: The validity of retrospective data. Annu Rev Anthropol. 13, 495–517 (1984).
Holme, P. & Litvak, N. Cost-efficient vaccination protocols for network epidemiology. arXiv preprint arXiv:1612.07425 (2016).
Cohen, R., Havlin, S. & Ben-Avraham, D. Efficient immunization strategies for computer networks and populations. Phys. Rev. Lett. 91, 247901 (2003).
Thompson, W. W., Comanor, L. & Shay, D. K. Epidemiology of seasonal influenza: use of surveillance data and statistical models to estimate the burden of disease. J. Infect. Dis. 194, S82–S91 (2006).
Rath, T. M., Carreras, M. & Sebastiani, P. Automated detection of influenza epidemics with hidden markov models. In International Symposium on Intelligent Data Analysis, 521–532 (Springer, 2003).
Vazquez, A., Racz, B., Lukacs, A. & Barabasi, A.-L. Impact of non-poissonian activity patterns on spreading processes. Phys Rev Lett. 98, 158702 (2007).
Acknowledgements
The authors thank J.-P. Eckmann for help with the Email data. This study was supported by the National Natural Science Foundation of China (61572226) and the National Natural Science Foundation of China (61373053).
Author information
Authors and Affiliations
Contributions
Y.B. and B.Y. conceived and designed the experiments; Y.B. and L.L. performed the experiments. Y.B., Z.D. and J.H. analysed the results. Y.B. and B.Y. wrote the first draft of the manuscript. P.H. provided help for the experiments and writing. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bai, Y., Yang, B., Lin, L. et al. Optimizing sentinel surveillance in temporal network epidemiology. Sci Rep 7, 4804 (2017). https://doi.org/10.1038/s41598-017-03868-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03868-6
This article is cited by
-
Influence maximization on temporal networks: a review
Applied Network Science (2024)
-
Risk-aware temporal cascade reconstruction to detect asymptomatic cases
Knowledge and Information Systems (2022)
-
Relevance of temporal cores for epidemic spread in temporal networks
Scientific Reports (2020)
-
Analysis of E-mail Account Probing Attack Based on Graph Mining
Scientific Reports (2020)
-
Inter-urban mobility via cellular position tracking in the southeast Songliao Basin, Northeast China
Scientific Data (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.