Abstract
This paper explores the phylogeny of the delphacid subfamily Delphacinae based on nuclear ribosomal and mitochondrial DNA sequences of four genetic loci (16S rDNA, 28S rDNA, Cytochrome oxidase I and Cytochrome b). Maximum likelihood and Bayesian analyses yield robust phylogenetic trees. The topologies support the monophyly of Delphacinae and its basal split into three tribes, and provisionally support subdividing Delphacini into three clades, including a more broadly defined Numatina. The tribe Tropidocephalini is divided into two clades. In addition, Paranectopia is transferred from Tropidocephalini to Delphacini and Harmalia syn. nov. is regarded as a junior synonym of Opiconsiva. The genera Bambusiphaga, Megadelphax and Muirodelphax are found to be paraphyletic. The estimated time to the most recent common ancestor of Delphacinae is roughly at 90 million years ago in Late Cretaceous.
Similar content being viewed by others
Introduction
The planthopper subfamily Delphacinae Leach is the most speciose and economically important group in Delphacidae Leach1, 2. It comprises approximately 80% of all delphacid species. Members in this subfamily are terraneous and phloem-feeding, with 55 species known as pests on 25 crops, such as rice, maize, wheat, barley, sugarcane and bamboo3. Of these, 30 species are known as plant pathogen vectors, mostly viruses, except phytoplasmas in the Saccharosydnini Vilbaste3,4,5,6,7,8. Delphacids may damage plants directly by feeding and oviposition, or less directly by the introduction of saliva9, potentially leading to “hopperburn”, or by the introduction of plant pathogens10, 11. The delphacid planthoppers on rice, the “ricehoppers”, are among the most devastating pests on cultivated rice in southern China and northern Indochina. For example, the continuous outbreak of the white-backed planthopper, Sogatella furcifera (Horváth), has caused immeasurable rice yield loss in this region over the last decade12, 13, and extensive outbreaks of the brown planthopper, Nilaparvata lugens (Stål) and the small brown planthopper, Laodelphax striatellus (Fallén), in recent decades have caused significant losses in rice yields in China14, 15.
The early classifications of Delphacinae were diagnostic groupings based on morphology. Early works divided Delphacinae into three tribes (Tropidocephalini Muir, Alohini Muir and Delphacini Leach)16,17,18, or treated it as a subfamily without tribal subdivision19. Wagner recognized no tribes in Delphacinae20. Asche provided the first cladistic investigation of the Delphacidae1, 21, and revised it into seven subfamilies, with the largest subfamily, Delphacinae, including over 80% of described species. Asche’s Delphacinae was comprised of tribes Saccharosydnini, Tropidocephalini, and Delphacini. Asche’s Delphacini was expansively defined to include the Achorotilinae Wagner, Alohini Muir, Chlorioinae Wagner, Megamelinae Haupt and Stirominae Wagner of previous authors, putatively supported by an articulated suspensorium1, 22.
Thereafter, three other studies treated the higher classification of Delphacidae using morphological features23,24,25. Yang and colleagues explored the phylogeny of Asiracinae Motschulsky and Tropidocephalini using phenetic and cladistic methods, including Ugyops Guerin-Meneville (in Asiracinae) and nine Asian genera of Tropidocephalini23. These analyses showed that Ugyops was the most basal group, sister to Tropidocephalini, with the genus Tropidocephala Stål placed sister to the remainder of the tribe. Emeljanov utilized features of immatures to define a more broadly inclusive Delphacinae24. Emeljanov portrayed the Delphacinae as being a clade paraphyletic within the Asiracinae, and comprised seven tribes (Vizcayini Asche, Kelisiini Wagner, Stenocranini Wagner, Plesiodelphacini Asche, Delphacini, Tropidocephalini and Saccharosydnini), four of which were treated as subfamilies by Asche. Hamilton generally followed Emeljanov except he treated Kelisiini as a subtribe of Stenocranini, and Saccharosydnini as a subtribe of Tropidocephalini25.
Molecular investigations began with Dijkstra et al.26, 27, who examined delphacid phylogeny using mitochondrial DNA nucleotide sequence data from COI (Cytochrome oxidase I) and 12S rDNA, respectively. Although a limited number of delphacine taxa were included, Dijkstra et al. provided insight into resolving the phylogenetic relationships of Delphacinae. More recently, the more comprehensive phylogeny of Urban et al. presented the evolution of Delphacidae, based on DNA nucleotide sequence data from four genetic loci (18S rDNA, 28S rDNA, wingless and Cytochrome oxidase I) and 132 coded morphological characters28. Urban et al.’s Delphacinae comprised 93 species (89 species in Delphacini, two species in each of Saccharosydnini and Tropidocephalini) and the topology generally supported the higher classifications of Delphacidae proposed by Asche, Emeljanov and Hamilton, and suggested a rapid diversification of the Delphacini associated with host shifts to, and within, Poaceae, and specifically from C3 to C4 grasses.
Despite these studies, the phylogeny of the Tropidocephalini remains essentially unexplored. The tribe currently contains more than 180 species in 37 genera widely distributed throughout the Palaearctic, Afrotropical, Indomalayan, Australian, Neotropical and Pacific Regions, but most diverse in the Indomalayan Region29,30,31. The Tropidocephalini primarily feed on bamboos, with a few species feeding on other grasses30, 31. The Chinese Tropidocephalini, comprising about 90 species in 23 genera, represents the richest species diversity of this tribe worldwide, with most taxa restricted to south China. Hou & Chen explored the relationships among three Belocera Muir species, and subsequently among four genera of Tropidocephalini, as inferred from partial 16S rDNA gene sequences. These studies resulted in partly resolved relationships within the Tropidocephalini32, 33.
Within the Delphacini, Emeljanov placed 22 genera into his new subtribe Numatina Emeljanov based on the presence of an articulated suspensorium of the phallobase, implying that all other genera of Delphacini should be assigned in the nominal subtribe Delphacina Leach34. While not directly tested, this subtribal classification was considered doubtful by Urban et al. because Numatina (represented by a single taxon) was nested deeply within the Delphacina. In results presented by Urban et al., the Delphacini was “divided consistently into three major clades (plus some ‘intermediate’ taxa)”. However, the relationships were only partially resolved, just as it stated “the resolution of relationships within Delphacini will require the addition of new data sources, such as sequence data from other genes with a higher rate of mutation”.
At present, the fossil record for Delphacidae is very incomplete. Fossils of delphacid planthoppers explicitly reported so far show that the known fossils of Delphacidae are from the Paleogene and Neogene of North America and Europe, including four species (Amagua fortis Cockerell, Chloriona stavropolitana (Becker-Migdisova), Delphax rhenana Statz and Delphax senilis Scudder) in Delphacini of the Delphacinae and one species in Ugyopini (Serafinana perperunae Gebicki & Szwedo) of the Asiracinae2, 35. The estimation of divergence dates among living delphacid taxa could provide insights into the evolution of the Delphacinae.
The purpose of this study is to examine the phylogeny of the speciose subfamily Delphacinae with a wide range of taxonomic subsampling based on nucleotide sequences. To reconstruct the generic relationships, three mitochondrial genes were selected with a high rate of mutation, and one nuclear ribosomal gene to avoid the side effect of maternal inheritance. All genera in Yang et al. and Hou & Chen’s studies were included to evaluate the support for the classification of Tropidocephalini23, 32, 33. Redivision of the tribe Delphacini was tested by including 23 species from Urban et al. and expanded species sampling. In addition, the positions of several controversial genera (i.e. Paranectopia Ding et Tian, Miranus Chen et Ding, Harmalia Fennah) were investigated. The divergence time of Delphacinae was estimated based on the concatenated dataset.
Results
Sequence characterization
The multiple sequence alignments of protein-coding genes cox1 (Cytochrome oxidase I) and cytb (Cytochrome b) were unambiguous, in which alignment of cox1 gene contained gaps, but none of that interrupted or shifted the reading frame. Unlike the sequences of protein-coding genes, the sequences of ribosomal genes varied in length across the sampled taxa. Highly variable regions of 16S rDNA and 28S rDNA that differed in length were excluded from phylogenetic analyses due to extreme ambiguity of alignment.
For 28S rDNA, the result of sequence variation analyses showed the sequences were 718 bp in length, including 185 variable sites, 533 conservative sites and 138 parsimony-informative sites, the percentage of A+T was 47.1%. The 16S rDNA comprised a sequence of 492 bp with gaps. Failed DNA extractions or PCR amplifications contributed to the missing data. The protein-coding genes alignment consisted of 1125 sites, including 537 bp of cox1. The average content of A+T is much higher than the content of C+G in mitochondrial genes, which is in accordance with Simon et al.36. The detailed descriptive statistics for sequence segments are listed in Table 1. After aligning with outgroups, the length of 28S rDNA and 16S rDNA became longer by including gaps. The sequence alignments of each partition are listed in Supplementary Tables S1–S4.
The results of chi-square tests for base compositional homogeneity of each gene showed homogeneity among taxa. Little saturation was detected in all four alignments both in assuming a symmetrical and an asymmetrical topology. All sequence segments are useful for tree reconstruction.
Phylogenetic analyses
The combined data consist of nuclear ribosomal gene 28S rDNA and mitochondrial genes of 16S rDNA, cox1, cytb, by which we obtained a robust phylogenetic tree of Delphacinae (Fig. 1). Delphacinae was recovered as monophyletic (Bootstrap support values = 95, Posterior probability values = 1) and divided into three clades represented by Saccharosydnini, Tropidocephalini and Delphacini, respectively. Saccharosydnini and Tropidocephalini were recovered as sister groups.
Phylogenetic tree of Delphacinae obtained from maximum likelihood (ML) analysis based on concatenated data of genes cox1, cytb, 16S rDNA and 28S rDNA. ML bootstrap values and Bayesian posterior probabilities are indicated at internal nodes. Bootstrap values under 50 are replaced by “*”. The species acquired from Urban et al. were indicated with A, B, C as belonging to clade 1, 2 and 3 in their research, respectively.
Within Tropidocephalini, all genera except Bambusiphaga Huang & Ding were recovered as monophyletic. In the ML topology, Belocera and a new genus were recovered as sister groups, with the combined clade sister to Arcofacies Muir. However, the new genus was more closely related to Arcofacies in BI tree. Two sister groups, Tropidocephala and Arcofaciella Fennah; Epeurysa Matsumura and Carinofrons Chen & Li were recovered.
In the tribe Delphacini, three clades were recovered. Several genera within Delphacini were recovered as paraphyletic, including Muirodelphax Wagner, Megadelphax Wagner, Harmalia Fennah and Opiconsiva Distant. Sister groups that were recovered within Delphacini include Cemus Fennah and Ecdelphax Yang; Sogata Distant and Miranus Chen & Ding; Paradelphacodes Wagner and Nilaparvata Distant; Ishiharodelphax Kwon and Falcotoya Fennah; Metadelphax Wagner and Toya Distant; Sogatella Fennah and Distantinus (Distant) (as Matutinus)37. The genus Paranectopia (formerly assigned to Tropidocephalini) was grouped with Mestus cruciatus Ren & Qin in the Delphacini. The three Palearctic genera, Megadelphax, Ribautodelphax Wagner and Muirodelphax formed a clade, with Ribautodelphax derived within Megadelphax.
Monophyletic tests
The log likelihood score of best ML tree was 66657.05. Ten taxa recognized as monophyletic groups were compared against the best ML tree by SH and AU tests. The results support the monophyly of Delphacidae, Delphacinae and Delphacini. Also, the paraphyly of Opiconsiva was corroborated. The monophyly of Tropidocephalini was statistically rejected by AU test (P < 0.05). However, after the transference of Paranectopia to Delphacini, the monophyly of Tropidocephalini and Delphacini were both supported. Both AU and SH tests failed to reject the monophyly of Bambusiphaga, Megadelphax and Harmalia, even though they were not recovered as monophyletic in the best ML topology.
Divergence time estimation
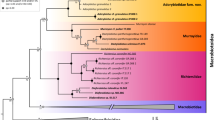
The divergence time chronogram of Delphacinae is presented in Fig. 2, with branch length as mean age. The divergence of Delphacidae from Cixiidae Spinola is approximately 211 Mya (million years ago). The estimated time to the most recent common ancestor of Delphacinae is roughly at 90 Mya. Saccharosydnini and Tropidocephalini split at some 82 Mya, slightly earlier than the origin of Tropidocephalini (75 Mya). The genus Tropidocephala originated at approximately 25 Mya.
Chronogram of Delphacinae divergence time estimated from the BEAST analysis with a Bayesian relaxed lognormal clock. The numbers at nodes indicate the mean ages and blue bars represent 95% highest posterior density intervals for the node ages. Two calibration points are indicated as *Cixiidae and *Delphacini.
Discussion
The present study reconstructed a relatively comprehensive phylogenetic tree of Delphacinae based on mitochondrial and nuclear gene sequences, which divide this subfamily into three distinct clades, representing the three tribes (Delphacini, Saccharosydnini and Tropidocephalini) of Delphacinae. In Tropidocephalini, the genus Lauriana Ren & Qin grouped with Malaxa Melichar, and thence to a clade contained Malaxella Ding & Hu, supporting the previous morphological studies of Ren et al.29. Belocera and Arcofacies are closely related and in accordance with the study of Yang23. The results also support the status of the Bambusiphaga fascia group, but failed recognize the monophyly of four other species groups included in the present study (B. nigropunctata group, B. citricolorata group, B. maculata group and B. mirostylis group)38, 39. In addition, B. maculata Chen & Li separated from the other members of the genus, sister to Specinervures basifusca Chen & Li. However, Bambusiphaga and Specinervures Kuoh & Ding are distinctly different morphologically (e.g., in Specinervures, the forewing with crossveins just proximad of midlength and trailing margin produced in distal half; ventrolateral margins of male pygofer strongly incised and gonostyli with inner angles reflected mesoventral apically40), it is unreasonable to transfer B. maculata to Specinervures based on current classification criteria. Thus, while the paraphyly of Bambusiphaga is here indicated, and further review of included taxa is needed to obtain a satisfactory classification. Furthermore, this study consistently divided the tribe Tropidocephalini into two major lineages, the first clade includes Malaxa, Lauriana, Specinervures, Bambusiphaga and Malaxella, another one contains the remaining genera in the tribe except Paranectopia. However, morphological diagnoses of these two clades require further study.
Although the monotypic genus Paranectopia was originally established in Tropidocephalini based on the post-tibial spur not having fine teeth along the posterior margin41, features of the male genitalia of Paranectopia lasaensis Ding et Tian show the aedeagus and anal segment are not in close functional contact, the suspensorium is present, and the aedeagal base is not twisted42, which meet the definition of the tribe Delphacini sensu Asche1. The assignment of Paranectopia in Tropidocephalini caused the monophyly rejected of Tropidocephalini in the AU test, and is accepted after deletion of Paranectopia. Here Paranectopia is transferred into Delphacini.
The subtribal classification of Delphacini can be evaluated using the ninety-one genera in Delphacini sampled here, including 23 species representing the “three clades” hypothesis in Urban et al.’s phylogeny28. Our results support the redivision of this tribe into three clades, but they compositionally differ from Urban et al. in that clade I comprises two genera, clade II includes 14 genera, and clade III holds 46 genera (Fig. 1). Furthermore, clade II has four (Bostaera Ball, Numata Busck, Cemus, Palego Fennah) of the 22 genera in subtribe Numatina as defined by Emeljanov34. In contrast, clades I and III do not include any Numatina. Therefore our results support the general concept of subtribe Numatina, although more broadly defined. However, the presence of Delphax Fabricius (the name-bearing genus of subtribe Delphacina) at the base of clade II (including the four genera in Numatina) presents difficulties. Hence, the resolution of relationships within Delphacini still requires more gene and taxon sampling.
This study included 23 species in Urban et al.’s phylogeny28, indicated by the letters A, B and C (in Fig. 1) to designate the three clades of that study. A includes Bakerella minuta Beamer, Pissonotus albovenosus Osborn, Pissonotus brunneus Van Duzee, Bostaera nasuta Ball and Delphax orientalis (Linnavuori) of clade 1 in Urban et al.’s study. B includes Liburniella ornata (Stål), Nothodelphax neocclusa (Muir & Giffard), Isodelphax basivitta (Van Duzee) and Javesella pellucida (Fabricius) of Clade 2 of Urban et al.’s study. C includes Caenodelphax teapae (Fowler), Muirodelphax arvensis (Fitch), Spartidelphax detecta (Van Duzee), Flavoclypeus nigrifacies (Muir), Spartidelphax penedetecta (Beamer), Delphacodes puella (Van Duzee), Dicranotropis hamata (Boheman), Harmalia ostorius Kirkaldy, Metadelphax propinqua (Fieber), Neomegamelanus elongatus (Ball), Opiconsiva sp., Prokelisia dolus Wilson, Prokelisia marginata (Van Duzee), Tagosodes orizicolus (Muir) of Clade 3 in Urban et al.’s study. This study placed Bakerella minuta in Urban et al.’s clade 2, consistent with Urban et al.’s MP analysis. Nothodelphax neocclusa was placed in clade 2 in Urban et al.’s analysis, but in the present study is allied with the Spartina-feeding clade within clade 3, rendering the clades 2 and 3 of Urban et al. paraphyletic. Therefore, sister group clades 2 and 3 of Urban et al. should be merged together as a monophyletic lineage.
Chen et al. established the genus Miranus Chen & Ding, for Stenocranus varians (Kuoh), formerly in Stenocraninae43, but was thereafter included in Delphacini44. This study revealed that M. varians (Kuoh) and M. serrulatus Dong & Qin formed a monophyletic group sister with Sogata Distant, supporting the assignment of Miranus species in Delphacini rather than in Stenocraninae. The genera Opiconsiva and Harmalia share many morphological synapomorphies and formed a clade (although in Opiconsiva the aedeagal base bears a dorsal longitudinally grooved structure), but Harmalia is deeply nested within Opiconsiva making it paraphyletic. Both the AU and SH test of monophyly for Opiconsiva are rejected. Harmalia is placed here as a junior synonym of Opiconsiva.
The monophyly of three genera Ribautodelphax, Muirodelphax and Megadelphax all established by Wagner in 1963 were tested. Each of these included two representative species. The topology among these taxa (Fig. 1) is (Megadelphax cornigera + Muirodelphax atratus) + (Megadelphax kangauzi + (Ribautodelphax bidentatus + Ribautodelphax tuvinus)), Muirodelphax arvensis is placed outside this clade sister to Elachodelphax metcalfi (Kusnezov), casting doubt on the monophyly of Muirodelphax, at least as comprised by Hamilton and Kwon45. The species R. bidentatus was once transferred to Megadelphax by Vilbaste (because the processes on the anal tube are not cruciate, a diagnostic feature of Ribautodelphax) but its placement in Ribautodelphax is supported in this study. Our analyses suggest that Ribautodelphax may be derived within Megadelphax, which is plausible given the many similarities among these genera. Nevertheless, we feel that the six species included in this study are not sufficient to solve the phylogenetic interrelationships among these genera, Hence, additional evidence and more taxa are needed to clarify their phylogeny.
The relaxed molecular clock analysis shows that the Delphacinae can be traced into the Late Cretaceous (90 Mya), and that the tribe Tropidocephalini also originated in the Late Cretaceous (75 Mya), with Saccharosydnini diverging from Tropidocephalini 82 million years ago. Moreover, these results indicate that the family Delphacidae can be traced into the Cretaceous (100 Mya), much before the earliest fossil record of this family (in the Paleogene). These observations are consistent with the idea that molecular estimates should be earlier than fossil ages. Nevertheless, Delphacidae originated in this study later than that estimated by Song and Liang (Early Cretaceous, around 129 Mya)46. The possible reasons for the disagreement of estimated divergence times include the delphacid fossil used as a calibration point to provide the minimum age for Delphacidae, and the greater taxon sampling of Delphacidae in this study. Furthermore, Delphacinae feed mostly on Poaceae, with a few species in Delphacini known as sedge feeders47, and a recent study has shown that the estimated age of the Poaceae ranged from 107 Mya to 129 Mya (in Cretaceous)48, earlier than the time estimation of Delphacinae in this study (90 Mya), which implies that the Delphacinae have undergone a rapid diversification after Poaceae evolved, indirectly supporting Wilson et al.’s hypothesis of delphacid adaptive radiation following the development of the grassland biome47. However, our divergence date estimates should be regarded as provisional, as more evidences, including fossil, morphological and molecular are still needed to support these results, particularly new fossil cross-calibrations.
A robust molecular phylogeny of Delphacinae was reconstructed in this study. The monophyly of Delphacinae was supported along with its basal split into three monophyletic tribes. The redivision of Delphacini into three clades, including a more broadly defined Numatina is also supported here. In addition, Paranectopia is placed in Delphacini and Harmalia syn. nov. is regarded as a junior synonym of Opiconsiva. Although the phylogenetic analyses elucidated many main points in Delphacinae, expanded data and greater taxon sampling are still needed to better understand the evolution of Delphacinae.
Materials and Methods
Taxon sampling
Specimens collected for this study were identified morphologically by the corresponding author and preserved in 100% ethanol at −20 °C in the Key Laboratory of Plant Protection Resources and Pest Management of Ministry of Education, Entomological Museum, Northwest A&F University (NWAFU). Sampling includes 123 ingroups and eight outgroups taxa (Supplementary Table S5). Ingroup sampling represents all three recognized tribes of this subfamily (Saccharosydnini, Tropidocephalini and Delphacini). Fifty-two ingroups nucleotide sequences were acquired from Genbank. Eight other species were selected as outgroups, including five species in Cixiidae, two species in Stenocraninae Wagner and one species in Kelisiinae Wagner.
Molecular data
DNA were extracted either from thoracic or leg muscle tissues using Qiagen DNEasy Kits (Qiagen, Inc., Valencia, CA, USA) or BioFlux Biospin Insect Genomic DNA Extraction Kit (Bioer, Inc., Hangzhou, China). The 28S rDNA, 16S rDNA, cox1 and cytb were amplified by using the previously reported oligonucleotide primers as Dietrich et al.49, Clary & Wolstenholme50, Simon et al.36 (Table 2). All polymerase chain reactions (PCR) were performed in 25 μl reaction volumes with the following cycling protocol: an initial denaturation step of 5 min at 94 °C, followed by 20–60 s at 94 °C, 35 cycles of 1 min at 49–52 °C, 1 min at 72 °C, ending with 7 min incubation at 72 °C. The PCR products were inspected in 1% agarose gel electrophoresis with ethidium-bromide staining. Sequencing was carried out with the same primers used for amplification on both strands. New sequences were submitted to GenBank (Supplementary Table S5).
Alignment
New sequences were proofread and aligned into contigs in BioEdit51. To detect contamination, each sequence was searched under BLAST on GenBank. Alignment was performed using MUSCLE as implemented in MEGA 652 with default options and checked manually.
The descriptive statistics for sequence segments were conducted in MEGA 6. Chi-Square test of homogeneity of each gene was performed in PAUP 4.0b1053. Substitution saturation of each sequence segment was tested using DAMBE 554 by comparing the index of substitution saturation (Iss) with critical values (Iss.c).
Phylogenetic analyses
One combined analysis of four segments was used for reconstructing phylogeny by using maximum likelihood (ML) and Bayesian inference (BI). Incomplete genes were included to increase the accuracy of phylogenetic analyses, especially for poorly supported nodes55, all gaps were treated as missing data.
ML analyses were implemented in PhyML 3.156 and RaxmlGUI 1.357 under the appropriate models. Tree topology search operations were conducted using SPR (Subtree Pruning Regrafting) moves. Bootstrap support values (BS) for nodes on the topology calculated were based on 1000 replicates.
Bayesian analyses were performed by simultaneously running two Monte Carlo Markov (MCMC) chains for 50 million generations in MrBayes 3.1.258. Each run has four chains, with trees sampled every 1000 generations. The average standard deviation of split frequencies and Potential Scale Reduction Factor (PSRF) were used for examining convergence. When stationarity was reached, the first 25% trees were discarded as burn-in and a consensus tree was obtained from the rest trees in the chain. Posterior probability values (PP) were considered as node support values. All tree topologies were displayed in Figtree 1.459, rooted using outgroups.
Models
PartitionFinder60 was used to determine the best-fitting model for each gene partition. The Bayesian information criterion (BIC) indicated that the GTR+I+G model was the best-fitting model for 28S rDNA, 16S rDNA and the position 1 and 2 of protein-coding genes. The most appropriate nucleotide substitution model for the position 3 was GTR+G. A mixed model Bayesian analysis was performed in Mrbayes, and GTR+I+G was conducted under ML analyses.
Monophyletic tests
Monophyletic constrained topologies of 10 taxa (Table 3) were reconstructed by the combined data using RaxmlGUI. Each topology was tested against the optimal ML tree calculated in RaxmlGUI respectively. The log-likelihoods of 10 topologies (shown in Table 3) were estimated with PAUP 4.0b10. The Shimodaira-Hasegawa (SH) test and the Approximately Unbiased (AU) test were done in the software CONSEL 0.1j61.
Divergence date estimation
BEAST 1.8.362 was used under an uncorrelated normal relaxed clock assumption and a speciation Yule process to estimate the divergence time. Two mitochondrial protein-coding genes were selected to ensure the equal mutation rates. Chains were run for 50 million generations, with sampling every 5000 generations. Tracer 1.6.0 was used to verify the posterior distribution and to ensure the effective sample sizes (ESSs) >200 from the Markov Chain Monte Carlo (MCMC) output. TreeAnnotator in the BEAST package was used to summarize tree data with ‘mean height’ and discarded the first 25% of trees as the “burn-in” period. The results were visualized in FigTree.
Fossils of Cixiidae and Delphacini were used as calibrations to estimate chronograms. A normal distribution at 210 ± 4 Mya was set for Cixiidae46, 63. In addition, the fossil of Delphax senilis dated from the lower Eocene Ypresian (56 Mya) to Lutetian (41.2 Mya) were used to calibrate the most recent possible origin of Delphacini, with a prior normally distributed around 48.6 ± 3.8 Mya35.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Asche, M. Zur Phylogenie der Delphacidae Leach, 1815 (Homoptera: Cicadina: Fulgoromorpha). Marburger Entomol. Publ 2, 1–910 (1985).
Bourgoin, T. FLOW (Fulgoromorpha Lists on the Web): a world knowledge base dedicated to Fulgoromorpha http://hemiptera-databases.org/flow (2016).
Wilson, S. W. & O’Brien, L. B. A survey of planthopper pests of economically important plants (Homoptera: Fulgoroidea) in: Proceedings of the 2nd International Workshop on Leafhoppers and Planthoppers of Economic Importance (eds. Wilson, M. R. & Nault, L. R.) 343–360 (Utah, 1987).
Arocha, Y. et al. Transmission of a sugarcane yellow leaf phytoplasma by the delphacid planthopper Saccharosydne saccharivora, a new vector of sugarcane yellow leaf syndrome. Plant Pathol. 54, 634–642 (2005).
Hogenhout, S. A., Ammar, E. D., Whitfield, A. E. & Redinbaugh, M. G. Insect vector interactions with persistently transmitted viruses. Annu. Rev. Phytopathol. 46, 327–359 (2008).
O’Brien, L. R. & Wilson, S. W. The systematics and morphology of planthoppers (Fulgoroidea) in: The Leafhoppers and Planthoppers (eds. Nault, L. R. & Rodriguez, J. G.) 61–102 (New York, 1985).
Wilson, S. W. Keys to the families of Fulgoromorpha with emphasis on planthoppers of potential economic importance in the southeastern United States (Hemiptera: Auchenorrhyncha). Fla. Entomol. 88, 464–481 (2005).
Yang, L., Chen, Y. & Chen, H. Notes on planthoppers infesting bamboo in Guizhou. J. Mt. Agri. Biol 18, 154–161 (1998).
Petrova, A. & Smith, C. M. Application of brown planthopper salivary gland extract to rice plants induces systemic host mRNA patterns associated with nutrient remobilization. PLoS One 10, e0141769 (2015).
Sōgawa, K. The rice brown planthopper: feeding physiology and host plant interactions. Annu. Rev. Entomol. 27, 49–73 (1982).
Backus, E. A., Serrano, M. S. & Ranger, C. M. Mechanisms of hopperburn: an overview of insect taxonomy, behavior, and physiology. Annu. Rev. Entomol. 50, 125–151 (2005).
Catindig, J. et al. Situation of planthoppers in Asia in: Planthoppers: New Threats to the Sustainability of Intensive Rice Production Systems in Asia (eds. Heong, K. L. & Hardy, B.) 191–220 (Los Baños, Philippines, 2009).
Cheng, J. A. Rice planthopper problems and relevant causes in China in: Planthoppers: New Threats to the Sustainability of Intensive Rice Production Systems in Asia (eds. Heong, K. L. & Hardy, B.) 157–178 (Los Baños, Philippines, 2009).
Cai, L. J. et al. Detecting rice stripe virus (RSV) in the small brown planthopper (Laodelphax striatellus) with high specificity by RT-PCR. J. Virol. Methods 112, 115–120 (2003).
Wang, Y. et al. Susceptibility to neonicotinoids and risk of resistance development in the brown planthopper, Nilaparvata lugens (Stål) (Homoptera: Delphacidae). Pest Manag. Sci. 64, 1278–1284 (2008).
Metcalf, Z. P. General Catalogue of the Hemiptera. Fascicle IV, Fulgoroidea, Part 3, Araeopidae (Delphacidae) 1–552 (Massachusetts, 1943).
Muir, F. A contribution towards the taxonomy of the Delphacidae. Can. Entomol. 47, 261–270 (1915).
Muir, F. On the classification of the Fulgoroidea. Ann. Mag. Nat. Hist 10, 461–478 (1930).
Haupt, H. Neueinteilung der Homoptera-Cicadina nach phylogenetisch zu wertenden Merkmalen. Zool. Jahrb. Abt. Syst. Oekol. Geogr. Tiere 58, 173–286 (1929).
Wagner, W. Dynamische Taxionomie, angewandt auf die Delphaciden Mitteleuropas. Mitt. Hambg. Zool. Mus. Inst 60, 111–189 (1963).
Asche, M. Vizcayinae, a new subfamily of Delphacidae with revision of Vizcaya Muir (Homoptera: Fulgoroidea)-a significant phylogenetic link. Bishop Mus. Occas. Pap 30, 154–187 (1990).
Bartlett, C. R. et al. Delphacid planthoppers of North America http://ag.udel.edu/enwc/research/delphacid/index.html (2016).
Yang, J. T., Yang, C. T. & Shao, K. T. Numerical taxonomic studies on the Asiracinae and Tropidocephalini (Fulgoroidea: Delphacidae) from Taiwan. Bull. Inst. Zool. Acad. Sin. 26, 215–230 (1987).
Emeljanov, A. F. On the question of the classification and phylogeny of the Delphacidae (Homoptera, Cicadina), with reference to larval characters. Entomol. Rev. 75, 134–150 (1996).
Hamilton, K. G. A. The planthopper genus Stenocranus in Canada: implications for classification of Delphacidae (Hemiptera). Can. Entomol. 138, 493–503 (2006).
Dijkstra, E., Rubio, J. M. & Post, R. J. Resolving relationships over a wide taxonomic range in Delphacidae (Homoptera) using the COI gene. Syst. Entomol. 28, 89–100 (2003).
Dijkstra, E., Slotman, M. A. & Post, R. J. Resolution of phylogenetic relationships of the major subfamilies of the Delphacidae (Homoptera: Fulgoroidea) using the mitochondrial ribosomal DNA. Insect Sci. 13, 167–177 (2006).
Urban, J. M., Bartlett, C. R. & Cryan, J. R. Evolution of Delphacidae (Hemiptera: Fulgoroidea): combined-evidence phylogenetics reveals importance of grass host shifts. Syst. Entomol. 35, 678–691 (2010).
Ren, F. J., Zheng, L. F., Huang, Y. X. & Qin, D. Z. Lauriana Ren & Qin, a new genus of the tribe Tropidocephalini (Hemiptera: Fulgoromorpha: Delphacidae) from China. Zootaxa 3784, 84–88 (2014).
Bartlett, C. R. A new genus of new world Tropidocephalini (Hemiptera: Delphacidae: Delphacinae), with the description of two new species. Entomol. News 120, 387–396 (2009).
Chen, X. S. & Tsai, J. H. Two new genera of Tropidocephalini (Hemiptera: Fulgoroidea: Delphacidae) from Hainan Province, China. Fla. Entomol. 92, 261–268 (2009).
Hou, X. H. & Chen, X. S. Evolutionary relationships among three Belocera species inferred from 16S rDNA sequences. Hubei Agr. Sci. 15, 3355–3357 (2012).
Hou, X. H. & Chen, X. S. Molecular identification of bamboo planthoppers (Hemiptera: Delphacidae) based on mitochondrial 16S rDNA gene sequences. Forest Res. 26, 65–69 (2013).
Emeljanov, A. F. A new genus of planthoppers of the family Delphacidae from Ethiopia (Homoptera, Cicadina). Zoosyst. Rossica 2, 93–96 (1993).
Szwedo, J., Bourgoin, T. & Lefebvre, F. Fossil Planthoppers (Hemiptera: Fulgoromorpha) of the world: An annotated catalogue with notes on Hemiptera classification. 1–199 (Polish Academy of Sciences, 2004).
Simon, C. et al. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 87, 651–701 (1994).
Bellis, G. A. & Donaldson, J. F. Reassessment of some of Kirkaldy’s Australian species of Delphacini (Hemiptera: Delphacidae: Delphacinae). Aust. J. Entomol. 1–14 (2016).
Chen, X. S. & Liang, A. P. Revision of the Oriental genus Bambusiphaga Huang and Ding (Hemiptera: Fulgoroidea: Delphacidae). Zool. stud. 46, 503–519 (2007).
Hou, X. H. & Chen, X. S. Oriental bamboo planthoppers: two new species of the genus Bambusiphaga (Hemiptera: Fulgoroidea: Delphacidae) from Hainan Island, China. Fla. Entomol. 93, 391–397 (2010).
Yang, J. T. & Yang, C. T. Delphacidae of Taiwan (1), Asiracinae and the tribe Tropidocephalini (Homoptera: Fulgoroidea). 1–79 (Taiwan Mus. Spec. Publ., 1986).
Ding, J. H. & Tian, L. X. Insects of Xizang. 229–232 (Science Press, 1981).
Chen, X. S. Notes on male Paranectopia lasaensis Ding et Tian (Homoptera: Delphacidae). Entomotaxonomia 21, 309–310 (1999).
Chen, X. S. & Li, Z. Z. Three new genera and four species of Delphacidae (Homoptera: Fulgoroidea). Acta Zootaxon. Sin. 26, 323–332 (2001).
Ding, J. H. Fauna Sinica Insecta Vol. 45, Homoptera Delphacidae. 529–530 (Science Press, 2006).
Hamilton, K. G. A. & Kwon, Y. J. Taxonomic supplement to “short-horned” bugs (Homoptera) of the Atlantic Maritime Ecozone in: Assessment of Species Diversity in the Atlantic Maritime Ecozone (ed. MacAlpine, D.) 421–431 (Ottawa, 2010).
Song, N. & Liang, A. P. A preliminary molecular phylogeny of planthoppers (Hemiptera: Fulgoroidea) based on nuclear and mitochondrial DNA sequences. PLoS One 8, e58400 (2013).
Wilson, S. W., Mitter, C., Denno, R. F. & Wilson, M. R. Evolutionary patterns of host plant use by delphacid planthoppers and their relatives in: Planthoppers: Their Ecology and Management (ed. Denno, R. F. & Perfect, T. J.) 7–113 (Springer US., 1994).
Prasad, V. et al. Late Cretaceous origin of the rice tribe provides evidence for early diversification in Poaceae. Nat. Commun. 2, 7 (2011).
Dietrich, C. H., Rakitov, R. A., Holmes, J. L. & Black, W. C. Phylogeny of the major lineages of Membracoidea (Insecta: Hemiptera: Cicadomorpha) based on 28S rDNA sequences. Mol. Phylogenet. Evol. 18, 293–305 (2001).
Clary, D. O. & Wolstenholme, D. R. The mitochondrial DNA molecule of Drosophila yakuba: nucleotide sequence, gene organization, and genetic code. J. Mol. Evol. 22, 252–271 (1985).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Swofford, D. L. PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods), Version 4. (Sinauer Associates, 2003).
Xia, X. DAMBE5: a comprehensive software package for data analysis in molecular biology and evolution. Mol. Biol. Evol. 92, 1–27 (2013).
Jiang, W., Chen, S. Y., Wang, H., Li, D. Z. & Wiens, J. J. Should genes with missing data be excluded from phylogenetic analyses? Mol. Phylogenet. Evol. 80, 308–318 (2014).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Silvestro, D. & Michalak, I. RaxmlGUI: a graphical front-end for RAxML. Org. Divers. Evol. 12, 335–337 (2012).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Rambaut, A. FigTree v1.3.1 http://tree.bio.ed.ac.uk/software/figtree (2009).
Lanfear, R., Calcott, B., Ho, S. & Guindon, S. PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 29, 1695–1701 (2012).
Shimodaira, H. & Hasegawa, M. CONSEL: for assessing the confidence of phylogenetic tree selection. Bioinformatics 17, 1246–1247 (2001).
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007).
Shcherbakov, D. E. Origin and evolution of the Auchenorrhyncha as shown by the fossil record in Studies on Hemiptera Phylogeny (ed. Say, T.) 31–45 (Lanham, 1996).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Nos 30970387, 31172126).
Author information
Authors and Affiliations
Contributions
D.Z.Q. and Y.X.H. conceived and designed the experiments. Y.X.H. and L.F.Z. performed the experiments. D.Z.Q. and Y.X.H. analyzed the data. D.Z.Q., Y.X.H. and C.R.B. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, YX., Zheng, LF., Bartlett, C.R. et al. Resolving phylogenetic relationships of Delphacini and Tropidocephalini (Hemiptera: Delphacidae: Delphacinae) as inferred from four genetic loci. Sci Rep 7, 3319 (2017). https://doi.org/10.1038/s41598-017-03624-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03624-w
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.