Abstract
Previous studies of the oligoacyllysyl (OAK) series acyl-lysyl-lysyl-aminoacyl-lysine-amide, suggested their utility towards generating robust linear lipopeptide-like alternatives to antibiotics, although to date, none exhibited potent broad-spectrum bactericidal activity. To follow up on this premise, we produced a new analog (C14KKc12K) and investigated its properties in various media. Mechanistic studies suggest that C14KKc12K uses a non-specific membrane-disruptive mode of action for rapidly reducing viability of Gram-negative bacteria (GNB) similarly to polymyxin B (PMB), a cyclic lipopeptide used as last resort antibiotic. Indeed, C14KKc12K displayed similar affinity for lipopolysaccharides and induced cell permeabilization associated with rapid massive membrane depolarization. Unlike PMB however, C14KKc12K was also bactericidal to Gram-positive bacteria (GPB) at or near the minimal inhibitory concentration (MIC), as assessed against a multispecies panel of >50 strains, displaying MIC50 at 3 and 6 µM, respectively for GPB and GNB. C14KKc12K retained activity in human saliva, reducing the viability of cultivable oral microflora by >99% within two minutes of exposure, albeit at higher concentrations, which, nonetheless, were similar to the commercial gold standard, chlorhexidine. This equipotent bactericidal activity was also observed in pre-formed biofilms of Streptococcus mutans, a major periodontal pathogen. Such compounds therefore, may be useful for eradication of challenging poly-microbial infections.
Similar content being viewed by others
Introduction
Facing the global crisis of multidrug resistant bacteria1,2,3,4,5,6, membrane active compounds (MACs) are earning a renewed attention for their potential to control infections7,8,9,10 by multiple mechanisms, including by affecting critical common bacterial processes such as communication11, 12 and virulence13, 14 at sub-inhibitory concentrations. Thus, although the molecular basis for these effects are relatively ill understood, various MACs are presently gaining interest for their potential to address antibiotic resistance challenges and promise to overcome infections while avoiding many of the known resistance mechanisms. Of particular interest are borderline-hydrophobic MACs which, at low micromolar concentrations instigate mild/transient membrane damages15,16,17, including partial loss of the transmembrane potential, believed to bare critical consequences on efflux function18 and expression of antibiotic resistance factors19. While borderline-hydrophobic MACs might exert a bacteriostatic mode of action at higher doses, outright-hydrophobic MACs tend to disrupt biological membranes structures abruptly20,21,22, which often culminates in a rapid bactericidal outcome, at or near the minimal inhibitory concentration (MIC)23, 24.
Many of the known antimicrobial peptides (AMPs) exert antibacterial activities over Gram-positive bacteria (GPB) and/or Gram-negative bacteria (GNB) through their MAC properties. Some, manage to breach the cytoplasmic membrane (CM) permeability barrier20, 25, 26 while others selectively increase the outer membrane (OM) permeability in GNB by perturbing the organization and function of the lipopolysaccharides (LPS) layer23, 27,28,29. Thus, owing to its hydrophilic attributes, the OM is often responsible for the low sensitivity of GNB to hydrophobic antimicrobials (that can be highly active on GPB), thereby further challenging the generation of broad-spectrum antibacterial compounds, particularly needed in poly-microbial infections, for instance.
A variety of strategies were proposed to generate chemical mimics of AMPs to alleviate complications associated with peptide drugs30, 31. AMP-mimics may also be beneficial in helping to improve understanding the complex molecular basis for AMPs function(s) owing to their relative molecular/structural simplicity. Namely, while charge and hydrophobicity represent most critical factors influencing AMP properties, it is often challenging to identify the respective optimal proportions for controlling potency or selectivity. For instance, excess hydrophobicity might promote self-assembly in aqueous solutions and consequently might reduce potency32, 33. Charge considerations are similarly intricate, namely due to the fact that the relative proportion of anionic phospholipids in bacterial membranes can reach 20–30% in GNB and up to nearly 100% in GPB34, 35. In this respect, the peptidomimetic approach using oligo-acyl-lysines (OAKs) seems particularly suitable for engineering high affinity antibacterial MACs22, 36,37,38 since OAKs composition consists exclusively of hydrophobic linear acyls and cationic lysine residues37,38,39, where the inherently simple and incremental nature of designed analogs provides a systematic tool for dissecting the relative importance of charge and/or hydrophobicity. In fact, OAK designs that concentrated on miniaturized sequences of the general formula: Acyl-lysyl-lysyl-aminoacyl-lysine-amide suggest its capacity to generate distinct MAC versions with rational antimicrobial behaviors. For instance, the inactive tetrapeptide H-KKc12K-NH2 became active upon conjugation of N-terminal acyls, gradually revealing increasing activity with increased acyl length. Thus, C8KKc12K or C10KKc12K remained essentially inactive40 unlike more hydrophobic analogs38, 41.
Here, we set out to verify the implied linear relationship (between potency and hydrophobicity) of this C-terminally amidated series by producing a previously untested intermediate analog, C14KKc12K (molecular structure depicted in Fig. 1a) and investigating its antibacterial activity, mode of action and potential application. Ultimately, the findings are discussed in the context of a series of analogs, CnKKc12K (n = 8, 10, 12, 14 and 16).
Molecular structure and organization in aqueous solution. (a) Structure of C14KKc12K (top), PMB (middle; R = 6-methyloctanoyl) and CHX (bottom). (b) Two fold dilutions of three OAKs were prepared in PBS and light scattering was measured after two hours incubation. Symbols: dotted line, C12KKc12K; solid line, C14KKc12K; dashed line, C16KKC12K. Results are from at least two independent experiments. Error bars represent the standard deviation. (c) Representative light scattering experiments showing the signal evolution immediately after adding 105 CFU/ml (top, S. aureus; bottom, E. coli), using the conditions described in panel (b) (OAK at 200 µM). Symbols: triangles, C12KKc12K; inverted triangles, C14KKc12K. The vast majority of data points standard deviations varied by <2%.
Results
As part of its biophysical characterization, the purified synthetic lipopeptide was first subjected to light scattering measurements in order to assess its potential for self-assembly in an aqueous environment, as compared with two analogs, C12KKc12K and C16KKc12K. Expectedly, the light scattered by these analogs at low concentrations and up to ~10 µM, displayed proportional amplitudes, where C14KKc12K was intermediate between C12KKc12K (lowest) and C16KKc12K (highest) (Fig. 1b) and started to deviate from linearity at a concentration range where both C12KKc12K and C16KKc12K were previously found to aggregate24, 41. At ~20 µM however, C14KKc12K revealed a sharp divergence since its light scattering pattern exhibited significantly higher amplitudes, likely to reflect its capacity to form supramolecular structures of larger sizes42. We also assessed the aggregation tendency in a more complex medium such as the supernatant of centrifuged saliva and found it to display an overall similar trend to that in PBS, only starting at somewhat lower concentrations (i.e., the respective critical aggregation concentrations values were 8 ± 1 and 23 ± 3 µM). Often, such self-assembly is deleterious to antibacterial potency of AMPs and OAKs24, 32, 33. Though, in the case of C12KKc12K and C16KKc12K, we previously used their respective unsaturated N-terminal acyls to show that the less hydrophobic counterparts (i.e., C12ω7KKc12K and C16ω7KKc12K)24, 43 displayed higher potencies, even as aggregates. The proposed explanation for this discrepancy was that the unsaturated analogs formed different structures where the self-assembled lipopeptides were not as tightly bound24, 43, thereby gaining the ability to dis-aggregate upon interaction with bacteria and the ability to exert their antibacterial activity, unlike the saturated counterparts. Consequently, we next verified the analogs propensities to disaggregate in presence of bacteria by monitoring their light scattering amplitudes. Figure 1c illustrates the rapidly reduced light scattered by C14KKc12K, indicating its greater tendency to disaggregate in presence of bacteria. While these findings raise interesting questions (such as whether higher bacterial concentrations and/or longer incubation periods will increase the OAK disassembly), the fact that C14KKc12K demonstrated a tendency for rapid disaggregation in presence of bacteria supports the rapid activity expected in certain applications (as exemplified later in this section). Collectively therefore, the data suggest that the OAK’s tendency for self-assembly in a biological milieu at high concentrations, might not interfere with its antibacterial activity.
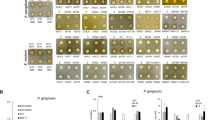
Next, we characterized its antibacterial activity by determining the MIC against a multispecies panel of 54 representative bacteria, including various medically relevant strains. Table 1 summarizes the MIC values obtained with 30 GPB (five species) and 24 GNB (seven species). With one exception (one of the Acinetobacter baumannii clinical isolates), the data reveal that C14KKc12K was active on most tested bacteria, although GPB appear generally more sensitive (MIC50 3 and 6 µM, respectively). Replacing LB with cation adjusted Mueller Hinton Broth resulted in essentially similar outcome. For instance, MICs of S. aureus 29213 or E. coli 25922 were 3 and 6 µM, respectively. Also note that C14KKc12K displayed an unchanged MIC value (3 µM) on both the wild type (AG100) and its efflux deletion-mutant (AG100A), unlike its analogs18 (this issue will be elaborated in the Discussion section).
To investigate the mode of action, we used standard methodologies for MAC characterization, including assessment of membrane damages and determination of time-kill kinetics over both Escherichia coli and Streptococcus mutans, respectively representing GNB and GPB. Figure 2a shows the OAK’s ability to affect viability of E. coli, reflecting a rapid bactericidal mode of action (e.g., >2-log-unit reduction within 2 hours exposure at ≥MIC).
Mode of action investigated against E. coli ML-35p as GNB representative. (a) Bactericidal kinetics upon exposure to C14KKc12K 0, 1, 2 and 4 MIC multiples (squares, circles, triangles and inverted triangles, respectively); dashed line, limit of detection (500 CFU/ml); asterisks denote lack of detected CFUs. (b,c) Membrane permeabilization, expressed as percentage of the positive control dermaseptin S4 (1–15) at 6.25 µM. Symbols: triangles, outer membrane; inverted triangles, cytoplasmic membrane. (c) Cytoplasmic membrane permeation to ethidium bromide. The inset shows representative permeation kinetics by the OAK at the MIC (triangles) and the positive control (inverted triangles); Min, minutes; F.U., fluorescence units (excitation: 535 nm, emission: 590 nm). Results are from at least two independent experiments performed in duplicate. Error bars represent the standard deviation.
To assess potential membrane damages, we used an assay capable of differentiating permeability changes in bacterial membranes by testing the leakage of small organic molecules. The assay employs the engineered E. coli strain, ML-35p, which is constitutive for cytoplasmic β-galactosidase, lacks lactose permease, and expresses a plasmid-encoded periplasmic β-lactamase44. The chromogenic β-galactosidase substrate ONPG is used to assess permeation of the CM, while OM permeability is assessed using nitrocefin, a chromogenic β-lactamase substrate. The data summarized in Fig. 2b suggest that both membranes were permeabilized at the MIC value (3 µM) albeit the OM appears more susceptible. To validate the CM damages at low concentrations, bacteria were subjected to another permeability assay, this time monitoring cytoplasmic access to the DNA binder, ethidium bromide (Fig. 2c). The fact that ethidium bromide accumulated in E. coli provides confirming evidence for the CM permeabilization at the MIC value. The inset shows two representative kinetic curves illustrating ethidium bromide’s rapid accumulation in bacterial cytoplasm. Combined, these findings support the view that bacterial death has resulted from the OAK’s capacity to abruptly disrupt both membranes. A similar mode of action was attributed to the natural bacteria-derived 11-residue cyclic lipopeptide, polymyxin B (PMB)45, an effect believed to stem from its high-affinity interaction with LPS. When compared, PMB and C14KKc12K exhibited a similar binding affinity to LPS originating from E. coli or Pseudomonas aeruginosa, as determined by their abilities to displace the binding of dansyl-PMB (Fig. 3). Under similar conditions46 the analogs C8, C10 and C12 exhibited a binding affinity that was lower than that of C14KKc12K but increased with increasing hydrophobicity40. This is further addressed below.
Dansyl-polymyxin binding assay. Interaction with LPS was assessed by incubation (1.5 hr) of C14KKc12K and polymyxin B with 2 μM pure monodansyl-polymyxin and 3 μg/ml LPS from E. coli (a) or P. aeruginosa (b). Symbols: triangles, C14KKc12K; inverted triangles, Polymyxin B; The Y axis represents fluorescence measurements (excitation: 340 nm, emission: 485 nm). Results are from two independent experiments performed in duplicate. Error bars represent the standard deviation.
Figure 4a shows the OAK’s ability to affect viability of S. mutans, reflecting, again, the OAK’s rapid bactericidal mode of action at low micromolar concentrations (e.g., ~2 log units reduction within 2 hours exposure to 1.56 µM). Membrane damages were evident from the rapid and massive leakage of protons and cytoplasmic accumulation of ethidium bromide (Fig. 4b and c, respectively).
Mechanistic studies (GPB). C14KKc12K mode of action was investigated against S. mutans ATCC 35668 as GPB representative. (a) Bactericidal kinetics upon exposure to 0, 1, 2 and 4 MIC multiples (squares, circles, triangles and inverted triangles, respectively); dashed line, limit of detection (500 CFU/ml); asterisks denote lack of detected CFUs. (b,c) Membrane damages instigated by C14KKc12K, expressed as percentage of the positive control dermaseptin S4(1–15) at 6.25 µM. Membrane depolarization (b) assessed by displacement of DiSC3(5), and membrane permeation (c) assessed by accumulation of EtBr. The insets in (b and c) show representative depolarization and permeation kinetics, of the OAK (triangles) and the positive control (inverted triangles) at 0.78 and 6.25 µM, respectively; Min, minutes; F.U., fluorescence units (excitation: 620 nm, emission: 680 nm in panel b; excitation: 535 nm, emission: 590 nm in panel c). Results are from at least two independent experiments performed in duplicate. Error bars represent the standard deviation.
To determine the applicative potential of C14KKc12K, we next aimed to exploit apparent OAK advantageous properties such as protease stability38, 39 and rapid bactericidal mode of action over a broad spectrum of bacteria. Hence, we performed a preliminary assessment for OAK’s ability to affect multiple bacterial species in saliva, given that the list of susceptible strains (Materials & Methods section) included various bacterial species known to promote periodontal diseases47, 48 whose treatment with an orally active antimicrobial has been the standard clinical approach. For this purpose, we elected to use S. mutans, a prototypical oral pathogen. Unlike other salivary floating bacteria, S. mutans can adhere to the oral cavity surfaces (particularly to teeth) and promote biofilm formation, which disturbs the balanced oral microbiome by pH reduction, culminating in dental complications such as caries and periodontal inflammations49,50,51.
Data shown in Fig. 5a confirms the OAK’s ability to maintain a rapid bactericidal activity in a complex medium, such as the supernatant from centrifuged human saliva52, albeit at the expense of higher doses compared with BHI medium. Furthermore, using whole saliva instead, the OAK has also maintained the capacity for rapid killing (within 2 minutes) of the multi-organism oral microflora, as efficiently as chlorhexidine (Fig. 5b). Chlorhexidine (CHX) is a cationic polybiguanide used since the 1970’s53 as a mouth-wash formulation (1–2 mM) to treat oral inflammations. CHX represents the gold standard reference in the field54 despite a few shortcomings such as dental/tissue discoloration and/or negative effects on taste55, 56. For these reasons its application is limited to short periods of time (about a week), and is usually employed as part of pre- or post-surgical interventions.
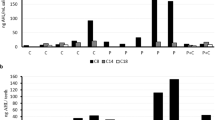
Potential oral application of C14KKc12K. (a) Bactericidal kinetics of C14KKc12K against S. mutans ATCC 35668 in the supernatant of centrifuged saliva. Symbols: squares, untreated control; triangles, C14KKc12K at 0.1 mM; inverted triangles, C14KKc12K at 1 mM; dashed line, limit of detection (500 CFU/ml); asterisks denote lack of detected CFUs. (b) Bacterial killing in whole saliva after 2 min treatment. Symbols: white bars, untreated control; gray bars, 0.1 mM, black bars, 1 mM; Con, untreated control; OAK, C14KKc12K; CHX, chlorhexidine. (c) Bactericidal kinetics against S. mutans ATCC 35668 upon exposure to chlorhexidine in BHI growth medium. Symbols: squares, untreated control; circles, 20 MIC; triangles, 40 MIC; inverted triangles, 60 MIC; dashed line, limit of detection (500 CFU/ml); asterisks denote lack of detected CFUs. (d) Bactericidal kinetics against S. mutans ATCC 35668 pre-formed biofilm upon exposure to 0.5 mM of C14KKc12K or chlorhexidine. Symbols: squares, untreated control; triangles, C14KKc12K; inverted triangles, chlorhexidine; dashed line, limit of detection (500 CFU/ml); asterisks denote lack of detected CFUs. Results are from at least two independent experiments performed in duplicate. Error bars represent the standard deviation.
In the literature, CHX MIC against S. mutans varies between 0.3 and 4 µM57 (0.3 µM, in our hands, representing a nearly 3-fold higher potency than C14KKc12K). However, when comparing their bactericidal kinetics (Figs 4a and 5c) CHX was bactericidal only at 40 multiples of its MIC (11.2 µM) i.e., at ~10 times the OAK’s bactericidal concentration (1.56 µM). While we are unable to explain this discrepancy, it is possible that these potency manifestations (i.e., the measure of inhibitory concentration versus time-kill kinetics) reflect an aptitude to compensate one for the other (likely related to mechanistic differences).
Moreover, noteworthy is the fact that, when compared at high concentrations (e.g., ≥0.5 mM) C14KKc12K and CHX were (again) equipotent in their ability to affect viability of S. mutans in a preformed biofilm, killing >99% of the initial massive inoculum (109 CFU/ml) within one hour of exposure (Fig. 5d).
Discussion
Antibacterial membrane-active lipopeptides are currently gaining extensive interest for their potential to affect critical bacterial processes ranging from communication8, 9 to antibiotic functions, in mono- and combination-therapy17. For example, PMB and daptomycin are two naturally occurring bactericidal cyclic lipopeptides, clinically used against GNB and GPB, respectively, with MIC values ranging between 0.12 and 8 µg/ml (PMB)58 or 0.015 and 32 µg/ml (daptomycin)59. In both cases, the acyl moiety plays a critical role45, 60.
Acyl conjugation was also shown to enhance the antimicrobial properties of various AMPs and synthetic mimics alike31, 61, 62. For instance, acylated derivatives of dermaseptin, a broad spectrum amphibian AMP, have drastically influenced its activity spectrum, enabling its conversion to specific activity on either GPB or GNB, depending on the acyl selected63, 64.
Other interesting studies reported synthetic ultra-short lipopeptides (3–5 residues) with potent activities against plant-related pathogenic fungi65 and bacteria66, though at higher doses67.
The present study provides evidence for the capacity of equivalent lipopeptides (generated via the OAK approach) to yield small molecules susceptible to be useful as simple investigation tools to help clarifying debated mechanistic aspects and potentially useful in biomedical applications. Namely, the findings reported herein, establish C14KKc12K as the shortest broad-spectrum antibacterial OAK known hitherto. The data also represent a quite remarkable yet ill-understood counter-intuitive outcome regarding the relationships between the hydrophobicity and self-assembly of this particular sequence. As expected from their respective elution time in HPLC using a hydrophobic column, C14KKc12K possesses an intermediate hydrophobicity value compared with C12KKc12K and C16KKc12K (Table 2). However, their tendencies for self-assembly, which presumably also depend on hydrophobicity, did not increase correspondingly, as evidenced by light scattering measurements (Fig. 1). Its sharp premature deviancy from linearity (as evidenced by high intensity) suggests that C14KKc12K forms drastically different aggregates in terms of three-dimensional organization. In contrast, our findings point to a dramatic increase in antibiotic efficiency since C14KKc12K achieved not only the lowest MIC value for this analog series (e.g., C14KKc12K MIC over S. mutans is 0.78 versus 1.56 µM for C12KKc12K or C16KKc12K) but has also upgraded its mode of action from bacteriostatic to bactericidal, including on GNB (Table 2). In this respect, the study expands and reinforces previous findings that illustrated the potential of N-terminal acyl conjugation to the core structure of AMPs63, 68, 69 or OAKs as a potent and versatile strategy for optimizing the hydrophobic/cationic balance required for antimicrobial properties.
The data also hint to a relationship between antibacterial potency and bacterial efflux function. Although so far, OAK properties were often rationalized strictly in terms of interactions with bacterial membranes, it is now clear that efflux70, 71 represents another decisive factor whose contribution to the mode of action must be accounted for. Previously, we proposed that related but borderline-hydrophobic analogs (e.g., C12ω7KKc12K18 or C10KKc12K40) are substrates of the AcrAB-TolC system, the resistance nodulation division (RND) efflux pump present in E. coli and various enteric bacteria. The fact that C12ω7KKc12K was potently active on isogenic mutant strains (whose efflux-pump components were deleted) supports the view that inactivity on normal GNB stems from the OAK’s rapid extrusion by these pumps18. Additional support for this view comes from the fact that bactericidal OAKs are also prone to deeper insertion within the CM17. In this respect, the fact that C14KKc12K is equally potent on both the wild type (AG100) and its deletion-mutant strain (AG100A) reinforces this hypothesis. Combined, our findings hint to a scenario implicating a simultaneous/competitive attraction of the OAK molecules in the periplasm, to the CM anionic phospholipids and the membrane-embedded efflux pumps. Consequently, borderline-hydrophobic OAKs would be more susceptible to extrusion, whereas outright-hydrophobic OAKs (e.g., C14KKc12K) are more likely to escape extrusion due to their tighter/deeper anchoring within the CM.
Considering the OAK properties in the context of a series of analogs (as outlined in Table 2), the resulting perspective provides an overall rational picture describing a bell-shaped continuum of effects that exacerbate (possibly through accumulation) with increasing hydrophobicity. Thus, the least hydrophobic among the tested lipopeptides (C8KKc12K) was clearly devoid of growth inhibitory activity against GNB but managed to induce mild damages, mostly to the OM40 without affecting bacterial proliferation. While these effects are readily reparable, they may cause some delay in initial bacterial doubling time upon minor hydrophobicity increase (as observed with C10KKc12K) where they start affecting the CM as well, as expressed by partial depolarization40. Additional increase in hydrophobicity (to yield C12KKc12K) further intensifies these effects to cause a variety of non-reparable damages to both GNB and GPB as expressed by a full-fledged growth inhibitory activity (i.e., MIC). These damages are nonetheless not severe enough to jeopardize bacterial viability, hence often leading to a bacteriostatic mode of action, as observed previously40 and in this study. Owing to its optimal hydrophobicity level, C14KKc12K was able to induce the most severe damages that culminated in rapid bactericidal outcomes. In contrast, excess hydrophobicity (e.g., in C16KKc12K24) leads to the formation of tight peptide aggregates, which in turn, inverse the MAC potency-trend by limiting OAK’s availability for optimal interactions with bacterial targets.
In conclusion, our results support the view that N-terminal acyl-manipulations of the core sequence KKc12K, exhibit straightforward structure-activity relationships. Thus, simply by switching the N-terminal acyl, the OAK properties became tunable, gradually evolving from lack of “visible” antibiotic activity on to exerting bacteriostatic activity over GPB only, and ultimately, exercising broad-spectrum bactericidal activity. The fact that C14KKc12K was equally potent on both wild type and efflux mutant strains suggests that stronger anchoring within the CM enables hydrophobic MACs to escape extrusion by RND pumps, thereby providing a rational for the observed increased potency.
Besides their potential role as investigation tools, such compounds may be useful in treating infections involving multiple microbial populations, such as oral mucositis. Our findings may have relevance to various biofilm-associated micro-environmental niches that hamper drug efficacy in infections or industry related issues. Future studies might clarify this issue. Interestingly, a 35-residue-long MAC currently in phase 2 clinical trials, C16G272, 73 displayed specific antistreptococcal bactericidal properties in saliva. In this respect, the OAK platform might present advantages in the capacity to generate superior anti-biofilm candidates, including in terms of biological robustness, simplicity and production costs.
Materials and Methods
Peptide synthesis
OAKs were synthesized in-house (433 A Peptide Synthesizer; Applied Biosystems, Foster City, CA, USA) by the solid-phase method using 9-fluorenylmethyloxycarbonyl (Fmoc) active-ester chemistry on 4-methylbenzhydrylamine (MBHA) resin. OAKs were then deprotected and cleaved from the resin using trifluoroacetic acid:H2O (95:5) and purified to >95% chromatographic homogeneity by reverse phase high performance liquid chromatography (RP-HPLC) using C18 column (Vydac), a flow rate of 2 ml/min and a linear acetonitrile gradient of 1%/min (Alliance; Waters, Milford, MA, USA). Peaks identity was verified by mass-spectrometry (Xevo G2 Tof; Waters, Milford, MA, USA). Purified OAKs were then lyophilized and kept as dry powder at −20 °C.
Organization in solution
To assess the OAK’s self-assembly in solution, serial two-fold dilutions of the OAK (initial concentration of 200 µM) were prepared in phosphate buffered saline (PBS; 10 mM Na2HPO4, 154 mM NaCl, pH = 7.4) and incubated for 2 hr at room temperature (RT). Light scattering at a 90° angle was measured through a 1 nm slit while holding both excitation and emission at 400 nm (Spectrophotometer Fluorolog-3 FL3–22; Horiba Jobin Yvon, Edison, NJ, USA).
To evaluate the disassembly of these aggregates upon bacterial exposure, an OAK solution (200 µM) was incubated (2 hr in PBS, at RT) after which, bacteria were added (105 CFU/ml) and the light scattering evolution of these suspensions was monitored as described above.
Bacteria
Gram-positive bacteria tested were: American Type Culture Collection (ATCC) strains Staphylococcus aureus 25923, 29213, MRSA 39592, 43300, BAA-43 (HSJ 216), BAA-1720 (252), S. epidermidis 12228, S. xylosus 29971, Enterococcus faecalis 29212, E. faecium 35667, Bacillus subtilis 33677, B. cereus 11778, Listeria grayi 19120, L. innocua 33090, L ivanovii 19119, L. monocytogenes 19115, L seeligeri 35967, L. welshimeri 35897, Streptococcus agalactiae 13813, 27956, S. bovis 9809, S. mutans 35668, 700610 (UA159), S. pneumoniae 49619, 6303, S. pyogenes 19615, S. sobrinus 27352 (6715) and clinical isolates MRSA 10017 (USA300), 15903, VRE Nu28.
Gram-negative bacteria tested were: ATCC strains Escherichia coli 25922, 43894, Pseudomonas aeruginosa 27853, 9027, Acinetobacter baumannii 19606, A. caloaceticus 31299, Salmonella Typhimurium 14028, Fusobacterium nucleatum 23726, Porphyromonas gingivalis 53977, clinical isolates E. coli 14182, 14384, U-16327, U-16329, P. aeruginosa 1278, Klebsiella pneumoniae 1287, K2–224, C2, A. baumannii 1279, 1280, 1281, F. nucleatum PK 1594, the engineered E. coli ML-35p and the isogenic K-12 pair AG100, AG100a (ΔacrAB). Dental related bacteria (generous gift of Prof. Doron Steinberg and Gilad Bachrach from the Hebrew University of Jerusalem) appear in bold characters.
Culture conditions
Staphylococci, Bacilli, Escherichia, Pseudomonas, Acinetobacter, Salmonella and Klebsiella species were grown in Luria Bertani broth (LB; 5 gr/l NaCl, 5 gr/l yeast extract, 10 gr/l tryptone). Enterococci and E. coli ML-35p were grown in Tryptic Soy Broth (TSB). Listeria and Streptococci were grown in Brain Heart Infusion (BHI). All bacteria were grown over-night at 37 °C with shaking. S. mutans 35668 plated on BHI agar for enumeration was grown for 48 hr. S. mutans UA159 was grown in 5% CO2 enriched atmosphere. Fusobacterium and Porphyromonas were grown in Wilkins-Chalgren growth medium and an anaerobic atmosphere.
Minimal inhibitory concentration (MIC) was determined using the microdilution assay. Mid-log-phase bacteria at 5 × 105 CFU/ml were incubated in a 96-well plate with serial two-fold dilutions of the tested compound for 18–24 hr at 37 °C (final volume of 200 µl). O.D. at 620 nm was measured (Synergy HT, BioTek Instruments, Winooski, VT, USA), and the MIC was determined as the lowest concentration for which no increase in O.D. was detected.
Bactericidal kinetics was assessed by incubating 5 × 105 CFU/ml of mid-log-phase bacteria with the OAK for 3 hr at 37 °C with shaking. Aliquots were taken at t = 0, 0.5, 1, 2 and 3 hr, subjected to serial 10-fold dilutions in saline (NaCl = 0.85%) and plated for enumeration after 24–48 hr incubation at 37 °C.
Cytoplasmic membrane permeation to ethidium bromide (EtBr) was evaluated as follows: Mid-log-phase bacteria at 1 × 108 CFU/ml were centrifuged for 5 min at 15,000 g. Pellet was washed twice with PBS containing 0.5% glucose (pH = 7.4), suspended in the same buffer and incubated for 10 min at 37 °C with shaking. 180 µl of the bacterial suspension were mixed in a 96-well plate with 25 µl of the tested compound and EtBr (EtBr final concentration = 1 µg/ml) and fluorescence was recorded immediately (excitation: 535 nm, emission: 590 nm) for up to 30 min at 37 °C with shaking (Synergy HT, BioTek Instruments, Winooski, VT, USA).
Outer and cytoplasmic membrane permeation in Gram-negative bacteria was assessed using the engineered E. coli ML-35p by monitoring the chromogenic hydrolysis of two indicators: ortho-nitrophenyl-β-galactoside (ONPG) and nitrocefin. Mid-log-phase E. coli ML-35p were centrifuged for 5 min at 15,000 g and the supernatant was removed. Pellet was washed three times with sodium phosphate buffer (SPB; 10 mM NaH2PO4, pH = 7) and suspended in the same buffer (O.D. at 620 nm = 1). Bacteria were then 10-fold diluted into SPB containing 3% TSB. 100 µl of bacterial suspension were placed in a 96-well plate with 100 µl of the tested compound and 20 µl of either ONPG (final concentration = 2.5 µM) or nitrocefin (final concentration = 25 µM). Hydrolysis of ONPG and nitrocefin was monitored immediately by recording the absorbance at 420 nm and 486 nm respectively, for 30 min at 37 °C with shaking (Synergy HT, BioTek Instruments, Winooski, VT, USA).
Cytoplasmic membrane depolarization was assessed by monitoring the displacement of the membrane binding fluorescent dye DiSC3(5) (3,3′-dipropylthiadicarbocyanine iodide) as follows. Mid-log-phase bacteria at 5 × 108 CFU/ml were centrifuged for 5 min at 15,000 g. Pellet was washed twice with 5 mM 4-(2-hydroxyethyl)piperazine-1-ethanesulfonic acid (HEPES) containing 5 mM glucose (pH = 7.2) and suspended in the same buffer. Bacteria were diluted 10-fold in the same buffer, DiSC3(5) was added to a final concentration of 4 µM and samples were incubated at RT in the dark for 1 hr. KCl was added to a final concentration of 100 µM and incubation continued for an additional hour. 180 µl of bacterial suspension were placed in a 96-well plate and fluorescence was recorded until signal stabilization (excitation: 620 nm, emission: 680 nm). Then, 20 µl of the tested compound were mixed into the wells and fluorescence was recorded immediately for up to 30 min at 37 °C with shaking (Synergy HT, BioTek Instruments, Winooski, VT, USA).
Dansyl-polymyxin displacement assay was assessed by displacement of dansyl-polymyxin B bound to lipopolysaccharide (LPS) as follows. Polymyxin B sulfate was covalently attached to dansyl chloride and mono-dansyl Polymyxin B (DPMB) was purified by RP-HPLC. 180 µl of 5 mM HEPES containing 3 µg/ml LPS (from E. coli or P. aeruginosa) and 2 µM mono-DPMB were incubated in a 96-well plate with 20 µl of the tested compound for 1.5 hr at RT and fluorescence (excitation: 340 nm, emission: 485 nm) was measured immediately (Synergy HT, BioTek Instruments, Winooski, VT, USA).
Bactericidal kinetics in saliva was assessed as follows. Whole saliva was pooled from two-three healthy volunteers at a time, after obtaining their informed consent. To test for specific activity against S. mutans, saliva was centrifuged for 5 min at 15,000 g and 4 °C, and the supernatant was spiked with 1 × 107 CFU/ml of mid-log-phase S. mutans 35668. Otherwise, activity against natural oral microflora was also performed on whole saliva. Saliva was mixed with the tested compound, incubated at 37 °C with shaking, and aliquots were taken at t = 0, 2, 5 and 10 min. Aliquots were subjected to serial 10-fold dilutions in saline and plated on BHI agar for enumeration.
Anti-biofilm activity was evaluated against established biofilms in 96-well plates as follows. 200 µl of mid-log-phase S. mutans 35668 at 5 × 105 CFU/ml in BHI containing 2% sucrose were placed in each well, and plates were incubated for 24 hr at 37 °C without shaking. Unattached cells were removed by decanting the plates and biofilms were washed three times with milliQ water. Solutions of the tested compounds were placed on the biofilms for up to 3 hr at 37 °C without shaking. Plates were decanted in 0.5 hr intervals and biofilms were washed three times with milliQ water to remove any compound residues. Biofilms were scraped from the bottom of the wells, suspended in BHI and sonicated for 5 min in a sonication bath, subjected to serial 10-fold dilutions in saline and plated on BHI agar for enumeration.
References
Schaberle, T. F. & Hack, I. M. Overcoming the current deadlock in antibiotic research. Trends in microbiology 22, 165–167, doi:10.1016/j.tim.2013.12.007 (2014).
Eckert, R. Road to clinical efficacy: challenges and novel strategies for antimicrobial peptide development. Future microbiology 6, 635–651, doi:10.2217/fmb.11.27 (2011).
Silver, L. L. Challenges of antibacterial discovery. Clinical microbiology reviews 24, 71–109, doi:10.1128/CMR.00030-10 (2011).
Fischbach, M. A. & Walsh, C. T. Antibiotics for emerging pathogens. Science 325, 1089–1093, doi:10.1126/science.1176667 (2009).
Vaara, M. New approaches in peptide antibiotics. Current opinion in pharmacology 9, 571–576, doi:10.1016/j.coph.2009.08.002 (2009).
Blair, J. M., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. Molecular mechanisms of antibiotic resistance. Nature reviews. Microbiology 13, 42–51, doi:10.1038/nrmicro3380 (2015).
Klitgaard, J. K., Skov, M. N., Kallipolitis, B. H. & Kolmos, H. J. Reversal of methicillin resistance in Staphylococcus aureus by thioridazine. The Journal of antimicrobial chemotherapy 62, 1215–1221, doi:10.1093/jac/dkn417 (2008).
Hurdle, J. G., O’Neill, A. J., Chopra, I. & Lee, R. E. Targeting bacterial membrane function: an underexploited mechanism for treating persistent infections. Nature reviews. Microbiology 9, 62–75, doi:10.1038/nrmicro2474 (2011).
Allen, R. C., Popat, R., Diggle, S. P. & Brown, S. P. Targeting virulence: can we make evolution-proof drugs? Nature reviews. Microbiology 12, 300–308, doi:10.1038/nrmicro3232 (2014).
Porter, E. A., Wang, X., Lee, H. S., Weisblum, B. & Gellman, S. H. Non-haemolytic beta-amino-acid oligomers. Nature 404, 565, doi:10.1038/35007145 (2000).
Daly, S. M. et al. omega-Hydroxyemodin limits staphylococcus aureus quorum sensing-mediated pathogenesis and inflammation. Antimicrobial agents and chemotherapy 59, 2223–2235, doi:10.1128/AAC.04564-14 (2015).
Strahl, H. & Hamoen, L. W. Membrane potential is important for bacterial cell division. Proceedings of the National Academy of Sciences of the United States of America 107, 12281–12286, doi:10.1073/pnas.1005485107 (2010).
Nizet, V. Understanding how leading bacterial pathogens subvert innate immunity to reveal novel therapeutic targets. The Journal of allergy and clinical immunology 120, 13–22, doi:10.1016/j.jaci.2007.06.005 (2007).
Sully, E. K. et al. Selective chemical inhibition of agr quorum sensing in Staphylococcus aureus promotes host defense with minimal impact on resistance. PLoS pathogens 10, e1004174, doi:10.1371/journal.ppat.1004174 (2014).
Hicks, D. B., Cohen, D. M. & Krulwich, T. A. Reconstitution of energy-linked activities of the solubilized F1F0 ATP synthase from Bacillus subtilis. Journal of bacteriology 176, 4192–4195, doi:10.1128/jb.176.13.4192-4195.1994 (1994).
Padan, E., Bibi, E., Ito, M. & Krulwich, T. A. Alkaline pH homeostasis in bacteria: new insights. Biochimica et biophysica acta 1717, 67–88, doi:10.1016/j.bbamem.2005.09.010 (2005).
Kaneti, G., Meir, O. & Mor, A. Controlling bacterial infections by inhibiting proton-dependent processes. Biochimica et biophysica acta 1858, 995–1003, doi:10.1016/j.bbamem.2015.10.022 (2016).
Goldberg, K. et al. Sensitization of gram-negative bacteria by targeting the membrane potential. FASEB journal: official publication of the Federation of American Societies for Experimental Biology 27, 3818–3826, doi:10.1096/fj.13-227942 (2013).
Kaneti, G., Sarig, H., Marjieh, I., Fadia, Z. & Mor, A. Simultaneous breakdown of multiple antibiotic resistance mechanisms in S. aureus. FASEB journal: official publication of the Federation of American Societies for Experimental Biology 27, 4834–4843, doi:10.1096/fj.13-237610 (2013).
Epand, R. F., Maloy, W. L., Ramamoorthy, A. & Epand, R. M. Probing the “charge cluster mechanism” in amphipathic helical cationic antimicrobial peptides. Biochemistry 49, 4076–4084, doi:10.1021/bi100378m (2010).
Westerhoff, H. V., Juretic, D., Hendler, R. W. & Zasloff, M. Magainins and the disruption of membrane-linked free-energy transduction. Proceedings of the National Academy of Sciences of the United States of America 86, 6597–6601, doi:10.1073/pnas.86.17.6597 (1989).
Epand, R. M., Rotem, S., Mor, A., Berno, B. & Epand, R. F. Bacterial membranes as predictors of antimicrobial potency. Journal of the American Chemical Society 130, 14346–14352, doi:10.1021/ja8062327 (2008).
Hancock, R. E. & Chapple, D. S. Peptide antibiotics. Antimicrobial agents and chemotherapy 43, 1317–1323 (1999).
Sarig, H., Rotem, S., Ziserman, L., Danino, D. & Mor, A. Impact of self-assembly properties on antibacterial activity of short acyl-lysine oligomers. Antimicrobial agents and chemotherapy 52, 4308–4314, doi:10.1128/AAC.00656-08 (2008).
Ruhr, E. & Sahl, H. G. Mode of action of the peptide antibiotic nisin and influence on the membrane potential of whole cells and on cytoplasmic and artificial membrane vesicles. Antimicrobial agents and chemotherapy 27, 841–845, doi:10.1128/AAC.27.5.841 (1985).
Zasloff, M. Antimicrobial peptides of multicellular organisms. Nature 415, 389–395, doi:10.1038/415389a (2002).
Vaara, M. et al. Novel polymyxin derivatives carrying only three positive charges are effective antibacterial agents. Antimicrobial agents and chemotherapy 52, 3229–3236, doi:10.1128/AAC.00405-08 (2008).
Zhang, L., Dhillon, P., Yan, H., Farmer, S. & Hancock, R. E. Interactions of bacterial cationic peptide antibiotics with outer and cytoplasmic membranes of Pseudomonas aeruginosa. Antimicrobial agents and chemotherapy 44, 3317–3321, doi:10.1128/AAC.44.12.3317-3321.2000 (2000).
Rosenfeld, Y. & Shai, Y. Lipopolysaccharide (Endotoxin)-host defense antibacterial peptides interactions: role in bacterial resistance and prevention of sepsis. Biochimica et biophysica acta 1758, 1513–1522, doi:10.1016/j.bbamem.2006.05.017 (2006).
Som, A., Vemparala, S., Ivanov, I. & Tew, G. N. Synthetic mimics of antimicrobial peptides. Biopolymers 90, 83–93, doi:10.1002/bip.20970 (2008).
Rotem, S. & Mor, A. Antimicrobial peptide mimics for improved therapeutic properties. Biochimica et biophysica acta 1788, 1582–1592, doi:10.1016/j.bbamem.2008.10.020 (2009).
Javadpour, M. M. & Barkley, M. D. Self-assembly of designed antimicrobial peptides in solution and micelles. Biochemistry 36, 9540–9549, doi:10.1021/bi961644f (1997).
Avrahami, D., Oren, Z. & Shai, Y. Effect of multiple aliphatic amino acids substitutions on the structure, function, and mode of action of diastereomeric membrane active peptides. Biochemistry 40, 12591–12603, doi:10.1021/bi0105330 (2001).
Ratledge, C. & Wilkinson, S. G. Microbial lipids. (Academic Press, 1988).
Yeaman, M. R. & Yount, N. Y. Mechanisms of antimicrobial peptide action and resistance. Pharmacological reviews 55, 27–55, doi:10.1124/pr.55.1.2 (2003).
Gaidukov, L., Fish, A. & Mor, A. Analysis of membrane-binding properties of dermaseptin analogues: relationships between binding and cytotoxicity. Biochemistry 42, 12866–12874, doi:10.1021/bi034514x (2003).
Livne, L. et al. Design and characterization of a broad -spectrum bactericidal acyl-lysyl oligomer. Chemistry & biology 16, 1250–1258, doi:10.1016/j.chembiol.2009.11.012 (2009).
Radzishevsky, I. S. et al. Structure-activity relationships of antibacterial acyl-lysine oligomers. Chemistry & biology 15, 354–362, doi:10.1016/j.chembiol.2008.03.006 (2008).
Radzishevsky, I. S. et al. Improved antimicrobial peptides based on acyl-lysine oligomers. Nature biotechnology 25, 657–659, doi:10.1038/nbt1309 (2007).
Jammal, J., Zaknoon, F., Kaneti, G., Goldberg, K. & Mor, A. Sensitization of Gram-negative bacteria to rifampin and OAK combinations. Scientific reports 5, 9216, doi:10.1038/srep09216 (2015).
Sarig, H. et al. A miniature mimic of host defense peptides with systemic antibacterial efficacy. FASEB journal: official publication of the Federation of American Societies for Experimental Biology 24, 1904–1913, doi:10.1096/fj.09-149427 (2010).
Kerker, M. In The Scattering of Light and Other Electromagnetic Radiation Ch. 7, 311–413 (Academic Press, 1969).
Epand, R. F., Sarig, H., Mor, A. & Epand, R. M. Cell-wall interactions and the selective bacteriostatic activity of a miniature oligo-acyl-lysyl. Biophysical journal 97, 2250–2257, doi:10.1016/j.bpj.2009.08.006 (2009).
Lehrer, R. I., Barton, A. & Ganz, T. Concurrent assessment of inner and outer membrane permeabilization and bacteriolysis in E. coli by multiple-wavelength spectrophotometry. Journal of immunological methods 108, 153–158, doi:10.1016/0022-1759(88)90414-0 (1988).
Storm, D. R., Rosenthal, K. S. & Swanson, P. E. Polymyxin and related peptide antibiotics. Annual review of biochemistry 46, 723–763, doi:10.1146/annurev.bi.46.070177.003451 (1977).
Moore, R. A., Bates, N. C. & Hancock, R. E. Interaction of polycationic antibiotics with Pseudomonas aeruginosa lipopolysaccharide and lipid A studied by using dansyl-polymyxin. Antimicrobial agents and chemotherapy 29, 496–500, doi:10.1128/AAC.29.3.496 (1986).
de Haar, S. F., Hiemstra, P. S., van Steenbergen, M. T., Everts, V. & Beertsen, W. Role of polymorphonuclear leukocyte-derived serine proteinases in defense against Actinobacillus actinomycetemcomitans. Infection and immunity 74, 5284–5291, doi:10.1128/IAI.02016-05 (2006).
Putsep, K., Carlsson, G., Boman, H. G. & Andersson, M. Deficiency of antibacterial peptides in patients with morbus Kostmann: an observation study. Lancet 360, 1144–1149, doi:10.1016/S0140-6736(02)11201-3 (2002).
Loesche, W. J. Role of Streptococcus mutans in human dental decay. Microbiological reviews 50, 353–380 (1986).
Bowen, W. H. & Koo, H. Biology of Streptococcus mutans-derived glucosyltransferases: role in extracellular matrix formation of cariogenic biofilms. Caries research 45, 69–86, doi:10.1159/000324598 (2011).
Klein, M. I., Hwang, G., Santos, P. H., Campanella, O. H. & Koo, H. Streptococcus mutans-derived extracellular matrix in cariogenic oral biofilms. Frontiers in cellular and infection microbiology 5, 10, doi:10.3389/fcimb.2015.00010 (2015).
Thanakun, S., Watanabe, H., Thaweboon, S. & Izumi, Y. An effective technique for the processing of saliva for the analysis of leptin and adiponectin. Peptides 47, 60–65, doi:10.1016/j.peptides.2013.06.010 (2013).
al-Tannir, M. A. & Goodman, H. S. A review of chlorhexidine and its use in special populations. Special care in dentistry: official publication of the American Association of Hospital Dentists, the Academy of Dentistry for the Handicapped, and the American Society for Geriatric Dentistry 14, 116–122, doi:10.1111/j.1754-4505.1994.tb01116.x (1994).
Jones, C. G. Chlorhexidine: is it still the gold standard? Periodontology 2000 15, 55–62, doi:10.1111/j.1600-0757.1997.tb00104.x (1997).
Greenstein, G., Berman, C. & Jaffin, R. Chlorhexidine - an Adjunct to Periodontal Therapy. J Periodontol 57, 370–377, doi:10.1902/jop.1986.57.6.370 (1986).
Fardal, O. & Turnbull, R. S. A review of the literature on use of chlorhexidine in dentistry. Journal of the American Dental Association 112, 863–869, doi:10.14219/jada.archive.1986.0118 (1986).
Gronroos, L. et al. Chlorhexidine susceptibilities of mutans streptococcal serotypes and ribotypes. Antimicrobial agents and chemotherapy 39, 894–898 (1995).
Zavascki, A. et al. Polymyxin B for the treatment of multidrug-resistant pathogens: a critical review. The Journal of antimicrobial chemotherapy 60, 1206–1215, doi:10.1093/jac/dkm357 (2007).
Steenbergen, J. N., Alder, J., Thorne, G. M. & Tally, F. P. Daptomycin: a lipopeptide antibiotic for the treatment of serious Gram-positive infections. The Journal of antimicrobial chemotherapy 55, 283–288, doi:10.1093/jac/dkh546 (2005).
Taylor, S. D. & Palmer, M. The action mechanism of daptomycin. Bioorganic & medicinal chemistry, doi:10.1016/j.bmc.2016.05.052 (2016).
Straus, S. K. & Hancock, R. E. Mode of action of the new antibiotic for Gram-positive pathogens daptomycin: comparison with cationic antimicrobial peptides and lipopeptides. Biochimica et biophysica acta 1758, 1215–1223, doi:10.1016/j.bbamem.2006.02.009 (2006).
Mangoni, M. L. & Shai, Y. Short native antimicrobial peptides and engineered ultrashort lipopeptides: similarities and differences in cell specificities and modes of action. Cellular and molecular life sciences: CMLS 68, 2267–2280, doi:10.1007/s00018-011-0718-2 (2011).
Rotem, S., Radzishevsky, I. & Mor, A. Physicochemical properties that enhance discriminative antibacterial activity of short dermaseptin derivatives. Antimicrobial agents and chemotherapy 50, 2666–2672, doi:10.1128/AAC.00030-06 (2006).
Marynka, K., Rotem, S., Portnaya, I., Cogan, U. & Mor, A. In vitro discriminative antipseudomonal properties resulting from acyl substitution of N-terminal sequence of dermaseptin s4 derivatives. Chemistry & biology 14, 75–85, doi:10.1016/j.chembiol.2006.11.009 (2007).
Makovitzki, A., Avrahami, D. & Shai, Y. Ultrashort antibacterial and antifungal lipopeptides. Proceedings of the National Academy of Sciences of the United States of America 103, 15997–16002, doi:10.1073/pnas.0606129103 (2006).
Makovitzki, A., Viterbo, A., Brotman, Y., Chet, I. & Shai, Y. Inhibition of fungal and bacterial plant pathogens in vitro and in planta with ultrashort cationic lipopeptides. Applied and environmental microbiology 73, 6629–6636, doi:10.1128/AEM.01334-07 (2007).
Nasompag, S. et al. Effect of acyl chain length on therapeutic activity and mode of action of the CX-KYR-NH2 antimicrobial lipopeptide. Biochimica et biophysica acta 1848, 2351–2364, doi:10.1016/j.bbamem.2015.07.004 (2015).
Dagan, A., Efron, L., Gaidukov, L., Mor, A. & Ginsburg, H. In vitro antiplasmodium effects of dermaseptin S4 derivatives. Antimicrobial agents and chemotherapy 46, 1059–1066, doi:10.1128/AAC.46.4.1059-1066.2002 (2002).
Radzishevsky, I. S. et al. Effects of acyl versus aminoacyl conjugation on the properties of antimicrobial peptides. Antimicrobial agents and chemotherapy 49, 2412–2420, doi:10.1128/AAC.49.6.2412-2420.2005 (2005).
Shafer, W. M., Qu, X., Waring, A. J. & Lehrer, R. I. Modulation of Neisseria gonorrhoeae susceptibility to vertebrate antibacterial peptides due to a member of the resistance/nodulation/division efflux pump family. Proceedings of the National Academy of Sciences of the United States of America 95, 1829–1833, doi:10.1073/pnas.95.4.1829 (1998).
Balthazar, J. T. et al. Lipooligosaccharide Structure is an Important Determinant in the Resistance of Neisseria Gonorrhoeae to Antimicrobial Agents of Innate Host Defense. Frontiers in microbiology 2, 30, doi:10.3389/fmicb.2011.00030 (2011).
Kaplan, C. W. et al. Selective membrane disruption: mode of action of C16G2, a specifically targeted antimicrobial peptide. Antimicrobial agents and chemotherapy 55, 3446–3452, doi:10.1128/AAC.00342-11 (2011).
Guo, L. et al. Precision-guided antimicrobial peptide as a targeted modulator of human microbial ecology. Proceedings of the National Academy of Sciences of the United States of America 112, 7569–7574, doi:10.1073/pnas.1506207112 (2015).
Acknowledgements
This work was supported by the Israel Science Foundation (grant 909/12) and in part by the Russell Berrie Nanotechnology Institute (Technion).
Author information
Authors and Affiliations
Contributions
O.M. synthesized reagents, performed research, analyzed data, wrote the paper; F.Z. synthesized reagents (mono-dansyl PMB); U.C. and A.M. designed the experiments, analyzed data, wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Meir, O., Zaknoon, F., Cogan, U. et al. A broad-spectrum bactericidal lipopeptide with anti-biofilm properties. Sci Rep 7, 2198 (2017). https://doi.org/10.1038/s41598-017-02373-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-02373-0
This article is cited by
-
Antimicrobial Peptides Targeting Streptococcus mutans: Current Research on Design, Screening and Efficacy
Current Microbiology (2024)
-
An efflux-susceptible antibiotic-adjuvant with systemic efficacy against mouse infections
Scientific Reports (2022)
-
Novel Miniature Membrane Active Lipopeptidomimetics against Planktonic and Biofilm Embedded Methicillin-Resistant Staphylococcus aureus
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.