Abstract
Multiple epidermal growth factor-like-domains 10 (MEGF10), a critical member of the apoptotic engulfment pathway, mediates axon pruning and synapse elimination during brain development. Previous studies indicated that synaptic pruning deficit was associated with autism-related phenotypes. However, the relationship between MEGF10 and autism remains poorly understood. Disease-associated variants are significantly enriched in the transcription regulatory regions. These include the transcription start site (TSS) and its cis-regulatory elements. To investigate the role of MEGF10 variants with putative transcription regulatory function in the etiology of autism, we performed a family-based association study in 410 Chinese Han trios. Our results indicate that three single nucleotide polymorphisms (SNPs), rs4836316, rs2194079 and rs4836317 near the TSS are significantly associated with autism following Bonferroni correction (p = 0.0011, p = 0.0088, and p = 0.0023, respectively). Haplotype T-A-G (rs4836316-rs2194079-rs4836317) was preferentially transmitted from parents to affected offspring (p permutation = 0.0055). Consistently, functional exploration further verified that the risk allele and haplotype might influence its binding with transcription factors, resulting in decreased transcriptional activity of MEGF10. Our findings indicated that the risk alleles and haplotype near the MEGF10 TSS might modulate transcriptional activity and increase the susceptibility to autism.
Similar content being viewed by others
Introduction
Autism is an early-onset neurodevelopmental disorder that is characterized by impaired social interaction and communication, repetitive behavior, and restricted interests1. The core symptoms of autism usually occur before three years of age. Epidemiological studies indicate that genetic factors strongly increase the risk of autism2, 3. Additionally, twin studies indicate that the estimated heritability of autism may be as much as 90%4. However, the pathogenesis of autism remains largely unclear.
Abnormal numbers and connectivity of synapses caused by synaptic pruning deficit might play a marked role in the pathogenesis of autism5, 6. Previous functional studies have revealed that deficient synaptic pruning is associated with autism-related phenotypes, such as deficits in social interaction and increased repetitive-behavior5, 7. In addition, previous genetic studies have reported that individuals affected with autism carry mutations or structural variants in several genes associated with synaptic formation and function, including NRXN1 (Neurexin1), NLGN3 (Neuroligin3), SHANK2 (SH3 and multiple ankyrin repeat domains 2), and SHANK3 (SH3 and multiple ankyrin repeat domains 3)8,9,10,11,12,13. These genes regulate neuronal activity by modulating synaptic number or strength14,15,16,17,18. Mice carrying mutations of these genes presented not only abnormal synaptic function or morphology but also abnormal social interaction19, 20. These lines of evidence suggest that synaptic dysregulation might contribute to the etiology of autism.
Previous studies have indicated that defects in synaptic pruning might cause altered synaptic density and/or shape of the autistic brain5, 21. Multiple epidermal growth factor-like-domains 10 (MEGF10), which is a critical member of the apoptotic engulfment pathway, modulates synaptic number and function via synaptic pruning and phagocytosis in the postnatal brain. A mouse model has suggested that MEGF10 might play a critical role in synapse elimination via astrocytes in the postnatal brain. Mice deficient in the Megf10 pathway displayed dysfunctional and weak synapses and failed to retain excess functional synapse during the postnatal period22. A recent study in mice also found that dysfunction of Megf10 impaired phagocytosis mediated by astrocytes in the developing cerebellum23. Moreover, previous studies have indicated that cell death abnormal 1 (CED-1) and draper (DRPR), which represent mammalian orthologues of MEGF10 in Caenorhabditis elegans and Drosophilia melanogaster respectively, might regulate the clearance of apoptotic cells, axon pruning and neuronal degeneration24,25,26. Axon pruning and neuronal degeneration mediated by CED-1 modulate the number of synapses and synaptic connectivity27, 28. Therefore, a critical role in the regulation of synaptic number and function suggests that MEGF10 is a candidate gene for autism.
MEGF10 is located on chromosome 5q33. Linkage analysis revealed that 5q33.1 was associated with motor/speech phenotype in autism29. An association study suggested that paired like homeodomain 1 (PITX1), which is located adjacent to MEGF10, might be an autism susceptibility gene30. Furthermore, other association studies have implicated that MEGF10 and other genes in the apoptotic engulfment pathway are significantly associated with schizophrenia31, 32. Autism and schizophrenia share several clinical features including impaired executive function and deficits in social function33, 34. Several lines of evidence indicate that schizophrenia and autism share similar candidate genes, at least partially, including CACNA1C 35, 36, DISC1 37, 38, SHANK3 9, 39, NRXN1 40, 41, and NTNG1 42, 43, which are related to the modulation of the synapses. However, the relationship between MEGF10 and etiology of autism remains poorly understood.
Recently, a systematic study revealed that disease-associated SNPs were significantly enriched in the functional regions annotated by the ENCODE consortium and Roadmap project44, 45. Several functional regions, including the transcription start site (TSS) and its cis-regulatory elements, play a critical role in modulating gene transcription. Moreover, SNPs in this region often represent significant expression Quantitative Trait Loci (eQTL) signals that are related to mRNA abundance46. On collectively considering both the role of MEGF10 and its regulation of synaptic pruning and the relationship of regulatory functional variants with disease, we hypothesized that variants in MEGF10 with putative transcriptional regulatory function might play a role in the pathogenesis of autism. To elucidate the genetic relationship between MEGF10 and autism, we primarily selected the three SNPs in the TSS and an additional two SNPs in the non-regulatory region. Then, we performed a family-based association study involving 410 Chinese Han family trios to investigate the association of five selected SNPs with autism. Furthermore, several functional assays were used to verify the regulatory function of three variants in the TSS of MEGF10.
Results
Quality control
According to the SNP selection criteria described in the Methods section of this article, seven SNPs (i.e., rs2408868, rs10519929, rs3812058, rs10519930, rs1363387, rs4836316 and rs2194079) were located in the active transcription start site and cis-regulatory elements. Of these SNPs, rs10519929, rs3812058 and rs10519930 that bind to TF were not selected due to their MAF having a value less than 0.05 in the CHB population of the Hapmap project (MAF = 0.018, 0.045 and 0.018 respectively). Rs4836316 and rs2194079 were completely linkage disequilibrium with rs1363387 and rs2408868 respectively. The number of binding TF motifs change for rs4836316 was more than that for rs1363387 (Supplementary Table S1). Moreover, the change of TF motif log-odds (LOD) score of rs2194079 was greater than that of rs2408868. Thus, rs4836316 and rs2194079 were preferentially selected as tag SNPs. Meanwhile, we selected rs4836317 to construct haplotype with rs4836316 and rs2194079. An additional two SNPs (i.e., rs9327438 and rs6866678) in the non-regulatory region were selected to compare their association with three selected SNPs near the TSS region. These selected SNPs covered 60.5% of the whole gene. The mean distance between each of the neighboring SNPs was 34.2 Kb.
A total of 41 missing genotypes consisted of three genotypes for SNP rs4836316; four for SNP rs2194079; 13 for SNP rs4836317; 16 for SNP rs9327438 and five for SNP rs6866678. The call rate of each genotype in the study subjects is greater than 95% which is for Sanger sequencing and the Agena Bioscience platform. SNPs frequencies are displayed in Table 1. None of the genotypic distributions of these SNPs in unaffected parents and affected offspring deviated from HWE (Supplementary Table S3). The genotyping concordance rate for the Agena Bioscience platform and Sanger sequencing in 10% of the samples exceeded 99%.
Putative regulatory function and eQTL of selected SNPs
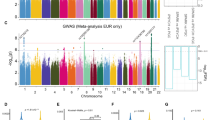
The rVarbase predicted that rs4836316, rs2194079 and rs4836317 were located in the active TSS region which might overlap with several cis-regulatory elements. In addition, HaploReg indicated that they disrupt binding motifs of several transcriptional regulatory factors (Supplementary Table S1). The eQTL analysis from the BRAINEAC database showed that all selected SNPs were associated with the expression level of MEGF10 in several brain regions (Supplementary Fig. S3). Remarkably, rs4836316 and rs4836317 were significantly associated with the expression of MEGF10 in the hippocampus after the p value was corrected by a false discovery rate (FDR) of <1% (p = 1.1e-6, p = 1.2e-7, respectively) (Supplementary Fig. S3). In the GTEx database, the multi-tissue (53 human tissues including 10 primary brain regions) eQTL comparison for five selected SNPs revealed that several brain regions, such as the cerebellum and frontal cortex, were predicted to show a strong eQTL effect (post-prob >0.9) (Supplementary Table S5).
SNP association and haplotype association analyses
Under an additive model, univariate (single marker) tests showed that the T allele of rs4836316, the A allele of rs2194079 and the G allele of rs4836317 were significantly preferentially transmitted from parents to the affected offspring (rs4836316: T > G, Z = 3.254, p = 0.0011; rs2194079: A > G, Z = 2.621, p = 0.0088; rs4836317: G > T, Z = 3.055, p = 0.0023) (Table 1). Moreover, the T allele of rs4836316 and the G allele of rs4836317 were preferentially transmitted from parents to offspring under a dominant model (rs4836316: T > G, Z = 2.762, p = 0.0057; rs4836317: G > T, Z = 2.752, p = 0.0059) (Supplementary Table S4). The statistical significance persisted even after Bonferroni correction (p = α/n = 0.05/5 = 0.01).
As shown in Fig. 1b, the r-square value indicated that five selected SNPs constructed two potential LD blocks. In addition, rs4836316, rs2194079 and rs4836317 were located in block 1, while others (i.e., rs9327438 and rs6866678) were located in block 2. Specific and global haplotype association tests were carried out only for the LD blocks. All haplotypes were constructed using the sliding window approach. We found that haplotype T-A-G (rs4836316-rs2194079-rs4836317; Z = 2.779, p = 0.0055, Global p = 0.0079) (Table 2) was transmitted more than haplotype G-G-T (Z = −3.087, p = 0.0020, Global p = 0.0079) (Table 2). These results remained significant after permutation testing (n = 10,000, p = 0.003). In addition, the haplotypes T-A (rs4836316-rs2194079) and the haplotype A-G (rs2194079-rs4836317) that was constructed from risk alleles also displayed preferential transmission from parents to affected offspring (Z = 2.595, p = 0.0094, Global p = 0.0058; Z = 2.780, p = 0.0054, Global p = 0.0020, respectively) (Table 2). However, the haplotype G-A that was composed of rs9327438 and rs6866678 did not show significant preferential transmission (Z = 1.907, p = 0.056, Global p = 0.014) (Table 2).
Linkage disequilibrium (LD) block constructed from 5 SNPs in MEGF10. (a) Markers with linkage disequilibrium (LD) (D’ < 1 and LOD >2) are shown in red through pink (color intensity decreases with decreasing D’ value). D’ value shown in each square represents pairwise LD relationship between the two polymorphisms. D prime values of 1.0 are never shown (the box is empty). The LD plot was generated using the Haploview program. (b) The value of r 2 is shown in each square. Black square indicated r 2 = 1 (i.e. perfect LD between a pair of SNPs). Markers with LD are shown in black to grey (color intensity decreases with decreasing r 2 value).
Regulatory effects of the haplotype constructed from three risk SNPs on the transcriptional activity
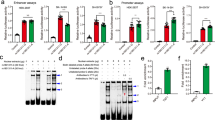
The 1.5 kb fragments that contained haplotype T-A-G (rs4836316-rs2194079-rs4836317) and G-G-T were independently cloned upstream of the luciferase reporter gene (pGL3-Basic). They were transfected into both 293T and C6 cells. The reporter construct containing the haplotype T-A-G showed decreased luciferase activity as compared with the reporter construct that contained the haplotype G-G-T in both cell lines (P < 0.05 for both cell lines) (Fig. 2). These results indicated that the haplotype T-A-G was less efficient in driving transcription than the haplotype G-G-T. Results of functional prediction were partially validated by this experiment.
Regulatory effects of the haplotypes constructed from rs4836316, rs2194079 and rs4836317 on the transcriptional activity. The transcriptional activity was decreased when the risk haplotype T-A-G in 293 and C6 cell lines. Bars are presented as fold-induction compared with promoter less vector. Error bars represent standard error of mean (SEM) (n = 5). *P < 0.05.
Protein binding ability of three risk SNPs
As shown in Fig. 3, band retardation was observed in the presence of nuclear proteins in lanes that were represented by rs2194079 and rs4836317. The binding activity of the A allele in rs2194079 was weaker than the G allele, while the alleles of rs4836317 were almost identical. The EMSA results showed that the A allele of rs2194079 is relatively weak in binding nuclear proteins as compared with the G allele of rs2194079.
Results of electrophoretic mobility shift assays for rs4836316, rs2194079 and rs4836317. Electrophoretic mobility shift assays were performed with biotinylated oligonucleotides and nuclear extracts isolated from C6 cell line to investigate the binding of associated SNP with nuclear extract. (a) The labeled probes containing variants of rs2194079, rs4836316, rs4836317 were incubated with the nuclear proteins. SNP rs4836316 showed no band shift. Both rs2194079 and rs4836317 showed band shifts. The allele G of rs2194079 showed preferential binding with the nuclear extract. (b) Specificity was determined by competition with 50-fold unlabeled probes. Protein-DNA complexes were observed for both alleles. However, preferential binding of G allele in rs2194079 was observed. The “+” and “−” indicate the presence and absence of nuclear extract or probe.
Expression pattern of MEGF10 in human brain and peripheral blood of autistic individuals
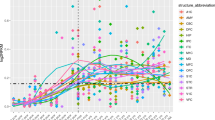
The expression of MEGF10 in 13 human brain regions was higher than that found in other human organs or tissues, and especially in the spinal cord, basal ganglia, substantia nigra and the hippocampus (Supplementary Figs S2 and S4). As shown in the figure from HBT database, during developmental stages, MEGF10 expression in the hippocampus and other brain regions was initially decreased and then progressively increased. After birth, its expression was stable and it was sustained at a relatively high level in different regions of the brain throughout life (Supplementary Fig. S5a). The dynamic expression of MEGF10 in 11 areas of the neocortex was similar to the expression that found in other brain regions (Supplementary Fig. S5b). In the GEO dataset from Alter et al.47, the normalized expression value of MEGF10 in autistic individuals was lower than that found in healthy controls (p < 0.05) (Supplementary Fig. S6).
Discussion
We studied the genetic association of five SNPs in MEGF10 with autism in a Chinese Han cohort. The results indicated that three SNPs (i.e., rs4836316, rs2194079 and rs4836317) in the transcription start site were significantly associated with autism. Our observations also indicated that haplotype T-A-G that was constructed from these SNPs might be a risk factor for autism.
A previous genome-wide association study (GWAS) of the Psychiatric Genomic Consortium (PGC) that included 4788 trio cases and 4788 trio pseudocontrols demonstrated that thirty SNPs in MEGF10 were nominally associated with autism in a CEU (Northern and Western European Ancestry in Utah) population35 (Supplementary Table S7). Of these SNPs, rs10519929, rs10519930 and rs17165105 were located in the same haplotype block which contained three positive SNPs (i.e., rs4836316, rs2194079 and rs4836317) that we tested (Supplementary Fig. S1). All six SNPs were mapped to haplotype spanning the transcription start site and were predicted to possess similar regulatory function. The rVarbase and HaploReg database also indicated that rare variants such as rs10519929 and rs10519930 which were located approximately 2Kb upstream of the risk haplotype T-A-G (rs4836316-rs2194079-rs4836317) might also disrupt transcriptional factor binding sites (Supplementary Table S1 and Supplementary Fig. S7). This evidence suggests that disrupted transcriptional regulation of MEGF10 contributes to autism risk. In our study, functional experiments verified that the risk alleles and haplotype might affect binding with transcription factors, which would result in decreased transcriptional activity of MEGF10. The HaploReg database indicated that the risk haplotype could lead to reduced transcription factor binding consistent with our results. Based on the putative regulatory function of these common and rare variants in the TSS proximal region of MEGF10, these variants would seem to increase autism risk through reducing MEGF10 dosage. A previous transcriptome study showed that MEGF10 is strongly expressed in glial cells, especially astrocytes48. Compared to haplotype T-A-G (rs4836316-rs2194079-rs4836317), haplotype G-G-T increased the reporter expression in the C6 cell-line, derived from rat glial cells. Meanwhile, the specific binding of nuclear factors to rs2194079 supported the differential function of each haplotype. This evidence suggests that the risk haplotype T-A-G might decrease the transcription of MEGF10 by disrupting binding with transcription factors found in human glial cells. Moreover, these three SNPs represented positive eQTL signals in primary brain regions and decreased expression of MEGF10 was observed in the peripheral blood of autistic individuals from a GEO profile dataset.
During the postnatal period, the low expression of MEGF10 in the brain might affect the modulation of synaptic number or function. Previous functional experiments conducted in Megf10 −/− mice indicated a critical role in synapse elimination in the developing and adult brain. In developing mice, Megf10-deficient astrocytes led to a failure in retaining excess functional synapses. During adulthood, astrocytes enriched in Megf10 continuously phagocytosed both excitatory and inhibitory synapses22. MEGF10 is evolutionarily conserved and is a key member of phagocytic pathways that contributes to axon pruning by glial cells in Drosophilia melanogaster 25. Further, functional investigation has indicated that synaptic pruning deficits caused by aberrant autophagy have contributed to the development of ASD-like social behavior5. Therefore, the dampened MEGF10 expression may trigger synaptic dysregulation and disrupt the rate of synapse formation and pruning, thus playing a role in the pathogenesis of autism.
In this study, we established a genetic relationship between MEGF10 and autism in the Chinese Han population. Further, for the risk haplotype T-A-G spanning the TSS, we explored its function in regulating the transcriptional activity of MEGF10. Notably, MEGF10 is intolerant of heterozygous loss of function (LoF) based on the probability of LoF intolerance (pLI) score of 0.99 from the ExAC database. This indicates that the inherited contribution to autism etiology of this gene would have to be explored through putatively hypomorphic variants only. Sequencing rare mutations of MEGF10 in autistic individuals is also needed in future studies. In individuals carrying the risk alleles of rs4836316, rs2194079 and rs4836317, the low level of MEGF10 expression should be further validated in the developing brain. Furthermore, functional studies of MEGF10 in mice are required to illustrate its relationship with autism.
In summary, our findings suggest that MEGF10, which mediates axon pruning and synapse elimination during brain development, is associated with autism in the Chinese Han population. In addition, the haplotype T-A-G composed of rs4836316, rs2194079 and rs4836317 that near the TSS seems to affect autism risk by modulating the transcriptional activity of MEGF10.
Methods
Ethics statement
All subjects (including singleton autistic disorder children and their unaffected biological parents) provided written informed consent to participate in this study. This study was approved by the Ethics Committee of the Institute of Mental Health, The Sixth Hospital, Peking University (P. R. China). All methods were performed in accordance with approved guidelines.
Subjects
All subjects were recruited at the Institute of Mental Health, The Sixth Hospital of Peking University, of the P. R. China. They were of Chinese Han descent for at least three generations (i.e. four Chinese Han grandparents). All patients fulfilled the criteria of the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV) for autistic disorders. The diagnosis of autism was established by two senior psychiatrists. The clinical features of patients were assessed using the Childhood Autism Rating Scale (CARS)49 and Autism Behavior Checklist (ABC)50. Children with CARS scores ≥35 and ABC score ≥53 were included in this study.
Children with fragile X syndrome, Asperger’s syndrome, Rett syndrome, tuberous sclerosis, or a previously identified chromosomal abnormality, dysmorphic features, or any other neurological condition were excluded from the study. Individuals with other severe psychiatric disorders or familial inherited diseases were also excluded from this study. Finally, 410 nuclear trios were included. Of the 410 probands affected by autism, 383 were male and 27 were female. The median age of the children at diagnosis was 5.5 years (range 3–17 years).
SNP selection
Information that was related to single nucleotide polymorphisms (SNPs) of MEGF10 in CHB (Chinese Han, Beijing) was downloaded from the dbSNP (http://www.ncbi.nlm.nih.gov/SNP/) and the international HapMap project in phase II and III (ftp://ftp.ncbi.nlm.nih.gov/hapmap). Minor allele frequency (MAF) and p-value derived from the Hardy-Weinberg Equilibrium (HWE) of all SNPs in the CHB population exceeded 5%, respectively. Tagger module in Haploview version 4.2 was used to optimize SNP selection.
Functional annotations and characteristics for all SNPs located in the haplotypes which included the TSS of MEGF10 were displayed in Supplementary Table S1. As shown in Supplementary Fig. S1, 18 SNPs were located in this haplotype block. Utilizing four online databases, there are three basic criteria that were required to select SNPs having potential transcriptional regulatory function: 1. based on the results from the rVarbase databases (http://rv.psych.ac.cn/)51, SNPs that were located proximal to the TSS and in annotated cis-regulatory elements were preferred to have regulatory function. The active TSS annotated by rVarbase spans from 908 bp upstream of MEGF10 TSS to 3305 bp downstream of that. SNPs in the annotated cis-regulatory elements, including TF-binding region and chromosome interactive region of this span, were preferentially selected. 2. For the SNPs that located in both active TSS and cis-regulatory elements annotated by rVarbase, their information about protein bound and motif change of TF that derived from HaploReg provided further evidence in support of their regulatory function (http://archive.broadinstitute.org/mammals/haploreg/)52. Based on the HaploReg database, disruption of TF binding motifs would be a major criterion for inclusion if SNPs were predicted to not bind the TF protein. SNPs that could disrupt more binding TF motifs were preferentially selected; if the number of binding TF motifs was identical, SNPs with greater change of motif LOD (log-odds) scores were selected. 3. The eQTL effects of SNPs on the brain regions derived from the Braineac and GTEx databases (http://www.braineac.org/; http://www.gtexportal.org/) were additive evidence to support their potential transcriptional regulatory function52, 53.
SNP genotyping
Genomic DNA was extracted from the blood using a Qiagen QIAamp DNA Mini Kit (Qiagen, Hilden, Germany). The Agena Bioscience platform (http://www.agenabio.com), which is based on the matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) primer extension assay, was used to genotype three SNPs (rs4836317, rs9327438 and rs6866678)54. We used the iPlex genotyping assay, which showed increased plexing efficiency and flexibility for the MassARRAY system via single base primer extension with mass-modified terminators55. All primers were designed according to the sequence of the forward strand from the dbSNP database (http://www.ncbi.nlm.nih.gov/SNP/). The remaining two SNPs (rs4836316 and rs2194079) were genotyped by direct Sanger DNA sequencing. The previous literatures reported that the genotyping accuracy of Agena Bioscience platform was up to 97.5%56. The raw accuracy of Sanger sequencing is 99.99%57. The primers that were required by the iPlex genotyping assay and Sanger sequencing are listed in Supplementary Table S2. The polymerase chain reaction (PCR) amplification in Sanger sequencing was performed in a 15 μL volume that contained 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, 200 mM of each dNTPs, 0.3 mM of each primer, 1 U of Taq DNA polymerase, and 40 ng of the genomic DNA. PCR amplification was conducted with an initial denaturation phase at 94 °C for 5 min, followed by 38 cycles at 94 °C for 30 sec, with annealing at 59 °C for 30 sec, and extension at 72 °C for 40 sec, followed by a final extension phase at 72 °C for 7 min. DNA sequencing was performed after cleaning the PCR product using a BigDye Terminator Cycle Sequencing Ready Reaction Kit with Ampli Taq DNA polymerase (PE Biosystem). The inner primers were used for cycle sequencing, and the fragments were separated by electrophoresis on an ABI PRISM 377-96 DNA Sequencer (Applied Biosystem, Foster city, USA). In addition, in 10% of the samples that were randomly selected, Sanger sequencing was used to confirm the genotyping results derived from the Agena Bioscience platform.
Statistical analyses
Deviation from the HWE for genotype frequency distributions was analyzed using the Chi-square goodness-of-fit test. Mendelian errors were detected by the family-based association test (FBAT) program version 2.0.3 (http://www.biostat.harvard.edu/fbat/default.html)58. Genotypes of families with Mendelian errors were reset to zero. Haploview version 4.2 (http://www.broad.mit.edu/mpg/haploview/) was used to calculate the values of pairwise D’ and r 2 for SNP pairs and construct haplotypes. SNP pairs were considered to be in strong linkage disequilibrium (LD) if r 2 > 0.80. We performed association tests using the FBAT program. The FBAT program implemented a generalized score statistic to perform a variety of transmission disequilibrium tests (TDT), including haplotype analysis. Additive and dominant inheritance models were examined in single marker association tests. The Bonferroni correction was used to reduce the rate of type I errors. The significance level was set to an alpha value of p < 0.01 (α/n = 0.05/5 = 0.01). Moreover, the global haplotype tests of association were performed under “multiallelic” mode in haplotype based association tests (HBAT). Meanwhile, the individual haplotype tests were conducted under “biallelic” mode in HBAT. A permutation test (n = 10,000) was used for multiple testing correction in HBAT, which is capable of recalculating the test statistic and reconstructing the distribution of the population. The empirical p value calculated by this test determines the significance level of results in the population. All p values calculated by the FBAT program were two-sided.
Dual-luciferase reporter assay
The dual-luciferase reporter assay was conducted to validate the potential regulatory effects of the risk haplotype on transcriptional activity. First, a 1.5 Kb fragment that included rs4836316, rs2194079 and rs4836317 was generated by PCR. The forward and reverse primers were 5′GAGCTCTCACGCAAGGACCCACATTAT 3′ and 5′CTCGAGCCCACTTCATGCTTCTGGTATA 3′, respectively.
We used the template DNA from individuals containing the haplotype T-A-G (rs4836316-rs2194079-rs4836317). The resulting fragments were cloned into the SacI and XhoI sites of the pGL3-basic vector (Promega, Madison, WI, USA). Multiple site-directed mutagenesis was then used to obtain the construct containing the haplotype G-G-T (rs4836316-rs2194079-rs4836317). All DNA constructs were verified by sequencing. The pGL3-basic plasmid for background luminescence derived from the empty vector served as the control. The pGL3-promoter plasmid containing the SV40 promoter was used as a positive control. Transfection was performed using Lipofactamine 2000 (Life Technologies, Carlsbad, CA, USA) in triplicate for each construct. The 293T and C6 cells (at a density of 2 × 105 each) were seeded into 12-well cell culture plates and co-transfected with pRL-TK (Promega, Madison, WI, USA). A pRL Renilla Luciferase Control Reporter Vector served as a normalizing control. After 24 h, cells were lysed with a passive lysis buffer and the luciferase activity was analyzed using the Dual-Luciferase Assay System (Promega, Madison, USA) on a luminometer (Turner Biosystems, Sunnyvale, USA). Reporter activity was measured as the ratio of firefly-to-Renilla luciferase activity. Cell-lines used in this assay were derived from the Cell Resource Center of Peking Union Medical College (PUMC). They were also confirmed not to be contaminated by mycoplasma. The value of fold induction over promoter, less the vector, in each group of different cells was compared by an independent t-test. Analyses above were conducted in SPSS v.20.0 statistical analysis software program. All p values calculated by these statistic tests were two-sided.
Electrophoretic mobility shift assay
To investigate the role of the three putative functional SNPs (rs4836316, rs2194079 and rs4836317) in protein-binding, an electrophoretic mobility shift assay (EMSA) was performed using 30 bp probes encompassing the common variants of each of these 3 SNPs. The sense strands of the double-stranded probes included 5′ biotin-labeled oligonucleotides, which were annealed with the antisense strand to form the probes. The C6 cell nuclear extracts were incubated with the probes at room temperature for 20 min. The Light Shift Chemiluminescent EMSA kit (Pierce, Holmdel, USA) was used. The reaction mixture was separated by 8% polyacrylamide gel electrophoresis (PAGE), and the products were detected by Stabilized Streptavidin-Horseradish Peroxidase Conjugate (Pierce, Holmdel, USA). Unlabeled probes at a 100-fold molar excess were added to the reaction for competition.
In silico analyses of MEGF10 expression in human brain and the peripheral blood of autistic individuals
Three online databases that included BRAINEAC and GTEx were utilized to determine the expression pattern of MEGF10 in humans. The GTEx databases revealed the expression of MEGF10 in 53 human tissues or cells (including 13 brain regions)53. The Human Brain Transcriptome (HBT) databases (http://hbatlas.org/pages/hbtd) provided the figure related to dynamic MEGF10 expression during development and adulthood in the cerebellar cortex, mediodorsal nucleus of the thalamus, striatum, amygdala, hippocampus and 11 areas of the neocortex59.
Meanwhile, a Gene Expression Omnibus (GEO) dataset from Alter et al. (https://www.ncbi.nlm.nih.gov/geoprofiles) provided the normalized expression value of MEGF10 that was counted by the affymetrix array in the peripheral blood of 69 autistic individuals and 77 healthy controls47, 60. This study was designed to compare gene expression profiles in peripheral blood lymphocytes of children with autism and controls. Quantile normalization adjusted all data so that they had similar distribution patterns. The normalized expression values between the two groups were compared by independent t-test. Analyses above were conducted with SPSS v.20.0 software. All p values calculated by these statistical tests were two-sided.
References
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders (Fourth edition) (American Psychiatric Association, Washington, DC, 1994).
Szatmari, P., Jones, M. B., Zwaigenbaum, L. & MacLean, J. E. Genetics of autism: overview and new directions. J Autism Dev Disord. 28, 351–368, doi:10.1023/A:1026096203946 (1998).
Constantino, J. N., Zhang, Y., Frazier, T., Abbacchi, A. M. & Law, P. Sibling recurrence and the genetic epidemiology of autism. Am J Psychiatry. 167, 1349–1356, doi:10.1176/appi.ajp.2010.09101470 (2010).
Bailey, A. et al. Autism as a strongly genetic disorder: evidence from a British twin study. Psychol Med. 25, 63–77, doi:10.1017/S0033291700028099 (1995).
Tang, G. et al. Loss of mTOR-dependent macroautophagy causes autistic-like synaptic pruning deficits. Neuron. 83, 1131–1143, doi:10.1016/j.neuron.2014.07.040 (2014).
Kim, H. J. et al. Deficient autophagy in microglia impairs synaptic pruning and causes social behavioral defects. Mol Psychiatry. (2016).
Zhan, Y. et al. Deficient neuron-microglia signaling results in impaired functional brain connectivity and social behavior. Nat Neurosci. 17, 400–406, doi:10.1038/nn.3641 (2014).
Jamain, S. et al. Mutations of the X-linked genes encoding neuroligins NLGN3 and NLGN4 are associated with autism. Nat Genet. 34, 27–29, doi:10.1038/ng1136 (2003).
Durand, C. M. et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet. 39, 25–27, doi:10.1038/ng1933 (2007).
Jiang, Y. H. et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am J Hum Genet. 93, 249–263, doi:10.1016/j.ajhg.2013.06.012 (2013).
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science. 316, 445–449, doi:10.1126/science.1138659 (2007).
Pinto, D. et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature. 466, 368–372, doi:10.1038/nature09146 (2010).
Glessner, J. T. et al. Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature. 459, 569–573, doi:10.1038/nature07953 (2009).
Kwon, H. B. et al. Neuroligin-1-dependent competition regulates cortical synaptogenesis and synapse number. Nat Neurosci. 15, 1667–1674, doi:10.1038/nn.3256 (2012).
Sudhof, T. C. Neuroligins and neurexins link synaptic function to cognitive disease. Nature. 455, 903–911, doi:10.1038/nature07456 (2008).
Varoqueaux, F. et al. Neuroligins determine synapse maturation and function. Neuron. 51, 741–754, doi:10.1016/j.neuron.2006.09.003 (2006).
Durand, C. M. et al. SHANK3 mutations identified in autism lead to modification of dendritic spine morphology via an actin-dependent mechanism. Mol Psychiatry. 17, 71–84, doi:10.1038/mp.2011.57 (2012).
Shcheglovitov, A. et al. SHANK3 and IGF1 restore synaptic deficits in neurons from 22q13 deletion syndrome patients. Nature. 503, 267–271, doi:10.1038/nature12618 (2013).
Zhou, Y. et al. Mice with Shank3 Mutations Associated with ASD and Schizophrenia Display Both Shared and Distinct Defects. Neuron. 89, 147–162, doi:10.1016/j.neuron.2015.11.023 (2016).
Tabuchi, K. et al. A neuroligin-3 mutation implicated in autism increases inhibitory synaptic transmission in mice. Science. 318, 71–76, doi:10.1126/science.1146221 (2007).
Martinez-Cerdeno, V. Dendrite and spine modifications in autism and related neurodevelopmental disorders in patients and animal models. Dev Neurobiol. 77, 393–404, doi:10.1002/dneu.22417 (2017).
Chung, W. S. et al. Astrocytes mediate synapse elimination through MEGF10 and MERTK pathways. Nature. 504, 394–400, doi:10.1038/nature12776 (2013).
Iram, T. et al. Megf10 Is a Receptor for C1Q That Mediates Clearance of Apoptotic Cells by Astrocytes. J Neurosci. 36, 5185–5192, doi:10.1523/JNEUROSCI.3850-15.2016 (2016).
Zhou, Z., Hartwieg, E. & Horvitz, H. R. CED-1 is a transmembrane receptor that mediates cell corpse engulfment in C. elegans. Cell. 104, 43–56, doi:10.1016/S0092-8674(01)00190-8 (2001).
MacDonald, J. M. et al. The Drosophila cell corpse engulfment receptor Draper mediates glial clearance of severed axons. Neuron. 50, 869–881, doi:10.1016/j.neuron.2006.04.028 (2006).
Ziegenfuss, J. S. et al. Draper-dependent glial phagocytic activity is mediated by Src and Syk family kinase signalling. Nature. 453, 935–939, doi:10.1038/nature06901 (2008).
Low, L. K. & Cheng, H. J. Axon pruning: an essential step underlying the developmental plasticity of neuronal connections. Philos Trans R Soc Lond B Biol Sci. 361, 1531–1544, doi:10.1098/rstb.2006.1883 (2006).
Schmitz, C., Kinge, P. & Hutter, H. Axon guidance genes identified in a large-scale RNAi screen using the RNAi-hypersensitive Caenorhabditis elegans strain nre-1(hd20) lin-15b(hd126). Proc Natl Acad Sci USA. 104, 834–839, doi:10.1073/pnas.0510527104 (2007).
Flax, J. F., Hare, A., Azaro, M. A., Vieland, V. J. & Brzustowicz, L. M. Combined linkage and linkage disequilibrium analysis of a motor speech phenotype within families ascertained for autism risk loci. J Neurodev Disord. 2, 210–223, doi:10.1007/s11689-010-9063-2 (2010).
Philippi, A. et al. Association of autism with polymorphisms in the paired-like homeodomain transcription factor 1 (PITX1) on chromosome 5q31: a candidate gene analysis. BMC Med Genet. 8, 74, doi:10.1186/1471-2350-8-74 (2007).
Chen, X. et al. MEGF10 association with schizophrenia. Biol Psychiatry. 63, 441–448, doi:10.1016/j.biopsych.2007.11.003 (2008).
Chen, X. et al. Apoptotic engulfment pathway and schizophrenia. PLoS One. 4, e6875, doi:10.1371/journal.pone.0006875 (2009).
Baron-Cohen, S. & Belmonte, M. K. Autism: a window onto the development of the social and the analytic brain. Annu Rev Neurosci. 28, 109–126, doi:10.1146/annurev.neuro.27.070203.144137 (2005).
Craddock, N. & Owen, M. J. Data and clinical utility should be the drivers of changes to psychiatric classification. Br J Psychiatry. 197, 158, author reply 158–159, doi:10.1192/bjp.197.2.158 (2010).
Cross-Disorder Group of the Psychiatric Genomics, C. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 381, 1371–1379, doi:10.1016/S0140-6736(12)62129-1 (2013).
Li, J. et al. Schizophrenia Related Variants in CACNA1C also Confer Risk of Autism. PLoS One. 10, e0133247, doi:10.1371/journal.pone.0133247 (2015).
Kilpinen, H. et al. Association of DISC1 with autism and Asperger syndrome. Mol Psychiatry. 13, 187–196, doi:10.1038/sj.mp.4002031 (2008).
Hennah, W. et al. Haplotype transmission analysis provides evidence of association for DISC1 to schizophrenia and suggests sex-dependent effects. Hum Mol Genet. 12, 3151–3159, doi:10.1093/hmg/ddg341 (2003).
Gauthier, J. et al. De novo mutations in the gene encoding the synaptic scaffolding protein SHANK3 in patients ascertained for schizophrenia. Proc Natl Acad Sci USA. 107, 7863–7868, doi:10.1073/pnas.0906232107 (2010).
Rujescu, D. et al. Disruption of the neurexin 1 gene is associated with schizophrenia. Hum Mol Genet. 18, 988–996, doi:10.1093/hmg/ddn351 (2009).
Kim, H. G. et al. Disruption of neurexin 1 associated with autism spectrum disorder. Am J Hum Genet. 82, 199–207, doi:10.1016/j.ajhg.2007.09.011 (2008).
Aoki-Suzuki, M. et al. A family-based association study and gene expression analyses of netrin-G1 and -G2 genes in schizophrenia. Biol Psychiatry. 57, 382–393, doi:10.1016/j.biopsych.2004.11.022 (2005).
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature. 485, 246–250, doi:10.1038/nature10989 (2012).
Schaub, M. A., Boyle, A. P., Kundaje, A., Batzoglou, S. & Snyder, M. Linking disease associations with regulatory information in the human genome. Genome Res. 22, 1748–1759, doi:10.1101/gr.136127.111 (2012).
Albert, F. W. & Kruglyak, L. The role of regulatory variation in complex traits and disease. Nat Rev Genet. 16, 197–212, doi:10.1038/nrg3891 (2015).
Nicolae, D. L. et al. Trait-associated SNPs are more likely to be eQTLs: annotation to enhance discovery from GWAS. PLoS Genet. 6, e1000888, doi:10.1371/journal.pgen.1000888 (2010).
Alter, M. D. et al. Autism and increased paternal age related changes in global levels of gene expression regulation. PLoS One. 6, e16715, doi:10.1371/journal.pone.0016715 (2011).
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J Neurosci. 28, 264–278, doi:10.1523/JNEUROSCI.4178-07.2008 (2008).
Schopler, E., Reichler, R. J., DeVellis, R. F. & Daly, K. Toward objective classification of childhood autism: Childhood Autism Rating Scale (CARS). J Autism Dev Disord. 10, 91–103, doi:10.1007/BF02408436 (1980).
Krug, D. A., Arick, J. & Almond, P. Behavior checklist for identifying severely handicapped individuals with high levels of autistic behavior. J Child Psychol Psychiatry. 21, 221–229, doi:10.1111/jcpp.1980.21.issue-3 (1980).
Guo, L., Du, Y., Qu, S. & Wang, J. rVarBase: an updated database for regulatory features of human variants. Nucleic Acids Res. 44, D888–893, doi:10.1093/nar/gkv1107 (2016).
Ramasamy, A. et al. Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat Neurosci. 17, 1418–1428, doi:10.1038/nn.3801 (2014).
Consortium, G. T. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 45, 580–585, doi:10.1038/ng.2653 (2013).
Storm, N., Darnhofer-Patel, B., van den Boom, D. & Rodi, C. P. MALDI-TOF mass spectrometry-based SNP genotyping. Methods Mol Biol. 212, 241–262 (2003).
Jurinke, C., van den Boom, D., Cantor, C. R. & Koster, H. The use of MassARRAY technology for high throughput genotyping. Adv Biochem Eng Biotechnol. 77, 57–74, doi:10.1007/3-540-45713-5 (2002).
Ho, D. W. H. et al. Genotyping Performance Assessment of Whole Genome Amplified DNA with Respect to Multiplexing Level of Assay and Its Period of Storage. PLoS One. 6, e26119, doi:10.1371/journal.pone.0026119 (2011).
Verma, M., Kulshrestha, S. & Puri, A. Genome Sequencing. Methods Mol Biol. 1525, 3–33, doi:10.1007/978-1-4939-6622-6_1 (2017).
Rabinowitz, D. & Laird, N. A unified approach to adjusting association tests for population admixture with arbitrary pedigree structure and arbitrary missing marker information. Hum Hered. 50, 211–223, 22918 (2000).
Pletikos, M. et al. Temporal specification and bilaterality of human neocortical topographic gene expression. Neuron. 81, 321–332, doi:10.1016/j.neuron.2013.11.018 (2014).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210, doi:10.1093/nar/30.1.207 (2002).
Acknowledgements
We thank all the subjects and their families for their sponsorship and participation in this study. This research was supported by grants from the National Natural Science Foundation of China (grant numbers 81471383, 81501183, 81471360, and 81671363), and Beijing Municipal Science & Technology Commission (No. Z131107002213100).
Author information
Authors and Affiliations
Contributions
D.Z., L.F.W. and J.L. conceived and designed the experiments. J.Q., Z.L.W., J.L., Y.Y., Y.L.M., L.F.W., L.Y.W., Y.Y.R. and T.L.L. performed the experiments. Z.L.W., J.Q., J.L. and W.H.Y. analyzed the data. M.X.J. collected the samples. Z.L.W., J.Q., L.F.W., J.L. and L.Y.W. wrote the paper.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, Z., Qin, J., You, Y. et al. Genetic variants in the transcription regulatory region of MEGF10 are associated with autism in Chinese Han population. Sci Rep 7, 2292 (2017). https://doi.org/10.1038/s41598-017-02348-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-02348-1
This article is cited by
-
Prenatal delta-9-tetrahydrocannabinol exposure is associated with changes in rhesus macaque DNA methylation enriched for autism genes
Clinical Epigenetics (2023)
-
eQTL colocalization analysis highlights novel susceptibility genes in Autism Spectrum Disorders (ASD)
Translational Psychiatry (2023)
-
Discovery of eQTL Alleles Associated with Autism Spectrum Disorder: A Case–Control Study
Journal of Autism and Developmental Disorders (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.