Abstract
The expanding global distribution of multi-resistant Klebsiella pneumoniae demands faster antimicrobial susceptibility testing (AST) to guide antibiotic treatment. Current ASTs rely on time-consuming differentiation of resistance and susceptibility after initial isolation of bacteria from a clinical specimen. Here we describe a flow cytometry workflow to determine carbapenem susceptibility from bacterial cell characteristics in an international K. pneumoniae isolate collection (n = 48), with a range of carbapenemases. Our flow cytometry-assisted susceptibility test (FAST) method combines rapid qualitative susceptible/non-susceptible classification and quantitative MIC measurement in a single process completed shortly after receipt of a primary isolate (54 and 158 minutes respectively). The qualitative FAST results and FAST-derived MIC (MICFAST) correspond closely with broth microdilution MIC (MICBMD, Matthew’s correlation coefficient 0.887), align with the international AST standard (ISO 200776-1; 2006) and could be used for rapid determination of antimicrobial susceptibility in a wider range of Gram negative and Gram positive bacteria.
Similar content being viewed by others
Introduction
The O’Neill Review on Antimicrobial Resistance (AMR) estimated that 700,000 people die from infections due to resistant organisms every year, and by 2050 AMR will surpass cancer as a cause of death1. The World Health Organization (WHO) recognises AMR as a serious threat to global health2, and singled out the emergence of carbapenem-resistant Klebsiella species as the leading priority in its first global report on AMR in 20143. Klebsiella species are the most prominent carbapenem-resistant Enterobacteriaceae (CRE) and cause an excess hospital mortality of 27% in patients with septicaemia and pneumonia4. The Indian Ocean Rim region has become one of the main foci of emerging carbapenemases, and has seen successive waves of diverse forms of carbapenem-resistant K. pneumoniae 5, 6.
Faster antimicrobial susceptibility test (AST) methods are an essential component of the multi-faceted measures needed to reduce inappropriate antibiotic use and combat the rise of AMR1, 7. Detection of multidrug-resistant bacteria currently relies on primary isolation followed by largely culture-dependent AST procedures, delaying the commencement of targeted treatment and infection prevention and control measures by 24–72 hours. Inappropriate, broad-spectrum antimicrobials are used in the absence of empirical laboratory results, prompting the search for faster methods5. Current rapid non-culture-based screening methods such as mechanism-specific PCR and the widely used Carba-NP test, can be unreliable8, which diminishes their value for predicting carbapenem susceptibility and thus their utility for the prescribing physician. MIC determination by broth microdilution is the internationally recognized standard for AST (ISO 200776-1, 2006)9. A categoric classification (susceptible, intermediate or resistant – SIR, or susceptible/non-susceptible – S/NS) can be made by comparing the MICs to species-specific breakpoints issued either by the European Committee on Antimicrobial Susceptibility Testing (EUCAST) or the Clinical & Laboratory Standards Institute (CLSI). All methods for antimicrobial susceptibility testing must be validated against broth microdilution before introduction into clinical practice10.
Flow cytometry has long been considered a candidate method for delivering rapid AST11. Early flow cytometry analyses of bacteria were limited by their low resolution to studies of cellular aggregation, but the introduction of bacterial viability dyes, improved flow cytometer resolution, and increased sophistication of multi-parameter analysis has prompted renewed attempts to establish a method for flow-assisted antimicrobial susceptibility analysis12,13,14,15,16. These studies produced a catalogue of complex interactions between membrane-permeable dyes and bacteria during sub-lethal damage, but the resolution limits of the best (previously) available hydrodynamic flow cytometers still constrained Flow cytometer-Assisted Susceptibility Test (FAST) methods17, 18 with little progress in making FAST methods widely accessible.
Here we report an acoustic flow cytometry (AFC) method capable of rapidly determining carbapenem MICs and assigning susceptibility categories. Furthermore, we compared the method against broth microdilution in a blinded, prospective validation using a collection of carbapenem-susceptible and carbapenem-resistant K. pneumoniae and K. oxytoca isolates, including internationally dominant ESBL- and carbapenemase-producing isolates.
Materials and Methods
Bacterial isolates
We assembled an internationally representative panel of carbapenemase-producing K. pneumoniae isolates from: the Western Australian Culture Collection (WACC, PathWest Laboratory Medicine, Western Australia which included American Type Culture Collection ATCC K. pneumoniae: ATCC BAA1705, ATCC BAA1706 and ATCC 700603); Public Health England’s Antimicrobial Resistance and Healthcare Associated Infections (AMRHAI) Reference Unit, (London, UK); the Norwegian Public Health Institute (Oslo, Norway); and the EUCAST development Laboratory (Växjö, Sweden) (Table 1). The carbapenem MICs for all referred isolates were known, as was the mechanism of carbapenem resistance for the majority of resistant isolates (Table 2). Prior to FAST and parallel broth microdilution MIC analysis, bacteria were recovered from frozen storage in accordance with ATCC guidelines to provide a standard cell density in suspension19, as illustrated in Fig. 1.
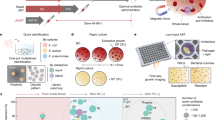
Schematic diagram of flow cytometry-assisted susceptibility test method. Demonstration of the workflow used. In brief, an isolate was retrieved from cryopreservation, plated onto blood agar to ensure purity, and inoculated into trypticase soya broth (TSB) overnight to simulate biological fluids. A 1 ml aliquot of this suspension was inoculated into Mueller-Hinton broth (MHB) and incubated at 37 °C to ensure bacteria were actively dividing. One aliquot was subjected to traditional microbroth dilution (MBD) susceptibility testing. Another aliquot was exposed to meropenem for 30 minutes, harvested, stained with SYTO®9, and then assayed with a flow cytometer. This figure is not covered by the CC-BY licence. [Credit to Life Technologies Corporation, a part of Thermo Fisher Scientific Inc. www.thermofisher.com. ©2016 Thermo Fisher Scientific Inc. Used under permission.] All rights reserved, used with permission.
Antimicrobial agents
Lyophilised meropenem (Ranbaxy, Haryana, India), imipenem, ertapenem and meropenem (Sigma-Aldrich, Missouri, USA) antibiotics were dissolved in sterile 0.85% saline to produce 5120 mg/L stock solutions, syringe-filtered at 0.1 µm and stored below −20 °C. FAST meropenem working stocks were made by serial 1:2 dilutions in filtered sterile Mueller-Hinton broth (MHB) to produce 1 mL aliquots ranging from 2560 mg/L to 2.5 mg/L.
Preparation of bacteria
Fluids for bacterial preparations and acoustic flow cytometer operation were filtered at 0.1 µm prior to use to minimise particulate contamination. A 1 mL aliquot of bacterial suspension was centrifuged, washed and resuspended in 1 ml filtered Hank’s Buffered Salt Solution (HBSS) and diluted in series to 1:1000. SYTO® 9 stain working stock solution (1 µl) was added to the final dilution at a final concentration of 5 µM and incubated for 5 minutes before determination of bacterial count by flow cytometer (Attune, ThermoFisher, Eugene, OR, USA), which was used to prepare a standardised inoculum density for susceptibility testing.
An aliquot of bacterial suspension was added to 50 mL centrifuge tubes (Corning, New York) containing 9 mL of pre-warmed (37 °C) filtered MHB to produce a final density of 5 × 105−1 × 106 bacteria per mL. This suspension was incubated at 37 °C with shaking at 100 RPM for 30 minutes to obtain an actively dividing culture. An aliquot of antibiotic working solution from the previously described dilution series (appropriate to the concentration tested) was then added to each tube before a further 30-minute incubation with shaking, during which time a microbroth dilution (MBD) plate was prepared for overnight incubation at 37 °C20. One millilitre of bacterial suspension from each antibiotic concentration was harvested by centrifugation at 7800 × g for 5 minutes, washed, resuspended and diluted 1:10 in filtered HBSS in a light-impermeable microcentrifuge tube, then stained with SYTO® 9 at a final concentration of 5 µM, and incubated for 5 min before AFC sampling. Hoechst 33342 dye (NucBlue, Thermo Fisher Scientific, Eugene, OR, USA) and a SYTO® 9/propidium iodide (PI) combination (Thermo Fisher Scientific) were used in a series of replicate experiments (data not shown).
Acoustic Flow Cytometer (AFC) operation and data analysis
The acoustic flow cytometer was calibrated at the beginning of each acquisition session in accordance with the manufacturer’s instructions (ThermoFisher Scientific). Flow cytometer settings were: Forward Scatter (FSC) voltage 3100, FSC threshold 4 × 1000 AND, blue laser 1 (BL1 – 530/30 nm) voltage 1900, BL1 threshold 1 × 1000 AND, high sensitivity, flow rate 25 µL/minute, and an acquisition volume of 125 µL. Acquisition halted after collection of 20,000 events across all gates, or after 3 minutes, with each sample acquired in technical triplicate.
Collected data were exported in the FCS 3.0 file format and analysed in Flow v10.0 (FlowJo LLC, Ashland, OR, USA) by a single user, blinded to the MIC results by broth microdilution (MICBMD).
Digital fluorescence microscopy
A 1 mL aliquot of growth from each antimicrobial concentration was harvested while conducting AFC measurements of antimicrobial-exposed bacteria, centrifuged at 7800 × g for 5 minutes, and resuspended in 10 µl of HBSS. A 0.1 µL aliquot of SYTO® 9 was added to each tube (final concentration 50 µM) and incubated for 5 minutes. A 2 µl aliquot was placed on a poly-L-lysine slide, sealed beneath a coverslip, and observed at 60x magnification by digital fluorescence microscopy. Samples were observed on the EVOS-FL digital fluorescence microscopy platform (Thermo Fisher, Eugene OR), with a representative field of view captured for each sample across the antimicrobial dilution series. High-resolution images were acquired using a Nikon Ts2R Eclipse inverted digital fluorescence microscope (Nikon, Tokyo, Japan).
Qualitative susceptibility testing
We sought to determine whether a qualitative susceptibility test could be developed using the FAST platform. Subsequent to quantitative MICFAST determination, the flow cytometer data were re-analysed in silico to produce a limited sub-set for qualitative susceptibility determination. The six antibiotic dilutions (0 mg/L, 0.25 mg/L, 1 mg/L, 2 mg/L, 4 mg/L and 16 mg/L) most relevant to qualitative susceptibility assessment were used. In silico analysis was restricted to the first technical triplicate of each recorded sample. Gating strategies remained consistent, with the addition of a gate restricting analysis to only those events recorded in the first 60 seconds of acquisition. Isolates were defined as meropenem susceptible (S) or non-susceptible (NS) using EUCAST clinical breakpoints for Enterobacteriaceae (S ≤2 mg/L, NS >2 mg/L).
Statistical analysis
Statistical software (Prism v 6.1, GraphPad, San Diego, CA, USA) was used to analyse both SIR categorization and quantitative MIC results. SIR results were analysed using a χ2 format. Clinical laboratory test performance measurements (sensitivity, specificity, positive predictive value, negative predictive value, Matthews Correlation Coefficient) were used to assess the ability of the FAST method to correctly determine carbapenem susceptibility. The correlation between MICBMD and MICFAST was analysed by calculating Spearman’s coefficient for non-parametric data. The MIC data were plotted on a log-log biaxial plot, using the microdilution results as the determinant.
Discrepancy investigation
We examined in more detail all isolates demonstrating anomalous S/NS categories (MICBMD vs. MICFAST), or MIC discrepancies outside the accepted tolerance of the microbroth dilution assay (+/− one two-fold dilution step). These isolates were subcultured once per day for three days to exclude the possibility of low prevalence contamination of cryo-preserved stocks by bacteria other than Klebsiella species. Any isolates displaying variable colony appearance on solid media had each observed colony morphotype sub-cultured separately, and their identity verified. We reconfirmed the molecular basis of carbapenem resistance using mechanism-specific PCR assays5, 8. In cases where contaminants or complex resistance mechanisms were identified, isolates were subjected to a further round of FAST following determination of identity and molecular basis of resistance.
Subpopulation investigation
Populations observed on bi-variate flow cytometry plots that seemed to segregate into two populations across a dilution series were observed in many isolates during validation. To investigate this, we selected a demonstrative example (K16, an IMP-4 producing isolate) and subjected it to our FAST assay. We referred to this as “Day one”. One mL of the 4 mg/L meropenem exposed culture was harvested, washed in fresh HBSS to remove the presence of meropenem, and inoculated into fresh TSB to provide input for a second round of FAST on “Day two”. MICFAST, population shapes, and progression to susceptibility-associated signature were compared between both experiments.
Results
A rapid flow-assisted susceptibility test for meropenem
Following our initial development process, we developed a new method (Fig. 1) by which susceptibility to meropenem can be assayed in K. pneumoniae. Using an acoustic flow cytometer to obtain optimal resolution of small particles, and a nucleic acid intercalating fluorophore to discriminate bacterial events from background debris, optimal results were achieved with SYTO®9. We used changes in size, shape, cytoplasmic volume and overall event numbers to predict susceptibility to meropenem in 1 hour, and MIC in 3 hours.
Defining meropenem susceptibility by AFC
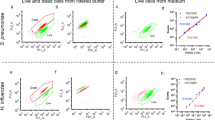
Susceptibility to meropenem was defined by careful pairing of observed shifts in FSC and BL1 fluorescence (530/30 nm – ideal collection for SYTO®9) in bi-axial AFC plots, and observation of bacterial structures consistent with meropenem compromise by fluorescence microscopy. Exposure of actively dividing meropenem-susceptible isolates to inhibitory concentrations of the drug has been demonstrated to produce a range of cellular morphotypes; cells elongate, swell, balloon, and eventually proceed to complete cell lysis as they become compromised21. When microscopy and biaxial AFC plots were compared, an increased prevalence of aberrant cell morphotypes (consistent with meropenem compromise) was found to correlate with an increase in FSC, increased BL1 fluorescence, and formation of populations that contour independently on biaxial plots. In a meropenem-susceptible isolate, these changes were observed at the lowest concentration tested (Fig. 2A), whereas in an isolate with a raised meropenem MIC, these changes were not observed at concentrations below the MICBMD (Fig. 2B). When a non-susceptible isolate was exposed to concentrations approaching or exceeding its elevated MICBMD, we observed forward scatter and BL1 changes associated with susceptibility. We refer here to this progressive change in morphotype approaching, and exceeding the MICBMD as the susceptibility-associated signature.
Standardised gating applied to raw data. (A) Collected events were gated to include only those with a SYTO®9 (BL1 - 530/30 nm) fluorescence of 104 arbitrary fluorescence units or higher. Doublets were removed via a FSC-A vs FSC-H plot. Background was removed by plotting specific SYTO®9 fluorescence (BL1 – 530/30) against an unused channel (BL3 – 640 LP). (B) In the antibiotic unexposed sample, 10% nearest-neighbour contouring was applied, and a gate (referred to as Unexposed Cell Morphotype) was set to include all clustered events. This gate was then applied to all samples across the antibiotic dilution series.
Standardised gating strategies result in reproducible quantitative end-point report
Events of interest were defined by SYTO® 9 fluorescence greater than 104 arbitrary fluorescence units. Aggregate and co-incident events were excluded by rational gating on FSC-A vs FSC-H bivariate plots, and background fluorescence (auto-fluorescence and electronic noise) was minimised by rational gating on populations of interest by BL1-H vs BL3-H plot (Fig. 3A). Using the technical triplicate of the unexposed bacteria with the median BL1-H geometric mean fluorescence intensity, the auto gate tool (FlowJo) defined a gate that bounded all contoured events on a bivariate contour plot of FSC-H vs BL1-H at the 10% threshold (Fig. 3B). This gate and the events it bounded were referred to as the Unexposed Cell Morphotype (UCM), and the gate was then applied consistently to all samples across the antimicrobial agent dilution series. The absolute count of event numbers in this gate was calculated to give a comparable measure of UCM for each sample, standardised by volume (events/µL). Changes in the prevalence of morphotypes when bacteria were exposed to meropenem at concentrations approaching or exceeding the MICBMD were evident as a distinct susceptibility-associated signature. Iterative comparisons between UCM event rates/µL in antibiotic-exposed samples and the unexposed control samples were used to determine the flow-associated susceptibility test MIC (MICFAST).
Susceptibility to meropenem can be identified by AFC by observing a susceptibility-associated signature. (A) Exposure of the K. pneumoniae susceptible type strain (ATCC 700603) increased forward scatter, SYTO®9 fluorescence, reduced overall event numbers, and formed a new contouring focus at the isolate’s MIC (0.25 mg/L). At 32 × MIC, a total of four contouring foci were observed, with an overall shift towards low forward scatter, low fluorescent debris. The progression of these features, when observed in combination, constitutes the susceptibility-associated signature. Colouring on biaxial plots indicates separate contouring foci. Fluorescence micrographs (acquired at 60x magnification) show reduced overall cell numbers and increase aberrant cell morphotypes as meropenem concentration increases. (B) Exposure of highly resistant clinical K. pneumoniae strain K8 to meropenem shows an absence of susceptibility-associated signature across clinically relevant meropenem concentrations by flow cytometer bi-axial plot, and an absence of aberrant cell morphotypes by fluorescence microscopy.
Prediction of MICBMD by FC
Our initial range-finding series demonstrated close correspondence between the meropenem concentrations that caused appearance of the susceptibility-associated signature in each of 10 isolates and their corresponding MICBMD values. We compared numbers of events bounded by the UCM gate per µL (UCMµ) in antibiotic-exposed samples with unexposed control samples, with a particular focus on cell numbers falling into and out of gated regions in those samples displaying the susceptibility-associated signature. We observed that the flow cytometer results accurately predicted meropenem MICBMD when a cut-off point was established as the first concentration in an antimicrobial dilution series in which two or more of the technical replicates had less than 30% of events falling into the UCM gate when compared to the unexposed control (Table 1 and Fig. 4A). We refer to this concentration as the MICFAST. Paired MICBMD and MICFAST results for the entire isolate collection are shown in Table 2. There was a strong positive correlation between MICBMD and MICFAST across the entire isolate collection (Spearman r = 0. 913, p < 0.0001, sensitivity 1.00, specificity 0.90, positive predictive value 0.86, negative predictive value 1.00 and Matthew’s correlation coefficient of 0.878) (Fig. 4B). MICFAST determination required 158 minutes from actively growing culture (35 minutes of incubation, 12 minutes for manual handling, 108 minutes for data acquisition, and 3 minutes of data interpretation from a pre-prepared workspace template). Qualitative meropenem susceptibility was assessed for the entire isolate collection from the previously described data subset. Three isolates were incorrectly determined; two isolates (KS1, OXA-181, and 500638, pAmpC) were incorrectly categorised as susceptible despite being non-susceptible (MICFAST 2, MICBMD 4) however, this two-fold inter-test variation is within the accepted tolerance of the broth microdilution assay. Isolate K16 (IMP-4, MICBMD 16; MICFAST 2) was the subject of extensive further investigation. Despite these isolates, the FAST susceptible/non-susceptible threshold/interpretive criterion was highly concordant with broth microdilution-derived susceptibility (χ2 = 37.03, df = 1, p < 0.0001, sensitivity 1.00, specificity 0.90, positive predictive value 0.875, negative predictive value 1.00 Matthew’s correlation coefficient 0.887). Based on the conditions selected for the data set assembly, the theoretical time-to-result for this qualitative test was 54 minutes from actively dividing culture (35 minutes of incubation, 11 minutes for manual handling, six minutes for data acquisition, and two minutes of data interpretation from prepared workspace template).
FAST accurately predicted MICBMD across our collection of K. pneumoniae isolates. Coloured regions represent the qualitative susceptibility (green)/non-susceptible (red) determination of (A) the initial 10 isolates tested (mean MICBMD/FAST and SEM), a strong positive correlation was observed (r = 0.899, p < 0.0001), represents four isolates, all with perfectly concordant MICBMD,FAST coordinates. (B) Full 48 isolate collection. Numerals indicate the number of isolates occupying the same MICBMD, MICFAST coordinates. Across the full collection of isolates, a strong positive correlation was observed (Matthew’s correlation co-efficient = 0.918).
FAST can be applied to other carbapenems
To examine the applicability of the FAST method to other carbapenems, carbapenem-susceptible (ATCC 700603) and -resistant (ATCC BAA 1705) control strains of K. pneumoniae were exposed to analytical grade meropenem, imipenem, ertapenem, and therapeutic grade meropenem. There was no difference between S/NS categorisation between MICBMD and qualitative FAST S/NS across all tested conditions (Table 3).
MICBMD vs MICFAST discrepancy analysis
Only five of 48 (10.4%) isolates showed discrepancies between MICBMD and MICFAST that resulted in a different meropenem S/NS assessment. The first, isolate 374, was found to contain a low-prevalence Staphylococcus aureus contaminant. Analysing the pure K. pneumoniae growth produced perfect concordance between MICBMD and MICFAST.
Three of these isolates, two with an OXA-48-family enzyme and one with an IMP-4 (3000763, KS11 and K23 respectively) had MICBMD and MICFAST values within the two-fold dilution tolerance of the BMD method, but straddled the EUCAST breakpoint. This is an error of classification, not an inaccuracy of our method. The remaining isolate (K16, IMP-4) initially produced a MICBMD of 16 mg/L and a MICFAST of 2 mg/L. This was sub-cultured once again to check for purity, whereupon smooth and rough colony variants were observed. Retesting of each colony type produced an MICFAST of 2 and 64 mg/L, respectively (Fig. 5). The MICBMD (16 mg/L) and MICFAST of the rough colony variant (64 mg/L), are within the resolution of the BMD test, and result in a concordant S/NS determination.
Differences in MICFAST were observed between colony variants of K. pneumoniae isolate K16. When subculturing IMP-4 producing K. pneumoniae isolate K16, a rough and smooth colony variant was observed. The smooth colony variant produced an MICFAST of 2 mg/L. The rough colony produced an MICFAST of 64 mg/L and, at 2 mg/L, was observed by AFC to contain a population consistent with a non-susceptible phenotype. Both MICFAST results were concordant with the MICBMD results.
Identification of subpopulation and persisting populations in isolates with discrepant MIC results
On Day 1, isolate K16, classified as susceptible by MICFAST, was found to contain a population of bacterial events consistent with unexposed cell morphotypes that persisted until 16 mg/L (Fig. 6 - Day 1). On Day 2 the bacterial population characteristics exhibited a different progression towards the susceptibility-associated signature across the same meropenem dilution range. Bacterial cells had a much higher forward scatter, without an associated BL1 increase on Day 2, and starting at 4 mg/L a subpopulation of cells again became evident (Figure Day 2). Subpopulations such as these have been observed across approximately one third of isolates assayed in our collection (n = 17).
Resistant sub-populations were observed in K. pneumoniae isolate K16 across two days of selective passage and FAST: Day One - IMP-4 producing K. pneumoniae isolate K16 was found, at 2 mg/L meropenem, to contain a minority population of cells with a phenotype consistent with unexposed cells (remaining within the Unexposed Cell Morphotype gate - indicated by arrow). This subpopulation persisted, at a diminished frequency, at 16 mg/L meropenem while the majority of cells display a compromised phenotype (shifted outside the gate). Day 2 – The 2 mg/L culture of K16 from Day 1 was subcultured and subjected to FAST on the following day. The isolate displayed an increased MIC (4 mg/L), delayed progression to the emergence susceptibility-associated signature, with most events consistent with a non-susceptible phenotype at 2 mg/L. Most events at 16 mg/L were consistent with a susceptible phenotype, however a small subpopulation remained inside the Unexposed Cell Morphotype gate.
Discussion
Antimicrobial susceptibility profiling of carbapenem-resistant K. pneumoniae by acoustic flow cytometer predicted both quantitative (MIC) and qualitative (susceptible/non-susceptible) carbapenem susceptibility. While flow cytometry has been used for antimicrobial susceptibility testing before11,12,13,14,15,16,17,18, our FAST assay is the first reported description of a validated method to generate a clinically-relevant quantitative end-point. Furthermore, our rapid phenotypic determination of antimicrobial susceptibility accurately predicts the qualitative result, and is therefore a significant step towards alignment of laboratory testing with clinical decision timelines. Broth microdilution is too labour-intensive for use in most clinical laboratories, which favour other methods of susceptibility determination. We present performance statistics for our qualitative susceptibility test but to demonstrate the power of single-cell level analysis rather than to expect immediate adoption of this assay in the clinical laboratory. To the prescribing physician, rapid qualitative susceptibility represents an ability to align the decision-making process of antibiotic prescribing to the best-practice ideals of effective anti-microbial stewardship7.
The FAST method is suitable for application as a rapid method to determine carbapenem resistance phenotype on the grounds of a strong correlation between MICBMD and MICFAST. MICFAST follows a pre-determined heuristic to generate quantitative results, rather than relying on potentially user-biased subjective end-points. Our use of workspace templates allowed replication of results by non-specialists after minimal instruction by a skilled operator using a proprietary software package (FlowJo). Furthermore, any flow cytometry software capable of generating a contouring output should be suitable. The FAST assay is underpinned by the reproducible flow cytometry model of a complex series of physiological interactions we established. Forward Scatter (FSC) is often used a surrogate for particle size, but this oversimplifies the dynamic behaviour of non-spherical particles17. There is much more information in this single measurement than the size and orientation of a particle passing through the flow cytometer. For example, changes in granularity and autofluorescence profiles also alter the absolute numbers of photons reaching the FSC detector, and in similar manner, photons absorbed and emitted by fluorescence signals can alter FSC measurements17, 18. Our choice of fluorescent dye (SYTO® 9) ensured that measurements in the BL1 channel (530/30 nm) contained information on DNA content, cytoplasmic volume and autofluorescence. Observed staining intensity profiles from a rigorously controlled experimental method offer additional insight into physiological properties such as membrane permeability and dye molecule efflux14. Isolates with the osmoporin ompK36 third eyelet insertion mutation (ins AA 134-135 GD5, 6) displayed a reduced BL1 intensity. This porin mutation excludes positively charged compounds such as SYTO®922 and has been shown to correlate with high-level meropenem resistance23, 24. The consistency of our observations across a collection of isolates from such diverse geographic origins and resistance mechanisms supports a conserved bacterial physiology.
The physiological response we detected by the FAST method after antimicrobial exposure resembles the range of carbapenem-induced morphotypes described previously21. Arrested cell division after inhibition of penicillin-binding proteins25, 26 leads to an overall increase in cellular DNA and increases the DNA-bound SYTO® detectable in BL1. The overall decrease in cell numbers by fluorescence microscopy and flow cytometry, and the corresponding increase in flow cytometer event populations with low forward scatter and varied BL1, is likely to reflect mixed cell debris from bacterial cell lysis during antimicrobial exposure. The broth microdilution MIC method relies on a subjective end-point and requires extended incubation27, allowing persistence of resistant sub-populations after inhibition of the susceptible majority of bacteria28,29,30. The FAST method measures the resistance phenotype of all bacterial cells in each aliquot, and adds to the evidence that carbapenem-resistant Enterobacteriaceae are phenotypically heterogeneous28,29,30. We postulate that broth microdilution over-simplifies the test method and overestimates the dose required to demonstrate antimicrobial efficacy. Highly resistant bacterial sub-populations have been implicated in failed meropenem monotherapy before28, 31. These bacteria may respond to meropenem combination therapy provided sufficient breakthrough growth has not occurred28, 31. Identification of these features of bacterial susceptibility in a shorter time could become the basis of more timely antimicrobial treatment guidance28, 29, 31. While our method eliminates the necessity of the secondary culture step required for either broth microdilution or other growth-dependent quantitative susceptibility determination27, further advances are needed to purify bacteria directly from patient samples so that laboratory results are available to the physician within a shorter time frame, particularly for patients with sepsis and other severe infections.
Discrepancies of ≥2 two-fold dilutions were observed between MICBMD and MICFAST for pAmpC- and IMP-4-producing isolates. These types of resistance cause inducible meropenem resistance30, 32. Induction of meropenem-resistant pAmpC-producing K. pneumoniae has been demonstrated after accumulation of transpeptidation by-products in the cytosol32,33,34,35. Selection of low-prevalence sub-populations with constitutive AmpC can also lead to rapid, time-dependent shifts in the overall resistance phenotype33, 34. Induction of expression does not occur within 30 minutes of antimicrobial exposure and may therefore contribute to discrepancies between MICBMD and MICFAST. In the case of IMP-4-producing isolates, high-level induced meropenem resistance is thought to be caused by intrinsically-resistant sub-populations30. The presence of persistent bacterial populations at higher meropenem concentrations in the UCM gate indicates a sub-population of inducible IMP-4-mediated meropenem resistant cells. Identification of inducible resistance is a challenge with any antimicrobial susceptibility test, but determination of the result shortly after the start of antimicrobial exposure should reduce the complex effects of prolonged antimicrobial exposure and improve the accuracy of test endpoints.
Our assay for MICFAST,has potential research applications such as resistant phenotype surveillance5,6,7, determining the impact of altered environmental/chemical conditions on resistance phenotype24,25,26,27,28,29,30,31,32,33,34,35,36,37,38, and modelling the interactions between mixed bacterial populations following antimicrobial insult. A potential use for this assay is its incorporation in a suite of orthogonal analyses that combine genomic and phenotypic investigations to assess the physiological features of a particular resistance phenotype24,25,26, 30,31,32,33,34,35,36.
The precision of our method for determining quantitative and qualitative susceptibility to meropenem in Klebsiella species compares favourably with the current international standard, while returning results in 1 (qualitative) to 3 (quantitative) hours after receipt of primary culture – in most cases a full 24 hours earlier than current standard practice. Transition from subjectively interpreted end-points to objectively-generated, single bacterial cell analysis can improve the resolution of an antimicrobial susceptibility test, without sacrificing either precision or specificity.
Change history
24 April 2018
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
The Review on Antimicrobial Resistance. Tackling drug-resistant infections globally: Final report and recommendations (ed. O’Neill, J.) 1–72 (HM Government, 2016).
World Health Organization. Antimicrobial resistance. Factsheet no. 194. World Health Organisation Media Centre. http://www.who.int/mediacentre/factsheets/fs194/en/HO AMR challenge (2014).
World Health Organization. WHO’s first global report on antibiotic resistance reveals serious, worldwide threat to public. World Health Organisation Media Centre health, http://www.who.int/mediacentre/news/releases/2014/amr-report/en/ (2014).
Hauck, C. et al. Spectrum of excess mortality due to carbapenem-resistant Klebsiella pneumoniae infections. Clin. Microbiol. Infect. 22, 513–519, doi:10.1016/j.cmi.2016.01.023 (2016).
Hall, J. M. et al. Molecular mechanisms of β-lactam resistance in carbapenemase-producing Klebsiella pneumoniae from Sri Lanka. J. Med. Microbiol. 63, 1087–92, doi:10.1099/jmm.0.076760-0 (2014).
Hall, J. M., Ingram, P. R., O’Reilly, L. C. & Inglis, T. J. J. Temporal flux in beta-lactam resistance among Klebsiella pneumoniae in Western Australia. J. Med. Microbiol. 65, 429–437, doi:10.1099/jmm.0.000242 (2016).
O’Neill, J. Rapid diagnostics: Stopping unnecessary use of antibiotics The Review on Antimicrobial Resistance 1–33 (HM Government, 2015).
Goire, N. et al. The implications of endemic IMP-4 carbapenemase for clinical laboratory susceptibility testing. J. Microbiol. Methods. 124, 10–2, doi:10.1016/j.mimet.2016.03.001 (2016).
ISO 20776-1:2006. Clinical laboratory testing and in vitro diagnostic test systems – Susceptibility testing of infectious agents and evaluation of performance of antimicrobial susceptibility test devices – Part 1: Reference method for testing the in vitro activity of antimicrobial agents against rapidly growing aerobic bacteria involved in infectious diseases (ISO, 2006).
Matuschek, E., Brown, D. F. J. & Kahlmeter, G. Development of the EUCAST disk diffusion antimicrobial susceptibility testing method and its implementation in routine microbiology laboratories. Clin. Microbiol. Infect. 20, O255–66, doi:10.1111/1469-0691.12373 (2014).
Martinez, O. V., Gratzner, H. G., Malinin, T. I. & Ingram, M. The effect of some β‐lactam antibiotics on Escherichia coli studied by flow cytometry. Cytometry. 3, 129–133, doi:10.1002/cyto.990030211 (1982).
Gant, V. A., Warnes, G., Phillips, I. & Savidge, G. F. The application of flow cytometry to the study of bacterial responses to antibiotics. J. Med. Microbiol. 39, 147–154, doi:10.1099/00222615-39-2-147 (1993).
Durodie, J. et al. Rapid detection of antimicrobial activity using flow Cytometry. Cytometry 21, 374–377, doi:10.1002/cyto.990210409 (1995).
Deere, D., Porter, J., Edwards, C. & Pickup, R. Evaluation of the suitability of bis-(1,3-dibutylbarbituric acid) trimethineoxonol, (diBA-C4(3)-), for the flow cytometric assessment of bacterial viability. FEMS Microbiol. Lett. 130, 165–9 (1995).
Mason, D. J. et al. Antibacterial action of ciprofloxacin. Antimicrob. Agents Chemother. 39, 2752–8, doi:10.1128/AAC.39.12.2752 (1995).
Novo, D. J., Perlmutter, N. G., Hunt, R. H. & Shapiro, H. M. Multiparameter flow cytometric analysis of antibiotic effects on membrane potential, membrane permeability, and bacterial counts of Staphylococcus aureus and Micrococcus luteus. Antimicrob. Agents Chemother. 44, 827–34, doi:10.1128/AAC.44.4.827-834.2000 (2000).
Caron, G. N., Stephens, P. & Badley, R. A. Assessment of bacterial viability status by flow cytometry and single cell sorting. J. Appl. Microbiol. 84, 988–98, doi:10.1046/j.1365-2672.1998.00436.x (1998).
Shapiro, H. M. & Nebe-von-Caron, G. Multiparameter flow cytometry of bacteria. Methods Mol. Biol. 263, 33–44, doi:10.1385/1-59259-773-4:033 (2004).
American Type Culture Collection. ATCC® Bacterial culture guide tips and techniques for culturing bacteria and bacteriophages 1–36 (American Type Culture Collection, 2015).
Clinical and Laboratory Standards Institute Performance Standards for Antimicrobial Susceptibility Testing: Informational Supplement. CLSI document M100-S25 (Clinical and Laboratory Standards Institute, 2015).
Choi, J. et al. A rapid antimicrobial susceptibility test based on single-cell morphological analysis. Sci Transl Med. 6, 10.1126/scitranslmed.3009650 (2014).
Dutzler, R. et al. Crystal structure and functional characterization of the OmpK36, the osmoporin of K. pneumoniae. Structure. 7, 425–434, doi:10.1016/S0969-2126(99)80055-0 (1999).
Landman, D., Bratu, S. & Quale, J. Contribution of OmpK36 to carbapenem susceptibility in KPC-producing Klebsiella pneumoniae. J. Med. Microbiol. 58, 1303–1308, doi:10.1099/jmm.0.012575-0 (2009).
Pages, J.-M. M., James, C. E. & Winterhalter, M. The Porin and the permeating antibiotic: a selective diffusion barrier in gram negative bacteria. Nat. Rev. Microbiol. 6, 893–903, doi:10.1038/nrmicro1994 (2008).
Kitano, K. & Tomasz, A. Triggering of autolytic cell wall degradation in Escherichia coli by beta-lactam antibiotics. Antimicrob. Agents Chemother. 16, 838–848, doi:10.1128/AAC.16.6.838 (1979).
Lakaye, B. et al. When drug inactivation renders the target irrelevant to antibiotic resistance: a case story with beta-lactams. Mol. Microbiol. 31, 89–101, doi:10.1046/j.1365-2958.1999.01150.x (1999).
Wiegand, I., Hilpert, K. & Hancock, R. E. W. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3, 163–175, doi:10.1038/nprot.2007.521 (2008).
Qureshi, Z. A. et al. Treatment outcome of bacteremia due to KPC-producing Klebsiella pneumoniae: superiority of combination antimicrobial regimens. Antimicrob. Agents Chemother. 56, 2108–2113, doi:10.1128/AAC.06268-11 (2012).
Adams-Sapper, S. et al. Rapid induction of high-level carbapenem resistance in heteroresistant KPC-producing Klebsiella pneumoniae. Antimicrob. Agents Chemother. 59, 3281–3289, doi:10.1128/AAC.05100-14 (2015).
Espedido, B., Iredell, J., Thomas, L. & Zelynski, A. Wide dissemination of a carbapenemase plasmid among gram-negative bacteria: implications of the variable phenotype. J. Clin. Microbiol. 43, 4918–4919, doi:10.1128/JCM.43.9.4918-4919.2005 (2005).
Nabarro, L. E. B. & Veeraraghavan, B. Combination therapy for carbapenem-resistant Enterobacteriaceae: increasing evidence, unanswered questions, potential solutions. Eur. J. Clin. Microbiol. Infect. Dis. 34, 2307–2311, doi:10.3389/fmicb.2016.02135 (2015).
Bartowsky, E. & Normark, S. Interactions of wild‐type and mutant AmpR of Citrobacter freundii with target DNA. Mol. Microbiol. 10, 555–565, doi:10.1111/mmi.1993.10.issue-3 (1993).
Jacobs, C. et al. Bacterial cell wall recycling provides cytosolic muropeptides as effectors for beta-lactamase induction. EMBO J. 13, 4684–4694 (1994).
Jacobs, C., Frere, J. M. & Normark, S. Cytosolic intermediates for cell wall biosynthesis and degradation control inducible beta-lactam resistance in gram-negative bacteria. Cell. 88, 823–832, doi:10.1016/S0092-8674(00)81928-5 (1997).
Nakano, R. et al. CFE-1, a novel plasmid-encoded AmpC beta-lactamase with an ampR gene originating from Citrobacter freundii. Antimicrob. Agents Chemother. 48, 1151–1158, doi:10.1128/AAC.48.4.1151-1158.2004 (2004).
Clancy, C. J. et al. Mutations of the ompK36 porin gene and promoter impact responses of sequence type 258, KPC-2-producing Klebsiella pneumoniae strains to doripenem and doripenem-colistin. Antimicrob. Agents and Chemother. 57, 5258–5265, doi:10.1128/AAC.01069-13 (2013).
Murray, J. L. et al. Mechanisms of synergy in polymicrobial infections. J. Microbiol. 52, 188–199, doi:10.1007/s12275-014-4067-3 (2014).
Roberts, A. E. L., Kragh, K. N., Bjarnsholt, T. & Diggle, S. P. The limitations of in vitro experimentation in understanding biofilms and chronic infection. J. Mol. Biol. 427, 3646–3661, doi:10.1016/j.jmb.2015.09.002 (2015).
Acknowledgements
This work is supported by grants from the Health Department of Western Australia (RTP 9), The Bill and Melinda Gates Foundation (Gates Foundation Grand Challenges, OPP 1150984), and equipment purchased by Rotarians and Lab Without Walls Inc., a not-for-profit pathology capability-building organisation. Reagents and technical operating advice were kindly provided free of charge by the Cellular Analysis Division, Thermo Fisher Scientific. Colleagues at the University of Western Australia, PathWest Laboratory Medicine WA, Public Health England, the EUCAST Development Laboratory, and the Norwegian Institute of Public Health provided assistance at different points in this work. The authors acknowledge the facilities, and the scientific and technical assistance of the Cytometry Core at the Centre for Microscopy, Characterisation & Analysis, The University of Western Australia, a facility funded by the University, State and Commonwealth Governments. M.J.E., K.L.H. and N.W. are part of PHE’s AMRHAI Reference Unit which has received financial support for conference attendance, lectures, research projects or contracted evaluations from numerous sources, including: Accelerate Diagnostics, Achaogen Inc., Allecra Therapeutics, Amplex, AstraZeneca UK Ltd., Basilea Pharmaceutica, Becton Dickinson Diagnostics, BioMérieux, Bio-Rad Laboratories, The BSAC, Cepheid, Check-Points B.V., Cubist Pharmaceuticals, Department of Health, Enigma Diagnostics, Food Standards Agency, GlaxoSmithKline Services Ltd., Henry Stewart Talks, IHMA Ltd., Kalidex Pharmaceuticals, Melinta Therapeutics, Merck Sharpe & Dohme Corp, Meiji Seika Pharma Co, Mobidiag, Momentum Biosciences Ltd., Nordic Pharma Ltd., Norgine Pharmaceuticals, Rempex Pharmaceuticals Ltd., Roche, Rokitan Ltd., Smith & Nephew UK Ltd., Trius Therapeutics, VenatoRx Pharmaceuticals and Wockhardt Ltd. We are particularly grateful to Drs Andrea Holme and Gunnar Kahlmeter for their guidance. Bill and Melinda Gates Foundation - OPP1150984 [Hall, Turnbull & Inglis]; Health Department of Western Australia, RTP9 [Inglis, Turnbull]; Rotary Club of Applecross, Western Australia [Inglis]; Lab Without Walls Inc is a registered NGO based in Western Australia [Inglis].
Author information
Authors and Affiliations
Contributions
K.T.M., J.M.H. and T.J.J.I. worked together on the first draft of the paper A.C., N.W., O.E., A.W., M.E., K.H. and U.N. provided critical input on the completed first draft and subsequent drafts. All authors reviewed the final edition of the manuscript. K.T.M. – optimisation of acoustic flow cytometer, dye selection, standard cytometry protocol development, cytometry bioinformatics and post hoc analysis, J.M.H. – Klebsiella isolate collection management, resistance mechanism confirmation, standard bacterial suspension protocol development, antimicrobial agent protocol development, statistical analysis of AST results, T.J.J.I. – project conception, initiation, preliminary scoping experiments, project planning, international isolate acquisition, confirmation of isolate identity, AST test performance analysis, X.H., N.B., and E.T. – operation of ACF for FAST determinations and broth microdilution MICs, ACF maintenance, A.C. – scrutiny of flow cytometry results, post hoc analysis and data presentation, M.E., K.H., and N.W. – AMRHAI isolate characterisation, curation and onward shipment, U.N., and A.W. - NPHI isolate characterisation, curation and onward shipment, (U.N. also method and data verification in Perth), O.E. - EUCAST isolate characterisation, curation and onward shipment, method and data verification in Perth, E.C. – curation, management and onward shipment of multi-resistant isolates from Sri Lanka. All authors contributed to writing the manuscript, reviewing its contents and approval of the final submitted version.
Corresponding author
Ethics declarations
Competing Interests
The corresponding author’s laboratory is funded by a Gates Foundation Grand Challenges award OP1150984 and a Research Translation Project grant from the Health Department of WA. Equipment has been purchased by the Health Department and a matching donation from Rotary Club of Applecross and Lab Without Walls, as stated in the manuscript.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mulroney, K.T., Hall, J.M., Huang, X. et al. Rapid susceptibility profiling of carbapenem-resistant Klebsiella pneumoniae . Sci Rep 7, 1903 (2017). https://doi.org/10.1038/s41598-017-02009-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-02009-3
This article is cited by
-
A 3D-printed microfluidic gradient concentration chip for rapid antibiotic-susceptibility testing
Bio-Design and Manufacturing (2022)
-
Determination of antibiotic susceptibility of bacteria by flow cytometric method
World Journal of Microbiology and Biotechnology (2022)
-
Optical microscopy reveals the dynamic nature of B. pseudomallei morphology during β-lactam antimicrobial susceptibility testing
BMC Microbiology (2020)
-
A fast impedance-based antimicrobial susceptibility test
Nature Communications (2020)
-
Microplate-based surface area assay for rapid phenotypic antibiotic susceptibility testing
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.