Abstract
Autosomal Dominant Polycystic Kidney Disease (ADPKD) is the most common inherited disorder of the kidneys. It is characterized by enlargement of the kidneys caused by progressive development of renal cysts, and thus assessment of total kidney volume (TKV) is crucial for studying disease progression in ADPKD. However, automatic segmentation of polycystic kidneys is a challenging task due to severe alteration in the morphology caused by non-uniform cyst formation and presence of adjacent liver cysts. In this study, an automated segmentation method based on deep learning has been proposed for TKV computation on computed tomography (CT) dataset of ADPKD patients exhibiting mild to moderate or severe renal insufficiency. The proposed method has been trained (n = 165) and tested (n = 79) on a wide range of TKV (321.2–14,670.7 mL) achieving an overall mean Dice Similarity Coefficient of 0.86 ± 0.07 (mean ± SD) between automated and manual segmentations from clinical experts and a mean correlation coefficient (ρ) of 0.98 (p < 0.001) for segmented kidney volume measurements in the entire test set. Our method facilitates fast and reproducible measurements of kidney volumes in agreement with manual segmentations from clinical experts.
Similar content being viewed by others
Introduction
Autosomal dominant polycystic kidney disease (ADPKD) is a systemic genetic disorder characterized by progressive enlargement of the kidneys caused by sustained development and expansion of bilateral renal cysts1. It is one of the leading causes of end-stage renal disease (ESRD)2 prompting dialysis or kidney transplantation in majority of the patients and is associated with extra-renal manifestations such as presence of cysts in the liver3. Renal Ultrasonography (US) is commonly performed for presymptomatic screening and assessment of ADPKD, however, imaging modalities such as Computed Tomography (CT) and Magnetic Resonance Imaging (MRI) offer higher spatial resolution thus facilitating the detection of smaller cysts4. Previous investigations have shown an association between total kidney volume (TKV) and renal function5, 6 and several studies have provided evidence for the use of TKV as an important imaging biomarker for assessment of disease severity as well as for predicting disease progression in ADPKD7,8,9,10. The European Medicines Agency (EMA) and the Food & Drug Administration (FDA) now acknowledge TKV as prognostic imaging biomarker for use in clinical trials11, 12. In order to evaluate the efficacy of novel therapies, it is therefore crucial to develop rapid and reliable methods for TKV quantification. However, segmentation of polycystic kidneys for quantifying kidney volumes is very challenging due to non-uniform renal cyst growth leading to high variability in kidney morphology. Polycystic kidneys are characterized by their markedly irregular shape and size in comparison to normal kidneys and sometimes surface irregularities are prominent due to the presence of surface cysts of different size. On both CT and MRI, most frequent clinical complications for an automated assessment of TKV include the presence of hepatic cysts (Fig. 1) which appear identical to kidney cysts as well as the presence of hemorrhagic renal cysts which appear rather dissimilar to other fluid filled cysts leading to high intensity variability within the kidney. Thus, development of a fully-automated segmentation method for fast and precise TKV estimation remains a challenging problem.
In ADPKD studies, traditional methods for TKV computation based on CT and MRI acquisitions are stereology13 and manual segmentation. For stereology, a square grid with user-specified cell positions and cell spacing is superimposed on every slice and the TKV is estimated by manually counting all grid points covering the kidney region. The accuracy of this method depends on user-specified parameters such as the cell spacing. Manual segmentation requires delineating the kidney on every slice using either an available free-hand contouring tool or an interactive segmentation method that guides the operator while delineating the region of interest. Both stereology and manual segmentation are tedious and require expert training in order to achieve an accurate assessment of TKV. As a consequence, several methods have been proposed with the goal of providing sufficiently accurate but less tedious means for TKV computation. The mid-slice method14 and the ellipsoid volume equations15, 16 have been proposed for estimating TKV from 1–2 slices. Even though these methods facilitate fast quantification, they provide only a rough estimate of TKV which could be convenient for clinical management but inadequate for clinical trials to detect smaller changes estimated to be approximately 5%7, 10 per year. Besides these approaches, segmentation of polycystic kidneys using semi-automated methods on MRI and CT have been reported previously. Among semi-automatic approaches on MRI, Daum et al.17 used 3D random walks while Racimora et al.18 proposed active contours and morphological operations for segmentation of polycystic kidneys. Moreover, methods based on region-growing and curvature motion have previously been suggested19, 20. Recently, geodesic active contours coupled with watershed edge detection have been used for semi-automatic segmentation on T2 weighted MR images21. On CT, Sharma et al.22 reported an approach based on random forests and geodesic distance volumes for 3D segmentation of polycystic kidneys from ADPKD patients with severe renal insufficiency. Among automatic approaches, Kline et al.23 described segmentation of follow-up MR images using initialization from previously performed manual segmentations of baseline MR acquisitions. In another recent work, Kim et al.24 reported a level set framework for automatic segmentation of polycystic kidneys on relatively small MRI dataset of ADPKD patients. Majority of these segmentation approaches focus on MRI and to the best of our knowledge, no study in ADPKD has evaluated a fully automated segmentation method both quantitatively and qualitatively on CT.
In recent years, Convolutional Neural Networks (CNNs) have shown superior performance in several computer vision tasks such as image classification, object detection and semantic segmentation. The main advantage of CNNs in comparison to many other machine-learning-based methods, such as random forests, is that they do not require hand-crafted features. Previous works have been reported on pixelwise labelling of objects for semantic segmentation using CNN25,26,27. In the domain of medical imaging, CNNs have previously been proposed for localization and segmentation of kidneys with mild morphological changes using patch-wise approaches on CT28, 29.
In this work, we present automated segmentation of ADPKD kidneys using fully convolutional neural networks, trained end-to-end, on slicewise axial-CT sections. We report the performance of our method quantitatively and qualitatively on 244 CT acquisitions of patients exhibiting different stages of ADPKD.
Results
For our experiments, baseline and follow-up CT acquisitions (training set = 165; test set = 79) of ADPKD patients (n = 125) with a wide range of TKV (321.2 mL–14,670.7 mL) and an estimated glomerular filtration rate (eGFR) ≥ 40 ml/min per 1.73 m 2 (Study 1) or 15 ≤ eGFR ≤ 40 ml/min per 1.73 m 2 (Study 2 and Study 3) were used. The main demographic and clinical characteristics of the patients are summarized in Table 1
. The results of study 2 and study 3 were combined together as patients in both studies show similar clinical characteristics.
Segmentation Similarity Analysis
In Fig. 2, segmentation predictions from the CNN of 4 different patients have been shown. The time required for segmentation prediction using automated segmentation method was only few seconds per patient CT acquisition, while the manual segmentation from experts required approximately 30 minutes per patient. The overall mean Dice Similarity Coefficient (DSC) between segmentations from the automated method and ground truth kidney segmentations from clinical experts was 0.86 ± 0.07 (mean ± SD) for the entire test set (n = 79). In particular, for study 1, consisting of patients with mild to moderate renal insufficiency (n = 26, Table 1), the mean DSC was 0.86 ± 0.06. Combining study 2 and study 3, consisting of patients with moderate/severe renal insufficiency (n = 53, Table 1), the mean DSC was 0.86 ± 0.08.
TKV Agreement Analysis
We performed volumetric measurement on kidney segmentations from the CNN and compared the automated TKV with the true TKV (obtained from ground truth annotations) in terms of accuracy and precision of the measurement. For study 1, as shown in Fig. 3 (top-left), there is substantial strength of association between the automated and true TKV with a concordance correlation coefficient (CCC) of 0.99 [95% Confidence Interval (CI): 0.97–0.99]. For the test set (n = 26, Table 1), the mean TKV error between automated and true measurements was −32.9 ± 170.8 mL and the mean percentage TKV error was 1.3% ± 10.3%. In addition, the mean absolute percentage error (MAPE) was 7.8% ± 6.7%. Bland Altman plots were used to further determine the agreement between the two methods. For study 1, as shown in Fig. 3 (top-right), the lower and upper limits of agreement (LOA) for percentage difference were −18.6% and 20.3%, respectively. The coefficient of variation (COV) between true and automated TKV was 6.5%.
Left: Concordance Correlation Coefficient (CCC) plots showing strength of association; Right: Bland-Altman plots showing agreement between TKV measurements. TKV measurements from automated segmentation method (main experiment) are compared with true TKV measurements from manual segmentations for study 1 (top, n = 26) and, studies 2 and 3 (bottom, n = 53).
For studies 2 and 3, in Fig. 3 (bottom-left), the test cases (n = 53, Table 1 with studies 2 and 3 combined) from automated and true TKV measurements show moderate strength of association with a CCC of 0.94 [95% CI: 0.91–0.96]. The mean TKV error between automated and true measurements was −44.1 ± 694.5 mL while the mean percentage TKV error was 6.5% ± 20.1%. The overall MAPE was 13.3% ± 16.4%. On the Bland-Altman plot, Fig. 3 (bottom-right), the lower and upper LOA were −29.6% and 38.9%. The overall COV between true and automated TKV was 17%.
The difference in the TKV measurements was found to be statistically insignificant for all the three clinical studies (p > 0.05). The automated TKV measurements showed high positive correlation with true TKV measurements for study 1 with mean correlation coefficient (Spearman’s rho) ρ = 0.97 (p < 0.001). Similarly, high positive correlations were observed for studies 2 and 3, with ρ = 0.98 (p < 0.001). In some cases with CT acquisitions containing severe liver cysts in close proximity of the kidney, the kidney volume was over-estimated due to inclusion of these cysts as false positive regions in the kidney segmentation. Example predictions mislabeled by the CNN have been shown in Fig. 4.
Cross Validation Analysis
In order to confirm the performance from the main experiment, we performed an additional 3-fold cross-validation. The details and results of the cross-validation analysis have been summarized in Table 2
. The DSC for cross-validation sets were 0.86 ± 0.1, 0.83 ± 0.8 and 0.87 ± 0.6, respectively. The MAPE ranged from 13.5 to 15. The COV for all three sets ranged from 14 to 15 while the root mean squared percentage error (RMSPE) ranged from 19 to 21.
Discussion
In this study, we present a novel method to automatically segment kidneys, and investigate its qualitative and quantitative accuracy and precision to measure TKV on a large dataset of CT acquisitions from ADPKD patients. The annual increase in TKV has been estimated to be around 5%7, 10 per year, suggesting that TKV measurement should be accurate to capture small changes over time. The most commonly used methods for kidney volume computation such as manual delineation and stereology13 are simple but time consuming and subject to intra/inter-observer variability. Alternatively, the mid-slice method14 and the ellipsoid equation15, 16 serve to provide quick TKV measurement but lead to low accuracy and precision compared to whole kidney segmentation.
On MRI, Racimora et al.18 proposed a segmentation approach yielding mean percentage TKV error of 22.0% ± 8.6% with an automated active contour algorithm that reduced to 3.2% ± 0.8% after manual post-editing efforts. Another semi-automatic approach with geodesic active contours and watershed edge detection21 achieved high accuracy with mean TKV difference of 0.19% ± 6.96%. Mignani et al.19 compared their results with stereology and reported mean percentage TKV error of −0.6 ± 5.8%, while, Turco et al.20 reported MAPE of 4.4% ± 4.1% and 4.2% ± 4.0% for the left and right kidneys, respectively. Other supervised segmentation methods based on stereology30 on MRI and random forests on CT22 have also been reported previously. However, these semi-automatic techniques are subject to intra/inter-observer variability and mostly require post-processing efforts to achieve higher accuracy leading to increase in overall processing time of TKV. Kline et al.23 proposed automatic segmentation on follow-up MR images, however, their method essentially requires previously performed manual segmentations of kidneys on baseline images as initialization for the segmentation process. In the work of Kim et al.24, a level set framework has been proposed for automatic segmentation in ADPKD. Even though their method shows good correlation between automated and manual TKV measurements, the results indicate high variability (LOA higher than ±25%) when compared with the manual method. Zheng et al.29 used patch-based CNN in combination with marginal space learning for localization of pathological kidneys prior to an active shape model for segmentation. Their results show good segmentation accuracy (DSC > 0.88) but there is substantial increase in segmentation error without CNN initialisation. Also, the presented dataset in their work appears to contain kidneys with milder morphological changes.
In this study, we trained and assessed the performance of the proposed automated segmentation method both quantitatively and qualitatively on a large CT dataset (n = 244) of patients at different stages of ADPKD, using manual segmentations from clinical experts as gold standard. In comparison with majority of the methods previously reported on TKV computation in ADPKD, our method has been evaluated on a larger TKV range (>300 mL and <15,000 mL). For study 1 with ADPKD patients at early stage of the disease and TKV range between 500 mL and 6,000 mL, the automated TKV shows very high strength of association (CCC = 0.99) with true TKV, however, for studies 2 and 3, with ADPKD patients at more advanced stage of the disease and the TKV range between 300 mL and 15,000 mL, there is moderate strength of association (CCC = 0.94) between the two methods. Similarly, the overall accuracy and precision of the TKV measurements from automated method is higher for study 1 (MAPE = 7.8% ± 6.7%; COV = 6.5%), compared to studies 2 and 3 (MAPE = 13.3% ± 16.4%; COV = 17.0%). The performance of the automated method is decreased particularly for very low TKV (<500 mL) and for extremely high TKV (>10,000 mL). This can be attributed to availability of very few instances of such small or very huge kidneys leading to poor predictions by the CNN during testing phase. However, the overall difference in TKV measurements was found to be statistically insignificant (p > 0.05) for all three clinical studies and the automated TKV measurements show high positive correlation with true TKV measurements (ρ = 0.98, p < 0.001). Moreover, the proposed method takes only few seconds for prediction of segmentation on each patient acquisition and avoids any intra/inter operator segmentation bias.
Despite the promising results, our study has some limitations. In some cases with several liver cysts in close proximity of the kidney (Fig. 4), the automated segmentation method over-estimated the kidney volume due to inclusion of liver cysts in the segmented kidney region. To potentially overcome this problem, the proposed method can be trained on 3D volumes of polycystic kidneys.
Regarding the importance visualization in Fig. 5, we see the importance of context for the segmentation especially for very rare subjects with extremely high TKV: In case of a typical patient, we see in Fig. 5 (top) that the largest change occurs when the kidneys themselves are occluded. This is not only intuitive but also confirms that the network is not confused by changes far away from the regions of interest. This highlights robustness against changes far away from the region of interest. Nonetheless, the visualized influence region extends over the object boundaries which indicates that not only the kidneys themselves, but also local context is used to find the final segmentation. For very rare cases with extremely high TKV (>13,000 mL) though, the spatial context changes entirely due to both kidneys occupying most of the abdominal region. As a consequence, the CNN cannot exploit context information leading to poor segmentation results which is also confirmed by the feature visualization experiment (Fig. 5 (bottom)): Consider particularly the upper areas (indicated by white arrows) in the annotated kidneys which exhibit low variation as kidney tissue does typically not appear in these areas indicating that the CNN is not expecting kidney tissues in this area.
Feature Visualization. Segmentation maps measuring change in DSC while occluding (shown with gray square) different parts of the image with respect to original unoccluded image as a measure of importance for the respective image region. The manually generated outline of the kidney is shown in red and black, respectively. Top: The largest change occurs when the kidneys themselves are occluded. Bottom: Same experiment for an ADPKD patient with high TKV (>13,000 mL).
Another limitation of our study is that we performed the analysis only on CT. Future work is needed to extend the proposed method to MRI, by training the CNN and specifically tune the parameters used during training for MRI images. Finally, despite using CT images with varying image quality to train the algorithm, the segmentation method turned out to be sensitive to image quality. In conclusion, we presented a fully automated method for the segmentation of polycystic kidneys from patients at different stages and severity of ADPKD using CT data. Our method can be reliably used on a TKV range of >500 and <10,000 mL, facilitating fast and reproducible measurements of kidney volumes in agreement with manual segmentations from clinical experts. The overall segmentation can be further improved by incorporating user interaction to correct mislabelled sections of CT. Particularly for high resolution CT images, this can significantly reduce the TKV computation time compared to manually tracing every section of the kidney and also, capture smaller changes in TKV over time. As a future work, the automated method can be trained on other affected organs such as the polycystic liver for computation of the liver volume in ADPKD.
Methods
Patients and CT Image Acquisition
The ADPKD patient dataset (244 CT acquisitions) used for our experiments was obtained from three different ADPKD clinical studies (Table 1): study 1 (SIRENA), study 2 (SIRENA 2), and study 3 (ALADIN 2).
The SIRENA study31 (ClinicalTrials.gov Identifier: NCT00491517) was designed to assess the risk/benefit of mTOR inhibitor therapy on disease progression in ADPKD. This randomized, crossover study compared 6-month treatment with sirolimus or conventional therapy alone on the increase in kidney volume and its compartments in adult patients with ADPKD and normal renal function or mild to moderate renal insufficiency (eGFR ≥ 40 ml/min per 1.73 m 2). The SIRENA 2 study32 (ClinicalTrials.gov Identifier: NCT01223755) was an academic, prospective, randomized, open label, blinded end point, parallel group trial that recruited adults with ADPKD and CKD stage 3b or 4. The aim of the study was to compare changes in GFR on 3-year treatment with sirolimus added on to conventional therapy or conventional treatment alone in ADPKD patients with moderate/severe renal insufficiency(15 ≤ eGFR ≤ 40 ml/min per 1.73 m 2). The ALADIN 2 Study (ClinicalTrials.gov Identifier: NCT01377246) is an ongoing multicenter, randomized, long-term (3-years) longitudinal study that recruited adult ADPKD patients with moderate/severe renal insufficiency (15 ≤ eGFR ≤ 40 ml/min per 1.73 m 2). The aim of this study is to assess the efficacy of 1 year treatment with long-acting somatostatin analogue (Octreotide LAR) compared with placebo in slowing kidney and liver growth rate in the ADPKD patients and to assess whether and to which extent this translates into slower renal function decline over 3-year follow-up. The CT images in all these clinical studies were acquired with a 64-slice CT scanner (LightSpeed VCT; GE Healthcare, Milwaukee, WI). A single breath-hold scan (120 kV; 150 to 500 mAs; matrix 512 × 512; collimation 2.5 mm; slice pitch 0.984; increment 2.5 mm) was initiated 80 seconds after the injection of 100 ml non-ionic iodinated contrast agent (Iomeron 350; Bracco, Italy) at a rate of 2 ml/s, followed by 20 ml physiologic solution at the same injection rate. The CT images from study 1 and study 2 were acquired at single centre in Bergamo, while CT images from study 3 were acquired at four different centres (Bergamo, Naples, Agrigento and Treviso) in Italy.
Data Annotation and Patient Selection
The left and the right kidneys were outlined manually by clinical experts and trained personnel using the ImageJ software (version 1.48v)33. In order to segment the kidneys, the image sequence for each CT acquisition was accessed using ImageJ software and manually delineated along the kidney border. The boundary delineation was performed using a standard protocol for all kidneys with respect to the hilum and liver cysts. To avoid inter-rater variability in the dataset, all manual segmentations were finally checked and corrected by a single operator (KS).
For our main experiment, patients were manually divided into the training and test set, trying to achieve a similar distribution in both sets based on the available TKV range (321.2 mL–14,670.7 mL), see Table 2.
Convolutional Neural Network Architecture
CNNs consist of several layers that learn hierarchy of features without relying on handcrafted features. Taking images as input, they convert raw pixels to final class scores by passing them through series of learnable convolutional filters. In order to perform tasks such as segmentation, it is necessary to obtain pixel-wise classification from the CNN. Our CNN architecture, shown in Fig. 6, follows the VGG-16 representation34 but employs only the first 10 layers comprising of convolution filters with a small receptive field of 3 × 3 and a spatial padding of 1 pixel for every convolution layer. As explained in the work of Ioffe et al.35, due to the internal covariance shift problem, it is rather difficult to train deep neural networks with saturating non-linearities. To alleviate this problem, batch normalization after every convolution layer allows proper initialization of the network by the forcing the activations to a standard Gaussian distribution and normalizing the inputs to zero mean and unit variance. In our experiments, we observed that application of batch normalization was crucial to improve the overall accuracy of the network. Each batch normalization layer is followed by a layer of neurons with Rectified Linear Unit (ReLU)36 as the activation function. In order to reduce the number of parameters, and thus the computation complexity of the network, max-pooling layers with a 2 × 2 pixel window and a stride of 2 pixels were used to progressively reduce the spatial size of the input to half the original size along both the height and the width. The described layers correspond to the feature extraction step, and to achieve pixelwise segmentation we employed series of deconvolution and unpooling layers for up-sampling the feature maps following the work of Zeiler et al.37 and Noh et al.38. The deconvolution layers essentially perform convolution like operation but in the opposite way leading to upsampling of coarse feature map into the reconstructed shape of the input. However, performing deconvolution step alone leads to a rather coarse reconstruction and therefore, the unpooling layers are required to refine the output. These unpooling layers reverse the operation of the pooling layers by preserving the locations of maximum activation that were extracted during the max-pooling step, and re-utilise these locations to place the maximum activations back to their original spatial position37. The unpooling layers and the deconvolution layers are connected to a 1 × 1 convolutional layer that maps the final feature vector to the desired foreground (kidney) and background (non-kidney) classes. The output of this final convolutional layer is connected to a multinomial logistic loss/cross-entropy loss to optimize the weights by penalizing the deviation between true and predicted labels.
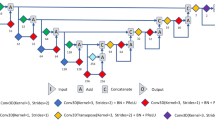
Fully Convolutional Neural Network Architecture. For feature extraction step, we used 10 layers of convolution filters with a receptive field of 3 × 3 and spatial padding of 1 pixel followed by max pooling layers with 2 × 2 pixel window and stride of 2 pixels to progressively reduce the spatial size of the input after convolution step. To achieve pixelwise segmentation, deconvolution and unpooling layers were used for upsampling the feature maps.
Data Augmentation, Training and Testing
In order to mitigate overfitting and to achieve a good generalisation, we performed two different augmentation methods on the training dataset; firstly by image shift in x-y direction and secondly by non-rigidly deforming the respective slice and applying a low frequent intensity variation. This increases the training dataset from 16000 CT slices to 48000 CT slices in case of main experiment. All this data is used for network training and allows it to learn desired invariances such as shift invariance or variable polycystic kidney shapes. Detailed information about the augmentation procedure is available as Supplementary information. All experiments were performed using the Caffe39 framework. In order to reduce the computational complexity of the network, the original size of the CT slices 512 × 512 was re-sampled to 224 × 224 and the image range was normalized to [0,255]. Additionally, we performed mean subtraction from the training dataset as a pre-processing step. Furthermore, slices were randomly shuffled before feeding to the CNN. Using a sampling size of 8 (batch-size) for each iteration, training was performed on a workstation with an Intel Xeon 8-core 2.40 GHz CPU and a NVIDIA GeForce TITANX (12 GB) GPU, taking approximately 24 hours to train the network. For optimization, we used the adaptive gradient (“AdaGrad”) method, with the learning rate set to 0.0001, and a weight decay of 0.0005, respectively. We initialized the weights in the convolution and the deconvolution layers using the Xavier initialization40.
For the test phase, the previously unseen 9000 CT slices (79 CT acquisitions) that were not included in the original training phase were passed through the trained network. The output prediction consisted of the foreground (kidney) and the background (non-kidney) pixels, where pixels with a probability higher than 0.5 were regarded as foreground (kidney) pixels. The threshold was selected based on the analysis of the Receiver Operating Characteristic (ROC) space. In conjunction to the ROC space, we also computed the Accuracy, Precision, F1 Score and the Youden-Index. Details about the threshold selection have been included in the Supplementary information.
Cross Validation
We performed an additional 3-fold cross validation on the dataset (Table 2). Out of the 244 acquisitions, 2 cases (TKV > 13,000 mL) were removed from the dataset as they are not representative over the patient population and thus do not provide adequate number of images for learning these rare cases. This was confirmed by a feature visualization experiment for one of these patients as shown in Fig. 5 (bottom). In order to perform the cross validation, the dataset was sorted according to ascending TKV range and then randomly partitioned into 3 subsets (n = 242: 80, 81, 81) allowing splitting of the dataset uniformly into the 3 cross-validation sets. For each experiment, 2 sets were used for training and the remaining set for testing. This process was repeated three times such that every data set was used once for testing.
Feature Visualization
To gain insight into the learned features and particularly the importance of context, we adapted the occlusion method from Zeiler and Fergus37 to the image segmentation domain. Their approach measures the change in classification while occluding the image with a square of fixed size and constant intensity value in a sliding window manner in order to derive the importance for each image region. Hereby in Fig. 5, we retrieve full segmentation maps from the network instead of classification scores. Thus, we compute the change in DSC with respect to the original, unoccluded image instead.
Total Kidney Volume Computation
All CT data sets were manually segmented by clinical experts and trained personnel in order to obtain ground truth annotations of the kidneys. Before computing TKV from individual segmentations obtained from the CNN, we first resampled the 224 × 224 segmentation predictions back to original size of 512 × 512 using bicubic interpolation method. Then, we performed a morphological closing operation to recover potential holes within predicted kidney regions and to remove any small isolated noise pixels wrongly predicted as foreground (kidney) pixels. Finally, TKVs were computed as the product of number of foreground pixels multiplied with the pixel spacing in x and y direction and the corresponding slice thickness.
Statistical analyses
In order to evaluate the performance of our automated segmentation method, the DSC41 was used as a statistical validation metric to assess the spatial overlap accuracy of the predicted and true manual segmentation labels. The CCC measure was used to evaluate the reliability and reproducibility of the automated method with respect to the standard manual method. Furthermore, a BlandAltman analysis was used to assess the agreement between TKV estimated from the automated segmentation method and the corresponding manual segmentation. Both absolute and relative differences were computed on the Bland-Altman plots. The COV for repeated measures42 (computed as the ratio of standard deviation to mean of the measurements) between true and automated TKV was also computed. The non-parametric Wilcoxon signed rank test for paired samples was used to measure the statistical significance of correlation between automated and true TKV measurements. Additionally, Spearman’s rank correlation coefficient (ρ) was employed as a non-parametric test to measure the strength of association between the automated and true TKV measurements. Finally, we computed the MAPE and RMSE with respect to TKV in order to provide error measures which are not sensitive to under-estimations and over-estimations cancelling out each other. All statistical analyses were performed using R studio43 version 0.98.953.
References
Grantham, J. J. The etiology, pathogenesis, and treatment of autosomal dominant polycystic kidney disease: recent advances. American journal of kidney diseases 28, 788–803, doi:10.1016/S0272-6386(96)90378-9 (1996).
Torres, V. E., Harris, P. C. & Pirson, Y. Autosomal dominant polycystic kidney disease. The Lancet 369, 1287–1301, doi:10.1016/S0140-6736(07)60601-1 (2007).
Pirson, Y. Extrarenal manifestations of autosomal dominant polycystic kidney disease. Advances in chronic kidney disease 17, 173–180, doi:10.1053/j.ackd.2010.01.003 (2010).
Pei, Y. Diagnostic approach in autosomal dominant polycystic kidney disease. Clinical Journal of the American Society of Nephrology 1, 1108–1114, doi:10.2215/CJN.02190606 (2006).
Fick-Brosnahan, G. M., Belz, M. M., McFann, K. K., Johnson, A. M. & Schrier, R. W. Relationship between renal volume growth and renal function in autosomal dominant polycystic kidney disease: a longitudinal study. American journal of kidney diseases 39, 1127–1134, doi:10.1053/ajkd.2002.33379 (2002).
Chapman, A. B. et al. Kidney volume and functional outcomes in autosomal dominant polycystic kidney disease. Clinical Journal of the American Society of Nephrology 7, 479–486, doi:10.2215/CJN.09500911 (2012).
Grantham, J. J. et al. Volume progression in polycystic kidney disease. New England Journal of Medicine 354, 2122–2130, doi:10.1056/NEJMoa054341 (2006).
Chapman, A. B. & Wei, W. Imaging approaches to patients with polycystic kidney disease. Seminars in Nephrology 31, 237–244, doi:10.1016/j.semnephrol.2011.05.003 (2011).
Alam, A. et al. Total kidney volume in autosomal dominant polycystic kidney disease: a biomarker of disease progression and therapeutic efficacy. American Journal of Kidney Diseases 66, 564–576, doi:10.1053/j.ajkd.2015.01.030 (2015).
Grantham, J. J. & Torres, V. E. The importance of total kidney volume in evaluating progression of polycystic kidney disease. Nature Reviews Nephrology 12, 667–677, doi:10.1038/nrneph.2016.135 (2016).
European Medicines Agency (EMA) qualification opinion: Total kidney volume (tkv) as a prognostic biomarker for use in clinical trials evaluating patients with autosomal dominant polycystic kidney disease (adpkd). www.ema.europa.eu/docs/en_GB/document_library/Regulatory_and_procedural_guideline/2015/11/WC500196569.pdf. (2015).
U.S. Food & Drug Administration (FDA) guidance for industry: Qualification of biomarker - total kidney volume in studies for treatment of autosomal dominant polycystic kidney disease. www.fda.gov/downloads/Drugs/Guidances/UCM458483.pdf. (2015).
Bae, K. T., Commean, P. K. & Lee, J. Volumetric measurement of renal cysts and parenchyma using mri: phantoms and patients with polycystic kidney disease. Journal of computer assisted tomography 24, 614–619, doi:10.1097/00004728-200007000-00019 (2000).
Bae, K. T. et al. Novel approach to estimate kidney and cyst volumes using mid-slice magnetic resonance images in polycystic kidney disease. American journal of nephrology 38, 333–341, doi:10.1159/000355375 (2013).
Irazabal, M. V. et al. Imaging classification of autosomal dominant polycystic kidney disease: A simple model for selecting patients for clinical trials. Journal of the American Society of Nephrology 26, 160–172, doi:10.1681/ASN.2013101138 (2014).
Higashihara, E. et al. Kidney volume estimations with ellipsoid equations by magnetic resonance imaging in autosomal dominant polycystic kidney disease. Nephron 129, 253–262, doi:10.1159/000381476 (2015).
Daum, V., Helbig, H., Janka, R., Eckardt, K.-U. & Zeltner, R. Quantitative Measurement of Kidney and Cyst Sizes in Patients with Autosomal Dominant Polycystic Kidney Disease(ADPKD). In Hornegger, J. et al. (eds.) 3rd Russian-Bavarian Conference on Biomedical Engineering vol. 1, 111–115 (Erlangen, 2007).
Racimora, D., Vivier, P.-H., Chandarana, H. & Rusinek, H. Segmentation of polycystic kidneys from MR images. In Karssemeijer, N. & Summers, R. M. (eds.) Medical Imaging 2010: Computer-Aided Diagnosis 76241W (SPIE-Intl Soc Optical Eng, 2010).
Mignani, R. et al. Assessment of kidney volume in polycystic kidney disease using magnetic resonance imaging without contrast medium. American journal of nephrology 33, 176–184, doi:10.1159/000324039 (2011).
Turco, D. et al. Reliability of total renal volume computation in polycystic kidney disease from magnetic resonance imaging. Academic Radiology 22, 1376–1384, doi:10.1016/j.acra.2015.06.018 (2015).
Kline, T. L. et al. Semiautomated segmentation of polycystic kidneys in t2-weighted mr images. American Journal of Roentgenology 207, 605–613, doi:10.2214/AJR.15.15875 (2016).
Sharma, K. et al. Semi-automatic segmentation of autosomal dominant polycystic kidneys using random forests. arXiv preprint arXiv:1510.06915 (2015).
Kline, T. L. et al. Automatic total kidney volume measurement on follow-up magnetic resonance images to facilitate monitoring of autosomal dominant polycystic kidney disease progression. Nephrology Dialysis Transplantation gfv314 (2015).
Kim, Y. et al. Automated segmentation of kidneys from mr images in patients with autosomal dominant polycystic kidney disease. Clinical Journal of the American Society of Nephrology 11, 576–584, doi:10.2215/CJN.08300815 (2016).
Long, J., Shelhamer, E. & Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 3431–3440 (2015).
Badrinarayanan, V., Kendall, A. & Cipolla, R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. arXiv preprint arXiv:1511.00561 (2015).
Pinheiro, P. H. & Collobert, R. Recurrent convolutional neural networks for scene labeling. Proceedings of the International Conference on Machine Learning 162, 82–90, doi:10.1016/j.vetpar.2009.02.011 (2014).
Thong, W., Kadoury, S., Piché, N. & Pal, C. J. Convolutional networks for kidney segmentation in contrast-enhanced CT scans. Computer Methods in Biomechanics and Biomedical Engineering: Imaging & Visualization 0, 1–6 (2016).
Zheng, Y., Liu, D., Georgescu, B., Xu, D. & Comaniciu, D. Deep learning based automatic segmentation of pathological kidney in ct: Local vs. global image context. In Deep Learning and Convolutional Neural Networks for Medical Image Computing (Springer, 2016).
Warner, J. D. et al. Supervised segmentation of polycystic kidneys: a new application for stereology data. Journal of digital imaging 27, 514–519, doi:10.1007/s10278-014-9679-y (2014).
Perico, N. et al. Sirolimus therapy to halt the progression of adpkd. Journal of the American Society of Nephrology 21, 1031–1040, doi:10.1681/ASN.2009121302 (2010).
Ruggenenti, P. et al. Effect of sirolimus on disease progression in patients with autosomal dominant polycystic kidney disease and CKD stages 3b-4. Clinical Journal of the American Society of Nephrology 11, 785–794, doi:10.2215/CJN.09900915 (2016).
Abràmoff, M. D., Magalhães, P. J. & Ram, S. J. Image processing with imagej. Biophotonics international 11, 36–42 (2004).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 (2014).
Ioffe, S. & Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. arXiv preprint arXiv:1502.03167 (2015).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. Imagenet classification with deep convolutional neural networks. In Advances in neural information processing systems 1097–1105 (2012).
Zeiler, M. D. & Fergus, R. Visualizing and understanding convolutional networks. In Proceedings of the European Conference on Computer Vision 818–833 (Springer, 2014).
Noh, H., Hong, S. & Han, B. Learning deconvolution network for semantic segmentation. In Proceedings of the IEEE International Conference on Computer Vision 1520–1528 (2015).
Jia, Y. et al. Caffe: Convolutional architecture for fast feature embedding. In Proceedings of the 22nd ACM international conference on Multimedia 675–678 (ACM, 2014).
Glorot, X. & Bengio, Y. Understanding the difficulty of training deep feedforward neural networks. In Proceedings of the International Conference on Artificial Intelligence and Statistics vol. 9, 249–256 (Society for Artificial Intelligence and Statistics, 2010).
Dice, L. R. Measures of the amount of ecologic association between species. Ecology 26, 297–302, doi:10.2307/1932409 (1945).
Jones, R. G. & Payne, R. B. Clinical investigation and statistics in laboratory medicine (American Association for Clinical Chemistry, 1997).
Studio, R. Rstudio: integrated development environment for r. RStudio Inc, Boston, Massachusetts (2012).
Acknowledgements
This work was funded by the TranCYST Marie Curie Initial Training Networks project, within the 7th European Community Framework Programme EU-FP7/2007–2013 grant 317246 (TranCYST). Maximilian Baust and Nassir Navab acknowledge the support of the Collaborative Research Centre SFB 824 (Z2) Imaging for Selection, Monitoring and Individualization of Cancer Therapies funded by the German Science Foundation. The authors would like to thank Prof. Giuseppe Remuzzi and Dr. Norberto Perico for their support on the clinical trials; Sergio Carminati and Davide Martinetti for their assistance on the system hardware and Michela Bozzetto for suggestions on preparing the manuscript. Also, sincere thanks to Christine Kroll and Dr. Seyed-Ahmad Ahmadi for fruitful discussions on deep learning.
Author information
Authors and Affiliations
Contributions
K.S., A.R., M.B. and N.N. conceived the experiments. K.S. and M.C.A. contributed to manual segmentation of the dataset. K.S. conducted the experiments, prepared the figures and tables, performed statistical analyses, and wrote the main manuscript. K.S., C.R., A.C. and M.B. analyzed the results and revised the manuscript. A.R., M.B. and N.N. supervised the work. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
41598_2017_1779_MOESM1_ESM.pdf
Supplementary Information Automatic Segmentation of Kidneys using Deep Learning for Total Kidney Volume Quantification in Autosomal Dominant Polycystic Kidney Disease
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sharma, K., Rupprecht, C., Caroli, A. et al. Automatic Segmentation of Kidneys using Deep Learning for Total Kidney Volume Quantification in Autosomal Dominant Polycystic Kidney Disease. Sci Rep 7, 2049 (2017). https://doi.org/10.1038/s41598-017-01779-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-01779-0
This article is cited by
-
Deep learning-based measurement of split glomerular filtration rate with 99mTc-diethylenetriamine pentaacetic acid renal scan
EJNMMI Physics (2024)
-
Automation of Wilms’ tumor segmentation by artificial intelligence
Cancer Imaging (2024)
-
AI-based segmentation of renal enhanced CT images for quantitative evaluate of chronic kidney disease
Scientific Reports (2024)
-
Fully-automated multi-organ segmentation tool applicable to both non-contrast and post-contrast abdominal CT: deep learning algorithm developed using dual-energy CT images
Scientific Reports (2024)
-
Multi-Scale and Spatial Information Extraction for Kidney Tumor Segmentation: A Contextual Deformable Attention and Edge-Enhanced U-Net
Journal of Imaging Informatics in Medicine (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.