Abstract
The current challenge in effectively treating atrial fibrillation (AF) stems from a limited understanding of the intricate structure of the human atria. The objective and quantitative interpretation of the right atrium (RA) in late gadolinium-enhanced magnetic resonance imaging (LGE-MRI) scans relies heavily on its precise segmentation. Leveraging the potential of artificial intelligence (AI) for RA segmentation presents a promising solution. However, the successful implementation of AI in this context necessitates access to a substantial volume of annotated LGE-MRI images for model training. In this paper, we present a comprehensive 3D cardiac dataset comprising 50 high-resolution LGE-MRI scans, each meticulously annotated at the pixel level. The annotation process underwent rigorous standardization through crowdsourcing among a panel of medical experts, ensuring the accuracy and consistency of the annotations. Our dataset represents a significant contribution to the field, providing a valuable resource for advancing RA segmentation methods.
Similar content being viewed by others
Background & Summary
Atrial fibrillation (AF) is a globally significant chronic disease, being the most common cardiac arrhythmia, and is associated with substantial morbidity and mortality1,2. The suboptimal clinical management of AF largely stems from a fundamental lack of understanding of atrial anatomy3. Recent advancements, particularly the widespread use of gadolinium-based contrast agents in assessing atrial fibrosis and scarring through late gadolinium-enhanced magnetic resonance imaging (LGE-MRI)4, have significantly improved the visualization of organ structures and related components5. Clinical investigations utilizing LGE-MRI in AF patients have highlighted that the extent and distribution of atrial fibrosis can reliably predict the success of ablation procedures6. Recent studies using LGE-MRI for atrial assessments have further emphasized the crucial role of atrial structure in comprehending and treating AF3,7. Therefore, a direct analysis of atrial structure is vital for effective AF treatment.
Atrial segmentation is a fundamental process involving the extraction of atrial cavity structures from LGE-MRI images. This process serves as a crucial preliminary step in enabling the objective evaluation and quantitative analysis of atrial structure within the context of AF. While extensive research has been conducted on the automatic and semi-automatic segmentation of the left atrium (LA), given its central role in AF studies, it is equally imperative to conduct comprehensive structural assessments of the right atrium (RA)8,9. Despite the relatively limited exploration of the pathological changes occurring in the RA within the context of AF, existing evidence strongly suggests that AF exerts its impact on both atria10. Therefore, it is imperative to delve into the intricate relationship between AF and the RA. This connection is primarily attributed to a complex interplay of structural, electrical, and metabolic remodeling processes that transpire within the RA11. Consequently, research endeavours dedicated to the segmentation of the RA from LGE-MRI scans are indispensable.
Manual segmentation is essential for precise analysis but can be time-consuming and labour-intensive, especially in the context of medical research. To enhance efficiency and accuracy, automated and semi-automated segmentation methods play a crucial role. In the 2018 Left Atrium Segmentation Challenge12, 15 teams utilized CNN-based segmentation methods, but 12 proposed CNN designs based on the popular U-Net architecture, achieving outstanding performance. The adoption of the popular U-Net architecture effectively enhanced the effectiveness of atrial segmentation. For instance, D. Borra et al.13 utilized a CNN-based U-SWNN for 3D left atrium segmentation, achieving a Dice score of 0.911. Xiong et al.14 employed a dual-branch multi-scale convolutional neural network, significantly improving segmentation results. In contrast, D. Borra et al.15 proposed a comprehensive two-stage workflow for automatic LA cavity segmentation, involving traditional automated segmentation algorithms for LA localization (first stage) and refined LA segmentation based on CNN outputs (second stage). Although algorithms for the LA are very advanced, there are currently no algorithms for the RA.
However, the development and evaluation of these automated approaches heavily rely on access to extensive datasets with comprehensive annotations. Presently, a noticeable gap exists in dedicated research focused on RA segmentation, and publicly available datasets catering to this specific need are limited. For instance, one available dataset is derived from the 2017 Multi-Modality Whole Heart Segmentation (MM-WHS) challenge16,17,18, which is based on non-contrast MRI scans. While non-contrast MRI yields precise images, contrast MRI, particularly LGE-MRI, offers superior clarity for detecting smaller tissue structures and assessing their extent within the surrounding tissues. LGE-MRI scans have proven invaluable for studying atrial fibrosis14. In the field of AF research, several datasets targeting LGE-MRI have been established, such as the 2018 Left Atrial Challenge12 and the 2022 Left Atrial and Scar Quantification and Segmentation Challenge19,20,21. However, datasets specifically dedicated to RA segmentation from LGE-MRI scans remain notably absent.
Thus, we introduce the RAS dataset22, a valuable resource comprising 50 high-resolution LGE-MRI scans, each with spatial dimensions of either 576 × 576 × 88 or 640 × 640 × 88 pixels. These scans have undergone meticulous pixel-wise manual annotation, performed by four highly trained graduate students and subsequently validated by three experienced advisors. The RAS dataset22 represents a significant contribution to the field, serving as a valuable resource for researchers engaged in developing and evaluating automatic RA segmentation algorithms. Furthermore, it has the potential to support the creation of image-based personalized models, thereby advancing our understanding and treatment of AF.
Methods
Data collection
The RAS dataset22 only provides labels for the right atrium, while the original data belongs to the 2018 Left Atria Challenge12 (https://www.cardiacatlas.org/atriaseg2018-challenge/atria-seg-data/) and has been made public. Each 3D MRI patient data in the dataset was acquired using a clinical MRI scanner, specifically a 1.5 Tesla Avanto or 3.0 Tesla Verio whole body MRI scanner. These scans were performed approximately 20–25 months after the injection of gadolinium contrast agent (Multihance, manufactured by Bracco Diagnostics Inc., Princeton, NJ).
Image annotation
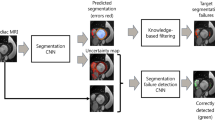
The annotation team consisted of a group of highly qualified individuals, including three experienced physicians and four postgraduates specializing in biomedical-related fields. These annotators underwent comprehensive training, which included online meetings and in-person guidance from the three experienced physicians. The primary focus of this training was to familiarize the annotators with the structure of the RA as it appears in LGE-MRI images. Each trained annotator was responsible for segmenting 25 LGE-MRI images using the pencil tool in Slicer 5.0.3, a software tool available at https://www.slicer.org/. This segmentation process involved pixel-wise annotation, where each pixel was carefully labelled to identify the RA structures, including the tricuspid valve (TV) and right atrial appendages (RAA) as well as the ostia of the superior/inferior vena cava (SVC/IVC). The resulting annotations were subjected to individual evaluations by the physicians. If an annotation was deemed unsatisfactory or inaccurate, the respective image was returned to the student for re-labelling (as illustrated in Fig. 1). During this annotation process, the following points should be followed: (1) ideally, there is a continuous relationship of adjacent pixels in the contour of each two-dimensional image, and there is a hierarchical relationship between the shape and size changes; and (2) Fibrosis in the right atrial wall appears bright white. When there is no obvious white border, the border is determined based on the difference in local gray values. The ground truths are binary, representing the presence or absence of RA structures, and are stored in the Near Raw Raster Data (NRRD) format.
In the following, we detail the process of labelling a 3D LGE-MRI image with spatial dimensions of either 576 × 576 × 88 (Fig. 2Ai) or 640 × 640 × 88 (Fig. 2Bi). This labelling procedure can be broken down into the following key steps:
-
Step 1 - Identifying SVC Region: To begin, we observed the SVC region, which typically exhibits a circular or oblate shape. This region was annotated in approximately the first 12 slices (Fig. 2Aii or Fig. 2Bii).
-
Step 2 - Marking RAA Area: Moving above the SVC area, the RAA, often appearing as a smaller ellipse or triangle, was annotated in the subsequent 4–6 slices (Fig. 2Aiii or Fig. 2Biii).
-
Step 3 - Defining the RA Region: The RAA area, connected to the SVC region, forms the broader RA region. This region was labeled in approximately the following 30 slices (Fig. 2Aiv or Fig. 2Biv). Throughout the annotation process, we relied on several anatomical landmarks to ensure precise labelling of the RA: a) Tricuspid Valve (TV): Serving as a reference point, the TV helped us delineate the boundary of the RA in specific image slices (e.g., Fig. 2Av or Fig. 2Bv). b) RV-LV Connection: The clear connection between the Right Ventricle (RV) and the Left Ventricle (LV) (e.g., Fig. 2Avi or Fig. 2Bvi) served as a visual guide for accurate RA labelling. c) RA-LA Wall: The wall separating the RA from the LA (e.g., Fig. 2Avii or Fig. 2Bvii) was another vital reference point used for precise identification and labelling of the RA. At this stage, we encountered approximately 25 slices with both RA and RV (e.g., Fig. 2Aviii or Fig. 2Bviii).
-
Step 4 - Transition to IVC Region: As the LA region gradually disappeared, the IVC region with a circular shape emerged below the RA region. Approximately 15 slices featured both RA and IVC regions (e.g., Fig. 2Aix or Fig. 2Bix). Subsequently, the following slices exclusively featured the IVC region.
-
Step 5 - Fine-Tuning Corrections: After labelling in the Z-axis direction, we conducted adjustments in the X- and Y-axis directions to ensure the smoothness and continuity of the ground truth (Fig. 3).
Data Records
All data records22 are available as files on the web page https://doi.org/10.5281/zenodo.10781134. In the unzipped folder, the “lgemri.csv” file describes the correspondence between the original LGE-MRI image and its RA cavity label file, and the “ras” folder contains the ground truth label corresponding to the RA cavity. The specific images in the “ras” folder are the ground truth for the corresponding images12 (https://www.cardiacatlas.org/atriaseg2018-challenge/atria-seg-data/), and their correspondences are described in the “lgemri.csv” file. Images in the “ras” folder contain pixels labeled 0 and 1, where 0 represents the background and 1 represents the RA cavity .
Technical Validation
In this study, each LGE-MRI image underwent annotation by one annotator, followed by refinement by a physician. Consequently, inter-annotator consistency warrants investigation. We selected 25 LGE-MRI images from the complete dataset to form an annotation set and assigned these images to two annotators (Dr1 and Dr2). We also evaluated the quality of labels predicted by the classical U-Net model23 compared to manual annotations performed by human experts. Dice and Jaccard indices can be used to represent the overlap of validation results, while recall and specificity can indicate the positive-to-negative ratio of validation results. Table 1 displays their respective Dice, Jaccard, recall, and specificity scores, namely AI vs. Dr1, AI vs. Dr2, and Dr1 vs. Dr2. We found that the results between artificial intelligence and humans (AI vs. Dr1 and AI vs. Dr2) were lower than those among humans (Dr1 vs. Dr2), indicating the challenge of automated segmentation for the right atrium. Specifically, for Dr1 vs. Dr2, the average Dice coefficient was calculated to be 93.85%, the Jaccard coefficient was 85.52%, the specificity coefficient was 99.95%, and the recall coefficient was 93.71%, indicating a very close agreement between the annotators.
Usage Notes
Users should cite this paper in their research output and acknowledge the contribution of this dataset in their study.
Code availability
No novel code used in the construction of RAS dataset22.
References
Peng, X. et al. Right atrial tachycardia after atrial fibrillation ablation: prevalence, clinical characteristics, electrophysiological mechanisms, and long-term outcomes. Journal of Interventional Cardiac Electrophysiology, 1–10 (2023).
Narayan, S. M. et al. Ablation of Focal Impulses and Rotational Sources: What Can Be Learned from Differing Procedural Outcomes? Current Cardiovascular Risk Reports 11, 27, https://doi.org/10.1007/s12170-017-0552-7 (2017).
Zhao, J. et al. Three-dimensional Integrated Functional, Structural, and Computational Mapping to Define the Structural “Fingerprints” of Heart-Specific Atrial Fibrillation Drivers in Human Heart Ex Vivo. J Am Heart Assoc 6, https://doi.org/10.1161/JAHA.117.005922 (2017).
McGann, C. et al. Atrial fibrillation ablation outcome is predicted by left atrial remodeling on MRI. Circulation: Arrhythmia and Electrophysiology 7, 23–30 (2014).
Hansen, B. J., Zhao, J. & Fedorov, V. V. Fibrosis and atrial fibrillation: computerized and optical mapping: a view into the human atria at submillimeter resolution. JACC: Clinical Electrophysiology 3, 531–546 (2017).
Oakes, R. S. et al. Detection and quantification of left atrial structural remodeling with delayed-enhancement magnetic resonance imaging in patients with atrial fibrillation. Circulation 119, 1758–1767 (2009).
Hansen, B. J. et al. Atrial fibrillation driven by micro-anatomic intramural re-entry revealed by simultaneous sub-epicardial and sub-endocardial optical mapping in explanted human hearts. European heart journal 36, 2390–2401 (2015).
Chen, Y. C., Voskoboinik, A., La Gerche, A., Marwick, T. H. & McMullen, J. R. Prevention of pathological atrial remodeling and atrial fibrillation: JACC state-of-the-art review. Journal of the American College of Cardiology 77, 2846–2864 (2021).
Shen, M. J., Arora, R. & Jalife, J. Atrial myopathy. JACC: Basic to Translational Science 4, 640–654 (2019).
Lang, R. M. et al. Imaging assessment of the right atrium: anatomy and function. European Heart Journal-Cardiovascular Imaging 23, 867–884 (2022).
Hiram, R. & Provencher, S. Pulmonary disease, pulmonary hypertension and atrial fibrillation. Cardiac Electrophysiology Clinics 13, 141–153 (2021).
Xiong, Z. et al. A global benchmark of algorithms for segmenting the left atrium from late gadolinium-enhanced cardiac magnetic resonance imaging. Medical image analysis 67, 101832 (2021).
Borra, D. et al. in Statistical Atlases and Computational Models of the Heart. Atrial Segmentation and LV Quantification Challenges: 9th International Workshop, STACOM 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 16, 2018, Revised Selected Papers 9. 329–338 (Springer).
Xiong, Z. et al. Fully automatic left atrium segmentation from late gadolinium enhanced magnetic resonance imaging using a dual fully convolutional neural network. IEEE transactions on medical imaging 38, 515–524 (2018).
Borra, D., Andalò, A., Paci, M., Fabbri, C. & Corsi, C. A fully automated left atrium segmentation approach from late gadolinium enhanced magnetic resonance imaging based on a convolutional neural network. Quantitative Imaging in Medicine and Surgery 10, 1894 (2020).
Zhuang, X. & Shen, J. Multi-scale patch and multi-modality atlases for whole heart segmentation of MRI. Medical image analysis 31, 77–87 (2016).
Zhuang, X. Multivariate mixture model for myocardial segmentation combining multi-source images. IEEE transactions on pattern analysis and machine intelligence 41, 2933–2946 (2018).
Luo, X. & Zhuang, X. X-Metric: An N-Dimensional Information-Theoretic Framework for Groupwise Registration and Deep Combined Computing. IEEE Trans Pattern Anal Mach Intell 45, 9206–9224, https://doi.org/10.1109/TPAMI.2022.3225418 (2023).
Li, L., Zimmer, V. A., Schnabel, J. A. & Zhuang, X. AtrialJSQnet: A New framework for joint segmentation and quantification of left atrium and scars incorporating spatial and shape information. Medical image analysis 76, 102303 (2022).
Li, L., Zimmer, V. A., Schnabel, J. A. & Zhuang, X. Medical image analysis on left atrial LGE MRI for atrial fibrillation studies: A review. Medical image analysis 77, 102360 (2022).
Li, L., Zimmer, V. A., Schnabel, J. A. & Zhuang, X. in Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part VI 24. 557–566 (Springer).
Bai, J. RAS Dataset. Zenodo. https://doi.org/10.5281/zenodo.10781134 (2024).
Çiçek, Ö., Abdulkadir, A., Lienkamp, S. S., Brox, T. & Ronneberger, O. in Medical Image Computing and Computer-Assisted Intervention–MICCAI 2016: 19th International Conference, Athens, Greece, October 17-21, 2016, Proceedings, Part II 19. 424–432 (Springer).
Acknowledgements
The work is funded by the Guangdong Basic and Applied Basic Research Foundation (2024A1515011886 and 2023A1515012833), National Natural Science Foundation of China under Grant 61901192, Guangzhou Municipal Science and Technology Bureau Guangzhou Key Research and Development Program under Grant 2024B03J1283 and 2024B03J1289, Guangzhou Science and Technology Planning Project under Grant 2023B03J1297 and China Scholarship Council. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Jinwen Zhu: Conceptualization, Manual segmentation, Writing, Reviewing, Visualization. Jieyun Bai: Conceptualization, Funding acquisition, Project administration, Reviewing. Zihao Zhou: Manual segmentation. Yaqi Liang: Manual segmentation. Zhiting Chen: Manual segmentation. Xiaoming Chen: Conceptualization, Reviewing. Xiaoshen Zhang: Conceptualization, Reviewing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhu, J., Bai, J., Zhou, Z. et al. RAS Dataset: A 3D Cardiac LGE-MRI Dataset for Segmentation of Right Atrial Cavity. Sci Data 11, 401 (2024). https://doi.org/10.1038/s41597-024-03253-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03253-9