Abstract
Drug repositioning is a faster and more affordable solution than traditional drug discovery approaches. From this perspective, computational drug repositioning using knowledge graphs is a very promising direction. Knowledge graphs constructed from drug data and information can be used to generate hypotheses (molecule/drug - target links) through link prediction using machine learning algorithms. However, it remains rare to have a holistically constructed knowledge graph using the broadest possible features and drug characteristics, which is freely available to the community. The OREGANO knowledge graph aims at filling this gap. The purpose of this paper is to present the OREGANO knowledge graph, which includes natural compounds related data. The graph was developed from scratch by retrieving data directly from the knowledge sources to be integrated. We therefore designed the expected graph model and proposed a method for merging nodes between the different knowledge sources, and finally, the data were cleaned. The knowledge graph, as well as the source codes for the ETL process, are openly available on the GitHub of the OREGANO project (https://gitub.u-bordeaux.fr/erias/oregano).
Similar content being viewed by others
Background & Summary
The rapid discovery of new drugs is a particularly topical issue. The Covid-19 health crisis that we have experienced has confirmed that. However, while current standard methods are time-consuming and very expensive, drug approvals are complicated to obtain. Before this approval, the therapeutic value of drugs must be proven in clinical trials that are organised in three phases1. Phase I focuses on toxicological tests on the molecule, while phase II is about the evaluation of the minimum dose to be administered to obtain an effective effect, as well as the listing of the various secondary effects. Phase III is designed to measure the effectiveness of the drug; the drug is authorised or not to be produced and sold at the end of this phase. This is a complex process, especially as almost 60% of new drugs tested in clinical trials do not pass Phase II2. This failure rate highlights drugs that are not effective enough and/or have too many side effects compared to their benefits. The time spent on clinical trials is therefore extremely time-consuming, taking on average 10 to 15 years from Phase I to Phase III. The development of new drugs is also very costly, so the pharmaceutical industry focuses its research on diseases that will make the heavy investments profitable. This need for cost-effectiveness is an obstacle to the discovery of certain treatments, especially for rare diseases.
Thus, alternatives to this costly traditional development process have emerged. This is the case of drug repurposing (aka drug repositioning)3, which aims to find a new use for existing drugs or compounds. The idea is to reuse drugs that have already been approved, as well as those that have not passed the final stages of clinical trials and have a better chance to do so. This possibility opens up prospects for the identification of drugs for rare diseases in particular4, as drugs that have already passed the first stages of clinical trials are less expensive to bring to market. Repositioning methods are categorised into either biological or computational methods5. Biological methods involve discovering new information about drugs and targets and also testing, in large studies, the possibility of binding them, mostly to proteins. However, computational methods predominate in this field, and these are diverse and varied.
Computational repositioning methods exploit knowledge about individual drugs to find possible new therapeutic targets6. The methods used differ according to the features describing the drugs that are taken into account. Indeed, knowledge about drugs is diverse, as compounds can be described by their chemical formula, their side effects, the targets they are aimed at, the diseases they treat and many other features. Numerous resources describe portions of this body of knowledge, many of them available online7. However, it is difficult to easily connect knowledge from these sources, primarily because they are not formatted in the same way and do not necessarily have cross-references allowing them to be interlinked with other knowledge sources. Similarly, the knowledge sources may not be produced for the same purpose and may not cover the same subject. Nevertheless, some projects aim to overcome these issues through different data fusion methods. The principle consists of linking the data and knowledge available in the various sources into a vast knowledge resource that is made available to all through a single endpoint. For example, the Linked Open Data cloud initiative brings together open source knowledge bases, linking them via nodes and links. In the area of drug repositioning, a key issue is the availability of these data and knowledge because drug information is not always made available by the companies working on the drugs.
It is possible to learn new knowledge and observations about drugs from those already available. Indeed, many studies implement learning algorithms that can, from the available data, determine whether a drug is potentially capable of binding to a new target8. The data used by learning algorithms must be represented in such a way that machines can learn a general model. One of our assumptions is that the more information there is to characterise a drug, the better the pattern. It is therefore necessary to integrate as much drug data as possible. One of the concerns that emerges in this context is that the data are very heterogeneous and it is difficult to characterise drugs in the same way. Indeed, drugs may have one or more targets, one or more side effects, and the targets are very diverse. In order to learn effectively on such heterogeneous data, it is necessary to find a way to represent the drug data and to leverage this representation. The type of representation that can be used at this stage is a knowledge graph, which has been defined by Hogan et al. as: “a graph of data intended to accumulate and convey knowledge of real world, whose nodes represent nodes of interest and whose edges represent links between nodes”9. A knowledge graph therefore accumulates knowledge of the real world in which the nodes represent notions of interest and whose edges represent links between them. A knowledge graph is thus a set of nodes (or entities) N and labelled links (or relations or predicates) L represented as triples of the form: (Nx, L1, Ny). The edges describe the binary links between two nodes and they are generally oriented and meaningful. In this case, the nodes are differentiated: the subject is the source node of the relationship, the object is the target node resulting in a triplet expressed as (subject, predicate, object).
From the perspective of defining a graph for drug repositioning, it is then possible to represent each drug, target, disease or other related entities by a node in the graph and then link these nodes together.
Biomedical data are well suited to be stored in knowledge graphs because they are scattered over many knowledge sources without being linked to each other. In this context, the “Semantic Web initiative” offers an idealised vision of the Web, with the idea that resources on the Web should be connected by semantic links (as opposed to hyperlinks) and that the meaning of these resources should be exploitable by machines10. Following this paradigm, various initiatives aiming to interconnect existing knowledge sources have emerge11,12.
The use of knowledge graphs for drug repurposing has been shown to be effective in recent years13,14,15. The identification of new drug-target relationships is the main goal of knowledge graph-based techniques for repositioning. The different methods focus on various aspects of the knowledge graph16,17. Some networks are not only composed of drugs and targets. Networks comprising other types of nodes can indeed participate in the discovery of new information for drug repositioning18,19,20,21,22. Most drug targets being proteins, protein-protein interaction networks are likely to provide very relevant information, as some proteins act indirectly on each other. Drug-target pairs can also have important consequences on other proteins, thus giving useful indications for possible repositioning23,24. Other studies investigated on predicting drug-disease links25. These studies were useful for identifying new avenues of research26. Graphs associating drugs and their side effects greatly assisted in the prediction of drug-target relationships27. These different pieces of information enrich the network and maximise the chances of finding drug-target pairs that are likely to lead to drug repurposing. The next logical step was to try to interconnect all this information into a single knowledge graph28,29,30,31.
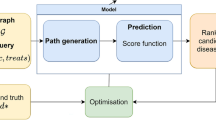
The OREGANO project falls within this approach. Its objective is to develop a holistic knowledge graph on drugs and related concepts for humans in order to identify possible repositionable molecules using machine learning (or more specifically deep learning) algorithms. In the context of drug repositioning, these algorithms aim to predict the probability that a link exists between two nodes in the knowledge graph based on existing links. Such algorithms have been applied in a preliminary work on a first version of the OREGANO knowledge graph32. The first results were promising, and the present work aims at extending and optimizing these results by means of an updated graph composed of well-integrated sources in a first step, then by optimizing the learning methods in a second step. The OREGANO project aims to fill the missing gap in terms of computational-based drug repositioning, and performs link prediction on a large knowledge graph of heterogeneous data for discovering missing molecule-target links.
Unlike similar studies, the OREGANO knowledge graph emphasizes the integration of natural compounds (i.e. herbal and plant remedies). Indeed, for three decades (1981–2010), more than 60% of drugs were developed from natural products, derivatives or natural product-like compounds33. Some prominent herbal prescriptions have been transformed into new drugs34 (e.g. Layla Tab in Korea35). Therefore, we hypothesize that there is a strong possibility to seek new indications for existing herbal compounds, which could lead to the development of new drugs through repurposing strategies.
To the best of our knowledge, there is no previous work on knowledge graphs incorporating together disease and drug information and natural compounds specifically used for drug repositioning. Only two studies do refer to natural compounds and graph based representations, but for other purposes36,37. Indeed, these studies predict links between herbs and diseases or between herbs and targets, but on networks that are not knowledge graphs.
Methods
This section describes the steps involved in building the knowledge graph (Fig. 1), i.e. the data selection, the actual construction and the data filtering performed via several stages in this work.
Overall workflow. After selecting the types of nodes which are of interest for drug repositioning, the labels of the links connecting these types are retrieved from the Relation Ontology73. The nodes and the way there are connected form the model of the knowledge graph. The sources to be integrated are chosen according to the nodes represented in the model and their free online availability. Each knowledge source has its data extracted by a dedicated wrapper. These data are processed by a binder and a manager to populate thereafter the model and thus generate the OREGANO knowledge graph.
The information to be acquired from the knowledge sources was selected using the model that was produced and according to their ability to be integrated (subsection A.). The first version of the OREGANO knowledge graph32 was built using the knowledge available in Bio2RDF12. The utility of this resource resides in its ability to establish links and cross-references among the various knowledge bases it incorporates. Nevertheless, Bio2RDF has not been updated since 2014. Consequently, the information contained in this resource does not align with the content present in the updated iterations of the knowledge bases. In the second version of the OREGANO knowledge graph, we have thus opted to incorporate the data directly from knowledge source repositories. This approach allows us to construct a knowledge graph enriched with up-to-date information. The integration process follows the Extract, Transform and Load (ETL) principle: the wrappers extract the data (subsection B.), which are then transformed by the binders (subsection C.) and managers (subsection D.).
A. Description of the selected knowledge sources
The process of building the OREGANO knowledge graph first required selecting the knowledge sources to be integrated. This choice was made in coherence with the previously designed OREGANO knowledge graph, which was built from data available on the Linked Open Data (LOD)32. The same set of sources was therefore chosen with the intention of reinforcing the integration process used to develop the first knowledge graph. The difference is that rather than retrieving the sources through the Bio2RDF proxy, we have included them in the current version as they were initially (official repository).
In this section, we introduce the four types of sources that were used in this work, i.e. those that describe: (i.) information about targets, (ii.) information about phenotypes, (iii.) information about natural compounds, and (iv.) information about drugs. In addition, we present: (v.) two additional sources, and (vi.) existing cross-references that are useful for integrating the aforementioned sources successfully.
i. Target-related sources
DrugBank is a free drug information source that was launched in 200638,39. It offers a wide range of information on drugs, their targets and interactions. The knowledge about drugs is both chemical and pharmaceutical, with different labels and dosages for drugs from around the world. The use of DrugBank is open to anyone, provided they create a profile to download the data. The version used in the current work is dated back April 1, 2023. It contains respectively 16,306 drugs and 4,939 targets.
SIDER contains information on drugs and their side effects40,41. The most recent version, which was released on October 21, 2015, comprises 139,756 pairs of 1,430 drugs and 5,868 side effects. This information was derived from data from clinical trials, in which participants were observed, and any side effects were noted. Prior to these human testing phases, animal testing also contributed to the data. The resource also lists the disease indications for the included drugs.
UniProt is a resource about proteins. It provides in particular their sequences, annotations and information about their functions in the organism42. The European Bioinformatics Institute (EMBL-EBI), the SIB Swiss Institute of Bioinformatics and the Protein Information Resource (PIR) have collaborated to create and maintain it. The version we used for OREGANO was released on September 13, 2023 and aggregated information on 570,157 proteins. More specifically, UniProt contains several data sources, including UniProtKB that provides information about proteins. The latter comprises data from Swiss-Prot43 and TrEMBL44. Since the TrEMBL protein dataset is not renewed, only the Swiss-Prot data are considered in the integration process.
Reactome is a knowledge source containing data on biological pathways45. Created in 2003, it contains 2,629 human pathways binding 1,114 drugs, 14,277 proteins and 2,004 small molecules through 14,628 reactions. In this work, we used release 85, which was published in June 2023.
PharmGKB is a pharmacogenetic resource combining knowledge of genetic variation and drug responses46,47. It aims at integrating knowledge to assist clinicians and researchers in their investigations. PharmGKB has grown since it was founded in 2000 and now contains 993 drug label annotations, 201 clinical guideline annotations, 181 curated pathways, and 428 annotated drugs. The release that we integrated was made available on September 5, 2023.
ii. Phenotype-related source
Human Phenotype Ontology (HPO) is a source which is created initially in 2007 to model an ontology based on the concepts described in the Online Mendelian Inheritance in Man (OMIM)48. Nowadays, HPO has evolved into an ontology for diseases and the relationships between diseases and phenotypes. It contains over 13,000 terms. The HPO diseases come from the following sources: OMIM49, Orphanet50 and DECIPHER51. The version we have integrated is part of the June 2023 release.
iii. Natural compound-related source
NPASS is the only source not present in Bio2RDF that was included in the first version of the OREGANO knowledge graph to complete data with knowledge about natural compounds.
NPASS contains information on natural compounds and their targets in organisms52,53. It comprises respectively 96,481 natural compounds and 7,753 biological targets linked by 958,866 relationships. The last update was on October 4, 2023.
iv. Drug-related source
Anatomical Therapeutic Chemical (ATC) classification has been developed by the WHO Collaborating Centre. It categorises active substances according to their chemical properties and their actions in the body. Each compound is assigned a 7-character code consisting of five levels that reflect the compound’s place in the classification according to the system in which it acts and its pharmaceutical and therapeutic effects.
v. Additional sources
Unified Medical Language System (UMLS) is a resource that gathers more than 150 knowledge sources (i.e. (bio)medical coding systems and terminologies) composed of terms and relationships between them54. Since OMIM did not provide external links to connect it to other knowledge sources, UMLS was used to ensure proper integration of OMIM.
Orphanet is an online resource about rare diseases50. Cross-references for these diseases are not available via HPO (in which Orphanet is included), so extraction of data from Orphanet was necessary to ensure optimal HPO integration.
vi. Existing cross-references
Finding the mappings that the knowledge sources make available is helpful to facilitate the integration process while building the knowledge graph, which involves combining the data. In Fig. 2, the different existing mappings between the seven knowledge sources are depicted. Some sources were directly connected, which made their integration simple. However, most of the information sources had indirect rather than direct mappings. Such indirect mapping entails that neither source gives cross-references to the other, but that they can still be connected by using one or more other cross-references that they both share. These mappings make it easier to find sources with related information. Since the information sources do not use the same nomenclatures, or even have the same meanings for some elements, it should be noted that establishing these cross-references requires a great deal of effort. For instance, issues can arise if duplicate proteins are found in poorly cleaned large protein libraries. In such cases, the use of an additional knowledge source as a pivot was required. As mentioned in the previous subsection, UMLS and Orphanet acted as pivots to enable the integration of OMIM and HPO, respectively.
Diagram showing the different sources to be integrated in the OREGANO knowledge graph and the mappings existing between them. Solid lines denote that the source provides mappings to another source in its data, and the direction of the arrow indicates that the target source is referenced in the original source’s data. Dashed lines correspond to indirect mappings that require an intermediate source.
B. Extraction of data from knowledge sources
First, data were extracted from the knowledge sources that have been selected for their potential utility in drug repositioning. Then, their structure was analysed to retrieve the relevant information from each source (Table 1). The formats of the sources could differ, but each of them underwent the same overall processing. The wrapper (top part of Fig. 3), being the module that handled this step, first scanned the resource and then extracted the data deemed of interest for drug repositioning. Then, the resulting data were formatted into triples of the form (subject, predicate, object).
Integration workflow of a knowledge source. Data from the knowledge sources are extracted by a wrapper. The data are of two types: links to be integrated into the knowledge graph and links to external sources. The external links are used to link the data via a binder that achieves a mapping table for each type of node. It is from this mapping table that the manager converts the triples to be integrated by replacing the source codes by the OREGANO codes. This integration process follows the traditional ETL principle.
The formats were quite similar, but the way the data were organised (e.g. columns for TSV or tag types and their structure for XML) and the initial data filters varied, so each knowledge source had its own wrapper.
Only information deemed pertinent to drug repositioning was included in the OREGANO knowledge graph, as the knowledge sources are overly large datasets. Since some sources provided information on the quality of their content, these were used to keep only the most relevant relations.
The NPASS data were filtered out because the targets come from a multitude of organisms (e.g. Angelica Gigas, Bos taurus or Saccharomyces cerevisiae). Therefore, only those relevant to humans were selected.
Only PharmGKB compounds associated with ATC codes with a length of seven digits were retained, as the other ATC codes correspond to drug classes (i.e. not to a given compound). Before integrating the links between drugs and diseases, another filtering process was applied. Indeed, the clinical annotations that highlight these links had different levels of evidence based on the associated research conducted to date. Only 1A, 1B, 2A and 2B levels were selected as they correspond to the links with the highest confidence levels (the detailed description of the different levels is available at: https://www.pharmgkb.org/page/clinAnnLevels).
For the integration of HPO, a filter was applied on the links between diseases and phenotypes. These links have a frequency of occurrence that is modelled by a calculation, as follows:
-
Excluded: present in 0% of cases.
-
Very rare: present in 1% to 4% of cases.
-
Occasional: present in 5% to 29% of cases.
-
Frequent: present in 30% to 79% of cases.
-
Very Frequent: present in 80% to 99% of cases.
-
Obligate: present in 100% of cases.
Below 30%, the frequency of occurrence of a symptom is thus occasional to rare. We thus made the choice to select only the phenotypes whose frequency of occurrence is greater than or equal to 30% in order to represent the diseases with the symptoms that most often describe these diseases. For the links that did not have such an indicator, the evaluation of the evidence of these links was used. The corresponding levels are: IEA (inferred from electronic annotation), PCS (published clinical study) and TAS (traceable author statement) (the detailed description of the different levels is available at: https://hpo-annotation-qc.readthedocs.io/en/latest/annotationFormat.html#phenotype-hpoa-format). Each of these levels describes how the associated link was obtained. The TAS and PCS levels have the highest confidence levels as they are derived from scientific works, while the IEA level corresponds to links inferred electronically. Therefore, only the TAS and PCS levels were kept when the frequency of occurrence was not available.
The graph integrates the HPO diseases with their OMIM codes (obtained through the UMLS). However, only the OMIM codes associated with HPO diseases were integrated to avoid overloading the knowledge graph.
UniProt is a very large knowledge source about genes and protein targets. Only the links between the nodes already present in the graph were conserved, in order to avoid introducing too many links that were not connected to the nodes of the graph.
The triples generated by the wrapper contained both edges of the knowledge graph and cross-references. The relationships were provided as input to the manager, while cross-references were directly transmitted to the binder.
C. Fusion of nodes from distinct knowledge sources
Knowledge sources did not necessarily provide mappings to other sources. In this case, a connection had to be established between the knowledge source to be integrated and the others. The connection could be created according to external data or knowledge sources and, if possible, a new source providing this link was added/used. This was the role of the binder.
The binder (middle part of Fig. 3) handled the creation and merging of nodes with the addition of new identifiers specific to the OREGANO knowledge graph into a global mapping table. To do this, a cross-reference match was sought to determine whether the node was new to the data. Depending on the cross-references, multiple candidates could exist for the same node. The goal of the binder algorithm was thus to be able to choose the best match for optimal data fusion. The correspondence of nodes was evaluated based on the number of common cross-references between two nodes. In the mapping table, each column corresponds to a knowledge source and each row represents a node with an OREGANO identifier in the first column. The binder also linked any OREGANO identifier to a new node if it was not already in the graph.
D. Constitution of the knowledge graph
The last algorithm of the integration process is performed by the manager algorithm. Indeed, the link manager (bottom part of Fig. 3) combined the links extracted by the wrapper and the mappings generated by the binder to format the links. Thus, the links with the original codes in the integrated knowledge sources were replaced by the OREGANO identifiers of the associated node in the mapping table.
The seven knowledge sources were integrated in a fully automatic way. The data were merged by evaluating whether the nodes were new or already present in the mapping table based on the cross-references. A node was not integrated if no linkage could be made for it.
Technical Validation
The technical validation of the OREGANO graph was carried out in four different ways: (i) a comparison with related knowledge graphs (subsection A.), (ii) a quality assessment according to criteria defined by Chen et al.57 (subsection B.), (iii) a practical assessment with the application of an embedding algorithm exhibiting the possibility of predicting links using the knowledge graph (subsection C.), and (iv) an example of a predicted link (subsection D.).
A. Comparison of OREGANO with other related knowledge graphs
Table 3 shows the biomedical knowledge graphs described in the Background & summary section, for which information about the number of nodes, edges, and integrated resources was available. We can see that the knowledge graphs from Zhu et al.22, PrimeKG31 and Hetionet29,30 have a significant number of nodes and relations, which is not surprising because the more knowledge sources the graph integrates and the more different types of nodes it includes, the larger the graph becomes. We can also notice that the OREGANO knowledge graph shares a similar set of data with some of these graphs, but none of them include data from PharmGKB and NPASS, and especially natural compound data. The OREGANO project thus offers a new angle to previous work by integrating this type of knowledge.
B. Evaluation regarding quality assessment
In the context of knowledge graph construction, there are four aspects that are usually neglected and need to be improved, according to Abu-Salih et al.58: (i) knowledge graphs must be accessible, (ii) construction methods must be explicit and detailed, (iii) knowledge sources must be of high quality, and finally (iv) the graph must prove its efficiency and usefulness in reality. The OREGANO knowledge graph meets the first three requirements, and the fourth will be addressed in future work.
Further, to evaluate the quality of the knowledge graph, we used the framework described by Zhu et al.57. The evaluation criteria of other works mentioned in the related works59,60 are more suitable for knowledge graphs with a hierarchical ontological structure, which is not the case of our knowledge graph. Zhu et al. exposed 18 criteria for evaluating the quality of a knowledge graph. These criteria are presented in Table 4 according to four levels of consistency for each of them, ranking from perfect consistency (+++) to inconsistency (−). This table shows that for most of the criteria, the graph is of good quality. More precisely, 15 out of 18 criteria are well addressed in this current version of the OREGANO knowledge graph, while three remain to be addressed.
C. Link prediction
The ultimate aim of the OREGANO project is to use its knowledge graph for drug repositioning as a knowledge base for discovering new links between molecules and targets. Node embedding algorithms can be used for link prediction over a knowledge graph61,62,63,64,65,66. By fitting such algorithms to a knowledge graph, they predict whether unknown edges have a high probability of existing. Each of the algorithms uses a different calculation to try to best predict the edges included in the knowledge graph (Supplementary data 1), and metrics are used to measure the probability of obtaining edges that actually exist in the graph. In this way, the knowledge graph was subjected to several node folding algorithms. To test the ability of the OREGANO knowledge graph to predict links, the whole graph was used (Supplementary data 2).
The statistical metrics used are the MRR and the Hit@N. Each one measures the possibility of obtaining a correct answer, in general for the MRR and in the first N suggestions for the Hit@N (details about these metrics are also provided in Supplementary data 1).
The PyKEEN library was used for each of the algorithms67. This is a Python package designed to train and evaluate knowledge graph embedding models. Each of the tested models was run for 50 epochs using the hyperparameters obtained after a parameter optimisation. Each model was run five times on five training sets from the knowledge graph. Each number is the average of the results over the five datasets.
We can see that link prediction over OREGANO performs best with the ComplEx algorithm (Table 5). The results show that with this algorithm, it is possible to obtain links with a high probability of existing at 42% of Hit@10. In other words, there is a 42% chance of obtaining a good result in the first 10 predictions.
D. Empirical predictions
Predictions were made on the entire knowledge graph using our best model (i.e., ComplEx as shown in Table 5). A set of link predictions was produced on the “has_target” link from the Compound entity in the knowledge graph. The top 10 predictions were retained for each compound. The results were then ranked from highest to lowest score. From the best results, we selected an example of a natural compound to illustrate the value of integrating them for drug repositioning (detailed statistics regarding the 22,676 natural compounds included in OREGANO can be found in Supplementary data 3).
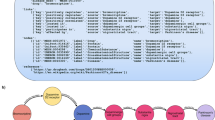
One of the best predictions was “COMPOUND:10025 has_target PROTEIN:4003”. Compound 10025 is epigallocatechin-gallate (EGCg) and protein 4003 is DNA polymerase k (Pol k). EGCg is the most abundant catechin in tea. In plants, catechins are secondary metabolites with antioxidant properties, belonging to the flavonoid subgroup of polyphenols68. Its medicinal uses are the subject of much research; EGCg has proven anti-cancer properties, notably against lung cancer69. Pol k is a member of the Y family of DNA polymerases, which have the ability to tolerate DNA damage during replication. Thanks to its ability to copy DNA, it plays a key role in maintaining genomic integrity70. It is also much studied for the role it plays in certain types of cancer. In many cancers, its expression is altered, inducing a disorder of genomic stability. The identification of Pol k inhibitors is therefore an important area of research; Pol k deregulation has been demonstrated in lung cancer71 in particular.
EGCg has been shown to inhibit all DNA polymerases except β72. By comparing all the data available on these two molecules, a more in-depth study could be envisaged to investigate the possibility of treating certain types of cancer whose Pol k regulation is altered with EGCg.
Code availability
The code for the integration and the knowledge graph are available on the GitHub of the OREGANO project in the Integration folder (https://gitub.u-bordeaux.fr/erias/oregano).
References
Mohs, R. C. & Greig, N. H. Drug discovery and development: Role of basic biological research. Alzheimer’s & Dementia: Translational Research & Clinical Interventions 3, 651–657, https://doi.org/10.1016/j.trci.2017.10.005 (2017).
Sun, D., Gao, W., Hu, H. & Zhou, S. Why 90% of clinical drug development fails and how to improve it? Acta Pharmaceutica Sinica B S2211383522000521, https://doi.org/10.1016/j.apsb.2022.02.002 (2022).
Jourdan, J.-P., Bureau, R., Rochais, C. & Dallemagne, P. Drug repositioning: a brief overview. Journal of Pharmacy and Pharmacology 72, 1145–1151, https://doi.org/10.1111/jphp.13273 (2020).
Sakate, R. & Kimura, T. Drug repositioning trends in rare and intractable diseases. Drug Discovery Today 27, 1789–1795, https://doi.org/10.1016/j.drudis.2022.01.013 (2022).
Pushpakom, S. et al. Drug repurposing: progress, challenges and recommendations. Nature Reviews Drug Discovery 18, 41–58, https://doi.org/10.1038/nrd.2018.168 (2019).
Jarada, T. N., Rokne, J. G. & Alhajj, R. A review of computational drug repositioning: strategies, approaches, opportunities, challenges, and directions. Journal of Cheminformatics 12, 46, https://doi.org/10.1186/s13321-020-00450-7 (2020).
Tanoli, Z. et al. Exploration of databases and methods supporting drug repurposing: a comprehensive survey. Briefings in Bioinformatics 22, 1656–1678, https://doi.org/10.1093/bib/bbaa003 (2021).
Yu, J.-L., Dai, Q.-Q. & Li, G.-B. Deep learning in target prediction and drug repositioning: Recent advances and challenges. Drug Discovery Today 27, 1796–1814, https://doi.org/10.1016/j.drudis.2021.10.010 (2022).
Hogan, A. et al. Knowledge graphs. ACM Computing Surveys 54, https://doi.org/10.1145/3447772 (2021).
Berners-Lee, T., Hendler, J. & Lassila, O. The Semantic Web. Scientific American 284, 34–43, https://doi.org/10.1038/scientificamerican0501-34 (2001).
Heath, T. & Bizer, C. Linked data: Evolving the web into a global data space. Synthesis Lectures on the Semantic Web: Theory and Technology 1, 1–136, https://doi.org/10.2200/S00334ED1V01Y201102WBE001 (2011).
Belleau, F., Nolin, M.-A., Tourigny, N., Rigault, P. & Morissette, J. Bio2RDF: Towards a mashup to build bioinformatics knowledge systems. Journal of Biomedical Informatics 41, 706–716, https://doi.org/10.1016/j.jbi.2008.03.004 (2008).
Lotfi Shahreza, M., Ghadiri, N., Mousavi, S. R., Varshosaz, J. & Green, J. R. A review of network-based approaches to drug repositioning. Briefings in Bioinformatics 19, 878–892, https://doi.org/10.1093/bib/bbx017 (2018).
MacLean, F. Knowledge graphs and their applications in drug discovery. Expert Opinion on Drug Discovery 16, 1057–1069, https://doi.org/10.1080/17460441.2021.1910673 (2021).
Bonner, S. et al. A review of biomedical datasets relating to drug discovery: a knowledge graph perspective. Briefings in Bioinformatics 23, bbac404, https://doi.org/10.1093/bib/bbac404 (2022).
Cheng, F. et al. Prediction of drug-target interactions and drug repositioning via network-based inference. PLOS Computational Biology 8, 1–12, https://doi.org/10.1371/journal.pcbi.1002503 (2012).
Mei, J.-P., Kwoh, C.-K., Yang, P., Li, X.-L. & Zheng, J. Drug–target interaction prediction by learning from local information and neighbors. Bioinformatics 29, 238–245, https://doi.org/10.1093/bioinformatics/bts670 (2013).
Dai, Y.-F. & Zhao, X.-M. A survey on the computational approaches to identify drug targets in the postgenomic era. BioMed Research International 2015, 1–9, https://doi.org/10.1155/2015/239654 (2015).
Chen, H.-R., Sherr, D. H., Hu, Z. & DeLisi, C. A network based approach to drug repositioning identifies plausible candidates for breast cancer and prostate cancer. BMC Medical Genomics 9, 51, https://doi.org/10.1186/s12920-016-0212-7 (2016).
Li, Z., Wang, R.-S. & Zhang, X.-S. Two-stage flux balance analysis of metabolic networks for drug target identification. BMC Systems Biology 5, S11, https://doi.org/10.1186/1752-0509-5-S1-S11 (2011).
Orth, J. D., Thiele, I. & Palsson, B. What is flux balance analysis. Nature Biotechnology 28, 245–248, https://doi.org/10.1038/nbt.1614 (2010).
Zhu, Q. et al. An integrative knowledge graph for rare diseases, derived from the Genetic and Rare Diseases Information Center (GARD). Journal of Biomedical Semantics 11, 13, https://doi.org/10.1186/s13326-020-00232-y (2020).
Azuaje, F. Drug interaction networks: an introduction to translational and clinical applications. Cardiovascular Research 97, 631–641, https://doi.org/10.1093/cvr/cvs289 (2013).
Fukuoka, Y., Takei, D. & Ogawa, H. A two-step drug repositioning method based on a protein-protein interaction network of genes shared by two diseases and the similarity of drugs. Bioinformation 9, 89–93, https://doi.org/10.6026/97320630009089 (2013).
Chen, H., Zhang, H., Zhang, Z., Cao, Y. & Tang, W. Network-based inference methods for drug repositioning. Computational and Mathematical Methods in Medicine 2015, 1–7, https://doi.org/10.1155/2015/130620 (2015).
Zhou, T., Ren, J., Medo, M. & Zhang, Y.-C. Bipartite network projection and personal recommendation. Physical Review E 76, 046115, https://doi.org/10.1103/PhysRevE.76.046115 (2007).
Ye, H., Liu, Q. & Wei, J. Construction of drug network based on side effects and its application for drug repositioning. PLoS ONE 9, e87864, https://doi.org/10.1371/journal.pone.0087864 (2014).
Tao, C., Sun, J., Zheng, W. J., Chen, J. & Xu, H. Colorectal cancer drug target prediction using ontology-based inference and network analysis. Database 2015, bav015, https://doi.org/10.1093/database/bav015 (2015).
Himmelstein, D. S. & Baranzini, S. E. Heterogeneous Network Edge Prediction: A Data Integration Approach to Prioritize Disease-Associated Genes. PLOS Computational Biology 11, e1004259, https://doi.org/10.1371/journal.pcbi.1004259 (2015).
Himmelstein, D. S. et al. Systematic integration of biomedical knowledge prioritizes drugs for repurposing. eLife 6, e26726, https://doi.org/10.7554/eLife.26726 (2017).
Chandak, P., Huang, K. & Zitnik, M. Building a knowledge graph to enable precision medicine. Scientific Data 10, 67, https://doi.org/10.1038/s41597-023-01960-3 (2023).
Boudin, M. Computational Approaches for Drug Repositioning: Towards a Holistic Perspective based on Knowledge Graphs. In Proceedings of the 29th ACM International Conference on Information & Knowledge Management, 3225–3228, https://doi.org/10.1145/3340531.3418510 (ACM, Virtual Event Ireland, 2020).
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. Journal of Natural Products 75, 311–335, https://doi.org/10.1021/np200906s (2012).
Li, J. et al. Traditional chinese medicine-based network pharmacology could lead to new multicompound drug discovery. Evidence-Based Complementary and Alternative Medicine 2012, 1–11, https://doi.org/10.1155/2012/149762 (2012).
Yoo, W.-H. et al. Efficacy and safety of PG201 (layla®) and celecoxib in the treatment of symptomatic knee osteoarthritis: a double-blinded, randomized, multi-center, active drug comparative, parallel-group, non-inferiority, phase III study. Rheumatology International 34, 1369–1378, https://doi.org/10.1007/s00296-014-2964-8 (2014).
Choi, W. et al. HerDing: herb recommendation system to treat diseases using genes and chemicals. Database 2016, baw011, https://doi.org/10.1093/database/baw011 (2016).
Wang, N. et al. Herb Target Prediction Based on Representation Learning of Symptom related Heterogeneous Network. Computational and Structural Biotechnology Journal 17, 282–290, https://doi.org/10.1016/j.csbj.2019.02.002 (2019).
Wishart, D. S. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Research 34, D668–D672, https://doi.org/10.1093/nar/gkj067 (2006).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Research 46, D1074–D1082, https://doi.org/10.1093/nar/gkx1037 (2017).
Kuhn, M., Campillos, M., Letunic, I., Jensen, L. J. & Bork, P. A side effect resource to capture phenotypic effects of drugs. Molecular Systems Biology 6, 343, https://doi.org/10.1038/msb.2009.98 (2010).
Kuhn, M., Letunic, I., Jensen, L. J. & Bork, P. The SIDER database of drugs and side effects. Nucleic Acids Research 44, D1075–D1079, https://doi.org/10.1093/nar/gkv1075 (2016).
The UniProt Consortium. et al. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Research 49, D480–D489, https://doi.org/10.1093/nar/gkaa1100 (2021).
Boutet, E., Lieberherr, D., Tognolli, M., Schneider, M. & Bairoch, A. UniProtKB/Swiss-Prot. In Edwards, D. (ed.) Plant Bioinformatics: Methods and Protocols, Methods in Molecular BiologyTM, 89–112, https://doi.org/10.1007/978-1-59745-535-0_4 (Humana Press, 2007).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Research 45, D158–D169, https://doi.org/10.1093/nar/gkw1099 (2017).
Jassal, B. et al. The reactome pathway knowledgebase. Nucleic Acids Research 48, D498–D503, https://doi.org/10.1093/nar/gkz1031 (2019).
Whirl-Carrillo, M. et al. Pharmacogenomics Knowledge for Personalized Medicine. Clinical Pharmacology & Therapeutics 92, 414–417, https://doi.org/10.1038/clpt.2012.96 (2012).
Whirl-Carrillo, M. et al. An Evidence-Based Framework for Evaluating Pharmacogenomics Knowledge for Personalized Medicine. Clinical Pharmacology & Therapeutics 110, 563–572, https://doi.org/10.1002/cpt.2350 (2021).
Köhler, S. et al. The Human Phenotype Ontology in 2021. Nucleic Acids Research 49, D1207–D1217, https://doi.org/10.1093/nar/gkaa1043 (2021).
Amberger, J. S., Bocchini, C. A., Schiettecatte, F., Scott, A. F. & Hamosh, A. OMIM.org: Online mendelian inheritance in man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Research 43, D789–D798, https://doi.org/10.1093/nar/gku1205 (2015).
Orphanet: an online rare disease and orphan drug data base. © INSERM 1999, http://www.orpha.net.
Firth, H. V. et al. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources. The American Journal of Human Genetics 84, 524–533, https://doi.org/10.1016/j.ajhg.2009.03.010 (2009).
Zeng, X. et al. NPASS: natural product activity and species source database for natural product research, discovery and tool development. Nucleic Acids Research 46, D1217–D1222, https://doi.org/10.1093/nar/gkx1026 (2018).
Zhao, H. et al. NPASS database update 2023: quantitative natural product activity and species source database for biomedical research. 51, D621–D628, https://doi.org/10.1093/nar/gkac1069.
Bodenreider, O. The Unified Medical Language System (UMLS): integrating biomedical terminology. Nucleic Acids Research 32, 267D–270, https://doi.org/10.1093/nar/gkh061 (2004).
Boudin, M., Diallo, G., Mougin, F. & Drancé, M. The OREGANO knowledge graph for computational drug repurposing. Figshare https://doi.org/10.6084/m9.figshare.23553114.v3 (2023).
Boudin, M., Diallo, G., Mougin, F. & Drancé, M. The OREGANO knowledge graph for computational drug repurposing. Zenodo https://doi.org/10.5281/zenodo.10103842 (2023).
Chen, H., Cao, G., Chen, J. & Ding, J. A practical framework for evaluating the quality of knowledge graph. In Zhu, X., Qin, B., Zhu, X., Liu, M. & Qian, L. (eds.) Knowledge Graph and Semantic Computing: Knowledge Computing and Language Understanding, vol. 1134, 111–122, https://doi.org/10.1007/978-981-15-1956-7_10 (2019).
Abu-Salih, B. et al. Healthcare knowledge graph construction: State-of-the-art, open issues, and opportunities, http://arxiv.org/abs/2207.03771 (2022).
Wang, X. et al. Knowledge graph quality control: A survey. Fundamental Research 1, 607–626, https://doi.org/10.1016/j.fmre.2021.09.003 (2021).
Zaveri, A. et al. Quality assessment for linked data: A survey: A systematic literature review and conceptual framework. Semantic Web 7, 63–93, https://doi.org/10.3233/SW-150175 (2015).
Bordes, A., Usunier, N., Garcia-Durán, A., Weston, J. & Yakhnenko, O. Translating embeddings for modeling multi-relational data. In Proceedings of the 26th International Conference on Neural Information Processing Systems - Volume 2, NIPS’13, 2787–2795, https://doi.org/10.5555/2999792.2999923 (Curran Associates Inc., Red Hook, NY, USA, 2013).
Wang, Z., Zhang, J., Feng, J. & Chen, Z. Knowledge graph embedding by translating on hyperplanes. In Proceedings of the Twenty-Eighth AAAI Conference on Artificial Intelligence, AAAI’14, 1112–1119, https://doi.org/10.5555/2893873.2894046 (AAAI Press, 2014).
Lin, Y., Liu, Z., Sun, M., Liu, Y. & Zhu, X. Learning entity and relation embeddings for knowledge graph completion. Proceedings of the AAAI Conference on Artificial Intelligence 29, https://doi.org/10.1609/aaai.v29i1.9491 (2015).
Sun, Z., Deng, Z.-H., Nie, J.-Y. & Tang, J. RotatE: Knowledge graph embedding by relational rotation in complex space. arXiv https://arxiv.org/abs/1902.10197 (2019).
Yang, B., Yih, W.-t., He, X., Gao, J. & Deng, L. Embedding entities and relations for learning and inference in knowledge bases. arXiv http://arxiv.org/abs/1412.6575 (2015).
Trouillon, T., Welbl, J., Riedel, S., Gaussier, E. & Bouchard, G. Complex embeddings for simple link prediction. In Balcan, M. F. & Weinberger, K. Q. (eds.) Proceedings of The 33rd International Conference on Machine Learning, vol. 48 of Proceedings of Machine Learning Research, 2071–2080, https://doi.org/10.5555/3045390.3045609 (2016).
Ali, M. et al. PyKEEN 1.0: A Python Library for Training and Evaluating Knowledge Graph Embeddings. Journal of Machine Learning Research 22, 1–6, https://doi.org/10.48550/arXiv.2007.14175 (2021).
Legeay, S., Rodier, M., Fillon, L., Faure, S. & Clere, N. Epigallocatechin Gallate: A Review of Its Beneficial Properties to Prevent Metabolic Syndrome. Nutrients 7, 5443–5468, https://doi.org/10.3390/nu7075230 (2015).
Chen, B.-H., Hsieh, C.-H., Tsai, S.-Y., Wang, C.-Y. & Wang, C.-C. Anticancer effects of epigallocatechin-3-gallate nanoemulsion on lung cancer cells through the activation of AMP-activated protein kinase signaling pathway. Scientific Reports 10, 5163, https://doi.org/10.1038/s41598-020-62136-2 (2020).
Stern, H. R., Sefcikova, J., Chaparro, V. E. & Beuning, P. J. Mammalian DNA Polymerase Kappa Activity and Specificity. Molecules 24, 2805, https://doi.org/10.3390/molecules24152805 (2019).
Pillaire, M.-J., Bétous, R. & Hoffmann, J.-S. Role of DNA polymerase ΰ in the maintenance of genomic stability. Molecular & Cellular Oncology 1, e29902, https://doi.org/10.4161/mco.29902 (2014).
Mizushina, Y. et al. Structural analysis of catechin derivatives as mammalian DNA polymerase inhibitors. Biochemical and Biophysical Research Communications 333, 101–109, https://doi.org/10.1016/j.bbrc.2005.05.093 (2005).
Huntley, R. P. et al. A method for increasing expressivity of Gene Ontology annotations using a compositional approach. BMC Bioinformatics 15, 155, https://doi.org/10.1186/1471-2105-15-155 (2014).
Breit, A., Ott, S., Agibetov, A. & Samwald, M. OpenBioLink: a benchmarking framework for large-scale biomedical link prediction. Bioinformatics 36, 4097–4098, https://doi.org/10.1093/bioinformatics/btaa274 (2020).
Walsh, B., Mohamed, S. K. & Nováček, V. BioKG: A knowledge graph for relational learning on biological data. In Proceedings of the 29th ACM International Conference on Information & Knowledge Management, 3173–3180, https://doi.org/10.1145/3340531.3412776 (ACM, 2020).
Hijazi, M., Smith, R., Rajeeve, V., Bessant, C. & Cutillas, P. R. Reconstructing kinase network topologies from phosphoproteomics data reveals cancer-associated rewiring. Nature Biotechnology 38, 493–502, https://doi.org/10.1038/s41587-019-0391-9 (2020).
Acknowledgements
The PhD thesis of Marina Boudin is supported within the framework of the Digital Public Health programme of PIA3 (Investment for the future). Project reference: 17-EURE-0019.
Author information
Authors and Affiliations
Contributions
M.B. conceived the experiment(s) and drafted the first version of the manuscript. M.B. and M.D. conducted the experiment(s). G.D. and F.M. designed the overall project and revised the manuscript. All authors analysed the results and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boudin, M., Diallo, G., Drancé, M. et al. The OREGANO knowledge graph for computational drug repurposing. Sci Data 10, 871 (2023). https://doi.org/10.1038/s41597-023-02757-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02757-0